Back to Journals » Infection and Drug Resistance » Volume 13
Molecular Mechanisms of Salmonella Effector Proteins: A Comprehensive Review
Authors Azimi T ![]() , Zamirnasta M, Sani MA, Soltan Dallal MM, Nasser A
, Zamirnasta M, Sani MA, Soltan Dallal MM, Nasser A
Received 11 September 2019
Accepted for publication 20 December 2019
Published 6 January 2020 Volume 2020:13 Pages 11—26
DOI https://doi.org/10.2147/IDR.S230604
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Taher Azimi,1,2 Maryam Zamirnasta,3 Mahmood Alizadeh Sani,4,5 Mohammad Mehdi Soltan Dallal,6 Ahmad Nasser3,6,7
1Pediatric Infections Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran; 2Students Scientific Research Center, Tehran University of Medical Sciences, Tehran, Iran; 3Clinical Microbiology Research Center, Ilam University of Medical Science, Ilam, Iran; 4Food Safety and Hygiene Division, Environmental health Department, School of Public Health, Tehran University of medical sciences, Tehran, Iran; 5Students Research Committee, Department of Food Sciences and Technology, Faculty of Nutrition and Food Sciences, Tabriz University of Medical Sciences, Tabriz, Iran; 6Department of Pathobiology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran; 7Department of Medical Microbiology, School of Medicine, Ilam University of Medical Science, Ilam, Iran
Correspondence: Ahmad Nasser
Clinical Microbiology Research Center, Ilam University of Medical Science, P.O. Box 14473-13324, Ilam, Iran
Tel +98 9166438900
Email [email protected]
Abstract: Salmonella can be categorized into many serotypes, which are specific to known hosts or broadhosts. It makes no difference which one of the serotypes would penetrate the gastrointestinal tract because they all face similar obstacles such as mucus and microbiome. However, following their penetration, some species remain in the gastrointestinal tract; yet, others spread to another organ like gallbladder. Salmonella is required to alter the immune response to sustain its intracellular life. Changing the host response requires particular effector proteins and vehicles to translocate them. To this end, a categorized gene called Salmonella pathogenicity island (SPI) was developed; genes like Salmonella pathogenicity island encode aggressive or modulating proteins. Initially, Salmonella needs to be attached and stabilized via adhesin factor, without which no further steps can be taken. In this review, an attempt has been made to elaborate on each factor attached to the host cell or to modulating and aggressive proteins that evade immune systems. This review includes four sections: (A) attachment factors or T3SS- independent entrance, (B) effector proteins or T3SS-dependent entrance, (c) regulation of invasive genes, and (D) regulation of immune responses.
Keywords: Salmonella, effector proteins, immune response, T3SS, pathogenesis
Introduction
Nearly all pathogens that attack the gastrointestinal tract spring from food. Pathogenic bacteria such as Salmonella spp., Campylobacter spp., Yersinia entrolitica, Shigella spp., and Enterotoxigenic Escherichia coli can invade the gastrointestinal lumen and cause diarrhea and other damage.1,2 One of the most important bacteria that penetrates the lumen out of different materials, such as dairy, vegetable, egg, etc., is Salmonella spp.3 Salmonella may cause death all around the world.4 Some species such as5 S. Typhi specifically show inclination towards humans and cause a higher rate of mortality; however, other species such as6 S. enteritidis cause self-limiting diarrhea; of note, the latter can be just as deadly as the former. Further, statistics have shown a quarter of mortality rates associated with the former type. However, all species should overcome a number of barriers, such as stomach and mucus, and evade an immune cell. Pathogenic Salmonella has a particular factor that differs from the non-pathogenic ones such as Type-3 Secretion System (T3SS) and Salmonella pathogenicity island (SPI).7,8 Interestingly, Salmonella has a two-cluster distinct T3SS, which is encoded by SPI-1 and SPI-2. Almost all effectors of SPI-1 and SPI-2 mediate cell invasion and intracellular survival, respectively.9,10 Having passed through stomach via food, Salmonella penetrates the intestine and causes enteritis. To this end, Salmonella needs to be attached to the host cell and cross the intestinal membrane via M cell or dendritic cell (DC).11 Following the attachment, (Part A) T3SS-independent entrance approach can be adopted by SiiE, RcK, PagN, and ShdA, or (Part B) T3SS-dependent entrance can be adopted by SipA, SipC, SopB, and SopE. The virulence factor is activated to modulate host cell life for the benefit of the striker. Regulation between activation of adhesion and virulence factor needs to be adjusted and activated (Part C) at a right moment. To ensure maximum coordination, this pathogenic gene is clustered into one genomic island. Finally, the immune response (Part D) is activated, and necessary actions are taken to put an end to this adventure.
