Back to Journals » Infection and Drug Resistance » Volume 13
Prevalence and Some Possible Mechanisms of Colistin Resistance Among Multidrug-Resistant and Extensively Drug-Resistant Pseudomonas aeruginosa
Authors Abd El-Baky RM , Masoud SM, Mohamed DS, Waly NGFM, Shafik EA, Mohareb DA, Elkady A , Elbadr MM, Hetta HF
Received 15 November 2019
Accepted for publication 11 January 2020
Published 3 February 2020 Volume 2020:13 Pages 323—332
DOI https://doi.org/10.2147/IDR.S238811
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Rehab M Abd El-Baky, 1, 2 Salwa M Masoud, 1 Doaa S Mohamed, 2 Nancy GFM Waly, 1 Engy A Shafik, 3 Dina A Mohareb, 4 Azza Elkady, 5 Mohamed M Elbadr, 6 Helal F Hetta 7, 8
1Department of Microbiology and Immunology, Faculty of Pharmacy, Minia University, Minia 61519, Egypt; 2Department of Microbiology and Immunology, Faculty of Pharmacy, Deraya University, Minia 11566, Egypt; 3Department of Clinical Pathology, South Egypt Cancer Institute, Assiut University, Assiut, Egypt; 4Department of Clinical Pathology, Faculty of Medicine, Assiut University, Assiut, Egypt; 5Sohag General Hospital, Sohag, Egypt; 6Department of Pharmacology, Faculty of Medicine, Assiut University, Assiut, Egypt; 7Department of Medical Microbiology and Immunology, Faculty of Medicine, Assiut University, Assiut, Egypt; 8Department of Internal Medicine, University of Cincinnati College of Medicine, Cincinnati, OH, USA
Correspondence: Helal F Hetta
Department of Internal Medicine, University of Cincinnati College of Medicine, PO Box 670595, Cincinnati, OH 45267-0595, USA
Email [email protected]
Background and Aim: The emergence of colistin-resistant strains is considered a great threat for patients with severe infections. Here, we investigate the prevalence and some possible mechanisms of colistin resistance among multidrug-resistant (MDR) and extensively drug-resistant (XDR) Pseudomonas aeruginosa (P. aeruginosa).
Methods: Antimicrobial susceptibility was performed using disc diffusion methods while colistin resistance was detected by agar dilution method. Possible mechanisms for colistin resistance were studied by detection of mcr-1 and mcr-2 genes by conventional PCR, detection of efflux mechanisms using Carbonyl Cyanide 3-Chlorophenylhydrazone (CCCP), studying outer membrane protein profile and Lipopolysaccharide (LPS) profile of resistant isolates.
Results: It was found that MDR and XDR represented 96% and 87% of the isolated P. aeruginosa, respectively, and colistin resistance represented 21.3%. No isolates were positive for mcr-2 gene while 50% of colistin-resistant isolates were positive for mcr-1. Efflux mechanisms were detected in 3 isolates. Protein profile showed the presence of a band of 21.4 KDa in the resistant strains which may represent OprH while LPS profile showed differences among colistin-resistant mcr-1 negative strains, colistin-resistant mcr-1 positive strains and susceptible strains.
Conclusion: The current study reports a high prevalence of colistin resistance and mcr-1 gene in P. aeruginosa strains isolated from Egypt that may result in untreatable infections. Our finding makes it urgent to avoid unnecessary clinical use of colistin.
