Back to Journals » International Journal of Nanomedicine » Volume 15
Synthesis, Characterization and DNA Binding Investigations of a New Binuclear Ag(I) Complex and Evaluation of Its Anticancer Property
Authors Zheng MH, Bigdeli F, Gao LX, Wu DZ, Yan XW, Hu ML, Morsali A
Received 27 July 2019
Accepted for publication 27 January 2020
Published 12 February 2020 Volume 2020:15 Pages 953—964
DOI https://doi.org/10.2147/IJN.S225038
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Prof. Dr. Anderson Oliveira Lobo
Mian-Hong Zheng,1,* Fahime Bigdeli,2,* Lan-Xing Gao,1,* Deng-Ze Wu,1 Xiao-Wei Yan,3 Mao-Lin Hu,3 Ali Morsali2
1College of Chemistry and Materials Engineering, Wenzhou University, Wenzhou 325035, People’s Republic of China; 2Department of Chemistry, Faculty of Sciences, Tarbiat Modares University, Tehran, Iran; 3College of Materials and Chemical Engineering, and Guangxi Key Laboratory of Calcium Carbonate Resources Comprehensive Utilization, Hezhou University, Hezhou, Guangxi 542800, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Deng-Ze Wu; Ali Morsali Email [email protected]; [email protected]
Aim: A new Ag(I) complex (A3) was synthesized and evaluated for its anticancer activity against human cancer cell lines.
Materials and Methods: The complex A3 was characterized by 1H, 13C, and 31P nuclear magnetic resonance (NMR), infrared (IR) spectra, elemental analysis, and X-ray crystallography. The interaction of the complex with CT-DNA was studied by electronic absorption spectra, fluorescence spectroscopy, and cyclic voltammetry; cell viability (%) was assessed by absorbance measurement of the samples.
Results: The interaction mode of the complex A3 with DNA is electrostatic, and this complex shows good potential in anticancer properties against HCT 116 (human colorectal cancer cells) and MDA-MB-231 (MD Anderson-metastatic breast) cell lines with 0.5 micromolar concentrations.
Conclusion: The Ag(I) complex could interact with DNA noncovalently and has anticancer properties.
Keywords: binuclear Ag(I) complex, X-ray crystallography, DNA, electrostatic binding, anticancer
Introduction
Cancer is a challenging disease and global deaths owing to it are increasing.1 Important methods for the treatment of cancer patients include surgery, chemical medication, radiotherapy, biological immunization, and chemotherapy. In chemotherapy, 5-fluorouracil (5-Fu) is the main anticancer drug, possessing an inhibitory capability against many cancers, particularly for gastrointestinal tumors.2,3 However, critical side effects have been observed associated with the clinical use of 5-Fu.4 In addition, 5-Fu possesses a short half-life and is swiftly removed after operation. Many studies have been carried out to combat these problems and increase the antitumor activity of 5-FU; some procedures include modulations of 5-FU in combination with other compounds and advanced delivery systems.2,5,6 Choi et al indicated that chloroquine can augment the antitumor effect of 5-FU by cell cycle blockage in human colon cancer cells.7 More research is needed to find new types of anticancer drugs. Metal complexes display good potential in cancer therapy due to their unique features, for example, various coordination routes, reactivity, and redox ability against organic materials. Therefore, based on these characteristics, it is possible to design metal complexes with the ability to interact with target biomolecules, and, as a result, to modify the cellular cycle.8 In some cases, metal complexes show better antitumor properties than the ligands. The complexes may act via their impact on electron transport, inhibition of DNA replication by the metal ion center, perturbation in the proteins and enzyme actions as well as redox balance in cells, reactive oxygen species (ROS) production and alteration of the cell membrane.9,10