Attachment Factors (T3SS-Independent Entrance)
Adhesin Proteins
SiiE
For Salmonella infection or invasion to occur, the first pathogen should reside in the site of infection. SiiE is a non-fimbrial adhesin of Salmonella that can be attached to the epithelial cell.12 This effector is transferred through T1SS and encoded by SPI-4. T1SS system is formed by three subunits: SiiF as an inner membrane and ATPase, SiiD as a transmembrane unit, and SiiC as an outer membrane protein.13 SPI-4 and T1SS, as well as its substrate SiiE, are required only to invade the polarized cell.14 HilA regulates the transcription of SPI-4 by a master regulator, SirA.15 The signal sequence of SiiE is located at terminal C and has a long linear structure to cross the LPS structure.16
Biofilm Association Protein (BapA)
Biofilm-associated protein (Bap) has a major role in the production of biofilm composed of cellulose and curli fimbriae. Bap secretes through T1SS and resides on the bacterial surface.17 Both components are under the regulation of CsgD regulator. CsgD activates csgBAC operon to produce curli pili.18 Active production of Bap is also regulated by CsgD regulator.17 As a curli fimbriae operon, CsgA can be up-regulated in many ways in gallstone.19
Resistance to Complement Killing (Rck)
The outer membrane protein, Rck, has a major role in invading the host cell. Rck generates a zipper-like structure by stimulating Cdc42, and Rac1 may produce actin formation.20 Furthermore, Rck can mediate complement resistance by inhibiting polymerization of C9 on the bacterial surface.21 Rck is attached to the Epidermal Growth Factor Receptor (EGFR) directly, and the attachment site differs from EGF, leading to the auto phosphorylation of the EGFR cytosolic tail.22 This phosphorylation leads to a signaling cascade and the activation of Src, which is a signaling molecule that finally causes bacterial internalization.23 Rck binds to the extracellular matrix (ECM). For more information, see Table 1.
 |
Table 1 Specifications of Salmonella Effector Proteins and Their Mechanisms |
PhoP Activated Gene N (PagN)
PagN is another outer membrane protein that interacts with the eukaryotic epithelial cell via haemagglutination property. This protein plays a role in adhesion to and invasion of the host cell. Heparan sulfate on the host cell acts as a receptor for this ligand.24,25 This gene is induced under cation-limited circumstances and in a PhoP-dependent manner.26 This condition may occur inside a macrophage.27 In the presence of cation, PhoQ inhibits the activation and expression of PhoP subset gene.27
Salmonella Typhi Invasion (STIV)
The next outer membrane protein involved in pathogenesis and invasion is STIV. Through the extracellular loop, STIV remains attached to the Met, the tyrosine kinase of the intestinal host cell.28 After attachment, the activation of Rac1 and Src via Met phosphorylation leads to actin polymerization and bacterial engulfment. This protein is required for systemic spread and intestinal colonization in S. Typhi.29 This protein can independently act in invasion and pathogenesis, which may encourage synergism with T3SS (Figure 1).29
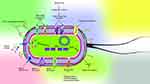 |
Figure 1 Interaction between Salmonella attachment factors (T3SS-independent entrance) and host proteins. |
Another Adhesin Factors
MisL is a protein of membrane insertion and secretion and is expressed in an outer membrane protein, which is encoded by SPI3, and binds to fibronectin. This attachment helps Salmonella invade epithelial cell and intestinal persistence.30 ShdA is also translocated via an auto-transporter system to the outer membrane and binds to fibronectin.31 The tip of flagella has a major role in the attachment of bacteria to the eukaryotic cell. Furthermore, flagella can actively cause bacteria to move towards a favorable attachment place.32,33 OmpD can interestingly be attached to the epithelial cell, and OmpD− mutant has mitigated affinity with the host epithelial cell.34
Virulence Factor of Salmonella
Salmonella Pathogenesis Island (SPI)
SPI encoder of T3SS directly translocates effector protein inside the host cell. SPI can be found in chromosome or in plasmid and its G+C differs from that in the surrounding regions. SPI is often associated with a mobile genetic element such as phage, insertion sequence (IS), and transposon.35 SPI-1 encodes SipA and SipB and, in a similar fashion, SPI-2 encodes SseF, Srf, and SseG; SPI-3 encodes MisL; SPI-4 encodes SiiE; SPI-5 encodes SopB; SPI-6 encodes Tae4.36 One of the important proteins is called SopB (Salmonella outer protein B) that has a major role in the secretion system and the recurrence of neutrophil.37 Further to that, SopB interacts with chloride channel, affects ion balance in the host cell, and manages Salmonella-containing vacuole to inhibit lysosome