Keywords: Pseudomonas aeruginosa, colistin resistance, mcr-1, mcr-2, toxA gene, XDR, MDR
Introduction
Pseudomonas aeruginosa (P. aeruginosa) is an opportunistic pathogen, commonly found in environment such as soil, water, plants and hospital environment, with known intrinsic resistance to many antimicrobials and the ability to cause life-threatening infections. It is considered the second common cause of sepsis in intensive care units (ICUs) and can cause ventilator-associated pneumonia, wound infections and urinary tract infections (UTI). Many studies reported the increase of mortality and morbidity of infections associated with P. aeruginosa, especially those showing multi-drug resistance patterns.1–3
The emergence of multidrug-resistant (MDR) or extensively drug-resistant (XDR) or pandrug-resistant (PDR) P. aeruginosa becomes a significant public health problem that can lead to delayed antimicrobial therapy or its failure and the increase in the mortality rate especially with the appearance of carbapenem-resistant P. aeruginosa. So, attention is required because these resistant strains may show resistance to all available antimicrobials or showed susceptibility only to toxic ones such as colistin or polymyxins leaving no choices for the health-care team in the treatment of severe infections associated with MDR P. Aeruginosa.4
Recently, emergence of resistance to polymyxins was observed among certain species of Enterobacteriaceae such as K. pneumoniae, E. coli, Enterobacter aerogenes and Enterobacter cloacae due to its wide use to control infections in veterinary medicine. Colistin resistance has become a major challenge for the treatment of life-threatening infections especially with the co-existence of mcr-1 genes with other multiple drug resistance genes as ESBL, MBL, NDM genes with the possibility of the emergence of pan-drug resistance.5,6
Colistin, known as polymyxin E, is one member of a family of cationic polypeptide known as polymyxins. This antibiotic family is characterized by the presence of a lipophilic fatty acyl side chain. Nowadays, colistin is reintroduced in medical therapy and considered the last resort for the treatment of severe infections caused by MDR and XDR stains. In general, the action of polymyxins on bacteria depends mainly on the electrostatic interaction between the positively charged antibiotic and the negatively charged phosphate group of lipid A localized on the outer membrane after its binding, it diffuses through the outer membrane, periplasmic space and interact with the inner membrane. Polymyxins cause destabilization to the outer membrane, pore formation, increase permeability, leakage to cytoplasmic content followed by cell lysis.7
Colistin resistance mainly occurs due to the chemical modification by the enzymatic addition of phosphoethanolamine at the 4ʹ- phosphate group of the lipid A moiety of the lipopolysaccharide decreasing the net-negative charge of the outer membrane resulting in decreasing polymyxin affinity. Resistance to colistin may be resulted from chromosomally encoded mutation as reported in K. pneumoniae or the horizontal transfer of resistance by means of plasmid carrying colistin-resistant gene (mcr-1).8–11
The emergence of colistin resistance in various countries in Asia, Europe and some countries in Africa has become one of the global concerns. As, colistin resistance dissemination indicates its ability to transfer horizontally by conjugative plasmids or vertically by chromosomal mutation.12,13 Also, being colistin one of the last lines of treatments to serious infections, making the emergence of colistin resistance isolates threatening the world by the appearance of untreatable infectious diseases.14 Detection of colistin resistance in Egypt, which is a country known by its high burden of infectious diseases and the presence of low or no restriction on the antimicrobial use in both veterinary and medicine, indicating the emergence of untreatable diseases in our area due to the possibility of transferring colistin resistance to highly resistant bacteria.15
In this study, we investigate the prevalence of colistin resistance among MDR and XDR P. aeruginosa isolated from patients suffering from a variety of infections in the intensive care unit (ICU) of Minia university hospital in Egypt.
Materials and Methods
Collection of Isolates
One hundred-seventy five clinical samples of different sources of infections were collected from patients admitted to ICU in Minia university Hospital, Minia, Egypt as part of routine hospital-laboratory procedures. All clinical samples were cultured on trypticase soy agar (Lab M, UK) at 37°C and 42°C for 24 hrs. One colony was sub-cultured on MacConkey agar plates and cetrimide agar. Isolated colonies were further identified according to colony morphology, lactose fermentation, biochemical reactions (including sulphide–indole motility, catalase, triple sugar iron, urease and oxidase tests), ability to grow on cetrimide agar and to grow at 42°C.16 P. aeruginosa colonies were purified by streaking, and pure colonies were stored at 4°C.
Antibiotic Susceptibility Tests
Antibiotic Susceptibility by Kirby-Bauer Disc Diffusion Method