Cisplatin is a compound that is currently used in cancer treatment. Despite some favorable results, the function of cisplatin is restricted to only some types of tumor, and its usage leads to toxicity and resistance in clinical application. Hence, it is essential to develop new types of anticancer compounds. Many researchers have focused on coinage metals (Au, Cu, and Ag) because the complexes of these metals exhibit a wide range of anticancer activity with less toxicity to non-cancerous cells.11 Among coinage metals, silver is of special importance since it is compatible with biological systems, and silver complexes display low toxicity to human cells, are selective against cancer cells, and act as antimicrobial agents.8,12 A few active anticancer complexes of Ag(I) such as carboxylate, phosphane, and bipyridine complexes have been reported.12–14 The presence of the ligands in the silver complexes leads to the improvement of the silver function.15 Moreover, the high number of Ag(I) center ions in a complex can improve the biological activity of silver complexes,16 and the cytotoxic effect of binuclear silver(I) complexes is significantly more in cancer cells in comparison with the normal cells.14 Haque et al showed that silver ions have a fundamental importance in the destruction of tumor cells.3,12
Anticancer agents mostly act through the DNA molecule.15 Molecules may interact with DNA by changing or preventing its operation, disordering gene sequencesand introde in transcription.17
The main binding types in DNA complexes are covalent and non-covalent. The mechanism of action of many anticancer drugs is by covalently binding to DNA. Covalent binding is observed at several binding sites including the nitrogen centers of the bases, and the phosphates. The antitumor activity of the covalent binding leads to blockage of DNA, irreversibly and completely. The non-covalent interactions with DNA consist of electrostatic bindings with the negatively charged phosphate, intercalation between adjoining base pairs, and groove interaction.18 Non-covalent interaction has advantages including reversibility and less cytotoxicity. This type of interaction can unwind and break the DNA double helix.19 Metal ions may play the role of a counter ion as a result of interaction with DNA noncovalently.20 Also, the groove binding of some octahedral complexes with DNA with base pairs has been reported.18,21–23
Silver compounds can interact with DNA via metalization of DNA. This is important in the development of biotechnological areas and also the ligands of the complexes can intercalate between bases of DNA. DNA may be metalized by the coordination of silver to bases of DNA15 or with the aid of electrostatic binding.24,25
This report describes the synthesis and characterization of a new binuclear Ag complex, [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3), where PPh3=triphenylphosphine, L1=5-fluorouracil-1-yl acetic acid, and its interaction with calf thymus DNA (CT-DNA). Also, we investigated the anticancer activity of the complex A3 toward human cancer cell lines.
Materials and Methods
Apparatus and Reagents
The ligand was prepared according to the method mentioned in this report, and all other chemicals and solvents were purchased from Aldrich Chemical Company or other commercial vendors. 1H, 13C, and 31P nuclear magnetic resonance (NMR) spectra were recorded on an AVANCE III AV 500 spectrometer using tetramethylsilane (TMS) as an internal standard and DMSO-d6 as the solvent at room temperature. IR spectra were recorded on a Nicolet IS10 spectrophotometer. Elemental analysis was carried on a EURO EA 3000 Elemental Analyzer. UV–vis absorption spectra were provided by a UV-6100 double beam spectrophotometer. Fluorescence measurements were made on an F-2700 FL spectrophotometer at room temperature. The electrochemical experiments were performed with a CHI 1030b electrochemical workstation (Shanghai Chenhua Co.). The absorbance (A) of wells in biological investigations was measured using a multiplate reader on a Multiskan Mk3, Thermo Scientific, Waltham, USA, at 450 nm. All cell lines were purchased commercially from the Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China).
X-Ray Crystallography
The specifications of the single crystal of the A3 class, orientation matrix, and cell dimensions were determined by the founded methods and Lorentz polarization and absorption corrections were performed. Empiric absorption corrections were provided by the SADABS program. Most of the non-hydrogen atoms were determined according to direct procedures, and the rest were located by subsequent Fourier synthesis. All non-hydrogen atoms were refined anisotropically, and all hydrogen atoms were maintained changeless and took in the final stage of full-matrix least squares refinement based on F2 by the SHELXS-97, SHELXL-97 and SHELXTL programs.
