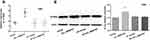Back to Journals » Cancer Management and Research » Volume 12
Effect of the Up-Regulation of Circular RNA Hsa_circ_0069767 Derived from C-KIT on the Biological Behavior of Multiple Myeloma Cells
Authors Chen F, Wang X, Fu S, Wang S, Fu Y, Liu Z, Zhang J
Received 8 May 2020
Accepted for publication 22 October 2020
Published 6 November 2020 Volume 2020:12 Pages 11321—11331
DOI https://doi.org/10.2147/CMAR.S259393
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Rudolph Navari
Fang Chen,1,2 Xiaohui Wang,1 Shuang Fu,1 Shaokun Wang,1 Yu Fu,1 Zhuogang Liu,2 Jihong Zhang1
1Hematology Laboratory, Shengjing Hospital of China Medical University, Shenyang 110022, People’s Republic of China; 2Department of Hematology, Shengjing Hospital of China Medical University, Shenyang 110022, People’s Republic of China
Correspondence: Jihong Zhang
Hematology Laboratory, Shengjing Hospital of China Medical University, No. 39 Huaxiang Road, Tiexi District, Shenyang 110022, People’s Republic of China
Tel/Fax +86 24-96615-24811
Email [email protected]
Purpose: Multiple myeloma (MM) is an incurable disease. This study focused on the expression of circular RNA circ_0069767 in MM and its influence on prognosis, in order to provide a potential target.
Patients and Methods: Totally 66 MM patients participated in this research. Using RT-PCR method to determine the expression level of circ_0069767 in 66 sorted samples from multiple myeloma patients and 21 normal control bone marrow samples, Kaplan–Meier was applied for survival analysis. We constructed stable over-expressing circ_0069767 and silenced circ_0069767 cell lines and used MTS experiment to detect cell viability, transwell experiment to detect cell migration and invasion ability and flow cytometry to detect cell apoptosis. Dual luciferase experiment, qRT-PCR experiment and Western blot were used to explore miRNA and downstream genes.
Results: The expression of circ_0069767 in MM was significantly higher than that of the normal control group. Patients with high expression of circ_0069767 had longer PFS and OS. Cell function experiments showed that overexpression of circ_0069767 in MM cells led to decreased proliferation, migration and invasion, but increased apoptosis; meanwhile, knockdown of circ_0069767 caused opposite biological behaviors. Circ_0069767 by sponging miR-636 in MM cells regulates the expression of K-RAS while the K-RAS gene remained unmutated.
Conclusion: Circ_0069767 plays an antitumor role and its expression can be used as a reliable prognostic indicator for MM patients.
Keywords: circular RNA, multiple myeloma, prognostic factor
Introduction
Multiple myeloma (MM) is a clonal plasma cell proliferative disorder as the second most common hematological malignant tumor, accounting for about 15% of all hematological malignancies. The disease is considered incurable,1 but due to the introduction of autologous stem cell transplant (ASCT), the availability of new drugs, and the application of novel immunotherapies, the therapeutic effect, prognosis of MM, and the 5-year overall survival (OS) has been significantly improved; nonetheless, most patients eventually suffer from relapse.2 Therefore, identifying factors that affect the prognosis of MM remains clinically important.
Circular RNA (circRNA) is a covalently closed endogenous non-coding RNA that is gradually recognized following miRNA and long non-coding RNA (lncRNA). CircRNA has neither the polarity of the 3ʹ and 5ʹ ends nor a polyA tail and is characterized by a stable structure (uneasily degraded by endonucleases)3–6 and tissue and disease-specific expression patterns. Moreover, circRNA plays multiple regulatory roles in carcinogenesis, and therefore has potential diagnostic and prognostic values in cancer evaluation.
With the development of sequencing technique, a large number of circRNAs have been discovered. Researches on this new type of non-coding RNA revealed that it is highly associated with molecular biology and molecular oncology. To date, a number of human circRNAs have been identified.7–13 With the in-depth study of structure and function of circRNAs, it was found to be involved in the occurrence and development of multiple diseases and is considered to be a key regulator in a wide range of biological processes.14–20 However, it is rarely reported in MM.