The antibiotic susceptibility against different classes of antibiotics was tested by the Kirby-Bauer disc diffusion method.17 Antibiotic discs used were amoxicillin/clavulanic (AMC) (20/10 μg), ampicillin/sulbactam (SAM) (20 μg), meropenem (MEM) (10 μg), imipenem (IPM) (10 μg), cefepime (FEB) (30 μg), cefoperazone (CEP) (75 μg), polymyxin B (PB) (300 μg), ciprofloxacin (CIP) (5 μg), levofloxacin (LEV) (5 μg), gentamicin (CN) (10μg), ceftazidime (CAZ) (30 μg), tigecycline (TGC) (15 μg), amikacin (AK) (30 μg), tobramycin (TOB) (10 μg), aztreonam (ATM) (30 μg), piperacillin (PRL) (30 μg), carbenicillin (CAR) (100 μg) (Oxoid; Basingstoke, UK). Isolates were classified as sensitive, intermediate and resistant according to inhibition zones' interpretation standards of Clinical Laboratory standards Institute (CLSI) 2018.18
MIC Determination of Colistin Antibiotic
Agar dilution method on Muller-Hinton agar was used to determine the colistin minimum inhibitory concentration.19 Resistance to colistin was considered if the MIC is ≥4μg/mL according to the standard guidelines of CLSI.18
According to the results of antibiotic susceptibility, isolates were classified to MDR, XDR and PDR according to the criteria previously reported.20
Combined Disc Diffusion Test (CDT)
All colistin-resistant isolates (MIC ≥4) were tested using 100 mM EDTA (Sigma-Aldrich; St.268 Louis, MO, USA) to inhibit the mcr-1 activity as this concentration showed no antimicrobial activity. The bacterial strains were cultured on Muller-Hinton agar (Lab M, UK) on which three discs were used. One disc was saturated with 10 μL of 100 mM EDTA to insure no inhibition of the bacterial growth by the used concentration of EDTA. The other two discs were 10μg colistin disc and 10 μg colistin plus 10 μL 100 mM EDTA disc. The isolates were observed for an increase of ≥3mm in the inhibition zone diameter of the colistin/EDTA disc comparing to the colistin disc.21
Alteration of Zeta Potential
The mcr genes encode phosphoethanolamine transferases enzymes which attach enzymatically a phosphoethanolamine (PEtN) moiety to the lipid A of the outer membrane of Gram-negative bacteria leading to reduction in its net negative charge conferring the colistin resistance.22
The bacterial cells have been allowed to grow in the presence and absence of 80 μg/mL EDTA. Then, the bacterial suspension was centrifuged at 5000 rpm for 5 min at 5°C then pellets were washed twice, after that pellets were suspended in 2 mL of sterile 1 mM NaCl solution adjusted to 0.5 McFarland standard solution turbidity. Samples were diluted to 1:4 using 1mM NaCl. Zeta potential was determined in 2 mL of the diluted sample. Alterations of Zeta potential induced by EDTA were calculated from the Zeta potential ratio (RZP=ZP+EDTA/ZP-EDTA), where ZP+EDTA and ZP-EDTA correspond to Zeta potential values obtained for bacterial suspensions grown in the presence or absence of 80μg/mL EDTA, respectively. RZP of ≥ 2.5 value considered as criteria for the identification of mcr-1 positive strains.21
DNA Extraction
The DNA template was extracted from an overnight culture of P. aeruginosa as previously described.23 A suspension of bacterial pellet was boiled for 10 min, then, centrifuged. Supernatant was used directly in the PCR assay.
PCR Analysis of the Tested Genes
Exotoxin A is an important virulence factor (a cytotoxic agent) of P. aeruginosa in clinical infections. This factor inhibits protein biosynthesis leading to great tissue and organ damage. The toxA gene, an inherent genetic sequence located on the P. aeruginosa chromosome, is used for P. aeruginosa confirmation by PCR.
PCR was performed in a total volume of 25 µL containing 1X PCR buffer, 1 µmol/L of each primer, 1 µL of genomic DNA (approximately150 ng), 200 µmol/L of dNTPs mix, 2 mmol/L of MgCl2, and 0.05 U/µL Taq DNA polymerase. PCR amplifications were performed for toxA FW:CTGCGCGGGTCTATGTGCC, RV:GATGCTGGACGGGTCGAG in an automated thermal cycler (Eppendorf, Hamburg, Germany) under the following conditions: 30 cycles of 1 min at 94°C, 1.5 min at 63°C, and 1 min at 72°C.24
The genes mcr-1 and mcr-2 were assayed by conventional PCR technique using the following primers: mcr-1 FW (5′-AGTCCGTTTGTTCTTGTGGC-3′), RV (5′-AGATCCTTGGTCTCGGCTTG-3′) and mcr-2 Fw (5ʹ-ATGACATCACATCACTCTTGG-3ʹ), Rv (5ʹ-TTACTGGATAAATGCCGCGC-3ʹ).25,26 The technique conditions were 34 cycles of 95°C for 1 min, 58°C for mcr-1 and 52°C for 30s, 72°C for 1min followed by final extension of 72°C for 5 mins.
Determination of Efflux Pumps Inhibition by MIC Reduction Using Efflux Pump Inhibitor (CCCP)
The agar dilution method was used for the determination of MICs using the Cation-adjusted Mueller-Hinton broth (Sigma-Aldrich, St Louis, USA). The MICs of CCCP (EPI) and colistin were determined for the tested isolates. Sub-MIC of CCCP was used in determining its effect on colistin MIC; the concentration of CCCP (0.5× MIC) was constantly kept at the MIC concentrations stated above whilst that of the antibiotic were serially increased. The MICs of the isolates to colistin in the absence and presence of CCCP were determined using a sub-MIC of CCCP (final concentration of 10 mg/L) as already described.27 The resulting MIC fold changes after the addition of CCCP were calculated as the ratio of the CCCP-free antibiotic’s MIC level to that of the CCCP-added antibiotic. As previously described by Osei Sekyere, Amoako28 who reported that the positive criterion for the presence of efflux pumps in isolates was a ≥8-fold decrease in colistin MIC after adding CCCP.
