Back to Journals » Cancer Management and Research » Volume 12
Predicting Prostate Cancer Upgrading of Biopsy Gleason Grade Group at Radical Prostatectomy Using Machine Learning-Assisted Decision-Support Models
Authors Liu H, Tang K, Peng E, Wang L, Xia D, Chen Z
Received 13 October 2020
Accepted for publication 25 November 2020
Published 22 December 2020 Volume 2020:12 Pages 13099—13110
DOI https://doi.org/10.2147/CMAR.S286167
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eileen O'Reilly
Hailang Liu,1 Kun Tang,1 Ejun Peng,1 Liang Wang,2 Ding Xia,1 Zhiqiang Chen1
1Department of Urology, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, Hubei, People’s Republic of China; 2Department of Radiology, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, Hubei, People’s Republic of China
Correspondence: Ding Xia; Zhiqiang Chen
Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, No. 1095 Jiefang Avenue, Wuhan 430030, Hubei, People’s Republic of China
Email [email protected]; [email protected]
Objective: This study aimed to develop a machine learning (ML)-assisted model capable of accurately predicting the probability of biopsy Gleason grade group upgrading before making treatment decisions.
Methods: We retrospectively collected data from prostate cancer (PCa) patients. Four ML-assisted models were developed from 16 clinical features using logistic regression (LR), logistic regression optimized by least absolute shrinkage and selection operator (Lasso) regularization (Lasso-LR), random forest (RF), and support vector machine (SVM). The area under the curve (AUC) was applied to determine the model with the highest discrimination. Calibration plots and decision curve analysis (DCA) were performed to evaluate the calibration and clinical usefulness of each model.
Results: A total of 530 PCa patients were included in this study. The Lasso-LR model showed good discrimination with an AUC, accuracy, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) of 0.776, 0.712, 0.679, 0.745, 0.730, and 0.695, respectively, followed by SVM (AUC=0.740, 95% confidence interval [CI]=0.690– 0.790), LR (AUC=0.725, 95% CI=0.674– 0.776) and RF (AUC=0.666, 95% CI=0.618– 0.714). Validation of the model showed that the Lasso-LR model had the best discriminative power (AUC=0.735, 95% CI=0.656– 0.813), followed by SVM (AUC=0.723, 95% CI=0.644– 0.802), LR (AUC=0.697, 95% CI=0.615– 0.778) and RF (AUC=0.607, 95% CI=0.531– 0.684) in the testing dataset. Both the Lasso-LR and SVM models were well-calibrated. DCA plots demonstrated that the predictive models except RF were clinically useful.
Conclusion: The Lasso-LR model had good discrimination in the prediction of patients at high risk of harboring incorrect Gleason grade group assignment, and the use of this model may be greatly beneficial to urologists in treatment planning, patient selection, and the decision-making process for PCa patients.
Keywords: prostate cancer, biopsy cores, Gleason grade group, upgrading, machine learning
Background
Despite first being introduced in 1966, the Gleason score (GS) remains the most widely used grading system for prostate cancer (PCa).1 The GS grading system was updated in 2005 and 2014 and a new five-tier grade group (GG) system was proposed and developed: GG 1 (GS≤6), GG 2 (GS3+4=7), GG 3 (GS 4+3=7), GG 4 (GS 8), and GG 5 (GS 9 and 10).2,3 Appropriate clinical management for PCa patients depends on accurate risk stratification, which is mainly reliant on the pretreatment prostate-specific antigen (PSA) level, Gleason grade group (GG) of positive biopsy cores, and tumor stage. However, up to 56% of high-grade patients at initial biopsy tend to be overestimated, compared with prostatectomy specimens due to the sampling error of biopsy and the multifocal nature of PCa.3,4 In addition, some low risk patients who embark on active surveillance (AS) will be upgraded to higher grade at RP so they are not suitable candidates for AS.5,6 The discordance between initial biopsy GG and radical prostatectomy (RP) GG can potentially impact over- and under-treatment.7 Therefore, for the management of PCa, it is of pivotal importance to identify PCa patients at a higher risk of upgrading at RP before making treatment-related decisions.