Syntheses
Synthesis of 5 FUAA Ligand 5-Fluorouracil-1-yl Acetic Acid (L1)
First, 31.2 g (240 mmol) 5-fluorouracil was added to 160 mL of aqueous solution of 51.2 g (92 mmol) potassium hydroxide and heated up to 40°C; then, 50 g (360 mmol) 80 mL bromoacetic acid solution was slowly added in the mixture for 120 min. Under this temperature mixing reaction was stirred overnight, cooled to room temperature, adjusted pH to 5.5 by concentrated hydrochloric acid and put in the refrigerator frozen for 2 h. After removing precipitate, sediment was filtered off and a solution of strong hydrochloric acid was added to it to bring the pH value to 2 then it was put in the refrigerator frozen for 6 h. After filtering sediment and washing in cold water three times, the compound was dried (Figure 1).26 Yield: 89%. M. p: 241ºC, Anal. Calc. for C6H5N2FO4 (%): C, 38.45; H, 2.48; N, 13.85. Found (%): C, 38.31; H, 2.68; N, 14.89. 1H NMR (500 MHz, DMSO): δ/ppm, 4.36 (2H, s), 8.07 (1H, d, J = 8.5 Hz), 11.91 (1H, d, J = 8.5 Hz), 13.22 (1H, s), 13C NMR (126 MHz, DMSO) δ130.70, 139.52, 149.85, 157.65, 169.45, IR: 3272.34; 3414.80; 1699.31; 1377.31; 1242.58 cm−1.
 |
Figure 1 Synthesis of ligand 5-fluorouracil-1-yl acetic acid (L1). |
Synthesis of Complex [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3)
AgBF4 (0.039 g, 0.3 mmol) was dissolved in (5 mL) CH3CN solution of PPh3 (0.052 g, 0.3 mmol) under ultrasonication; then, 42 μL Et3N (0.3 mmol) was added, and a white cloudy solution was obtained. Then, 5-FUAA (0.038 g, 0.2 mmol) was added to the resulting solution, the suspension was sealed and heated to 70°C for 20 h. After cooling to room temperature, the solution was filtered off. After slow evaporation of the colorless and transparent solution, colorless crystals were obtained.
Yield: 85.23%. M.p:171.1ºC-173.7ºC. 1H NMR, 13C NMR, and 31P NMR are given in Supporting Information, Figure 1S–3S.
DNA Binding and Anti-Cancer Activity Experiments
The binding of A3 with CT-DNA was investigated by electronic absorption spectra, fluorescence spectroscopy, and cyclic voltammetry studies. Biological activity tests of complex A3 as a drug, with a molecular size of 14.46×20.05Å2 as measured using Diamond Software, have been performed in the solution and on a molecular scale. The size of complex A3 is suitable for delivery and penetration as a drug in the body.27 To determine the anticancer activity of the complex, the cell viability (%) was investigated by absorbance measurement of the samples.
Preparation of DNA-Modified Electrode
For preparation of the electrode-modified DNA, gold disk electrodes were glossed by the alumina powder (1.0, 0.5, and 0.05 μm). Then, the Au electrode was refined and left in fresh piranha solution (30% H2O2 and 70% H2SO4) to remove adsorbed impurities, then it was placed in an ultrasound bath for 5 min.
In order to prepare a neat Au electrode, the surface of the electrode was activated electrochemically through sweeping from −0.3 to +1.5 V in 0.1 M H2SO4 until a stable cyclic voltammogram was created. After washing with water, the Au electrode was modified by pouring a droplet of 10 μL of 1.0 μg μL−1 CT-DNA solution onto its surface, then drying in the air overnight. Then, the DNA-modified electrode (named CT-DNA/Au all over) was immersed in water for about 4 h to eliminate any unadsorbed CT-DNA.
The Preparation of the Samples for Cytotoxic Measurements
The cell viability of A3 was measured in cell lines of HCT 116 (human colorectal cancer cells) and MDA-MB-231 (MD Anderson-metastatic breast) using the cell counting kit-8 (CCK-8) assay. Cancer cells were seeded in 96-well plate with a cell density of 10,000 cells/well and incubated in a humidified 37°C incubator with an atmosphere of 5% CO2/95% air. After 24 h, A3 with or without NIR irradiation (808 nm laser, 0.2 W/cm2 for 1 h) was diluted to 0.5, 1, 2, 4, and 6 μM, respectively, and added into the plate. For comparison, free 5-Fu with the same concentrations was also added into the plate. 24 h post-treatment, the medium in each well was refreshed with the culture medium (200 μL with 10 μL CCK-8 reagent). After 2 h, the absorbance (A) of each well was measured at 450 nm.
