Back to Journals » OncoTargets and Therapy » Volume 14
Targeting c-MET to Enhance the Efficacy of Olaparib in Prostate Cancer
Authors Wang Z, Dai Z , Wang B, Gao Y, Gao X, Wang L, Zhou S, Yang L, Qiu X, Liu Z
Received 13 March 2021
Accepted for publication 26 May 2021
Published 4 August 2021 Volume 2021:14 Pages 4383—4389
DOI https://doi.org/10.2147/OTT.S291267
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr William C. Cho
Zhenwei Wang,1,2 Zhihong Dai,1,* Bingwei Wang,2 Yuren Gao,1 Xiang Gao,1 Liang Wang,1 Sihai Zhou,1 Liqin Yang,1 Xiaofu Qiu,2,3,* Zhiyu Liu1
1Department of Urinary Surgery, Second Hospital of Dalian Medical University, Dalian, Liaoning, People’s Republic of China; 2Department of Urology, Guangdong Second Provincial General Hospital, Guangzhou, Guangdong, People’s Republic of China; 3The Second School of Clinical Medicine, Southern Medical University, Guangzhou, Guangdong, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Zhiyu Liu Tel +86 15541190788
Email [email protected]
Purpose: Prostate cancer is the second leading cause of cancer death in men worldwide. Olaparib is clinically approved for the treatment prostate cancer, but cytotoxicity and off-target effects including DNA damage limit its clinical applications. In the current study, new strategies to improve the therapeutic efficacy of olaparib for the treatment of prostate cancer were investigated.
Methods: Two prostate cancer cell lines were exposed to the c-MET inhibitor PHA665752 and/or the PARP inhibitor olaparib. Cell counting kit-8, colony formation assays, and transwell assays were conducted to evaluate the cytotoxicity of olaparib alone or in combination with PHA665752 in prostate cancer cell lines. Western blotting, immunofluorescence staining, and the comet assay were used to assess the effects of PHA665752 on olaparib-induced DNA damage.
Results: Combined inhibition of c-MET and PARP resulted in effective and synergistic blocking of the growth of prostate cancer cell lines. Invasion and migration were significantly suppressed when the agents were combined. Mechanistically, dual blocking of PARP and c-MET in prostate cancer cell lines was associated with an impaired DNA damage response. Interestingly, immunofluorescence staining analysis of RAD51 protein indicated that the c-MET inhibitor PHA665752 significantly impaired homologous repair via downregulated translocation of RAD51 into the nucleus in prostate cancer cells.
Conclusion: The combination of the c-MET inhibitor PHA665752 and the PARP inhibitor olaparib may be a promising therapeutic strategy in patients with prostate cancer.
Keywords: combination therapy, PARP inhibitor, PHA665752, prostate cancer
Introduction
Prostate cancer (PCa) is the most common cancer diagnosed in men and is the second leading cause of cancer death in men after lung cancer in the United States.1 Improved therapeutic options for patients with advanced PCa in the past decade have dramatically increased survival rates, but castration-resistant PCa (CRPC) remains a lethal disease.2 After the emergence of CRPC, docetaxel chemotherapy is evidently therapeutically efficacious, but in a recent study, the median increase in survival was only 4 months.3 Thus, there is a need for targeted drugs for the treatment of advanced PCa.
The oral PARP-1 and PARP-2 inhibitor olaparib (Lynparza, AstraZeneca Pharmaceuticals, LP) increases DNA damage in cancer cells by inhibiting base excision repair. After being approved by the USA Food and Drug Administration (FDA) in 2014, olaparib has been used in BRCA-mutated advanced ovarian cancer and breast cancers.4 In a Phase II clinical trial, it exhibited durable antitumor activity in sporadic cases of metastatic CRPC patients who had defects in DNA-repair genes.5 The FDA recently approved olaparib for use in patients with deleterious or suspected deleterious germline or somatic homologous recombination repair gene-mutated metastatic CRPC who had previously been administered a second-generation hormonal agent. In addition, the Gleason score is higher, overall prognosis is poor, and overall survival is lower in PCa patients carrying germline mutations in the Homologous repair (HR) DNA repair gene.6 There are several DNA repair genes including BRCA1/BRCA2, ATM, FANC A/F, CHK2, RAD51B/C, and CDK12 that can predict PARP-inhibitors sensitivity in prostate cancer cells.7 Notably, however, a substantial fraction of PCa patients without HR relative gene deficiency do not respond to PARP inhibitors (PARPi). Increasing the response rate of PCa to PARP inhibitors and enhancing their therapeutic potential would expand their application and increase their effectiveness in PCa.