Outer Membrane Protein Pattern
A single colony of the tested P. aeruginosa isolates was cultured in 5 mL of LB broth at 37°C for 2 days with shaking at 200 rpm. Cells were centrifuged at 8000 rpm for 5 mins. Bacterial pellets were suspended in 1 mL of lysis buffer (0.05 M Tris HCL, 2% SDS, 10% glycerol), heated at 95°C for 10 mins. Then, the samples were centrifuged for 10.000 rpm for 30 min. About 50 µL of extracted protein was mixed with sample buffer (4 mL deionized water, 1 mL of 0.5 M Tris HCL, 1.6mL 10% SDS, 0.4 mL 2-mercaptoethanol, 0.2 mL of 1% (w/v) Bromophenol blue) (1:1) and separated by 12% sodium dodecyl sulfate-polyacrylamide gel (SDS-PAGE).29
Lipopolysaccharide SDS-Polyacrylamide Gel Profile for Colistin Sensitive and Colistin-Resistant Isolates
LPS of the tested isolates were extracted and purified by hot aqueous-phenol method using Westphal, Jann30 and analyzed the purified material using SDS-PAGE, followed by carbohydrate-specific silver staining.31
Results
Pseudomonas aeruginosa Isolation and Antibiotic Susceptibility
Out of 175 samples collected from patients suffering from different infections, 75 samples (42.8%) were positive phenotypically for P. aeruginosa and positive for toxA gene.
Antimicrobial susceptibility testing revealed that the isolated P. aeruginosa were completely resistant to amoxicillin/clavulanic acid and high resistance was observed against ampicillin/sulbactam (68%), ceftazidime (63%) and azetreonam (60%). Moderate resistance was observed against both tobramycin and tigecycline (50% each). Furthermore, low resistance was shown against imipenem (6%) and meropenem (5.3%) (Figure 1). According to the antibiotic susceptibility results the resistant isolates were classified to MDR (96%), XDR (87%) and no isolate was classified as PDR. In addition, it was found that out of 75 isolates, 16 isolates (21.3%) showed resistance to colistin antibiotic with MIC ≥ 4μg/mL (ranged from 8 to 256 μg/mL).
 |
Figure 1 Antibiotic resistance pattern of all isolated P. aeruginosa isolates. |
Determination of Mcr-1 and Mcr-2 Genes
Mcr-1 gene was detected phenotypically in the colistin-resistant isolates by CDT where the differences between the diameters of inhibition Zones of colistin/EDTA and colistin discs were measured to be ≥ 3mm. The results showed that 6 isolates (37.5%) showed an increase in the diameter of the colistin/EDTA disc by 3 to 10 mm in comparison to colistin disc alone (Figure 2).
Alteration of Zeta Potential
On the other hand, alteration of Zeta potential assay was held as a phenotypic detection to MCR genes, but results showed no significant change in the zeta potential except in 2 isolates.
Detection of Resistance Genes
The genetic detection of mcr genes using conventional PCR technique revealed that 8 (50%) isolates were positive for mcr-1, 6 of them were positive for CDT, while 100% (16 isolates) were negative for mcr-2.
Antibiotic Susceptibility of Colistin-Resistant Isolates
The susceptibility of the colistin-resistant isolate against other antibiotics was determined by Kirby-Bauer disc diffusion method, the results showed that 100% of isolates were resistant to Amoxicillin/clavulanic, while resistance to Ampicillin/sulbactam, Cefepime and Tobramycin was 78.12%, 71.87% and 68.75% respectively. The most effective drugs were meropenem, imipenem and ciprofloxacin (Figure 3)
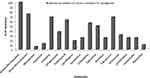 |
Figure 3 Antibiotic resistance pattern of colistin-resistant isolates. |
Determination of Efflux Pumps Inhibition by MIC Reduction Using CCCP
By studying the effect of 0.5 MIC of CCCP on the MIC colistin, it was found that only 3/16 isolates (P6, P8 & P16) (18.75%) showed a reduction in the MICs of colistin ≥ 8 fold (Table 1) in the presence of CCCP. From previous results, the isolate no. 16 was found to have efflux mechanism and mcr-1 gene.
 |
Table 1 Colistin-Resistant Isolates, Some Possible Mechanisms of Resistance to Colistin and Their Susceptibility to Other Antibiotics |
Outer Membrane SDS-PAGE Profile
Table 2 and Figure 4 show that five bands with molecular weights of 66.7, 56.06, 47.8, 40.18 and 23.6 KDa were stable in sensitive and resistant isolates while one band with a molecular weight of 21 KDa was found only in colistin-resistant strains which were P1 (mcr-1 positive) and P12 (mcr-1 negative).