In our previous studies, MM patients with high C-KIT (CD117) expression were observed of better prognosis compared with patients with low C-KIT (CD117) expression.21 According to the circBase database, there are 12 circular RNAs derived from the C-KIT gene, and circ_0069767 was found in the K562 cells, which belong to the same blood tumor as multiple myeloma. So I tried qRT-PCR detection of circ_0069767 first, and the pre-experiment results showed that there was a difference in expression of MM compared with the normal control group. So we chose circ_0069767 as the target gene for follow-up research. To further explore the effect of circ_0069767 on tumor cells, we built stable overexpressed circ_0069767 and silenced circ_0069767 cell lines based on MM cell KM3 and explored its function through in-vitro experiments which demonstrated that KM3 cells altered biological behaviors including decreased cell viability, migration and invasion capacities, and increased cell apoptosis.
Patients and Methods
Patients and Clinical Samples
A total of 66 bone marrow samples from MM patients and 21 healthy control samples (healthy donors for bone marrow transplants) were collected from patients treated at Shengjing Hospital of China Medical University between January 2014 and December 2018. All patients were diagnosed and newly treated by the updated International Myeloma Working Group (IMWG) criteria and evaluated for staging based on the International Staging System (ISS) and Durie-Salmon (DS). All patients enrolled in this study did not receive any treatment before sampling and were treated with bortezomib-based combination chemotherapy after admission. The study protocol was reviewed and approved by the Ethics Committee of Shengjing Hospital of China Medical University, and all participants provided informed consents.
Magnetic-Activated Cell Sorting
All bone marrow samples were residual samples from routine clinical laboratory tests, from which MM mononuclear cells (MNCs) were extracted using magnetic-activated cell sorting (Miltenyi, Germany) following the manufacturer’s instruction. The extracted bone marrow MNCs were added to Trizol reagent (Takara, Japan) and cryopreserved at −80°C for future testing.
Total RNA Extraction and qRT-PCR
The Trizol method was used to extract total RNAs from the sorted MM cells following the manufacturer’s instructions. The extracted total RNA was reverse transcribed using PrimeScript™RT reagent Kits, and real-time PCR was performed using TAKARA SYBR ® Premix Ex Taq ™ II kits (TaKaRa, Dalian, China) on the ABI 7500HT. The process of the thermal cycler was as follows: 95°C5min→(95°C5s, 60°C34s)×40cycles→95°C15s →60°C1min→ 95°C15s.
All reactions were performed in triplicate. A reverse primer was designed to amplify the head-to-tail splice junction of the circ_0069767, the sequences were F:5ʹ-GTAATCGTAGCTGGCATGAT-3ʹ, R:5ʹ-GAATGCTTCATATTCAACAATC-3ʹ, with the GAPDH as an internal reference gene, the primer sequences was F: 5ʹ-TGTTCGTCATGGGTGTGAAC-3ʹ, R: 5ʹ-ATGGCATGGACTGTGGTCAT-3ʹ. The relative expression of circ_0069767 was calculated using the 2-ΔΔCT method. We used the median value of circ_0069767 expression as a boundary, and divided 66 patients into high expression group and low expression group.
Cell Culture and Construction of Stably Transfected Cell Lines
MM cell line KM3 was selected for subsequent experiments for positive expression of CD117. The expression of CD117 in KM3 lines is about 82% (Figure S1). MM cell line KM3 and human embryonic kidney cell line HEK-293T were purchased from Cell Bank of the Chinese Academy of Sciences (Shanghai, China) and ATCC. Cells were kept in RPMI 1640 or Dulbecco’s Modified Eagle Medium (DMEM) containing 10% fetal bovine serum (HyClone, USA) in humidified 37°C incubators with 5% CO2.
The lentivirus vector mediating the overexpression of circRNA_0069767 was provided by Liaoning Baihao Biotechnology Co., Ltd. (Benxi, China), which was constructed by cloning circRNA_0069767 cDNA into PLCDH vector. A reverse indirect linker sequence of the silenced sh-circRNA specifically targeting circRNA_0069767 was designed with an shRNA sequence of:
AAGTTTACAGATAAAGGATTCATTTTGGATCCAAAATGAATCCTTTATCTGTAAC.