Machine learning (ML) is the semi-automated extraction of knowledge and insight from data.8 Developed within the fields of statistics, computer science and artificial intelligence, it allows the training of algorithms that can discover and identify complex patterns and relationships faster than conventional statistical models that focus on only a handful of patient variables.8 Owing to the ability of ML algorithms to improve the accuracy of predicting diseases and subsequent outcomes over the use of traditional statistical models, they have been applied extensively in the field of clinical research.9,10 In the present study, we apply ML algorithms to the dataset, with the goal of identifying those patients at high risk of harboring upgrading at RP before making treatment decisions, and to determine the best predictive models.
Additionally, there is still no consensus on how to choose a “case level” biopsy GG (GS) for patients regarding the reporting of the “worst/highest” and “global/overall” GG (GS).7,11 Considering that the biopsy global GG is more likely be in line with RP GG, we selected global GG together with other preoperative clinical parameters to construct predictive models to calculate the probability of upgrading for each PCa patient.
Methods
Patient Selection and Study Parameters
Patients who underwent radical prostatectomy at Tongji Hospital of Tongji Medical College, Huazhong University of Science and Technology between January 2015 and December 2019 were retrospectively enrolled in this study. This retrospective study was approved by the Ethics Committee of Tongji Hospital, Huazhong University of Science and Technology. The need for informed consent from all patients was waived due to its retrospective nature. All patient information was strictly confidential and our procedures were carried out according to the Declaration of Helsinki. The inclusion criteria were as follows: 1) multi-parametric magnetic resonance imaging (mp-MRI) performed for all patients before surgery; 2) standard systematic (12-core) transrectal ultrasonography (TRUS)-guided biopsy prior to surgery performed for all patients; and 3) final pathological results of each patient including a detailed description of Gleason grade group. The following exclusion criteria were applied: 1) neoadjuvant therapy prior to MRI examination; 2) patients with incomplete clinical data; and 3) MRI images of unsatisfactory quality. The clinical parameters included patient age, total prostate-specific antigen (TPSA), prostate volume (PV=Height×Width×Length×0.52), PSA density (PSAD), %fPSA (free PSA/TPSA), maximum diameter of index lesion (D-max), the Prostate Imaging Reporting and Data System (PI-RADS) score, clinical and pathological T stage (T1-2, T3a, T3b and T4), apical involvement at MRI, global biopsy GG, number of positive cores, number of cores with clinically significant PCa (csPCa, defined as cores with GG≥2), maximum tumor length at biopsy core, percentage of tumor in total biopsy cores, and GG at RP. The MRI findings were re-reported and scored by the same dedicated radiologist on a five-point scale using the modified PI-RADS version 2 criteria.12 If multiple tumor foci existed on the MRI, only the highest PI-RADS score of index lesions and the maximum diameter of the largest tumor were included in the analysis. GG of the biopsy specimen was assigned following the 2014 International Society of Urological Pathology (ISUP) criteria.13 The global GG of the biopsy was defined as the most prevalent GG among all positive cores. The upgrades from biopsy to RP represented at least one grade difference in the GG.
Development, Validation, and Performance of ML-Based Models
The dataset was randomly split into two datasets: 70% for model training and 30% for model testing. For model training, data from the training set were used to approximate model parameters. Four ML algorithms were performed to build predictive models: logistic regression (LR), logistic regression optimized by least absolute shrinkage and selection operator (Lasso) regularization (Lasso-LR), random forest classifier (RF), and support vector machine (SVM) integrated with recursive feature elimination (RFE).
Model evaluation was carried out by examining discrimination and calibration. The receiver operating characteristic (ROC) curve analysis was used to evaluate the discrimination ability of predictive models in both the training dataset and testing dataset; the discrimination ability of each model was quantified by the area under the ROC curve (AUC). Moreover, discrimination metrics including accuracy, sensitivity, specificity, Youden index (YI), positive predictive value (PPV), and negative predictive value (NPV) were also applied to assess the discriminative power of predictive models. Comparisons between ROC Curves were performed using the method described by DeLong et al.14 As logistic regression analysis was one of the most widely used statistical methods in binary data classification, we used the LR model as the reference in the pairwise comparison of AUC. Calibration plots were used to investigate the extent of over- or under-estimation of predicted probabilities relative to the observed probabilities. Decision curve analysis (DCA) was conducted to determine the clinical net benefit associated with the use of the predictive models at different threshold probabilities in the patient cohort.
