Back to Journals » Infection and Drug Resistance » Volume 13
Successful Treatment of a Multidrug-Resistant Tuberculosis Patient with a Negative Xpert MTB/RIF Test for Rifampicin-Resistant Tuberculosis in Guizhou Province of China: A Case Report
Authors Zhao ZL, Chen L , Zhang H
Received 8 January 2020
Accepted for publication 26 April 2020
Published 8 May 2020 Volume 2020:13 Pages 1351—1355
DOI https://doi.org/10.2147/IDR.S245219
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Zhao-Liang Zhao,1 Ling Chen,1 Hong Zhang1,2
1Tuberculosis Division of Respiratory and Critical Care Medicine, Affiliated Hospital of Zunyi Medical University, Zunyi, Guizhou 563003, People’s Republic of China; 2Z-BioMed, Inc., Rockville, MD 20855, USA
Correspondence: Ling Chen; Hong Zhang Tel +86-851-28609237
Fax +86-851-28608239
Email [email protected]; [email protected]
Abstract: The Xpert MTB/RIF (Xpert) assay recommended by the World Health Organization (WHO) can be used to simultaneously detect Mycobacterium tuberculosis complex (MTBC) and rifampicin (RIF) resistance associated mutations. However, if Xpert testing results are negative for RIF resistance because mutations outside the RIF resistance-determining region (RRDR) are not detectable by the assay, patients with RIF-resistant/multidrug-resistant tuberculosis (RR/MDR-TB) will be treated inappropriately for several weeks prior to obtaining the drug susceptibility testing (DST) results. Here, we report a rare case of TB in Guizhou Province of China that was identified as RIF-susceptible by the Xpert MTB/RIF assay, but later was confirmed as MDR-TB by DST, and its successful treatment with effective second-line anti-TB drugs. We detected a rare rpoB mutation (Ile572Phe) in clinical samples of this patient, highlighting the importance of using other methods such as PCR and sequencing to complement the Xpert MTB/RIF assay for the routine diagnosis of RR/MDR-TB because of the limited scope of the assay. These complementary methods allow for the detection of rare rpoB mutations outside the RRDR and can be beneficial when used in geographical locations where such rpoB mutations are frequently reported. However, these methods may not be feasible for resource-limited settings.
Keywords: commercial tests, MTB/RIF Xpert assays, Ile572Phe, Ile491Phe, rifampicin/multidrug-resistant tuberculosis
Introduction
Drug-resistant tuberculosis (DR TB) is a major global public health threat. In 2018, about half a million new cases of rifampicin (RIF)-resistant TB (RR-TB) were reported to WHO. Rifampicin resistance was considered as a predictor of multidrug-resistant TB (MDR-TB) because 78% of RR-TB were MDR-TB.1 To prevent the spread of RR/MDR-TB through quick diagnosis and effective treatment, WHO recommended the use of the Xpert MTB/RIF assay (Xpert, Cepheid, USA) for the rapid diagnosis of Mycobacterium tuberculosis complex (MTBC) and uncovering RIF resistance by detecting common mutations in the 81 bp RIF resistance-determining region (RRDR) of the rpoB gene.1 Usually, when Xpert testing results are positive for MTBC and RR, effective second-line regimens are immediately initiated for TB patients weeks before the drug susceptibility testing (DST) results become available. However, rpoB mutations outside the RRDR are undetectable by commercial molecular assays such as the Xpert MTB/RIF assay.2 If Xpert MTB/RIF testing results are negative for RR due to the limited scope of the assay, but later confirmed by DST as RR/MDR-TB, patients with RR/MDR-TB would be treated inappropriately for several weeks prior to obtaining the DST results. During that period, RR/MDT-TB patients treated with ineffective regimens could potentially spread the pathogen to many people in public places, causing heavy economic and social burdens. Here, we report a rare case of TB, which was identified as RIF-susceptible by the Xpert MTB/RIF assay, but was confirmed as MDR-TB by DST, and its successful treatment with effective second-line anti-TB drugs.
Case Report
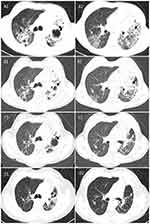
A 32-year-old Chinese woman with a five-year history of pulmonary TB presented to our hospital in February 2017 with symptoms of cough, sputum, and shortness of breath. Physical examination revealed that her heart and abdomen were normal, and no dry and wet rales were heard in her lungs. Computed tomography (CT) scans of the chest showed bilateral pulmonary TB with cavities, bilateral pleural thickening, and enlarged lymph nodes in the upper mediastinum (Figure 1, A1 and A2). Her sputum smear results revealed the presence of acid-fast bacilli (AFB) graded as 2+. No obvious abnormality was found in routine biochemical tests. The patient claimed to have no history of hypertension, diabetes, hepatitis, typhoid and other infectious diseases. While waiting for the DST results, a treatment with four first-line anti-TB drugs was initiated, which included isoniazid (INH, 400 mg daily), RIF (450 mg daily), pyrazinamide (PZA, 1.5 g daily), and ethambutol (EMB, 0.75 mg daily).