The MET receptor tyrosine kinase system regulates important biological processes, including cell scattering, invasion, and survival as well as epithelial remodeling and angiogenesis.8 Hepatocyte growth factor/scatter factor and its receptor MET may play important roles in CRPC progression. MET has been suggested as a promising target because its expression is detected in 75% to 100% of metastatic prostate tumors9 and has been associated with the emergence of castration-resistant tumor growth. The role of c-MET in HR-mediated repair is not clear. Medova et al10 reported that blocking MET with the small-molecule PHA665752 reduced HR in the DNA damage response pathway of cells via effects on the formation of the RAD51-BRCA2 complex, which is crucial for error-free HR repair of double-stranded DNA lesions. Furthermore, the combination of c-MET and PARP-1 inhibitors has synergistic therapeutic effects in the treatment of triple-negative breast cancer in vitro, as well as in breast cancer and lung cancer xenografted tumor models.11
To date, the combination of c-MET and PARP inhibitors in PCa has not been investigated. In the current study, the effects of c-met inhibitors and olaparib on the growth of PCa cells were investigated, as were the underlying mechanisms involved.
Materials and Methods
Cell Lines and Cell Culture
The human PCa cell lines LNCaP and DU-145 were purchased from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). DU-145 cells were cultured in Hyclone DMEM medium, and LNCaP cells were cultured in Gibco RPMI-1640 medium. All culture media were supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin. All cells were maintained in a humidified incubator at 37°C and 5% CO2.
Proliferation Assays
Cells were seeded into 96-wells plates (3000 cells/well). After a 24-h incubation, the cells were divided into four groups: control, PHA665752, olaparib, and olaparib combined with PHA665752. Cell viability was determined 3 days later via the cell counting kit-8 assay (Dojindo Molecular Technologies) as previously described.12 Combination indexes were calculated using the Calcusyn software program (Biosoft).13,14
Colony Formation Assays
Cells were seeded into 6-well plates (1000 cells/well). After a 24-h incubation, the cells were divided into four groups: control, PHA665752 (1 μM), olaparib (3 μM), and PHA665752 (1 μM) + olaparib (3 μM). Medium was replaced every 3 days. Ten days later, the cells were treated with cold methanol, stained with crystal violet (Sigma–Aldrich), then extracted with 10% glacial acetic acid. Optical density was measured at 570 nm by EnSpire® Multimode Plate Readers (PerkinElmer).
Transwell Assays
For migration assays, a total of 5×104 cells treated with the specified drugs were seeded into transwell inserts (BD Biosciences) and cultured for an additional 24 h. For invasion assays 105 cells treated with the specified drugs were seeded into a thick layer of Matrigel in a transwell insert. Cells that adhered to the lower surface of the filter were washed with phosphate-buffered saline, stained with 0.1% crystal violet solution, then examined via light microscopy at 100× magnification (Carl Zeiss, Inc.).
Comet Assays
Cells were treated with drugs for 12 h, then detached mechanically and added to low-melting-point agarose. After lysis, the cells were subjected to electrophoresis and the resulting “comets” were stained with ethidium bromide and viewed using a fluorescence microscope at 100× magnification (Carl Zeiss, Inc.). For each condition, a minimum of 50 cells were analyzed using Pect Comet Assay software (CaspLab).