vacuole fusion.38 After translocation, SopB activates Rho GTPase in the host cell. Furthermore, SopB has a lipid phosphatase activity and hydrolyzes phosphatidylinositol biphosphate that causes failure in Na+/H+ activity exchange and diarrhea.39 Phosphatase activity can reduce phosphatidylinositol biphosphate PI (3, 4) and induce Akt (Protein kinase B that regulates cell survival) to ensure the best bacterial growth inside the cell and inhibit apoptosis.40–42 The reduction of the phosphatidylinositol biphosphate leads to a decrease in negative charge on the surface of Salmonella-containing vacuole (SCV), resulting in the prevention of lysosomal enzyme fusion.43 After the invasion and lipid activity of SopB, RhoH, RhoD, RhoB, and RhoJ recurred within the location of Salmonella invasion. RhoB and RhoH activate Akt, and Rhoj has a role in the invasion of Salmonella; finally, RhoD has a role in membrane trafficking and actin reorganization.44 SopB can also subvert signaling in the host cell and cause actin rearrangement in membrane cell.44,45 On the other hand, SopB increases the possibility of the return of Rab5 to the SCV and, thus, causes the aggregation of phosphatidylinositide-3- phosphate on SCV.46 Tae4 encoded by SPI-6 can help Salmonella overcome colonization resistance. By inducing bacterial lysis, Tae4 (an antimicrobial amidase) can confer a benefit to the Salmonella in the gut lumen establishment.47 Another effector in Sop family is SopC that mediates invasion, neutrophil recruitment, and fluid secretion. This effector is found in S. dublin and some strains of S. typhimurium.48
Vi Antigen (Capsule)
One of the important factors present in specific Salmonella such as S. Typhi is a polysaccharide capsule, called Vi antigen. Because of polysaccharide cover, the O polysaccharide may explain the resistance of bacteria to a specific antibody against O polysaccharide. Vi inhibits the deposition of C3b in the bacterial cell.49 Another noteworthy point to consider is that natural IgM secreted by B-cell cannot be attached to the bacterial surface, which is an important step for phagocytosis and ROS as a mediator of neutrophil killing.50 Vi antigen is encoded by SPI-7, which also encodes the T3SS effector SopE and pilus type IVb.51–53 Vi is encoded in viaB locus and TviA is a positive regulator for this locus. TviA can downregulate SPI-1 and flagellar genes to stop the invasion and motility process. TviA can repress flagellar regulator flhDC, the master regulator (which finally activated promotors),54 and alter the expression of T3SS and Vi in response to osmolarity. In fact, HilA controls the expression of T3SS, and TivA can repress the expression of HilA.55 On the other hand, SPI-1 and flagella are expressed to facilitate the Salmonella invasion,56 because Vi mask of the cell surface can reduce inflammatory cytokine-like IL-8 by attaching it to the prohibitin molecule on the surface of the intestinal cell.57 Toll-like receptors (TLR) can recognize specific molecules on the pathogens. After identifying the pathogen via TLR, chemoattractants such as IL-8 are secreted. Furthermore, Vi antigen (masking) can reduce IL-8 production in a TLR-dependent manner.58,59
Lipopolysaccharide (LPS)
One of the important factors on the surface of Salmonella is LPS. This factor has an important role in intestinal colonization, macrophage resistance, and modulation of the humoral immune response.60,61 LPS consists of three parts: Lipid A as an inner part, oligosaccharide as a middle part, and O antigen as a variable-length outer part.62 TLR4-MD2-CD14 can recognize the inner part of LPS, Lipid A; however, O antigen stands against humoral immune response.63 Furthermore, LPS inhibits the penetration of lipophilic antibiotic such as macrolide.64 However, if O antigen is too long or is modified via molecules that mimic the host cell, such a sialic acid modulates the immune system.65 Factors that affect elongated O antigen production include temperature and iron. Data have shown that iron limitation in medium causes a change in the elongation of LPS. A decrease in temperature can change phosphate and amine in an LPS structure that protects Salmonella from bacteriophage.66,67 S. paratyphi A expressed O2 antigen with a very long O-antigen chain that prevents both antibody binding and Neutrophil respiratory burst.68 The model of LPS is presented in Salmonella, where O12 is the backbone in all three types, O2 in S. Paratyphi A, and O9 or O4 in S. Typhi and S. typhimurium. The rfbE gene connects cytidine diphosphate-paratose to the cytidine diphosphate-tyvelose. In the case of S. Paratyphi A, rfbE is a pseudogene and is the reason why paratose remains (very long O2 antigen) on the bacterial surface.69,70 Therefore, non-opsonic paratose in a very long O2 helps S. Paratyphi A remain unrecognizable to human IgM.