Results
Structural Studies
The complex [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3) was formed by reaction of AgBF4 with CH3CN solution of PPh3, Et3N and 5-FUAA. Slow evaporation of the resulting solution led to the formation of colorless and transparent crystals. After preparation of A3 it was characterized by Fourier-transform infrared spectroscopy (FT-IR) spectroscopy, 1H NMR, 13C NMR, 31P NMR, and X-ray crystallography. The structural data and description of the diffraction experiment are listed in Table 1 and a schematic representation of the complex is given in Figure 2. The structural details are discussed in the Discussion section.
 |
Table 1 Data Collection and Refinement Parameters for A3 |
Anal.Calc.for C102H83O4N4F2P5Ag2 (%): C, 64.38; H, 4.37; N, 2.95. Found (%): C, 64.45; H, 4.32; N, 2.95.
1H NMR (500 MHz, DMSO) δ 7.46 (t, J = 7.1 Hz, 15H), 7.40–7.29 (m, 60H), 4.10 (s, 4H).
13C NMR (126 MHz, DMSO) δ 133.45 (d, J = 16.7 Hz), 132.05 (s), 131.85 (s), 130.45 (s), 128.98 (d, J = 9.5 Hz). 31P NMR (202 MHz, DMSO) δ 7.89 (s). IR: 1763, 1377.95, 1057, 444.05, 428.14, 408.85 cm−1.
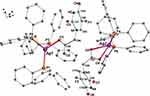 |
Figure 2 Molecular diagram of [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3). |
Electronic Absorption Spectra Studies
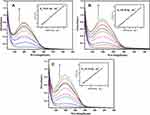
UV-vis absorption spectroscopy is one of the prevalent methods for studying the interactions between DNA and complexes. Absorption titration experiments were carried out at a constant concentration of complex A3 (10 μM) with changing concentrations of (0–0.045 mg mL−1) from CT-DNA in buffer containing KH2PO4/NaOH (at pH = 6.2, 7.2, and 8.2). Absorption spectra were measured at each increase in the CT-DNA solution (Figure 3).
Fluorescence Studies
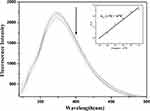
A simple method to check the interaction characteristics between the complex and DNA is fluorescence spectroscopy. Ethidium bromide, 3,8-diamino-5-ethyl-6-phenyl phenanthridium bromide (EtBr), is widely used to study small molecule–DNA interactions because of its high fluorescence when binding to DNA.28 Emission spectra were carried out with EtBr (0.05 mM) connected to CT-DNA (0.12 mg mL−1) in the absence and presence of A3 in buffer KH2PO4/NaOH (pH = 7.2) with a range of concentrations of 0–0.1 mM (Figure 4).
Electrochemical Characterization
Cyclic Voltammetry
Voltammetric measurements were performed in a prevalent three-electrode cell contains a bare gold or CT-DNA/Au as the working electrode, a saturated calomel electrode (SCE), and a platinum wire auxiliary electrode. The CV experiments were performed using a 5 mM solution of K3[Fe(CN)6]/K4[Fe(CN)6] (1:1) in 0.05 M KH2PO4/NaOH buffer at pH 7.2 with 0.1 M KCl at room temperature (25ºC). All solutions were deaerated with pure nitrogen, and the electrochemical experiments were carried out at a scan rate of 0.1 V s−1.
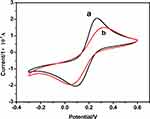
A typical cyclic voltammogram of the bare Au electrode (curve a) and CT-DNA/Au (curve b) in 5.0 mM Fe(CN)6 3-/4-solution at a scanning rate of 0.1 V s −1 is observed in Figure 5. As seen from Figure 5, two redox peaks were obtained in both cases. In addition, the peak current at CT-DNA/Au was lower than that in the bare Au electrode. This is because DNA acts as an electron and mass transfer blocker layer, which prevents the motion of Fe(CN)63-/4-toward the electrode surface. The results also demonstrate the good modification of the DNA on the surface of Au electrode. The CV of the DNA-modified Au electrode remained stable after 20 scans in the KH2PO4/NaOH buffer solution, representing electrochemical stability of the DNA films.