3D Sphere Culture
A total of 3×103 cells were suspended in serum-free medium mixed with 2% fetal bovine serum and 2% Matrigel. The media with drugs were replaced every 3 days. Two weeks later, floating cells were counted manually under a microscope.
Immunofluorescence
Cells treated with the specified drugs were cultured on coverslips. After 48 hours, they were fixed with 4% paraformaldehyde and blocked with a 5% bovine serum albumin-phosphate-buffered solution. They were then stained with rabbit anti-RAD51 (Santa Cruz Biotechnology). After washing with phosphate-buffered saline, they were treated with secondary antibodies. Slides were mounted in Vectashield with DAPI. Analyses were performed using an immunofluorescence microscope (Leica).
Statistical Analysis
Numerical results are presented as means ± S.D. The t-test was used to compare means in different groups via GraphPad Prism 6.0 software (San Diego, CA, USA). In all statistical analyses p < 0.05 was considered statistically significant.
Results
PHA665752 and Olaparib Synergistically Inhibited the Growth of PCa Cells
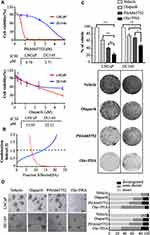
In cell proliferation assays, the respective IC50s of PHA665752 in LnCAP cells and DU145 cells were 0.70 μM and 3.71 μM, and the respective IC50s of olaparib in LnCAP cells and DU145 cells were 13.55 μM and 22.21 μM (Figure 1A). To assess the combined effects of the drugs on cell viability, PCa cells were treated with increasing concentrations of olaparib and PHA665752. In the majority of concentrations tested, the combination treatment resulted in a synergistic increase in viability inhibition at combination index values and 0.5 fractions affected (Figure 1B). In colony formation assays, the combination treatment strongly decreased the long-term growth of LNCaP and DU145 cells (Figure 1C). In 3D sphere culture experiments investigating the combined effects of the drugs on PCa cells,15 inhibition of c-MET and PARP induced dramatic LNCaP and DU145 cell death (Figure 1D).
Combined Inhibition of c-MET and PARP Synergistically Inhibited PCa Cell Migration and Invasion
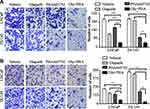
In transwell assays, the combined use of PHA665752 and olaparib reduced the migratory and invasive capacities of LNCaP and DU145 cells compared with the corresponding parameters in vector control or single-drug cells (Figure 2). These results suggested that combined treatment of c-MET and PARP may have a therapeutic value for PCa.
Combined Inhibition of c-MET and PARP Synergistically Induced DNA Damage and Apoptosis in PCa Cells
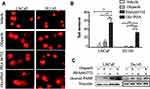
In comet assays, treatment with a combination of agents resulted in higher tail intensities in two PCa cell lines than treatment with single agents (Figure 3A and B). Treatment with a combination of agents also induced an abundance of cleaved PARP in LNCaP cells and DU145 cells (Figure 3C). These data indicate that combined treatment may synergistically influence PCa cell lines by altering DNA damage and apoptosis.
Combined Inhibition of c-MET and PARP Synergistically Resulted in DNA Damage via Inhibition of Nuclear Translocation of RAD51 in PCa Cell Lines
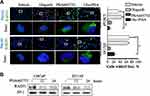
Adding c-MET inhibitors to PCa cell line cultures markedly decreased nuclear levels of RAD51 (Figure 4A). In Western blotting analyses nuclear localization of RAD51 was significantly impaired in PCa cells after treatment with PHA665752 (Figure 4B). Previous studies indicate that nuclear transport of RAD51 is an integral part of the cellular response to DNA damage.16 The above evidence suggests that combined treatment may synergistically influence DNA damage by altering the nuclear localization of RAD51.