Salmonella-Containing Vacuole (SCV)
SPI-2 encodes different types of secretion system (Type 3) that are important for maintaining the intracellular life of Salmonella.71 The effector protein may mimic the structure of the main cellular proteins to interact with modulator enzyme.72 To internalize the first vesicle, budding should be formed and, then, transferred to the inside of the cell; finally, its tethering and fusion take place. In brief, the membrane proximal coat binds to the specific membrane-associated GTPase; then, SNAREs and transmembrane cargo protein joined together in place of assembling coat.73 Finally, following the transfer of vesicle, uncoating occurs through the inactivation of GTPase and vesicle merges with the acceptor compartment by means of Rab5.74 Following the internalization of Salmonella into the host cell, bacteria engulf the vacuole by an early marker, which is to be later replaced by lysosomal-associated protein.75 In other words, at the first hour, EEA1 (early endosomal antigen 1) and Rab5 are present on the SCV; however, after this time, they are replaced with LAMP (lysosome-associated membrane proteins).76 Regulator Rab protein mediates transport and fusion of vesicle between the receptor and donor component.77 Phosphatidylinositol-3-phosphate develops an interaction between EEA1 and Rab5 on the SCV.78 Salmonella-induced filaments (SIF) are membrane tubules in the eukaryotic cell that can provide Salmonella with access to nutrition.79 Rab5 by means of phosphatidylinositol-3-phosphate facilitates the attachment of EEA1 to SCV, which, in turn, produces fusion with endosome.80 After an hour, an exchange of the marker occurs by replacing Rab5 by Rab7 and lysosomal-associated membrane protein (LAMP) on the SCV surface.81 LAMP and Rab7 are centralized to SIF and, in this stage, SIFs are free of hydrolysis enzyme.82 In fact, Rab7 causes the recurrence of the attachment of LAMP to the SCV and positioning in a perinuclear region.81 Rab interacting lysosomal protein (RILP), the effector of Rab7, can use a microtubule motor to manage the perinuclear position of SCV.83 In this stage, sorting nexin 1 (SNX1) removes the cation-independent mannose-6- phosphate receptor from SCV and prevents lysosomal hydrolases.84 The most important factors absent in SCV include cathepsin D and mannose-6-phosphate receptor.85 SNX1 induces tubulation in an early endosome in a dose-dependent fashion.86 At the first hour of Salmonella invasion, SNX1 spreads out in the ruffle membrane that engulfs bacteria.84 After a few hours, v-ATPase acidifies the SCV that results in the activation of SPI-2.87 SPI-2 and its effector mediate SIF. SIF is a double-membrane network, and the inner tubular space contains a portion of host cytosol; however, the outer tubular space contains an SCV material whose structure may show the elongation of vacuole-containing bacteria.88–90 Nutrients can be exchanged between SIF lumen and endolysosomal system, and pathogen from SCV can reach out to nutrient through SIF.91 Furthermore, Salmonella subverts the secretory vesicle to SCV to obtain nutrient.92 In the final stage, SCV may combine with endolysosomal vesicle; yet, after a short while, endolysosomal substance separates from SCV.93 Vesicle-associated membrane protein 7 (VAMP7) is a major factor in lysosomal fusion and has a role in SIF formation and recurrence of late lysosomal to SIF.94 To inhibit the degradation of vesicle via the lysosomal enzyme, SopD2 as an effector protein from T3SS-2 should disrupt Rab7 activity, leading to the inhibition of nucleotide exchange and disruption of the interaction between Rab7 and dynein.95 Altogether, this process facilitates the attachment of lysosome without hydrolytic enzyme to SCV and, thus, provides a safe place for Salmonella replication.96 SopA mimics the ubiquitin ligase of host and may induce intestinal inflammation.97 However, SspH2 is encoded by phage aid to downregulate proinflammatory response to the bacteria inside the SCV.98 For vacuole stability, Salmonella translocates effector Ssej to the eukaryotic cell to interact with oxysterol binding protein 1 (OSBP1). OSPB1 is demonstrated to be involved in translocating sterols from lysosomes to the nucleus and exchanging sterol with phosphatidylinositol-4-phosphate, thus leading to the membrane stability of SCV.99 SseF is an effector that secretes SPI-2 and mediates SCV position close to Golgi network. The persistence of SCV near the Golgi required the recurrence of dynein on the SCV surface.100 SseG interacts with SseF to localize SCV in the Golgi region and convert the monolayer membrane to the double-membrane SIF.88,101 SseL mediates macrophage killing and inhibits autophagy induced by the formation of ubiquitinated aggregates.102 Salmonella translocates effector A (SteA) standing on the Sif and SCV and is involved in regulating SCV membrane dynamics. This attachment occurs when SteA binds to the phosphatidylinositol-4- phosphate in the eukaryotic cell.103 SteD can block MHC of Class 2, deplete surface MHCII, and inhibit T-cell activation.104 SteC regulates intracellular replication and interacts with MAP kinase to induce actin formation.105 SpiC inhibits endosomal trafficking and is involved in translocating effector proteins such as SseB and SseC.106 Salmonella-induced filaments A and B (SifA and SifB) represent other effector proteins necessary for SIF formation. SifA interacts with Rab7 to block the interaction between RILP and Rab7, leading to the stability of SCV in a perinuclear position.107 SifA also can block the interaction between Rab9 and SKIP (SifA and Kinesin interacting protein), which has a major role in the recurrence of Lamp1 to the SCV and membrane stability.108 SifA binds to the SKIP and Rab9 and forms a stable complex that affects the return of mannose-6-phosphat receptor to the SCV surface.85 Salmonella invasion protein A (SipA) mediates localization of SifA on SCV.109
Similar to SifA, SifB is translocated through SPI-2 T3SS and shares subcellular localization with SseJ on the SCV.110 SifB is encoded outside of the SPI-2 and, similar to SifA, it mediates SIF formation, recurrent exocytic vesicle to the SCV, and prenuclear position of SCV.98 For the dormant intracellular life, SPI-2 effector is needed and induced by the two-component system (OmpR-EnvZ). However, among these effectors, compared to SifB, SseG and SseJ are required for the dormant life.111 The presence of two types of homologous proteins such as SifA and SifB may show their overlapping functions in the cells or synergism between these proteins.112
Another effector called PipB2 can promote SIF extension and cause the recurrence of Kinesin to the SCV membrane.98,113 With the recurrence of Kinesin to the SIF, PipB2 may play a role in extending nascent filament to the outside of SCV.113 PipB2 helps Salmonella with intra-macrophage survival.114 For more information, see Table 2.