The electrochemical behaviors of Fe(CN)63-/4- at CT-DNA/Au were further investigated by a change in scan rate in KH2PO4/NaOH buffer solution (pH 7.14) containing 5 mM Fe(CN)63-/4-. From Figure 6, it can be seen that the anodic peak currents increased with increasing the scan rate, and there is a good linear relation between peak current and scan rate in the range of 0.01–0.13 V s −1. These results show that the electrochemical kinetic process is a typical surface adsorption-regulated electrochemical process.
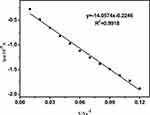 |
Figure 6 The relationship between anodic peak current and the scanning rate (0.01–-0.13 V s −1) for CT-DNA/Au. |
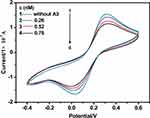
Since complex A3 is a non-electroactive organic small molecule, Fe(CN)63-/4- was employed as a redox probe to study the interactions of non-electroactive of the complex with CT-DNA. Cyclic voltammogram changes of Fe(CN)63-/4- at different concentrations of A3 at CT-DNA/Au are observed in Figure 7.
As observed in Figure 7, both the reduction and oxidation peak currents gradually decrease with seven increasing concentrations from 0.26 mM to 2.50 mM of A3 (some concentrations have been omitted in the Figures). This process can be ascribed to the fact that DNA films cause the redox activation of Fe(CN)63/4- to be harder at the Au electrode because of the physical blockage and possible electrostatic repulsion. After adding the prodrugs to the solution, they interact with DNA, resulting in the formation of a denser DNA film; therefore, the migration of Fe(CN)6 3-/4- ions via the film becomes harder and the redox peak current of Fe(CN)6 3-/4- is decreased.
As shown in Figure 8, both the peak currents of the cyclic voltammograms decreased with increasing the concentrations of drugs and reached a saturation value according to Langmuir adsorption behavior.
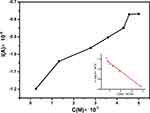 |
Figure 8 Adsorption isotherm of drug A3 on CT-DNA/Au. The solid line corresponds to the Langmuir model. Inset: The relationship between 1/[DRUG] and 1/ΔIP. |
Biological Activity
In order to perform a quantitative comparison of the binding strength of A3 with CT-DNA, the binding constant (K) was calculated according to the Langmuir formula in Equation (A.1).29 Based on the procedure of Qu et al30 it is considered that DNA and DRUG only create a single complex DNA.DRUGm.
(A)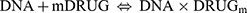
Using Equations 1S-8S in Supporting Information, Equation (A.1) is obtained.
(A.1)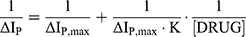
where ΔIP =IP0-IP, IP and IP0 show the oxidation peak current of Fe(CN)63-/4- in the presence and absence of the drugs, respectively; ΔIP, max is the maximum difference of the oxidation peak current; and [DRUG] shows the concentration of the drug. Therefore, the binding constant (K) is 3.13 ×10 3 L mol −1.
Anti-Cancer Activity
Anticancer potential of A3 on both HCT 116 (human colorectal cancer cells) and MDA-MB-231 cancer cell lines was evaluated by cell counting kit-8 (CCK-8) assay. The results were reported as percentage inhibition of cell proliferation (±SD). The relative cell viability was calculated by the following equation:
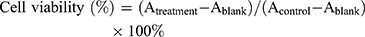
Ablank indicates the absorbance of the well without adding CCK-8 reagent and Acontrol indicates the absorbance of the well with the addition of saline buffer instead of any drugs. The compounds A3+NIR and A3 in counteranions (0.5–6µM) were found to be well active (Figures 9 and 10). At the various concentrations, significant differences in the activity of compounds 5-A3+NIR and A3 were observed. Increasing the concentrations led to an enhancement of compounds' activities.