Three months later, chest CT scans of the patient showed increased lesions in her left lung (data not shown) and her sputum smear was still graded as 2+. Results from the Xpert MTB/RIF assay were positive for MTBC but negative for RIF resistance, suggesting that the M. tuberculosis isolate was susceptible to RIF. However,results from the traditional DST showed that the clinical isolate collected from this patient three months after admission were resistant to INH, RIF, EMB, streptomycin (STR), levofloxacin (LFX), ofloxacin (OFX) and gatifloxacin (GFX), but susceptible to moxifloxacin (MFX), amikacin (AM), prothionamide (PTO), capreomycin (CM), kanamycin (KM) and para-aminosalicylic acid (PAS). Based on the DST results and WHO treatment guidelines for drug-resistant TB, a new treatment regimen was subsequently selected for this confirmed MDR-TB patient. This regimen contained up to five TB medicines from different groups: group A (fluoroquinolones), MFX; group B (second-line injectable agents), AM; group C (other core second-line agents), PTO and cycloserine (CS); and group D1 (add-on agents), pyrazinamide (PZA).3 The 20-month treatment plan included six months with five medicines (MFX, 400 mg daily; AM, 600 mg a day intramuscularly and five times weekly; CS, 500 mg daily; PTO, 600 mg daily; and PZA, 1.5 g daily), and 14 months with four medicines (MFX, 400 mg daily; CS, 500 mg daily; PTO, 600 mg daily; and PZA, 1.5 g daily).
During the revised treatment period, patient’s sputum smears for AFB were negative seven times while sputum cultures were negative four times during testing at two, four and 11 months. Chest CT scans of the patient performed two months after starting the new treatment showed decreased lesions in bilateral lungs (Figure 1, B1 and B2), and those performed at eight months after starting the new treatment showed that lesions and cavities in bilateral lungs disappeared while pulmonary fibrosis remained unchanged, indicating the treatment was successful (Figure 1, C1 and C2). After the lung lesions of the patient stabilized and her symptoms abated, treatments with second-line anti-TB medicines were discontinued after the 20 months. Follow-up chest CT scans performed seven months after the completion of the 20-months of treatment showed that patient’s lung lesions and fibrosis remained unchanged (Figure 1, D1 and D2), and sputum cultures were negative for M.tuberculosis.
To understand why the Xpert MTB/RIF results were inconsistent with the DST results, we extracted genomic DNA from the clinical isolate (No. s20170731y)4 collected from the first sputum sample of this patient when she was admitted to the hospital, and sequenced the entire RNA polymerase β-subunit (rpoB) gene. We detected an A→T nucleotide substitution at the Escherichia coli codon position 572 of the rpoB gene (ATC572TTC) leading to a missense mutation (Ile572Phe), which was the same as the M. tuberculosis codon position 491 (Ile491Phe).5 In addition, sequencing results showed no other mutations in the rpoB coding region including the 81 bp RRDR.
Discussion
Resistance of M. tuberculosis to RIF is mainly caused by mutations in the rpoB gene, and the majority of mutations are located in the RRDR.6 Mutations outside the RRDR, such as Val146Phe and Ile572Phe, were also found to be associated with RIF resistance in M. tuberculosis isolates.7,8 Currently, commercial molecular assays such as the Xpert MTB/RIF and Genotype MTBDRplus VER 2.0 are unable to detect rpoB gene mutations outside the RRDR.2 Therefore, a novel rapid PCR method was developed to detect the Ile491Phe mutation and proposed to complement commercial assays for the diagnosis of RR-TB, particularly in countries where this specific mutation is frequent.9,10
In this case report, results from the Xpert MTB/RIF assay indicated that the M. tuberculosis was susceptible to RIF, but results from the traditional DST showed that the clinical isolate collected from this patient was MDR-TB. Our sequencing results demonstrated that the inconsistency was caused by a point mutation at the codon 572 of the rpoB gene (Ile572Phe), which was detected previously in RIF-resistant clinical M. tuberculosis isolates.8,11 It has been reported that rpoB mutation profiles of RR-TB isolates are variable depending on the geographical locations,8 and the same is true for the Ile572Phe mutation. For example, the Ile572Phe mutation was identified in one out of 33 RR-TB isolates (3.0%) in Australia,11 in one out of 32 RR-TB isolates (3.1%) in Guizhou Province of China,8 in one out of 50 RR-TB isolates (2.0%) in Hong Kong,7 and in one out of 53 RR-TB isolates (1.9%) in Pakistan.12
Even though the Ile572Phe (Ile491Phe in the M. tuberculosis numbering system) mutation is relatively uncommon, assuming 2% to 3% prevalence in the 500,000 new cases of RR-TB reported globally in 2018, thousands of RR-TB patients may have been misidentified as RIF-susceptible using current commercial molecular assays. Delayed appropriate treatment of RR/MDR-TB patients could lead to extended treatment periods, increases in unsuccessful treatment outcomes, and outbreaks of M. tuberculosis strains with rare mutations such as the Ile572Phe mutation. In this case report, the patient required effective drug regimens consisting of up to five TB medicines from different groups over 20 months in order to be successfully treated, highlighting the importance of rapid and appropriate diagnosis and treatment of RR/MDR-TB.