Statistical analyses were performed using R software (Version 3.6.0; https://www.R-project.org) with the following packages: “rms“, “glmnet”, “caret”, “rpart”, “randomForest”, “gplots“, “e1071“, “kernlab”, “pROC”,and “MachineShop”. P<0.05 was considered statistically significant.
Results
Baseline Characteristics and Pathological Results
Table 1 lists patient characteristics and pathological results in the total population, training dataset and testing dataset. The median patient age of the overall cohort was 69 (interquartile range [IQR]=63–75) years. The median TPSA value was 21.0 (IQR=10.9–42.2) ng/mL. The median D-max on MRI was 1.9 (IQR=1.3–2.7) cm. Most patients had clinical stage T1–2 (57.9%, n=307) and PI-RADS score 5 (58.6%, n=310). Table 2 details the concordance between the biopsy global GG and the final RP GG, and the corresponding downgrades and upgrades for GG 1–4. The most prevalent GGs assigned on biopsy were GG 1 (31.3%, n=166) and GG 2 (26.0%, n=138). Overall, 262 patients (49.4%) experienced upgrading at final pathology. The overall incidence of biopsy GG 1 upgrading was 120 (72.3%) of 166 patients, of which most were to GG 2 (50.0%, n=83), followed by GG 3 (12.7%, n=21), GG 4 (6.6%, n=11), and GG 5 (3.0%, n=5). Biopsy GG 3 (47.7%) and GG 4 (44.3%) showed the highest agreements when compared with RP GG. Patients with lower biopsy GG were more likely to harbor upgrading at RP.
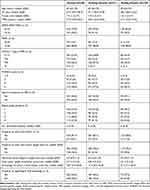 |
Table 1 Characteristics of Total Population, Training Dataset, and Testing Dataset |
 |
Table 2 Global Grade Groups on Biopsy and Radical Prostatectomy and Change in Grade |
ML-Assisted Models
In multivariable analysis, %fPSA (>0.16 vs ≤0.16) (OR=0.52; 95% CI=0.27–0.995; P=0.048), apical involvement (No vs Yes) (OR=1.80; 95% CI=1.02–3.19; P=0.042) on MRI and biopsy GG 1 (P<0.001) were significantly associated with upgrading at RP (Table 3). According to their respective coefficients, the LR model was constructed using the following formula: 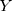 =2.29
=2.29 0.65
0.65 (%fPSA)
(%fPSA) 0.59
0.59 (apical involvement)
(apical involvement) 0.97
0.97 (biopsy GG) (where
(biopsy GG) (where 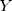 is the output value of predictive models).
is the output value of predictive models).
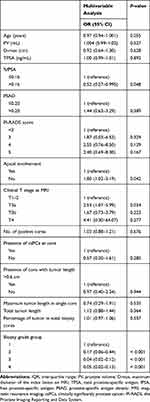 |
Table 3 Factors Associated with Upgrading on Multivariable Logistic Regression Analyses |
Based on the results of Lasso analysis, those clinical features with coefficients >0.1 were selected as the parameters included in the construction of the Lasso-LR model. Finally, %fPSA, apical involvement, PI-RADS score, clinical T stage, and biopsy GG were the selected features (Figure 1). The Lasso-LR was constructed by using the following formula: 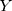 =1.81
=1.81 0.41
0.41 (%fPSA)
(%fPSA) 0.25
0.25 (apical involvement)
(apical involvement) 0.81
0.81 (biopsy GG)
(biopsy GG) 0.15
0.15 (clinical T stage)
(clinical T stage) 0.11
0.11 (PI-RADS score) (where
(PI-RADS score) (where 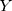 is the output value of predictive model).
is the output value of predictive model).
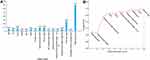
In the RFE-SVM analysis, 10 clinical parameters were selected as the final candidates for constructing the predictive model without impacting the prediction accuracy of the model, including biopsy GG, apical involvement, maximum tumor length in single core, %fPSA, PSAD, presence of core with tumor length >0.6 cm, presence of csPCa at core, PI-RADS score, D-max, and clinical T stage (Figure 2A). As depicted in Figure 2B, with the selected features being added to the SVM model one by one, the AUC value of model also increased little by little.