 |
Table 2 Salmonella Adhesins and Their Effects |
Salmonella Toxins
Cytotoxin
Salmonella can induce pyroptosis via T3SS effector protein (needle protein) in a flagellin manner. This protein can be detected by NLRC4 (Nod-Like Receptor) inflammasome; then, inflammasome activates caspase-1 that leads to the secretion of IL-1β and programs cell death called pyroptosis.115 Inflammasome is composed of multiple proteins that resist the pathogen.116 To activate NLRC4, flagellin or needle protein of T3SS should be recognized by NLR family apoptosis inhibitory protein (NAIPs).117 On the other hand, Salmonella can downregulate the flagellin expression and evade NLRC4 activation.118 In the final step, Salmonella secretes flagellin into the cytosol and activates NLRC4 to induce pyroptosis and cell death, leading to Salmonella release.119 Salmonella induces interferon type I and, therefore, leads to inducing necroptosis controlled by a receptor-interacting protein (RIP).120 Interestingly, not all of the macrophage dies result from infection, and a particular phenotype that remains alive acts as a reservoir for Salmonella.121 This phenotype is called M2 and in spleen called hemophagocytes macrophage.122 Spv locus and its effector SpvB are required to induce apoptosis in macrophage. This locus has a major role in inducing systemic disease.123 SpvB has an ADP- ribosylation role in the activation of caspase and inducing apoptosis and actin polymerization during the intracellular life.124 SpvC is another effector that is secreted through T3SS with a phosphothreonine lyase activity and inhibits the activation of MAP kinase and de-phosphorylate ER, leading to the downregulation of proinflammatory cytokines.125,126 Another effector that mediates apoptosis in epithelial cells is SrfT, which is encoded in SPI-2.127 Moreover, SrfH mediates actin remodeling and is located in prophage. SrfD, SrfE, SdrI, SrfK, SrfL, and SrfM also are encoded by Phage. Among these effectors, SrfJ mediates apoptosis in host cells and, by glycoside hydrolase activity, modifies SCV membrane lipid to mediate the increase of salmonella virulence.128
Toxins
SopE is a nucleotide exchange factor that affects Rho GTPase family such as Cdc42 and Rac1. This family is involved in activation (GTP form) and inactivation (GDP form) of Rho that is controlled by a guanine nucleotide exchange factor.129 Only the activated form can affect the downstream of cascade and make a signal that causes cell responses such as gene transcription and actin rearrangement.130 SopE acts as a guanine exchange factor in activating both Rac1 and Cdc42, leading to actin rearrangement and infiltration of bacteria into the host cell.131 This event results from SopE and can be detected via NOD1 and proinflammatory response.132 Another effector or toxin that interacts with cytoskeleton is an SptP, acting as a GTPase-activating protein, and deactivates Rac1 and Cdc42 to return cytoskeletal change to the relax form; this factor goes in contrast to the SopE factor.133 Furthermore, SptP can suppress MAP kinase activation. MAP kinase is involved in the activation of cellular responses such as proinflammatory response.134 In fact, the tyrosine phosphatase activity of SptP interacts with Raf-1 to inhibit the activation of the extracellular-signal-regulated kinase (ERK).135 SptP toxin is not effectively presented in Salmonella Typhi and, because of the changes in some amino acids, chaperon cannot be attached to the effector and stabilize it.136,137 SipA can activate caspase-3 and actin polymerization.138 Note that SipA cannot induce caspase-3 in polymorphonuclear.138 Caspase-3 can degrade SipA to C-terminal and N-terminal: the C-terminal is responsible for actin polymerization and the N-terminal contains an SNARE motif.139 The latter is responsible for the attachment and recurrence of host syntaxin 8 to SCV. SNARE is involved in endocytosis and syntaxin in rapid endocytosis and vesicle mobilization.140 Altogether, this effect results in the recurrence of syntaxin8 to SCV and its fusion with early endosome to inhibit vesicle maturation.139 (Figure 2)
 |
Figure 2 An overview of the entrance, Salmonella-containing vacuole formation, Sif formation, and replication inside the host vacuole. |