Discussion
Olaparib, a first-in-class PARP inhibitor, kills cancer cells by exploiting tumor DNA repair pathway deficiencies. In May 2020, The FDA approved olaparib for use in adults with metastatic CRPC carrying a germline or somatic HR gene mutation (specifically, BRCA1/2, ATM, BARD1, BRIP1, CDK12, CHEK1, CHEK2, FANCL, PALB2, RAD51B, RAD51C, RAD51D, or RAD54L), who have progressed following prior treatment with enzalutamide or abiraterone. Notably, however, HR gene mutations are not common in PCa, and HR wild-type patients do not respond well to olaparib. It is therefore important to identify an effective approach to rendering non-HR gene-mutant patients adequately responsive to PARPi and/or enhancing the efficacy of olaparib in HR gene-mutated patients.
The c-MET receptor tyrosine kinase was originally discovered as an oncoprotein, and has been implicated in the proliferation and progression of PCa and a wide variety of other human malignancies.9,17 High c-MET expression is observed in the late stages and metastases of PCa.18,19 In previous studies, c-MET inhibition increased sensitivity to olaparib in triple-negative breast cancer cells with acquired resistance to PARPi,20 and in breast cancer, lung cancer,11 and ovarian cancer.21 However, it is unclear whether the combination of c-MET and PARP-1 inhibitors results in synergistic suppression of the growth of PCa. In the current study, c-MET inhibition by PHA665752 significantly increased sensitivity to olaparib in both the CRPC cell line DU145 and the hormone-sensitive PCa cell line LNCaP. Combined inhibition of c-MET and PARP effectively blocked PCa cell proliferation, migration, invasion, and survival in vitro.
RAD51, a marker associated with the competency of HR repair, is needed for meiotic HR in eukaryotes and mitotic HR events such as double-strand breaks repair.22 Biochemical studies suggest that only the RAD51-ssDNA nucleoprotein filament can catalyze DNA joint formation,23 which supports the premise that HR in cells is triggered by recruitment of RAD51 to ssDNA produced by nucleolytic processing of DSBs, or ssDNA that is associated with stalled or damaged DNA replication forks. Hence, inhibition of RAD51 function may reduce HR and constitute an effective approach to cancer intervention.24 RAD51 mainly spreads in cis around DSBs. Interestingly, in the current study, the c-MET inhibitor PHA665752 nearly abolished RAD51 translocated from the cytosol to the nucleus in two PCa cell lines. Although the primary mechanism involved is not clear, it can evidently weaken the function of RAD51.
In conclusion, to the best of our knowledge, the present study is the first to indicate that the combined inhibition of c-MET and PARP can synergistically inhibit proliferation, migration, invasion, and survival of PCa cells in vitro. Mechanistically, c-MET—which contributes to reduced translocation of RAD51 into the nucleus—significantly increased sensitivity to PARP via an exacerbated DNA damage response.
Funding
This study was supported by the Science and Technology Planning Project of Guangzhou City, Guangdong, China, under grant 201904010035, the Natural Science Foundation of Guangdong Province, China, under grant 2018A030313905, and the Medical Science Foundation of Dalian, China, under grant 2012020.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. doi:10.3322/caac.21590
2. Rini BI, Small EJ. Hormone-refractory prostate cancer. Curr Treat Options Oncol. 2002;3(5):437–446. doi:10.1007/s11864-002-0008-1
3. Petrylak DP, Tangen CM, Hussain MHA, et al. Docetaxel and estramustine compared with mitoxantrone and prednisone for advanced refractory prostate cancer. N Engl J Med. 2004;351(15):1513–1520. doi:10.1056/NEJMoa041318
4. Kaufman B, Shapira-Frommer R, Schmutzler RK, et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J Clin Oncol. 2015;33(3):244–250. doi:10.1200/JCO.2014.56.2728
5. Mateo J, Carreira S, Sandhu S, et al. DNA-repair defects and olaparib in metastatic prostate cancer. N Engl J Med. 2015;373(18):6981–6996. doi:10.1056/NEJMoa1506859
6. Marshall CH, Fu W, Wang H, et al. Prevalence of DNA repair gene mutations in localized prostate cancer according to clinical and pathologic features: association of Gleason score and tumor stage. Prostate Cancer Prostatic Dis. 2019;22(1):59–65.