The process of feature selection by RF model and the importance of features are illustrated in Figure 3. Based on different combinations of clinical parameters, each tree in the forest votes for the major classification, and the final classification of the RF model is derived from the majority of these votes (Figure 3A). The best number of trees and the best number of variables tried at each split were 131 and 4, respectively. The out of bag (OOB) estimate of error rate was 33.42%, suggesting that the generalization error was quite unsatisfactory.
Comparison Between ML-Based Models
Among these models, the Lasso-LR model had the highest AUC (0.776, 95% confidence interval [CI]=0.729–0.822), followed by SVM (AUC=0.740, 95% CI=0.690–0.790), LR (AUC=0.725, 95% CI=0.674–0.776) and RF (AUC=0.666, 95% CI=0.618–0.714) (Figure 4A). Similarly, in the testing dataset, the Lasso-LR model had the highest AUC (0.735, 95% CI=0.656–0.813), followed by SVM (AUC=0.723, 95% CI=0.644–0.802), LR (AUC=0.697, 95% CI=0.615–0.778), and RF (AUC=0.607, 95% CI=0.531–0.684) (Figure 4B). The Lasso-LR model illustrated an accuracy of 0.712, a sensitivity of 0.679, and a specificity of 0.745, indicating that this model correctly identified 67.9% of PCa patients who experienced upgrading at RP and 74.5% of PCa patients who did not experience upgrading at RP (Table 4). In addition, the Lasso-LR model had the highest YI (0.424) compared with other models. Due to the fact that the YI was calculated as a summation of the sensitivity and specificity minus 1, the highest YI indicated that both the sensitivity and specificity of the Lasso-LR model are reasonably good relative to other models. Pairwise comparison of ROC curves showed that the AUC of the Lasso-LR model was significantly higher than that of LR (P=0.002), while the AUCs of SVM and RF were not significantly different to that of LR (P>0.05) (Figure 4A).
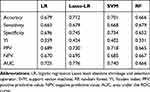 |
Table 4 Discrimination of Prediction Models |
The calibration of ML-based models was evaluated graphically by the formulation of calibration curves (Figure 5). The green line represented the fit of the model. Deviations from the 45° line indicated miscalibration. Part of the green line below the 45° line indicated that higher predicted probabilities might overestimate the true outcome, and part of the green line upon the 45° line indicated that lower predicted probabilities might under-predict the true probability of upgrading. The SVM model was well-calibrated (Figure 5D), followed by Lasso-LR (Figure 5B), RF (Figure 5C), and LR (Figure 5A). Moreover, the bootstrapped DCA also suggested higher net benefits of the three predictive models with threshold probabilities between 30% and 78% for LR, threshold probabilities between 50% and 75% for SVM and threshold probabilities between 30% and 90% for Lasso-LR (Figure 6).
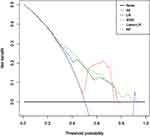 |
Figure 6 Decision curve analysis for ML-based models. Abbreviations: LR, logistic regression, Lasso least absolute shrinkage and selection operator; SVM, support vector machine; RF, random forest. |
Discussion
Accurate risk stratification, which mainly depends on PSA level, biopsy grade group, and stage classification, plays a pivotal role in guiding treatment management for PCa patients. However, it has been demonstrated that prostate biopsy often underestimates the cancer and incorrectly assigns National Comprehensive Cancer Network (NCCN) risk stratification.3,4 There are several reasons accounting for the discrepancies between the biopsy and RP grades: sampled cores and indications for biopsy, differences in biopsy techniques, erroneous diagnostic interpretation, tumor heterogeneity, sampling error on biopsy, clinician interpretation of the biopsy GG (global vs highest/composite/overall score), and practice variations regarding biopsy grade assignment.7,16 The incorrect risk stratification may impact treatment planning, patient selection, and decision-making processes. Therefore, it is extremely important to identify risk factors associated with upgrading to avoid under-treatment, especially among those PCa patients who are considered appropriate candidates for AS. Unfortunately, there are currently no widely accepted predictive models to accurately predict the final individualized GG at RP and the discrimination ability of various models remains modest.17–20 ML has been previously used for predicting outcomes in other fields of medicine, including the identification of lung cancer based on routine blood indices and the in-hospital rupture of type A aortic dissection.21,22 Given the excellent performance of machine learning algorithms in classification, four machine learning algorithms were employed in our study to determine relevant risk factors; we then developed and validated four novel prediction models to identify those PCa patients at high risk of harboring upgrading at RP before making treatment decisions.