 |
Table 2 Molecular Weights and Amount % of Extracted Outer Membrane Proteins of Colistin Resistant and Colistin Sensitive P. Aeruginosa |
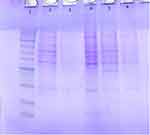 |
Figure 4 Outer membrane SDS-PAGE of colistin resistant and sensitive strains. Lane 1: Protein Marker, Lane 2 and Lane 3: colistin-resistant strains (P1 & P12), Lanes 4–6: colistin sensitive strains. |
Lipopolysaccharide (LPS) SDS-PAGE
Lipopolysaccharide silver-stained SDS-PAGE showed that colistin-resistant mcr-1 negative isolates (P3, P6 and P10) showed no LPS bands pattern (O-antigen repeats or LPS core) that revealed the possibility of their loss and the resistance of these isolates to colistin. On the other hand, Colistin-resistant mcr-1 positive strain showed O-antigen repeats (Figure 5, Lane 5) that differs from O-antigen repeats pattern of Colistin sensitive strain (Figure 5, Lane 4) while both showed LPS core. These results may indicate the presence of modified LPS in the mcr-1 positive strain.
Discussion
Recently, multidrug-resistant pathogenic bacterial strains appear where most of the available antibiotics are not effective against them.6,32–36 The polymyxins considered the last resort for treatment of multi-drug resistant bacterial infections, so studying the emergence of colistin-resistant was a must. Polymyxins showed their activity through their electrostatic interaction between them and the negatively charged moieties on the lipid A of Gram-negative bacteria resulting in the destabilization of the outer membrane and the leakage of the cytoplasmic content and lysis.37,38
It was found that the most common cause of polymyxin resistance is LPS modification by the addition of 4-amino- 4-deoxy-L-arabinose (Lara4N) and phosphoethanolamine (encoded by mcr-type genes) or galactosamine to lipid A of LPS core. As a result, a decrease in the net-negative charge of phosphate residues affects the affinity of polymyxin to the membrane or due to the effect of two-component regulatory systems (TCSs) pmrA/pmrB and phoP/phoQ.39
In our study, we detected colistin resistance according to the results of MICs followed by their testing for the presence of mcr-1 phosphoethanolamine transferase using phenotypic methods and the detection of mcr-1gene. Phenotypic methods depend on that mcr-1 phosphoethanolamine is zinc metalloprotein. So, any decrease in zinc will decrease MICs of colistin in isolates positive for mcr-1. Being mcr-1 encoding enzyme, a zinc metalloprotein permits using EDTA as a metal chelator to decrease zinc in media and affect colistin MICs and the zeta potentials of mcr-1 positive isolates.40
Our study showed high prevalence of P. aeruginosa (42.8%). MDR P. aeruginosa corresponded to 96% of total isolates and 87% was XDR. High prevalence of colistin-resistant P. aeruginosa (21.3%) was detected which may be a result of insufficient infection control measures and misuse of bactericidal antibiotics in the intensive care units of our country hospitals. In addition, Colistin is widely used in our countries in the growth promotion of food-producing animals, especially in poultry industry while carbapenems used in emergency cases.15 So, carbapenems showed observable activity against the tested organisms in comparison to colistin. On the other hand, our results were observed to be higher than those reported by Liassine et al25 who reported that one isolate of 300 isolates of different bacterial species was identified as P aeruginosa showing resistance to colistin and harboring mcr-1 gene.
Combined disc diffusion test (CDT) and the alteration of zeta potential induced by EDTA were used as phenotypic methods41,42 for the detection of mcr-1 gene. The results showed that no isolates were positive for mcr-2 and 8 (50%) isolates of colistin-resistant isolates were mcr-1 positive while 2 isolates of these isolates showed RZP > 2.5. Out of 8 mcr-1 positive isolates, 6 isolates were positive for CDT while two mcr-1 positive (strain No. P15 and P16) were negative for CDT which may be due to co-production of additional mechanism of colistin resistance that interferes with the effect of EDTA.21 As, it was found that isolate no. P16 (mcr-1 positive and CDT negative) was positive for efflux.21,43–45 In addition, colistin-resistant isolates that were negative for mcr-1 may have mutations due to the long-term use of antimicrobials.