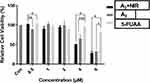 |
Figure 9 Cell viability of A3+NIR, A3 and 5-Fu in HCT 116 cell line. Significant differences are indicated as ***P<0.001, **P<0.01, *P<0.05, n=3. |
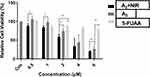 |
Figure 10 Cell viability of A3+NIR, A3 and 5-Fu in MDA-`MB-231 cell line. Significant differences are illustrated as **P<0.01, *P<0.05, n=3. |
Discussion
Structural Characterization
Characteristic absorption bands of the ligands PPh3 and L1 are observed in the FT-IR spectra for complex [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3) with the some extent shift except of absorption band of OH (3414 cm−1) that is related to the carboxylic functional group due to coordination bonding of it as an O-donor moiety in A3. Single X-ray crystal analysis shows that there are two Ag(I) ions with different coordination environments consisting of AgOP3 and AgO2P2. Ag1 is coordinated by one O atom of carboxylate: Ag1-O1: 2.3787(64) Å and three P atoms of PPh3 molecules: Ag1-P1: 2.5316(22) Å, Ag1-P2: 2.5817(15) Å, Ag1-P3: 2.5855(15) Å. The coordination sphere of Ag2 includes two O atom belonging to a carboxylate group: Ag2-O5: 2.4990(55) Å, Ag2-O6: 2.4507(69) Å and two P atoms of PPh3 molecules, Ag1-P4: 2.4209(21) Å, Ag1-P5: 2.4550(17) Å; therefore, four coordinated and binuclear complexes are formed in which the Ag−Ag distance of the two silver atoms Ag1 and Ag2 is 8.5318(13) Å. The structure of A3 crystallizes in the triclinic space group Pī with lattice constants a = 13.806(3) Å, b =14.652(3), c= 27.796(5), Z = 2 (Figure 2).
The Interaction of the Complex with the DNA Molecule
The binding mode between DNA and complexes can be determined by absorption spectroscopy based on the shifts and absorption strength. According to Figure 3 the absorption spectra of the complex incubated with increasing concentrations of CT-DNA. The absorption bands of A3 undergo hyperchromism in molar absorptivity, which is ascribed to the electrostatic interaction between the metal ions of the complex and phosphate group of DNA.31 The higher Kb value for b in comparison to a (or c) indicates that neutral environments provide more favorable conditions for this interaction.
In the other investigation, the study of complex binding to DNA was carried out using the fluorescence feature of EtBr. As given in Figure 4, with an increase in the concentration of the complex, the intensity of the emission band in the EtBr-DNA system decreased as indicated by the replacement of the EtBr with the complex at DNA binding sites. The fluorescence quenching efficiency is evaluated by the Stern–Volmer constant (KSV) based on the classical Stern–Volmer equation: F0/F = 1 + KSV[Q], in which F0 and F are related to the fluorescence intensities of EtBr-DNA in the absence and presence of the complex, respectively, and [Q] is the concentration of the complex. KSV is a linear Stern–Volmer quenching constant achieved from the linear regression of F0/F with [Q].
Voltammetric measurements display good electrochemical stability of the DNA on the Au electrode and the correlation between the peak currents of the cyclic voltammograms and concentrations of drugs shows a Langmuir adsorption behavior.
Anti-Cancer Activity
According to the literature, silver complexes exhibit anticancer activity by various mechanisms.3,16 Herein silver(I) ions interfere with the function of cancer cells by interacting with the phosphate group of DNA molecules.
The cell viability of A3 with or without NIR irradiation was evaluated in both HCT 116 (human colorectal cancer cells) and MDA-MB-231 (MD Anderson-metastatic breast) cell lines by cell counting kit-8 (CCK-8) assay. According to Table 2 and the IC50s (half-maximal inhibitory concentrations), the comparison of the anticancer activities of compounds is as follows: A3+NIR˃A3˃5-Fu. The results demonstrate a significant influence of the silver complex on the anticancer activity.
 |
Table 2 IC50s of A3+NIR, A3 and 5-Fu (µm). IC50s Were Calculated Based on CCK-8 Results by IBM SPSS Statistics Software |
Conclusion
In summary, a new Ag(I) complex, [(PPh3)3Ag(L1)].[(PPh3)2Ag(L1)] (A3), L1=5-fluorouracil-1-yl acetic acid, was prepared and the crystal structure of it was studied by single crystal X-ray analysis. The binding of the complex to CT-DNA was studied by electronic absorption spectra, fluorescence emission, and cyclic voltammetry, and the results suggested an electrostatic interaction with DNA. The prepared binuclear complex, because it includes ion metal centers (Ag+) and ligand L1 (5-fluorouracil-1-yl acetic acid) with a molecular size of 14.46×20.05 Å2, can be regarded as a new agent for evaluation of anticancer properties. The results of the biological study indicated that complex A3 possesses a good potential in antiproliferative activity against human cancer cell lines, with the 0.5 micromolar concentrations.