After lentivirus infection, antibiotics screenings were performed to select stably transfected cells which were overexpressing circ_0069767 cells (circ_0069767), empty vector cells (vector), silenced circ_0069767 cells (sh-circ_0069767), and silenced empty vector cells (sh- vector).
Cell Viability Assay
The MTS assay (Promega) was used to test the viability of all cells. The four groups of cells were centrifuged, the supernatant was discarded, and approximately 3mL of RPMI 1640 was added then mixed. The cells were seeded in 96-well plates with 1 * 104 cells per well. Cell colony formation was detected in 5 replicates for each group (24h, 48h, 72h, and 96h, respectively). MTS (20 µl) was added to each well and incubated for 2 hours before detection.
Cell Migration and Invasion Assays
The cells were centrifuged, the supernatant was discarded, and approximately 5 mL of culture solution was added. The cells were seeded in cell chamber slides with 1×105 cells per chamber. A layer of matrigel was coated onto the chambers’ polycarbonate membrane for the invasion assay. Culture solution that contained a semi-solid medium made from agarose (500 µl) was added to each chamber (3 duplicate chambers were set). After a cultivation period of 24h images were obtained under a microscope. The cells in the lower chamber were counted with a cell counter.
Apoptosis Assay
The cells were centrifuged at 1000 rpm for 5 min, washed twice with PBS and then centrifuged at 1000 rpm for 5 min. Then, the cells were collected then added 100 µL binding buffer to prepare a single-cell suspension with a density of 2×105/mL. 5 µL Annexin V-APC and 5 µL 7-AAD (Becton Dickinson) were added per tube and mixed thoroughly. Each mixture was then incubated at room temperature for 10 min and then detected using flow cytometry. Empty cell controls were set and the above procedures were repeated.
Immunophenotyping
Immunophenotypic analysis was performed using a FACS Calibur 4-color flow cytometer (Becton, Dickinson and Company, USA). Monoclonal mouse anti-human antibodies were labeled using fluoresceins including CD45-Percp (Peridinin-Chlorophyll-Protein), CD117-PE (phycoerythrin), and CD38-APC (allophycocyanin). The antibodies (Becton Dickinson), after binding to antigen, were kept in a dark place for 15 mins. BD FACS Lysing Solution was added for hemolysis for 10 min. Following antibody labeling, 50,000 cells were analyzed with CellQuest software on a flow cytometer.
Luciferase Reporter Assay
We predicted the targeted miRNAs that may be bound to circ_0069767 by three online tools (https://omictools.com/circnet-tool, http://regrna2.mbc.nctu.edu.tw/, https://circinteractome.nia.nih.gov/). The full length of circ_0069767 and 3ʹ-UTR of K-RAS containing wild-type or mutant miR-636 binding site were synthesized and cloned into pmirGLO REPORT vector, followed by co-transfection with miRNAs mimics into HEK-293T cells using Lipofectamine™ 3000 (Invitrogen). The relative luciferase activity was detected by the Dual-Luciferase Reporter Assay Kit purchased from Promega company (Madison, WI, USA).
Western Blot
Cells from each group were collected and protease and phosphatase inhibitors were added to extract total protein, which was then dyed and electrophoresed on a 10% SDS-PAGE gel. After electrophoresis, the total protein was transferred to polyvinylidene difluoride (PVDF) parts, and then blocked with 5% skimmed milk powder. Thereafter, the membrane was incubated with an anti-K-RAS (# 180772, Abcam) primary antibody at 4° C overnight and with an anti-rabbit secondary antibody for 1h at room temperature the next day. The membrane was developed using a chemiluminescent solution in a dark room.
Next-Generation Sequencing
The next-generation sequencing technology used Illumina/Sol-exa GenomeAnalyzer (Illumina, USA) to complete K-RAS gene sequencing.
Statistical Analyses
All statistical analyses were performed using SPSS 19.0 software (IBM, USA), and the data reported as mean ± SD from at least three independent experiments. Differences between groups were analyzed by Student’s t-test and Kruskal–Wallis test. P value less than 0.05 was considered statistically significant. *p<0.05, **p<0.01, ***p<0.001.