Furthermore, we tested colistin-resistant isolates for the presence of efflux mechanisms using CCCP (an efflux pump inhibitor) and the difference in outer membrane protein and LPS SDS-PAGE profile among sensitive and resistant isolates. Our results revealed the presence of efflux mechanism among 3 isolates while one of them was mcr-1 positive. Outer membrane protein profile showed one band with a molecular weight of 21 KDa in the resistant isolates P1 (mcr-1 positive) and P12 (colistin-resistant mcr-1 negative). In addition, it was found that colistin-resistant mcr-1 negative strains showed no LPS bands pattern (O-antigen repeats or LPS core) but mcr-1 positive (P1) and colistin sensitive isolates showed LPS core but different O-antigen repeats pattern. Machado et al20 studied the role of efflux pump in colistin resistance in Acinetobacter baumanni and found that efflux activity contributes to the heteroresistance of A. baumanni in absence of mutation. Marjani et al43 showed that 22.5% of the isolated P. aeruginosa were resistant to colistin which is close to our results and more than 50% of colistin-resistant isolates were positive for efflux pumps.
Although the exact mechanism of bacterial killing by colistin or polymyxins is not clearly known, it is known that their binding to the positively charged peptides and the negatively charged Lipid A is a critical step. So, we tested their LPS SDS-PAGE profile and a significant difference among the tested strains was observed. In a study done by Moffatt et al,46 it was reported that the loss of LPS resulted in the emergence of A. baumanii colistin resistant which occurs due to the inactivation of lipid A biosynthesis genes (lpxA, lpxC, or lpxD). Outer membrane protein patterns showed the presence of a band of molecular weight which is 21 KDa in colistin-resistant isolates which may correspond to OprH according to that reported by Nicas and Hancock47 who reported that OprH expression plays a role in the resistance of Pseudomonas to polymyxins and EDTA because OprH replaces divalent cations in the outer membrane resulting in the blocking of polycationic antibiotic uptake. The previous finding may explain why strain no. P1 (mcr-1 positive) was negative for CDT.
Conclusions
The present study showed a high prevalence of MDR and XDR P. aeruginosa showing colistin resistance among patients admitted to ICU suffering from different infections. Also, it showed the presence of different mechanisms that can result in colistin resistance. This indicates the urgent need of changing the antibiotic-treatment strategies for both humans and animals.
Acknowledgment
We would like to thank Dr. Enas Daef and all members of the Medical Research Center, Faculty of Medicine, Assiut University, Egypt for providing the necessary laboratory facilities for carrying out the experiments.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Gad GF, El-Domany RA, Zaki S, Ashour HM. Characterization of Pseudomonas aeruginosa isolated from clinical and environmental samples in Minia, Egypt: prevalence, antibiogram and resistance mechanisms. J Antimicrob Chemother. 2007;60(5):1010–1017. doi:10.1093/jac/dkm348
2. Farhan SM, Ibrahim RA, Mahran KM, Hetta HF, El-Baky RMA. Antimicrobial resistance pattern and molecular genetic distribution of metallo-β-lactamases producing Pseudomonas aeruginosa isolated from hospitals in Minia, Egypt. Infect Drug Resist. 2019;12:2125. doi:10.2147/IDR.S198373
3. El-Mokhtar MA, Hetta HF. Ambulance vehicles as a source of multidrug-resistant infections: a multicenter study in Assiut City, Egypt. Infect Drug Resist. 2018;11:587. doi:10.2147/IDR.S151783
4. Hosseininassab Nodoushan SA, Yadegari S, Moghim S, et al. Distribution of the strains of multidrug-resistant, extensively drug-resistant, and pandrug-resistant pseudomonas aeruginosa isolates from burn patients. Adv Biomed Res. 2017;6:74. doi:10.4103/abr.abr_239_16
5. Baron S, Hadjadj L, Rolain JM, Olaitan AO. Molecular mechanisms of polymyxin resistance: knowns and unknowns. Int J Antimicrob Agents. 2016;48(6):583–591. doi:10.1016/j.ijantimicag.2016.06.023
6. Al-Kadmy IM, Ibrahim SA, Al-Saryi N, Aziz SN, Besinis A, Hetta HF. Prevalence of genes involved in colistin resistance in acinetobacter baumannii: first report from Iraq. Microb Drug Resist. 2019. doi:10.1089/mdr.2019.0243
7. Wanty C, Anandan A, Piek S, et al. The structure of the neisserial lipooligosaccharide phosphoethanolamine transferase A (LptA) required for resistance to polymyxin. J Mol Biol. 2013;425(18):3389–3402. doi:10.1016/j.jmb.2013.06.029