Overall, up to 49.4% included patients were upgraded at RP, especially biopsy GG 1 patients, with the proportion of upgrading being 72.3%. Similarly, Altok et al23 reported that 70.9% of biopsy GG 1 patients in their study cohort were upgraded at RP, and most were upgraded to GG 2. These observations explain why some patients with GG 1 disease at biopsy suffer metastases or die of prostate cancer and suggest that a substantial proportion of biopsy GG 1 patients who embark on active surveillance are not, in fact, suitable candidates.24 Given the known risk of underestimation in biopsy specimens, the prediction of GG upgrade plays a major role when considering individualized therapy for PCa patients, especially AS.25 In our series, %fPSA (>0.16 vs ≤0.16), apical involvement at MRI (No vs Yes), and biopsy grade group (GG 4, GG 3, GG 2 vs GG 1) were independent factors in multivariable LR analysis. %fPSA, apical involvement at MRI, biopsy grade group, and clinical T stage at MRI were significantly associated with upgrading in Lasso-LR and SVM models. However, in a comparable study, Alshak et al15 demonstrated that only the PI-RADS score was a significant predictor of upgrading. Besides, Gandaglia et al26 reported that preoperative PSA level, GG at MRI-targeted biopsy, and clinically significant PCa at systematic biopsy were independent risk factors of upgrading at RP. The differences in results between our study and the latter two studies might be due to the fact that the latter two studies did not include detailed core biopsy information, which has been successfully shown to contain huge potential predictive value.
In our study, imaging factors such as apical involvement at MRI and clinical T stage at MRI, were more important predictors than clinical parameters according to the results of ML-based feature ranking analyses, except for RF analysis. This implied that mp-MRI had great potential in predicting upgrading, irrespective of its important role in detecting csPCa and assigning accurate risk stratification for PCa patients. The routine mp-MRI examination for patients with suspected PCa before biopsy was indeed beneficial and helpful.27 Among those biopsy-related variables, biopsy GG was always the strongest predictor. In the LR and Lasso-LR model, the number of positive cores, presence of csPCa at core, presence of a core with a tumor length >0.6 cm, maximum tumor length in a single core, total tumor length, and percentage of tumor in total biopsy cores demonstrated almost no value in the prediction of upgrading at RP, while the number of positive cores, total tumor length and percentage of tumor in total biopsy cores ranked ahead in the RF model. This is not in line with the findings of Pepe et al.28 As these features, including D-max, were reliable proxies of tumor volume, the size of tumor should not be considered relevant to the presence of upgrading.29 Nonetheless, Corcoran et al16 reported that tumor volume of PCa was a significant predictor of upgrading in multivariable analysis, and the measurement of surrogate of tumor volume might predict those at greatest risk of Gleason score upgrade. One thing to be noted was that the patient cohort in the study of Corcoran et al16 did not include those patients with biopsy GG 3 and 4. %fPSA outperformed TPSA and PSAD in the prediction of upgrading in the LR, Lasso-LR, and SVM models. On the contrary, in the mean decrease accuracy and mean decrease Gini evaluation of RF models, TPSA and PSAD ranked higher than %fPSA. Apart from those variables included in our models, Ferro et al30–32 have successfully demonstrated that the serum total testosterone level and biomarkers including prostate cancer antigen 3, prostate health index, and sarcosine play an important role in predicting GG upgrading. Regrettably, our center has not yet launched these tests in the management of PCa.