In conclusion, as the Xpert MTB/RIF assay is increasingly used for simultaneous detection of MTBC and RR, other methods such as PCR and sequencing should be used to complement commercial assays such as the Xpert MTB/RIF for the routine diagnosis of RR/MDR-TB because of the limited scope of the assay. These complementary methods allow for detection of rare rpoB mutations outside the RRDR and can be beneficial when used in geographical locations where such mutations (Val146Phe and Ile572Phe) are frequently reported. However, these methods may not be feasible for some resource-limited settings.
Consent
Written informed consent was obtained from the patient for publication of this report. The collection of M. tuberculosis isolates was a part of the routine hospital laboratory procedures and therefore the institutional approval was not required to publish the case details.
Acknowledgments
This study was supported by the National Natural Science Foundation of China (NSFC no. 81760003), the 2nd 2011 Collaborative Innovation Center for TB Prevention and Cure in Guizhou Province (Incubation Project), and the 4th Talented Individual Base for Infectious Disease Prevention and Control in Guizhou Province (Qian 2013–15). The sponsors played no role in the collection, analysis, and interpretation of data; in writing the manuscript; and in the decision to submit the report for publication. We thank Derek T. Zhang for reading, editing and revising the manuscript prior to submission.
Disclosure
HZ is employed by and has shares in Z-BioMed, Inc., which is involved in infectious disease research. The authors report no other conflicts of interest in this work.
References
1. World Health Organization. Global Tuberculosis Report 2019. Geneva: World Health Organization; 2019. Licence: CCBY-NC-SA3.0IGO.
2. Torrea G, Ng KCS, Van Deun A, et al. Variable ability of rapid tests to detect Mycobacterium tuberculosis rpoB mutations conferring phenotypically occult rifampicin resistance. Sci Rep. 2019;9(1):11826. doi:10.1038/s41598-019-48401-z
3. World Health Organization. WHO treatment guidelines for drug-resistant tuberculosis. Geneva, Switzerland; 2016. Available from: http://apps.who.int/medicinedocs/en/d/Js23097en/.
4. van Soolingen D, Hermans PW, de Haas PE, Soll DR, van Embden JD. Occurrence and stability of insertion sequences in Mycobacterium tuberculosis complex strains: evaluation of an insertion sequence-dependent DNA polymorphism as a tool in the epidemiology of tuberculosis. J Clin Microbiol. 1991;29(11):2578–2586.
5. Andre E, Goeminne L, Cabibbe A, et al. Consensus numbering system for the rifampicin resistance-associated rpoB gene mutations in pathogenic mycobacteria. Clin Microbiol Infect. 2017;23(3):167–172. doi:10.1016/j.cmi.2016.09.006
6. Zaw MT, Emran NA, Lin Z. Mutations inside rifampicin-resistance determining region of rpoB gene associated with rifampicin-resistance in Mycobacterium tuberculosis. J Infect Public Health. 2018;11(5):605–610. doi:10.1016/j.jiph.2018.04.005
7. Siu GK, Zhang Y, Lau TC, et al. Mutations outside the rifampicin resistance-determining region associated with rifampicin resistance in Mycobacterium tuberculosis. J Antimicrob Chemother. 2011;66(4):730–733. doi:10.1093/jac/dkq519
8. Chen L, Gan X, Li NN, et al. rpoB gene mutation profile in rifampicin-resistant Mycobacterium tuberculosis clinical isolates from Guizhou, one of the highest incidence rate regions in China. J Antimicrob Chemother. 2010;65(6):1299–1301. doi:10.1093/jac/dkq102
9. André E, Goeminne L, Colmant A, Beckert P, Niemann S, Delmee M. Novel rapid PCR for the detection of Ile491Phe rpoB mutation of Mycobacterium tuberculosis, a rifampicin-resistance-conferring mutation undetected by commercial assays. Clin Microbiol Infect. 2017;23(4):
10. Al-Mutairi NM, Ahmad S, Mokaddas E, Eldeen HS, Joseph S. Occurrence of disputed rpoB mutations among Mycobacterium tuberculosis isolates phenotypically susceptible to rifampicin in a country with a low incidence of multidrug-resistant tuberculosis. BMC Infect Dis. 2019;19(1):3. doi:10.1186/s12879-018-3638-z
11. Yuen LK, Leslie D, Coloe PJ. Bacteriological and molecular analysis of rifampin-resistant Mycobacterium tuberculosis strains isolated in Australia. J Clin Microbiol. 1999;37(12):3844–3850. doi:10.1128/JCM.37.12.3844-3850.1999
12. Javed H, Bakuła Z, Pleń M, et al. Evaluation of genotype MTBDRplus and MTBDRsl assays for rapid detection of drug resistance in extensively drug-resistant Mycobacterium tuberculosis isolates in Pakistan. Front Microbiol. 2018;9:2265. doi:10.3389/fmicb.2018.02265
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.