8. Cannatelli A, D’Andrea MM, Giani T, et al. In vivo emergence of colistin resistance in Klebsiella pneumoniae producing KPC-type carbapenemases mediated by insertional inactivation of the PhoQ/PhoP mgrB regulator. Antimicrob Agents Chemother. 2013;57(11):5521–5526. doi:10.1128/AAC.01480-13
9. Paterson DL, Harris PN. Colistin resistance: a major breach in our last line of defence. Lancet Infect Dis. 2016;16(2):132–133. doi:10.1016/S1473-3099(15)00463-6
10. Hasman H, Hammerum AM, Hansen F, et al. Detection of mcr-1 encoding plasmid-mediated colistin-resistant Escherichia coli isolates from human bloodstream infection and imported chicken meat, Denmark 2015. Euro Surveill. 2015;20:49. doi:10.2807/1560-7917.ES.2015.20.49.30085
11. Zhi C, Lv L, Yu LF, Doi Y, Liu JH. Dissemination of the mcr-1 colistin resistance gene. Lancet Infect Dis. 2016;16(3):292–293. doi:10.1016/S1473-3099(16)00063-3
12. Schwarz S, Johnson AP. Transferable resistance to colistin: a new but old threat. J Antimicrob Chemother. 2016;71(8):2066–2070. doi:10.1093/jac/dkw274
13. Ye H, Li Y, Li Z, et al. Diversified mcr-1-harbouring plasmid reservoirs confer resistance to colistin in human gut microbiota. MBio. 2016;7(2):e00177. doi:10.1128/mBio.00177-16
14. Liu YY, Wang Y, Walsh TR, et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis. 2016;16(2):161–168. doi:10.1016/S1473-3099(15)00424-7
15. Khalifa HO, Ahmed AM, Oreiby AF, Eid AM, Shimamoto T, Shimamoto T. Characterisation of the plasmid-mediated colistin resistance gene mcr-1 in Escherichia coli isolated from animals in Egypt. Int J Antimicrob Agents. 2016;47(5):413–414. doi:10.1016/j.ijantimicag.2016.02.011
16. Fazeli H, Akbari R, Moghim S, Narimani T, Arabestani MR, Ghoddousi AR. Pseudomonas aeruginosa infections in patients, hospital means, and personnel’s specimens. J Res Med Sci. 2012;17(4):332–337.
17. Hudzicki J. Kirby-Bauer Disk Diffusion Susceptibility Test Protocol. 2009.
18. Wayne P. CLSI. Performance Standards for Antimicrobial Susceptibility Testing. CLSI Supplement M100.
19. Wiegand I, Hilpert K, Hancock RE. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc. 2008;3(2):163–175. doi:10.1038/nprot.2007.521
20. Machado D, Antunes J, Simoes A, et al. Contribution of efflux to colistin heteroresistance in a multidrug resistant Acinetobacter baumannii clinical isolate. J Med Microbiol. 2018;67(6):740–749. doi:10.1099/jmm.0.000741
21. Esposito F, Fernandes MR, Lopes R, et al. Detection of colistin-resistant MCR-1-positive escherichia coli by use of assays based on inhibition by EDTA and zeta potential. J Clin Microbiol. 2017;55(12):3454–3465. doi:10.1128/JCM.00835-17
22. Liu YY, Chandler CE, Leung LM, et al. Structural modification of lipopolysaccharide conferred by mcr-1 in gram-negative ESKAPE pathogens. Antimicrob Agents Chemother. 2017;61:6. doi:10.1128/AAC.00580-17
23. Wilson R, Pitt T, Taylor G, et al. Pyocyanin and 1-hydroxyphenazine produced by Pseudomonas aeruginosa inhibit the beating of human respiratory cilia in vitro. J Clin Invest. 1987;79(1):221–229. doi:10.1172/JCI112787
24. Faraji F, Mahzounieh M, Ebrahimi A, Fallah F, Teymournejad O, Lajevardi B. Molecular detection of virulence genes in Pseudomonas aeruginosa isolated from children with Cystic Fibrosis and burn wounds in Iran. Microb Pathog. 2016;99:1–4. doi:10.1016/j.micpath.2016.07.013
25. Liassine N, Assouvie L, Descombes M-C, et al. Very low prevalence of MCR-1/MCR-2 plasmid-mediated colistin resistance in urinary tract Enterobacteriaceae in Switzerland. Int J Infect Dis. 2016;51:4–5. doi:10.1016/j.ijid.2016.08.008
26. Rebelo AR, Bortolaia V, Kjeldgaard JS, et al. Multiplex PCR for detection of plasmid-mediated colistin resistance determinants, mcr-1, mcr-2, mcr-3, mcr-4 and mcr-5 for surveillance purposes. Euro Surveill. 2018;23(6). doi:10.2807/1560-7917.ES.2018.23.6.17-00672
27. He F, Fu Y, Chen Q, et al. Tigecycline susceptibility and the role of efflux pumps in tigecycline resistance in KPC-producing Klebsiella pneumoniae. PLoS One. 2015;10(3):e0119064. doi:10.1371/journal.pone.0119064
28. Osei Sekyere J, Amoako DG. Carbonyl Cyanide m-Chlorophenylhydrazine (CCCP) reverses resistance to colistin, but not to carbapenems and tigecycline in multidrug-resistant enterobacteriaceae. Front Microbiol. 2017;8:228. doi:10.3389/fmicb.2017.00228
29. Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227(5259):680–685. doi:10.1038/227680a0
30. Westphal O, Jann K. Bacterial lipopolysaccharides extraction with phenol-water and further applications of the procedure. Methods Carbohydr Chem. 1965;5:83–91.