Results
Increased Circ_0069767 Expression in MM
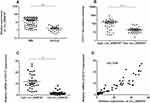
QRT-PCR analysis was performed on the 66 sorted MM bone marrow samples with purity of 90.76% (Figure S2) and 21 normal control bone marrow samples which confirmed that circ_0069767 expressed higher in MM samples than in normal control samples (Figure 1A). By the median expression of circ_0069767 in MM patients (median = 8.81), patients were divided into “high” circ_0069767 and “low” circ_0069767 groups. The results showed that low circ_0069767 was closely correlated with advanced Durie-Salmon (DS) stage (Table 1). Our present study results indicated low expression circ_0069767 patients had more frequent p53 deletion than high expression circ_0069767 patients (Table 2). QRT-PCR analysis showed that the higher expression of mRNA of CD117 was related to higher expression of circ_0069767 (Figure 1C), also there was a strong positive correlation between circ_0069767 and mRNA expression of CD117 in MM tissues (Figure 1D). And the average fluorescence intensity of CD117 protein encoded by the KIT gene showed that patients with stronger CD117 fluorescence intensity had higher expression of circ_0069767 (Figure 1B). At the same time, we used qRT-PCR to test some rechecked patient samples, including 6 CR patients, 7 VGPR patients, 8 PR patients and 13 PD patients. The results showed that the expression of circ_0069767 in PD patients was the lowest (Figure S3).
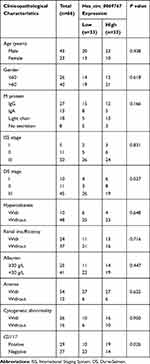 |
Table 1 Difference in the Hsa_circ_0069767 Expression in Multiple Myeloma Patients Grouped by Clinicopathological Characteristics |
 |
Table 2 The Hsa_circ_0069767 Expression as Grouped by FISH |
Kaplan–Meier Survival Analysis Results Indicated That High Circ_0069767 Led to a Longer PFS and OS
All 66 patients were treated with bortezomib-based chemotherapy. Kaplan-Meier survival analyses of the high and low circ_0069767 groups indicated that the high circ_0069767 group had a longer progression-free survival (median, 15 vs 10 months; p<0.05) and overall survival (median, 35 vs 21 months; p<0.001; Figure 2).
Circ_0069767 Overexpression Led to Reduced Cell Viability
To clarify the biological function of circ_0069767 in MM, we built stable circ_0069767 overexpressed and silenced cell lines (Figure 3A) and performed a series of functional assays. Multiple myeloma cell lines were used for viability assays. MTS assay results suggest that the cells with overexpressed circ_0069767 had decreased viability, while sh-circ cells had restored viability (Figure 3B).
The Cells Overexpressing Circ_0069767 Had Decreased Migration and Invasion Capacities
We performed transwell assays to detect the invasion and migration capacities of the four cell lines (circ_0069767, vector, sh-circ_0069767, sh-vector). The stable cell line overexpressing circ_0069767 showed decreased invasion and migration capacities (Figure 4A and B).
The Cells Overexpressing Circ_0069767 Showed Increased Apoptosis
Flow cytometry was used to test apoptosis of circ_0069767, vector, sh-circ_0069767, and sh-vector cell lines. Originally there were few apoptotic cells in each group, but after treated with bortezomib (IC50 values 100 ng/mL, 24h), the apoptosis of cells overexpressing circ_0069767 was significantly increased compared to other groups (Figure 5).