Another effector called SipC directly binds to actin and causes actin polymerization. In addition to translocated effector, SipC can interact with cell vesicle trafficking and reset the vesicle onto the cell surface.141 Further, SipC interacts with p53 effector related to PMP-22 (PERP) and, thus, facilitates the formation of exocyst, which is transferred to the cell membrane. The accumulation of exocyst leads to the recurrence of vesicle to the cell surface, and this accumulation in membrane results in the rippling membrane that permits Salmonella entrance.142,143 Spi-D is used to translocate effector proteins to the eukaryotic cell.144 In the infected host cell, caspase-1 is activated to make pyroptosis; however, this effect also leads to lysophosphatidylcholine release that forces Salmonella to react in the form of secretion of SIPs.145 In other words, Salmonella can detect lysophosphatidylcholine and release SIPs effector. Lysophosphatidylcholine also stimulates the invasive power of bacteria. Due to the presence of lipid in plasma, Salmonella can be hyperinvasive through it.146 AvrA is another effector protein that is injected into the intracellular antigen with acetyltransferase activity. Avra inhibits the activation of c-Jun kinase (JNK) and NF-κB (nuclear factor kappa-light-chain-enhancer of activated B cells) in the epithelial cell to modulate apoptosis and protect pathogen strategy.147 Cytolethal Distending Toxin (CDT) breaks the DNA of the host cell and, in turn, provoke DNA repair response.148 Interestingly, the cell surviving the DNA repair response causes genomic instability and becomes exposed to cancer progression.149 This event leads to the persistence of host and reduces the inflammatory response and the long durability of bacteria.150 Typhoid toxin consists of two A subunits (enzymatic activity) and five B subunits; this toxin is transferred from SCV to the extracellular milieu and affects cell other than the infected cell.151 Part A consists of PltA and CdtB; the former has an ADP- ribosylation transferase activity and the latter has a deoxyribonuclease activity, which is damaging to the cell.152 Subunit A of Typhoid toxin is homologous to the CDT and pertussis toxin.153
How to Activate Pathogenic Genes
SPI-1 mediates invasion and is controlled by HilA, InvF,154 and HilD. Environmental conditions such as pH and osmolarity activate transcriptional regulatory proteins of HilC and HilD so that they can bind to the upstream of master regulatory hilA.155 HilC and HilD counteract global repressor H-NS on the hilA promotor.156 Following the activation of HilA, InvF is subsequently activated.157 InvF activates the gene inside and outside of SPI-1. HilC and HilD also induce the expression of hilC and hilD genes that ultimately activate hilA.158 HilD first induces hilA that is located in SPI-1 and, then, induces ssrAB located in SPI-2. Following the internalization of bacteria to the acidic vacuole, another effector from SPI-2 should be secreted to modify the circumstance. First, EnvZ/OmpR should be considered; the inner membrane protein regulatory system senses acid and activates ssrA/ssrB and PhoQ/P systems.159 To this end, first, EnvZ interacts with OmpR and, then, OmpR activates ssrA transcription and produces SsrA and SsrB. Interestingly, by suppressing cadC/BA operon, OmpR blocks a neutralization reaction to acidity.87 CadA is a lysine decarboxylase that produces cadaverine and, then, is translocated to cadB to export out the bacterial cell and modulate acidification.160 However, OmpR blocking this event leads to the maintenance of acidification of bacterial cytoplasm, which is essential to the secretion of SPI-2 effector protein. In the presence of SsrA kinase, phosphorylated SsrB induces SPI-2; however, in neutral PH, SsrA kinase is very low and SsrB is unphosphorylated; yet, biofilm gene is expressed.159 In addition to the acid response, ssrB can induce biofilm formation and switch between intracellular and extracellular lifestyles.161 When the acidification of bacterial cytoplasm increases, supercoiling of DNA may decrease and OmpR binding site is exposed.162 For more information, see Figure 3. However, HilD distinctively regulates both SPI-1 and SPI-2. HilD can counteract the repressive effect of H-NS in the regulatory region of ssrAB operon.163 In summary, following the activation of SPI-1 and invasion against the cell and entrance, SPI-2 activates and suppresses the expression of SPI-1 through SsrB. SsrB directly binds to the regulatory region of hilA and hilD and suppress it.164 Recently, it has been observed that the intestinal butyrate derived from clostridia can inhibit the expression of hilD, encoding the T3SS-1 gene regulator.165 Salmonella utilizes butyrate using β-oxidation and, in turn, provides fitness advantages during pathogen growth. Interestingly, gene of β-oxidation ydi QRSTD is not presented in S. Typhi, leading to reduced invasion against the intestinal lumen.166 This issue may explain why S. Typhi does not induce inflammation in the first phase, and this provides an opportunity to spread through another organ without inflammation.