For the performance of ML-based models, the Lasso-LR model showed the best discriminative power with an AUC of 0.776 (95% CI=0.729–0.822), followed by SVM (AUC=0.740; 95% CI=0.690–0.790), LR (AUC=0.725, 95% CI=0.674–0.776), and RF (AUC=0.666; 95% CI=0.618–0.714). The nomogram developed by He et al33 achieved an AUC of 0.753 in the prediction of upgrading, which was higher than that of LR but lower than Lasso-LR in the present study. Also, Moussa et al34 constructed a nomogram for predicting the possibility of upgrading, with a concordance index of 0.68. Additionally, all of the ML-based models except for RF outperformed the predictive models constructed by Kulkarni et al35 and Athanazio et al7, with AUC values of 0.71 and 0.699 in the respective studies. Of note, in a study consisting of 2,982 PCa patients treated with RP, the model for predicting upgrading based on LR analysis showed a predictive accuracy of 0.804; in contrast, in our study, the Lasso-LR model presented the best predictive accuracy of 0.712.19 Despite the better predictive accuracy of the model in the study of Chun et al,19 it was still difficult to determine the model with best performance when compared with our ML-based models as there was no other discrimination metrics such as AUC, sensitivity, specificity, PPV, and NPV in their study. It should be noted that the good performance of our ML-based models might be related to the inclusion of mp-MRI information and detailed biopsy information.
Limitations
Despite several strengths, our study has certain limitations. First, the data on PCa patients who underwent RP enrolled in our study cohort were retrospectively collected at a single institution, which may have resulted in selection bias. Second, the case-level highest Gleason grade group was more commonly assigned to patients undergoing systematic TRUS-guided biopsy in our country; hence, we should also construct predictive models to identify risk factors associated with upgrading using a comparison between the highest biopsy GG and final RP samples. Moreover, we did not include the radiomic features of PCa in the construction of prediction models. Considering that the radiomic features play an important role in the detection of PCa, the inclusion of radiomic features may significantly improve the performance of models.
Conclusions
In summary, we developed four ML-based models to help clinicians identify the individualized risk of upgrading for PCa patients after prostate needle biopsy. The Lasso-LR model had the best discriminative power according to the results of pairwise comparisons. We believe that our research findings can make a significant difference in the process of treatment decision-making by more accurately identifying patients at high risk of harboring upgrading at RP. Of course, further validation in multiple institutions with a large sample size is warranted.
Abbreviations
AS, Active surveillance; AUC, Area under the ROC curve; CI, Confidence interval; csPCa, Clinically significantly prostate cancer; D-max, Maximum diameter of the index lesion on MRI; fPSA, Free prostate-specific antigen; GG, Gleason grade group; IQR, Interquartile range; Lasso, least absolute shrinkage and selection operator; LR, Logistic regression; ML, Machine learning; mp-MRI, Multi-parametric MRI; MRI, Magnetic resonance imaging; NCCN, National Comprehensive Cancer Network; NPV, Negative predictive value; OOB, Out of bag; PCa, Prostate cancer; PI-RADS, The Prostate Imaging Reporting and Data System; PPV, Positive predictive value; PSA, Prostate-specific antigen; PSAD, Prostate-specific antigen density; PV, Prostate volume; RF, Random forest; RFE, recursive feature elimination; SVM, Super vector machine; TPSA, Total prostate-specific antigen; TRUS, Transrectal ultrasonography; YI, Youden index.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest for this work.