Circ_0069767acts as a Sponge of miR-636 in MM Cells
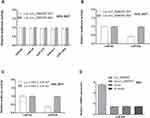
We predicted the targeted miRNAs capable of binding to circ_0069767 using different online tools, the overlapped results that met the criteria were miR-29, miR-370, miR-491, miR-636 and miR-1264. The full length of circ_0069767 wild-type was synthesized and cloned into pmirGLO REPORT vector, followed by co-transfection with miR-29, miR-370, miR-491, miR-636 and miR-1264 mimics into HEK-293T cells using Lipofectamine™ 3000. The relative luciferase activity was detected by the Dual-Luciferase Reporter Assay Kit. The results showed that overexpression of miR-636 was able to dramatically reduce the luciferase activity of wild-type circ_0069767 reporter, whereas other miRNAs had no effects on the wild-type circ_0069767 reporter. Then the full length of circ_0069767 mutant-type of miR-636 binding site and miR-636 mimics were co-transfected into HEK-293T cells, and the results showed that there was no interaction between them (Figure 6A and B). Then, we used the online miRWalk database to identify the downstream target of miR-636, and found that K-RAS, the well-known oncogene, may be bound by miR-636. To confirm such prediction, we conducted a luciferase reporter assay in HEK-293T cells. As shown in Figure 6C, miR-636 overexpression significantly decreased the luciferase activity of wild-type K-RAS 3ʹ-UTR reporter, while this effect was not observed in the mutant reporter. We then performed qRT-PCR assay in KM3 cells, and the results displayed that miR-636 expression was significantly decreased after circ_0069767 overexpression (Figure 6D).
The Cells Overexpressing Circ_0069767 Had Increased K-RAS Expression
RT-PCR analysis showed that relative expression of K-RAS was higher in cells overexpressing circ_0069767 (Figure 7A). Western blot analysis was repeated triple times and the results demonstrated that K-RAS protein expression was increased in cells overexpressing circ_0069767 (Figure 7B).
There Was No Mutation of K-RAS Gene
We performed second-generation gene sequencing on cells overexpressing circ_0069767, sh-circ_0069767, vector and sh-vector, and there was no mutation of K-RAS gene.
Discussion
The development of sequencing technique has led to the identification of circRNAs that are considered to be widely expressed in human cells. By joining the 3ʹ and 5ʹ ends together, circRNA forms a covalently closed continuous loop that confers excellent stability. Therefore, circRNA is increasingly expected to be ideal indicator for tumor diagnosis.22,23 An increasing number of studies have reported that circRNA is associated with the occurrence and development of tumors; for instance, the up-regulation or knockdown of circRNA_0020123 significantly affect the growth and metastasis of tumor cells in non-small cell lung cancer.24 Reducing the expression of hsa_circ_0000064 has been shown to inhibit cell proliferation, promote apoptosis, and regulate the cell cycle in lung cancer cells. Similarly, in liver cancer cells, circ_0000064 was revealed to promote the progression of the malignancy.25,26 In diffuse large B-cell lymphoma (DLBCL), circ-Foxo3 is closely related to cell cycle progression-the intervention with circ-Foxo3 accelerates the cell cycle and enhances proliferation capacity, while its overexpression has been shown to suppress cell cycle progression. Researchers also suggested that circ-Foxo3 inhibits cell cycle progression by forming a complex with cyclin-dependent kinase CDK2 and cyclin-dependent kinase inhibitor P21.27 These studies suggest that circRNAs are involved in the occurrence and development of tumors. In our study, based on information from the circBase database, circ_0069767 is derived from the transcript of the C-KIT protein (CD117) gene, with a sequence length of 722bp and located on chromosome 4 chr4: 55573263–55593490. Our results showed that circRNA circ_0069767 was differentially expressed in human MM cells and normal bone marrow cells and it was significantly correlated with the prognosis of MM patients. High expression of circ_0069767 in MM patients leads to improved PFS and OS, suggesting that circ_0069767 may be an eligible indicator to MM prognosis evaluation. To accurately assess the prognosis of MM and provide clinically reasonable and effective prognostic information, stable expression of biomarkers is particularly practical. For that circRNA has a closed-loop structure, it is very stable and easy to be detected, which makes it suitable as a biomarker for prognosis evaluation than linear RNA. In addition, MM cells overexpressing circ_0069767 demonstrated decreased proliferation, migration, and invasion capacities, while the MM cells with circ_0069767 knockdown demonstrated increased proliferation, migration, and invasion capacities. After bortezomib treatment, the apoptosis rate of each group was up-regulated, and the apoptosis of cells overexpressing circ_0069767 was significantly increased than other groups. It may be that the pathway of apoptosis activated by circ_0069767 and bortezomib was the same. And this is what our research group will study in the future. Cell function test results showed that circ_0069767 played a role of tumor suppressor gene in MM, explained why the higher circ_0069767 expression linked to better PFS and OS.