 |
Figure 3 A comprehensive review of Salmonella mechanisms in the regulation of pathogenic genes. |
Regulation of Immune Response
One of the important immune cells that controls Salmonella is DC and is responsible for cytokine secretion, presenting antigen to a T cell, and activating natural killer cells (NK cells).167 Following the phagocyte of bacteria, the DC process presents bacterial antigen to the T cell. However, if bacterial cell survives, it can migrate through DC to another organ.168 Salmonella can prevent DC migration through SseI protein that secretes via T3SS.169 This effector inhibits chemotaxis of DC toward the T cell zone by following chemokine (C-C motif) ligand 19 (CCL19).170 Salmonella SseI interacts with the downstream of CC-chemokine receptor 7 (CCR7) that responds to CCL19.168 Chemotaxis towards CCR7 is controlled by MAP kinase.171 SseI can interact with filamins172 and cytoskeletal regulatory protein, IQ motif-containing GTPase-activating protein (IQGAP1). The migration of DC and macrophage requires the IQGAP1 factor, and SseI exactly interacts with it to suppress the migration and prevent T CD4+ response.173 Deletion or pseudogenization of SseI causes systemic infection.169 Macrophage via TLR2 and 4 is able to recognize Salmonella; however, this signaling also induces Salmonella replication. This event can be explained by acidification in Salmonella-containing phagosome that activates SPI-2.174 TLR controls DC maturation and, thus, activates the adaptive immune response.175 Although other TLRs such as TLR2, TLR5, and TLR9 are involved in detecting salmonella, TLR4 has a major role in apoptosis.176 It should be noted that, in normal macrophage, TLR4 also activates NF-κB and MAP kinase with an anti-apoptosis effect.177 Mucus property influenced by the gel-forming composition is called musin. Musin is composed of glycosylated and non-glycosylated domains with a peptide backbone.178 At the end of the oligosaccharide branch of musin, sialic acid or sulfate can be found. Salmonella can be attached to the mannose and sialic acid, and it goes through the mucus via sialidase.179 This enzyme can cleave the α-ketosidic bond in the terminal of sialic acid residue.180 Interestingly, IFNγ has a dual role in mucosal defense; first, signaling of IFNγR in Goblet cell (mucosal secretion cell) leads to the loss of mucus; on the other hand, IFNγR restricts the growth of pathogen in macrophage.181 After passing the mucus layer and entering the lamina propria, TLR is activated via flagelin and mediates phagocyte by DC and macrophage.182 Another host sensor, NLRC4, can recognize an intracellular antigen such as flagellin.118 This recognition leads to the secretion of proinflammatory cytokines such as IL-1β and IL-18. NLRP3 is activated through reactive oxygen species (ROS), or membrane damage leading to activated caspase-1 and pyroptosis.183 The activation of pyroptosis secures the release of Salmonella from macrophage and swallowing by Neutrophil and degradation by ROS.119 However, Salmonella can degrade superoxide and limit peroxynitrite formation by superoxide dismutase.184 Salmonella forces macrophage and epithelial cell to release IL-18 and IL-23 secretion from the dendritic cell.185 Then, IL-18 stimulates Th1 to release IFNγ and IL-23, thus stimulating Neutrophil, TH 17, and Tγδ to produce IL-22 and IL-17.186,187 The activation of TH 17 expresses CXC chemokine such as CXCL and CXCL2 and results in the recurrence of Neutrophil to the infection site.188 IL-22 can cause Neutrophil and epithelial cell to secrete Lipocalin-2 and calprotectin; these molecules inhibit Salmonella access to iron and manganese.189 However, oxidative stress can stimulate Salmonella to upregulate the sitABCD manganese transport system (high-affinity transporter) that effectively helps the bacteria overcome growth inhibition, which started with calprotectin.190 Manganese acts as a cofactor in the superoxide dismutase enzyme. Furthermore, in the periplasmic space, Dsb proteins can act as an oxidase reductase, and this protein with four components of DsbA, DsbB, DsbD, and DsbC also works as follows: DsbA is tasked with creating a disulfide bond; DsbB, DsbD are electron donors; DsbC proofreads the disulfide bond formation.191
Conclusion
Following the attachment of Salmonella to the epithelial cell, two routes can be selected: (a) one entrance to the cell in a T3SS-dependent fashion, thus mediating effector protein and forming SCV; (b) entrance in a T3SS-independent fashion. The formation of SCV management with T3SS-1 secretes invasive protein; however, afterwards, there are two ways ahead: either maintaining in SCV or exiting it. For the first one, Salmonella requires to be transferred from SPI-1 (invasive) to SPI-2 (maintaining). After maintenance in the SCV, the next step is replication that requires a nutrient; for this reason, a long appendage is extended into the cytoplasm called SIFs. Salmonella can exit SCV, replicates in the cytoplasm, and targets autophagy system; however, it can stop autophagy or kill the macrophage and is released afterwards. Following its release, Salmonella is engulfed by Neutrophil and degraded with ROS; like other bacteria, it can resist ROS (ref. staph). After the activation of macrophage and DC that migrated to the lymph node, the antigen, which was modified earlier inside the macrophage and DC, was presented to the T-cell existing in the paracortex. Salmonella that induces enteritis utilizes the butyrate via β-oxidation and provides energy for the duplication, and this in turn made the inflammation as derived from the duplication. On the contrary, S. Typhi did not activate inflammation and, thus, is the reason for the lack of immune attention, giving S. Typhi time to spread.