References
1. Gleason DF, Mellinger GT. Prediction of prognosis for prostatic adenocarcinoma by combined histological grading and clinical staging. J Urol. 1974;111(1):58–64. doi:10.1016/s0022-5347(17)59889-4
2. Epstein JI, Allsbrook WC, Amin MB, Egevad LL; ISUP Grading Committee. The 2014 International Society of Urological Pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma: definition of grading patterns and proposal for a new grading system. Am J Surg Pathol. 2016;40(2):244–252. doi:10.1097/PAS.0000000000000530
3. Alchin DR, Murphy D, Lawrentschuk N. Risk factors for Gleason score upgrading following radical prostatectomy. Minerva Urol Nefrol. 2017;69(5):459–465. doi:10.23736/S0393-2249.16.02684-9
4. Müntener M, Epstein JI, Hernandez DJ, et al. Prognostic significance of Gleason score discrepancies between needle biopsy and radical prostatectomy. Eur Urol. 2008;53(4):767–775. doi:10.1016/j.eururo.2007.11.016
5. Hamdy FC, Donovan JL, Lane JA, et al. 10-year outcomes after monitoring, surgery, or radiotherapy for localized prostate cancer. N Engl J Med. 2016;375(15):1415–1424. doi:10.1056/NEJMoa1606220
6. Qi F, Zhu K, Cheng Y, et al. How to pick out the “Unreal” Gleason 3 + 3 patients: a nomogram for more precise active surveillance protocol in low-risk prostate cancer in a Chinese population. J Invest Surg. 2019;6:1–8. doi:10.1080/08941939.2019.1669745
7. Athanazio D, Gotto G, Shea-Budgell M, et al. Global Gleason grade groups in prostate cancer: concordance of biopsy and radical prostatectomy grades and predictors of upgrade and downgrade. Histopathology. 2017;70(7):1098–1106. doi:10.1111/his.13179
8. Wong NC, Lam C, Patterson L, et al. Use of machine learning to predict early biochemical recurrence after robot-assisted prostatectomy. BJU Int. 2018;123(1):51–57. doi:10.1111/bju.14477
9. Chen JH, Asch SM. Machine learning and prediction in medicine - beyond the peak of inflated expectations. N Engl J Med. 2017;376:2507–2509. doi:10.1056/NEJMra1814259
10. Rajkomar A, Dean J, Kohane I. Machine learning in medicine. N Engl J Med. 2019;380:1347–1358. doi:10.1056/NEJMra1814259
11. Trpkov K, Sangkhamanon S, Yilmaz A, et al. Concordance of “case level” global, highest, and largest volume cancer grade group on needle biopsy versus grade group on radical prostatectomy. Am J Surg Pathol. 2018;42(11):1522–1529. doi:10.1097/PAS.0000000000001137
12. Weinreb JC, Barentsz JO, Choyke PL, et al. PI-RADS prostate imaging - reporting and data system:2015, version 2. Eur Urol. 2016;69(1):16–40. doi:10.1016/j.eururo.2015.08.052
13. Epstein JI, Zelefsky MJ, Sjoberg DD, et al. A contemporary prostate cancer grading system: a validated alternative to the Gleason score. Eur Urol. 2016;69(3):428–435. doi:10.1016/j.eururo.2015.06.046
14. DeLong ER, DeLong DM, Clarke-Pearson DL. Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics. 1988;44(3):837–845. doi:10.2307/2531595
15. Alshak MN, Patel N, Gross MD, et al. Persistent discordance in grade, stage, and NCCN risk stratification in men undergoing targeted biopsy and radical prostatectomy. Urology. 2020;135:117–123. doi:10.1016/j.urology.2019.07.049
16. Corcoran NM, Hovens CM, Hong MK, et al. Underestimation of Gleason score at prostate biopsy reflects sampling error in lower volume tumours. BJU Int. 2012;109(5):660–664. doi:10.1111/j.1464-410X.2011.10543.x
17. Kuroiwa K, Shiraishi T, Naito S. Gleason score correlation between biopsy and prostatectomy specimens and prediction of high-grade Gleason patterns: significance of central pathologic review. Urology. 2011;77(2):407–411. doi:10.1016/j.urology.2010.05.030
18. Epstein JI, Feng Z, Trock BJ, et al. Upgrading and downgrading of prostate cancer from biopsy to radical prostatectomy: incidence and predictive factors using the modified Gleason grading system and factoring in tertiary grades. Eur Urol. 2012;61(5):1019–1024. doi:10.1016/j.eururo.2012.01.050
19. Chun FK, Steuber T, Erbersdobler A, et al. Development and internal validation of a nomogram predicting the probability of prostate cancer Gleason sum upgrading between biopsy and radical prostatectomy pathology. Eur Urol. 2006;49(5):820–826. doi:10.1016/j.eururo.2005.11.007