Our previous studies showed that patients with high expression of C-KIT protein (CD117) had better PFS and OS,21 and circ_0069767 was derived from the C-KIT gene, which may further explain why patients with positive CD117 have improved PFS and OS. Some research showed that circRNA was produced by co-transcription and competed with conventional splicing during the production process. Therefore, the biogenesis of circRNA lead to a decrease in mRNA synthesis from the same locus.28,29 But our research showed that circ_0069767 is positively correlated with the mRNA expression of CD117, so there may be other regulatory pathways. However, it is not yet clear how circ_0069767 affects the cellular biological behaviors, which will be the focus of subsequent studies.
We focused on the sponging function of circular RNA and bioinformatically predicted the targeted miRNAs and downstream mRNAs that circ_0069767 may competitively bind with and confirmed through experiments, the results of which demonstrated that circ_0060767 regulates the expression of K-RAS through miR-636. In other words, the circ_0069767/miR-636/K-RAS regulatory axis exists in the KM3 cells. We also noticed an interesting phenomenon regarding an association between high K-RAS mRNA expression and increased protein expression (p<0.001) in circ_0069767 overexpressing cells. As it is known that only when the K-RAS gene is mutated can it activate downstream signaling pathways, promote cell growth and proliferation, and inhibit cell senescence and death, eventually causing cell to become cancerous.30,31 In our study, the expression of K-RAS gene and protein increased after overexpression of circ_0069767 while the K-RAS gene remain unmutated. This phenomenon further supported the hypothesis that the K-RAS gene does not cause disease and affect tumor cell behaviors unless mutated. Therefore, circ_0069767 affects the biological behavior of MM cells through other pathways rather than the K-RAS pathway.
The mystery of circRNA is gradually being unveiled, but its structural specificity also foretells the diversity of its roles. As the current study suggests, circ_0069767 plays an antitumor role and its expression can be used as a reliable prognostic indicator of MM patients.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article was submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Funding
This research was supported by grants from National Natural Science Foundation of China (Grant No. 81300420) and the Liaoning Province Natural Science Foundation of China (Grant No. 20170540996).
Disclosure
The authors report no conflicts of interest for this work and declared no potential conflicts of interest with respect to the research, authorship, or publication of this article.
References
1. Raab MS, Podar K, Breitkreutz I, et al. Multiple myeloma. Lancet. 2009;374(9686):324–339. doi:10.1016/S0140-6736(09)60221-X
2. Brigle K, Rogers B. Pathobiology and diagnosis of multiple myeloma. Semin Oncol Nurs. 2017;33(3):225–236. doi:10.1016/j.soncn.2017.05.012
3. Li P, Chen S, Chen H, et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015;444:132–136. doi:10.1016/j.cca.2015.02.018
4. Hentze MW, Preiss T. Circular RNAs: splicing’s enigma variations. EMBO J. 2013;32(7):923–925. doi:10.1038/emboj.2013.53
5. Jeck WR, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–157. doi:10.1261/rna.035667.112
6. Suzuki H, Zuo Y, Wang J, et al. Characterization of RNase R-digested cellular RNA source that consists of lariat and circular RNAs from pre-mRNA splicing. Nucleic Acids Res. 2006;34(8):e63. doi:10.1093/nar/gkl151
7. Salzman J, Gawad C, Wang PL, et al. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7(2):e30733. doi:10.1371/journal.pone.0030733
8. Cooper DA, Cortés-López M, Miura P. Genome-wide circRNA profiling from RNA-seq data. Methods Mol Biol. 2018;1724:27–41.