31. Davis MR
32. Ahmed S, Ahmed S, Mohamed W, et al. Nosocomial vancomycin and methicillin resistant staphylococcal infections in intensive care units in Assiut University Hospitals. Egypt J Med Microbiol. 2011;20:2.
33. Farhan SM, Ibrahim RA, Hetta HF, Mahran KM, Abdelbaky RM. Prevalence of Oxa-23 in Multidrug Resistance Acinetobacter Baumannii Isolated from Different Infections at Minia University Hospital. N. Egypt. J. Microbiol. 2018; 50:34–43.
34. Abd NE, Abdel-Aleem JA, Abdo MN, Abou-Ghadir OF, Zahran KM, Hetta HF. Efficacy of ketoconazole gel-flakes in treatment of vaginal candidiasis: formulation, in vitro and clinical evaluation. Int J Pharm. 2019;567:118472. doi:10.1016/j.ijpharm.2019.118472
35. Al-Saryi N, Ibrahim SA, AL-Kadmy IM, Hetta HF. Whole genome sequencing of Streptococcus pneumoniae serotype 33C causing fatal sepsis in a hospitalized patient with nephrotic syndrome. Gene Reports. 2019;100434:e2774.
36. El-Baky RMA, Sandle T, John J, Abuo-Rahma GE-DA, Hetta HF. A novel mechanism of action of ketoconazole: inhibition of the NorA efflux pump system and biofilm formation in multidrug-resistant Staphylococcus aureus. Infect Drug Resist. 2019;12:1703–1718. doi:10.2147/IDR.S201124
37. Falagas ME, Kasiakou SK. Colistin: the revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin Infect Dis. 2005;40(9):1333–1341. doi:10.1086/429323
38. Jeannot K, Bolard A, Plesiat P. Resistance to polymyxins in Gram-negative organisms. Int J Antimicrob Agents. 2017;49(5):526–535. doi:10.1016/j.ijantimicag.2016.11.029
39. Aires CAM, da Conceicao-neto OC, Tavares EOTR, et al. Emergence of the plasmid-mediated mcr-1 gene in clinical KPC-2-producing klebsiella pneumoniae sequence Type 392 in Brazil. Antimicrob Agents Chemother. 2017;61(7). doi:10.1128/AAC.00317-17
40. Stojanoski V, Sankaran B, Prasad BVV, Poirel L, Nordmann P, Palzkill T. Structure of the catalytic domain of the colistin resistance enzyme MCR-1. BMC Biol. 2016;14(1):81. doi:10.1186/s12915-016-0303-0
41. Hinchliffe P, Yang QE, Portal E, et al. Insights into the mechanistic basis of plasmid-mediated colistin resistance from crystal structures of the catalytic domain of MCR-1. Sci Rep. 2017;7:39392. doi:10.1038/srep39392
42. Sun J, Xu Y, Gao R, et al. Deciphering MCR-2 colistin resistance. MBio. 2017;8(3):e00625–00617. doi:10.1128/mBio.00625-17
43. Marjani MFA, Mohammed NR, Abd SY, Mansour RF. Efflux pumps in colistin resistant pseudomonas aeruginosa isolates in Baghdad. Int J. 2015;3(11):680–685.
44. Carattoli A, Villa L, Feudi C, et al. Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Eurosurveillance. 2017;22(31). doi:10.2807/1560-7917.ES.2017.22.31.30589
45. Yin W, Li H, Shen Y, et al. Novel plasmid-mediated colistin resistance gene mcr-3 in Escherichia coli. MBio. 2017;8(3):e00543–00517. doi:10.1128/mBio.00543-17
46. Moffatt JH, Harper M, Harrison P, et al. Colistin resistance in Acinetobacter baumannii is mediated by complete loss of lipopolysaccharide production. Antimicrob Agents Chemother. 2010;54(12):4971–4977. doi:10.1128/AAC.00834-10
47. Nicas TI, Hancock RE. Outer membrane protein H1 of Pseudomonas aeruginosa: involvement in adaptive and mutational resistance to ethylenediaminetetraacetate, polymyxin B, and gentamicin. J Bacteriol. 1980;143(2):872–878. doi:10.1128/JB.143.2.872-878.1980
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.