Acknowledgments
Support of this investigation by the National Natural Science Foundation of China (nos. 21371137 and 21571143) and Tarbiat Modares University is gratefully acknowledged. Also, the authors thank Huacheng He (Hezhou University) for his valuable guidance on testing activities and providing high-quality figures.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Dewangan J, Srivastava S, Mishra S, Divakar A, Kumar S, Rath SK. Salinomycin inhibits breast cancer progression via targeting HIF-1α/VEGF mediated tumor angiogenesis in vitro and in vivo. Biochem Pharmacol. 2019;164:326–335. doi:10.1016/j.bcp.2019.04.026
2. Li M, Liang Z, Sun X, Gong T, Zhang Z. A polymeric prodrug of 5-fluorouracil-1-acetic acid using a multi-hydroxyl polyethylene glycol derivative as the drug carrier. PLoS One. 2014;9:e112888.
3. Iqbal MA, Haque RA, Nasri SF, et al. Potential of silver against human colon cancer:(synthesis, characterization and crystal structures of xylyl (Ortho, meta, & Para) linked bis-benzimidazolium salts and Ag(I)-NHC complexes: in vitroanticancer studies). Chem Cent J. 2013;7:27. doi:10.1186/1752-153X-7-27
4. Bounous G, Pageau R, Regoli D. Enhanced 5-FU mortality in rats eating definded formula diets. Int J Clin Pharmacol Biopharm. 1978;16:265–267.
5. Shen L, Hu J, Wang H, Wang A, Lai Y, Kang Y. Synthesis and biological evaluation of novel uracil and 5-fluorouracil-1-yl acetic acid-colchicine conjugate. Chem Res Chin Univ. 2015;31:367–371. doi:10.1007/s40242-015-4445-3
6. Zhou W-M, He -R-R, Ye J-T, Zhang N, Liu D-Y. Synthesis and biological evaluation of new 5-fluorouracil-substituted ampelopsin derivatives. Molecules. 2010;15:2114–2123. doi:10.3390/molecules15042114
7. Choi JH, Yoon JS, Won YW, Park BB, Lee YY. Chloroquine enhances the chemotherapeutic activity of 5‐fluorouracil in a colon cancer cell line via cell cycle alteration. Apmis. 2012;120:597–604. doi:10.1111/j.1600-0463.2012.02876.x
8. Ndagi U, Mhlongo N, Soliman ME. Metal complexes in cancer therapy–an update from drug design perspective. Drug Des Devel Ther. 2017;11:599. doi:10.2147/DDDT.S119488
9. Cuin A, Massabni AC, Pereira GA, et al. 6-Mercaptopurine complexes with silver and gold ions: anti-tuberculosis and anti-cancer activities. Biomed Pharmacother. 2011;65:334–338. doi:10.1016/j.biopha.2011.04.012
10. Zhang P, Sadler PJ. Advances in the design of organometallic anticancer complexes. J Organomet Chem. 2017;839:5–14. doi:10.1016/j.jorganchem.2017.03.038
11. Tan SJ, Yan YK, Lee PPF, Lim KH. Copper, gold and silver compounds as potential new anti-tumor metallodrugs. Future Med Chem. 2010;2:1591–1608. doi:10.4155/fmc.10.234
12. Haque RA, Nasri SF, Iqbal MA. A new dinuclear Ag (I)–N-heterocyclic carbene complex derived from para-xylyl linked bis-imidazolium salt: synthesis, crystal structure, and in vitro anticancer studies. J Coord Chem. 2013;66:2679–2692. doi:10.1080/00958972.2013.813492
13. Eloy L, Jarrousse AS, Teyssot ML, et al. Anticancer activity of silver–N‐heterocyclic carbene complexes: caspase‐independent induction of apoptosis via mitochondrial Apoptosis‐Inducing Factor (AIF). ChemMedChem. 2012;7:805–814. doi:10.1002/cmdc.201200055
14. Iqbal MA, Umar MI, Haque RA, Ahamed MBK, Asmawi MZB, Majid AMSA. Macrophage and colon tumor cells as targets for a binuclear silver(I) N-heterocyclic carbene complex, an anti-inflammatory and apoptosis mediator. J Inorg Biochem. 2015;146:1–13. doi:10.1016/j.jinorgbio.2015.02.001