9. Memnzak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7741):333–338.
10. Westhohn JO, Miura P, Olson S, et al. Genomewide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 2014;9(5):1966–1980. doi:10.1016/j.celrep.2014.10.062
11. Zhang Z, Qi S, Tang N, et al. Discovery of replicating circular RNAs by RNA-seq and computational algorithms. PLoS Pathog. 2014;10(12):
12. Ivanov A, Memczak S, Wyler E. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2015;10(2):170–177. doi:10.1016/j.celrep.2014.12.019
13. Zhang Y, Xue W, Li X, et al. The biogenesis of nascent circular RNAs. Cell Rep. 2016;15(3):611–624. doi:10.1016/j.celrep.2016.03.058
14. Zhong ZY, Huang MG, Lv MX, et al. Circular RNA MYLK as a competing endogenous RNA promotes bladder cancer progression through modulating VEGFA/VEGFR2 signaling pathway. Cancer Lett. 2017;403:305–317. doi:10.1016/j.canlet.2017.06.027
15. Liang HF, Zhang XZ, Liu BG, et al. Circular RNA circ-ABCBIO promotes breast cancer proliferation anti progression through sponging miR-1271. Am J Cancer Res. 2017;7(7):1566–1576.
16. Cao S, Wei D, Li X, et al. Novel circular RNA expression profiles reflect progression of patients with hypopharyngeal squamous cell carcinoma. Hepatology. 2017;8(28):45367–45379.
17. Hsiao KY, Lin YC, Gupta SK, et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 2017;77(9):2339–2350. doi:10.1158/0008-5472.CAN-16-1883
18. Ye M, Hou H, Shen M, et al. Circular RNA circFOXM1 plays a role in papillary thyroid carcinoma by sponging miR-1179 and regulating HMGB1 expression. Mol Ther Nucleic Acids. 2019;19:741–750. doi:10.1016/j.omtn.2019.12.014
19. Xiao W, Zheng S, Zou Y, et al. CircAHNAK1 inhibits proliferation and metastasis of triple-negative breast cancer by modulating miR-421 and RASA1. Aging. 2019;11(24):12043–12056. doi:10.18632/aging.102539
20. Verduci L, Strano S, Yarden Y, et al. The circRNA–microRNA code: emerging implications for cancer diagnosis and treatment. Mol Oncol. 2019;13(4):669–680. doi:10.1002/1878-0261.12468
21. Chen F, Hu Y, Wang X, et al. Expression of CD81 and CD117 in plasma cell myeloma and the relationship to prognosis. Cancer Med. 2018;7(12):5920–5927. doi:10.1002/cam4.1840
22. Lasda E, Parker R. Circular RNAs: diversity of form and function. RNA. 2014;20:1829–1842. doi:10.1261/rna.047126.114
23. Jeck WR, Sharpless NE. Detecting and characterizing circular RNAs. Nat Biotechnol. 2014;32(5):453–461. doi:10.1038/nbt.2890
24. Qu D, Yan B, Xin R, et al. A novel circular RNA hsa_circ_0020123 exerts oncogenic properties through suppression of miR-144 in non-small cell lung cancer. Am J Cancer Res. 2018;8(8):1387–1402.
25. Luo Y-H, Zhu X-Z, Huang K-W, et al. Emerging roles of circular RNA hsa_circ_0000064 in the proliferation and metastasis of lung cancer. Biomed Pharmacother. 2017;96:892–898. doi:10.1016/j.biopha.2017.12.015
26. Wu L, Xu XM, Li Y, et al. Circ_0000064 adsorption of microRNA-143 promotes malignant progression of hepatocellular carcinoma. Eur Rev Med Pharmacol Sci. 2019;23(21):9321–9330.
27. Du WW, Yang W, Liu E, et al. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nat Struct Mol Biol. 2016;44:2846–2858.
28. Ashwal-Fluss R, Meyer M, Pamudurti NR, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55–66. doi:10.1016/j.molcel.2014.08.019
29. Patop IL, Wüst S, Kadener S. Past, present, and future of circRNAs. EMBO J. 2019;38(16):e100836. doi:10.15252/embj.2018100836
30. Hayes TK, Der CJ. Mutant and WT Ras: co-conspirators in cancer. Cancer Discov. 2013;3(1):10. doi:10.1158/2159-8290.CD-12-0521
31. Cox AD, Der CJ. Ras history: the saga continues. Small GTPases. 2010;1:2–27.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.