20. Thomas C, Pfirrmann K, Pieles F, et al. Predictors for clinically relevant Gleason score upgrade in patients undergoing radical prostatectomy. BJU Int. 2012;109(2):214–219. doi:10.1111/j.1464-410X.2011.10187.x
21. Wu J, Qiu J, Xie E, et al. Predicting in-hospital rupture of type A aortic dissection using random forest. J Thorac Dis. 2019;11(11):4634–4646. doi:10.21037/jtd.2019.10.82
22. Wu J, Zan X, Gao L, et al. A machine learning method for identifying lung cancer based on routine blood indices: qualitative feasibility study. JMIR Med Inform. 2019;7(3):e13476. doi:10.2196/13476
23. Altok M, Troncoso P, Achim MF, et al. Prostate cancer upgrading or downgrading of biopsy Gleason scores at radical prostatectomy: prediction of “regression to the mean” using routine clinical features with correlating biochemical relapse rates. Asian J Androl. 2019;21(6):598–604. doi:10.4103/aja.aja_29_19
24. Yang DD, Mahal BA, Muralidhar V, et al. Pathologic outcomes of Gleason 6 favorable intermediate-risk prostate cancer treated with radical prostatectomy: implications for active surveillance. Clin Genitourin Cancer. 2018;16(3):226–234. doi:10.1016/j.clgc.2017.10.013
25. Morlacco A, Cheville JC, Rangel LJ, et al. Adverse disease features in Gleason score 3 + 4 “favorable intermediate-risk” prostate cancer: implications for active surveillance. Eur Urol. 2017;72(3):442–447. doi:10.1016/j.eururo.2016.08.043
26. Gandaglia G, Ploussard G, Valerio M, et al. The key combined value of multiparametric magnetic resonance imaging, and magnetic resonance imaging-targeted and concomitant systematic biopsies for the prediction of adverse pathological features in prostate cancer patients undergoing radical prostatectomy. Eur Urol. 2020;77(6):733–741. doi:10.1016/j.eururo.2019.09.005
27. Pepe P, Garufi A, Priolo GD, et al. Is it time to perform only magnetic resonance imaging targeted cores? Our experience with 1032 men who underwent prostate biopsy. J Urol. 2018;200(4):774–778. doi:10.1016/j.juro.2018.04.061
28. Pepe P, Fraggetta F, Galia A, et al. Is quantitative histologic examination useful to predict nonorgan-confined prostate cancer when saturation biopsy is performed? Urology. 2008;72(6):1198–1202. doi:10.1016/j.urology.2008.05.045
29. Gandaglia G, Ploussard G, Valerio M, et al. A novel nomogram to identify candidates for extended pelvic lymph node dissection among patients with clinically localized prostate cancer diagnosed with magnetic resonance imaging-targeted and systematic biopsies. Eur Urol. 2019;75(3):506–514. doi:10.1016/j.eururo.2018.10.012
30. Ferro M, Lucarelli G, Bruzzese D, et al. Low serum total testosterone level as a predictor of upstaging and upgrading in low-risk prostate cancer patients meeting the inclusion criteria for active surveillance. Oncotarget. 2017;8(11):18424–18434. doi:10.18632/oncotarget.12906
31. Ferro M, Lucarelli G, Cobelli O, et al. Circulating preoperative testosterone level predicts unfavorable disease at radical prostatectomy in men with International Society of Urological Pathology Grade Group 1 prostate cancer diagnosed with systematic biopsies. World J Urol. 2020. doi:10.1007/s00345-020-03368-9
32. Ferro M, Lucarelli G, Bruzzese D, et al. Improving the prediction of pathologic outcomes in patients undergoing radical prostatectomy: the value of prostate cancer antigen 3 (PCA3), prostate health index (phi) and sarcosine. Anticancer Res. 2015;35(2):1017–1023.
33. He B, Chen R, Gao X, et al. Nomograms for predicting Gleason upgrading in a contemporary Chinese cohort receiving radical prostatectomy after extended prostate biopsy: development and internal validation. Oncotarget. 2016;7(13):17275–17285. doi:10.18632/oncotarget.7787
34. Moussa AS, Kattan MW, Berglund R, et al. A nomogram for predicting upgrading in patients with low- and intermediate-grade prostate cancer in the era of extended prostate sampling. BJU Int. 2010;105(3):352–358. doi:10.1111/j.1464-410X.2009.08778.x
35. Kulkarni GS, Lockwood G, Evans A, et al. Clinical predictors of Gleason score upgrading: implications for patients considering watchful waiting, active surveillance, or brachytherapy. Cancer. 2007;109(12):2432–2438. doi:10.1002/cncr.22712
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.