7. Criscuolo D, Morra F, Giannella R, et al. Identification of novel biomarkers of homologous recombination defect in DNA repair to predict sensitivity of prostate cancer cells to PARP-inhibitors. Int J Mol Sci. 2019;20(12):12. doi:10.3390/ijms20123100
8. Knudsen BS, Vande Woude G. Showering c-MET-dependent cancers with drugs. Curr Opin Genet Dev. 2008;18(1):87–96. doi:10.1016/j.gde.2008.02.001
9. Knudsen BS, Edlund M. Prostate cancer and the met hepatocyte growth factor receptor. Adv Cancer Res. 2004;91:31–67.
10. Medova M, Aebersold DM, Zimmer Y. MET inhibition in tumor cells by PHA665752 impairs homologous recombination repair of DNA double strand breaks. Int J Cancer. 2012;130(3):728–734. doi:10.1002/ijc.26058
11. Du Y, Yamaguchi H, Wei Y, et al. Blocking c-Met–mediated PARP1 phosphorylation enhances anti-tumor effects of PARP inhibitors. Nat Med. 2016;22(2):194–201. doi:10.1038/nm.4032
12. Junttila TT, Akita RW, Parsons K, et al. Ligand-independent HER2/HER3/PI3K complex is disrupted by trastuzumab and is effectively inhibited by the PI3K inhibitor GDC-0941. Cancer Cell. 2009;15(5):429–440. doi:10.1016/j.ccr.2009.03.020
13. Chou T-C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 2010;70(2):440–446. doi:10.1158/0008-5472.CAN-09-1947
14. Speit G, Rothfuss A. The comet assay: a sensitive genotoxicity test for the detection of DNA damage and repair. Methods Mol Biol. 2012;920:79–90.
15. Muranen T, Selfors LM, Worster DT, et al. Inhibition of PI3K/mTOR Leads to Adaptive Resistance in Matrix- Attached Cancer Cells. Cancer Cell. 2012;21:227–239
16. Gildemeister OS, Sage JM, Knight KL. Cellular redistribution of Rad51 in response to DNA damage: novel role for Rad51C. J Biol Chem. 2009;284(46):31945–31952. doi:10.1074/jbc.M109.024646
17. Bottaro DP, Rubin J, Faletto D, et al. Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science. 1991;251(4995):802–804. doi:10.1126/science.1846706
18. Humphrey PA, Zhu X, Zarnegar R, et al. Hepatocyte growth factor and its receptor (c-MET) in prostatic carcinoma. Am J Pathol. 1995;147(2):386–396.
19. Knudsen BS, Gmyrek GA, Inra J, et al. High expression of the Met receptor in prostate cancer metastasis to bone. Urology. 2002;60(6):1113–1117. doi:10.1016/S0090-4295(02)01954-4
20. Chu YY, Yam C, Chen MK, et al. Blocking c-Met and EGFR reverses acquired resistance of PARP inhibitors in triple-negative breast cancer. Am J Cancer Res. 2020;10(2):648–661.
21. Sahin ID, Jonsson J-M, Hedenfalk I. Crizotinib and PARP inhibitors act synergistically by triggering apoptosis in high-grade serous ovarian cancer. Oncotarget. 2019;10(65):6981–6996. doi:10.18632/oncotarget.27363
22. Bonilla B, Hengel SR, Grundy MK, et al. RAD51 gene family structure and function. Annu Rev Genet. 2020;54(1):25–46. doi:10.1146/annurev-genet-021920-092410
23. Sung P, Robberson DL. DNA strand exchange mediated by a RAD51-ssDNA nucleoprotein filament with polarity opposite to that of RecA. Cell. 1995;82(3):453–461. doi:10.1016/0092-8674(95)90434-4
24. Sak A, Stueben G, Groneberg M, et al. Targeting of Rad51-dependent homologous recombination: implications for the radiation sensitivity of human lung cancer cell lines. Br J Cancer. 2005;92(6):1089–1097. doi:10.1038/sj.bjc.6602457
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.