15. Kasyanenko N, Qiushi Z, Bakulev V, Osolodkov M, Sokolov P, Demidov V. DNA binding with acetate bis (1, 10-phenanthroline) silver(I) monohydrate in a solution and metallization of formed structures. Polymers. 2017;9:211. doi:10.3390/polym9060211
16. Asif M, Iqbal MA, Hussein MA, et al. Human colon cancer targeted pro-apoptotic, anti-metastatic and cytostatic effects of binuclear Silver(I)–N-Heterocyclic carbene (NHC) complexes. Eur J Med Chem. 2016;108:177–187. doi:10.1016/j.ejmech.2015.11.034
17. García-Giménez JL, Hernández-Gil J, Martínez-Ruíz A, et al. DNA binding, nuclease activity, DNA photocleavage and cytotoxic properties of Cu (II) complexes of N-substituted sulfonamides. J Inorg Biochem. 2013;121:167–178. doi:10.1016/j.jinorgbio.2013.01.003
18. Topala T, Bodoki A, Oprean L. R. Oprean, experimental techniques employed in the study of metal complexes-DNA–interactions. Exp Tech. 2014;62:6.
19. LePecq J-B, Paoletti C. A fluorescent complex between ethidium bromide and nucleic acids: physical—chemical characterization. J Mol Biol. 1967;27:87–106. doi:10.1016/0022-2836(67)90353-1
20. Hossain Z, Huq F. Studies on the interaction between Ag+ and DNA. J Inorg Biochem. 2002;91:398–404. doi:10.1016/S0162-0134(02)00454-3
21. Benjamin Garbutcheon-Singh K, Grant MP, Harper BW, et al. Transition metal based anticancer drugs. Curr Top Med Chem. 2011;11:521–542. doi:10.2174/156802611794785226
22. Łęczkowska A, Vilar R. Interaction of metal complexes with nucleic acids. Ann Rep Section” A”(Inorganic Chemistry). 2013;109:299–316. doi:10.1039/c3ic90029k
23. Gago F. Stacking interactions and intercalative DNA binding. Methods. 1998;14:277–292. doi:10.1006/meth.1998.0584
24. Espinosa Leal LAS, Karpenko A, Swasey S, et al. The role of hydrogen bonds in the stabilization of silver-mediated cytosine tetramers. J Phys Chem Lett. 2015;6:4061–4066. doi:10.1021/acs.jpclett.5b01864
25. Swasey SM, Gwinn EG. Silver-mediated base pairings: towards dynamic DNA nanostructures with enhanced chemical and thermal stability. New J Phys. 2016;18:045008. doi:10.1088/1367-2630/18/4/045008
26. Qu J-Q, Qu L, Jia X-F. 5-Fluoro-uracil-1-acetic acid. Acta Crystallogr Sect E. 2006;62:o1504–o1506. doi:10.1107/S1600536806009676
27. Enriquez GG, Rizvi SA, D’Souza MJ, Do DP. Formulation and evaluation of drug-loaded targeted magnetic microspheres for cancer therapy. Int J Nanomedicine. 2013;8:1393. doi:10.2147/IJN.S37465
28. Olmsted J
29. Chakravarty S, Dureja V, Bhattacharyya G, Maity S, Bhattacharjee S. Removal of arsenic from groundwater using low cost ferruginous manganese ore. Water Res. 2002;36(3):625–632. doi:10.1016/S0043-1354(01)00234-2
30. Qu X, Trent JO, Fokt I, Priebe W, Chaires JB. Allosteric, chiral-selective drug binding to DNA. Proc Natl Acad Sci. 2000;97(22):12032–12037. doi:10.1073/pnas.200221397
31. Kyros L, Banti C, Kourkoumelis N, Kubicki M, Sainis I, Hadjikakou S. Synthesis, characterization, and binding properties towards CT-DNA and lipoxygenase of mixed-ligand silver(I) complexes with 2-mercaptothiazole and its derivatives and triphenylphosphine. J Biol Inorganic Chem. 2014;19:449–464. doi:10.1007/s00775-014-1089-6
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.