Acknowledgment
We would like to thank “School of Public Health, Tehran University of Medical Sciences, Tehran, Iran” for their kind cooperation. The authors received no specific funding for this work.
Author Contributions
All authors made substantial contributions to the conception and design, acquisition of data, or interpretation of data. They played an active role in drafting the article or revising it critically to achieve important intellectual content. They gave the final approval of the version to be published and agreed to be accountable for all aspects of the work.
Disclosure
All of the authors declare that there are no commercial, personal, political, and any other potentially conflicting interests related to the submitted manuscript.
References
1. Khosravani M, Dallal MMS, Norouzi M. marA efflux pump gene expression in Salmonella enteritidis strains treated with Artemisia tournefortiana hydroalcoholic extract and comparison with commercial efflux pump inhibitor, carbonyl cyanide 3-chlorophenylhydrazone (CCCP). Arch Med Lab Sci. 2019;4:1.
2. Soltan Dallal M, Khalilian M, Masoumi Asl H, et al. Molecular epidemiology and antimicrobial resistance of Salmonella spp. Isolated from resident patients in Mazandaran Province, Northern Iran. J Food Qual Hazards Control. 2016;3(4):146–151.
3. Sayadnouri S, Dallal MS, Akbarzadeh S, Fard RMN. Evaluation of class 1 and 2 integrons and antibiotic resistance pattern in Salmonella enterica isolated from diarrheal food-borne outbreaks in Iran. J Food Qual Hazards Control. 2019. doi:10.18502/jfqhc.6.3.1384
4. Nasser A. Prevalence of sulfonamide resistance genes in Salmonella spp. isolated from Milad hospital of Tehran. Iran J Public Health. 2014;43(2):87.
5. Crump JA, Mintz ED. Global trends in typhoid and paratyphoid fever. Clin Infect Dis. 2010;50(2):241–246. doi:10.1086/649541
6. Rivera-Chavez F, Baumler AJ. The pyromaniac inside you: Salmonella metabolism in the host gut. Annu Rev Microbiol. 2015;69:31–48. doi:10.1146/annurev-micro-091014-104108
7. Kaur J, Jain SK. Role of antigens and virulence factors of Salmonella enterica serovar Typhi in its pathogenesis. Microbiol Res. 2012;167(4):199–210. doi:10.1016/j.micres.2011.08.001
8. Fardsanei F, Dallal MMS, Douraghi M, et al. Antimicrobial resistance, virulence genes and genetic relatedness of Salmonella enterica serotype Enteritidis isolates recovered from human gastroenteritis in Tehran, Iran. J Global Antimicrob Resist. 2018;12:220–226. doi:10.1016/j.jgar.2017.10.005
9. Figueira R, Holden DW. Functions of the Salmonella pathogenicity island 2 (SPI-2) type III secretion system effectors. Microbiology. 2012;158(5):1147–1161. doi:10.1099/mic.0.058115-0
10. Galan JE. Salmonella interactions with host cells: type III secretion at work. Annu Rev Cell Dev Biol. 2001;17:53–86. doi:10.1146/annurev.cellbio.17.1.53
11. Rescigno M, Urbano M, Valzasina B, et al. Dendritic cells express tight junction proteins and penetrate gut epithelial monolayers to sample bacteria. Nat Immunol. 2001;2(4):361. doi:10.1038/86373
12. Kiss T, Morgan E, Nagy G. Contribution of SPI-4 genes to the virulence of Salmonella enterica. FEMS Microbiol Lett. 2007;275(1):153–159. doi:10.1111/fml.2007.275.issue-1
13. Wagner C, Polke M, Gerlach RG, et al. Functional dissection of SiiE, a giant non-fimbrial adhesin of Salmonella enterica. Cell Microbiol. 2011;13(8):1286–1301. doi:10.1111/cmi.2011.13.issue-8
14. Gerlach RG, Claudio N, Rohde M, Jackel D, Wagner C, Hensel M. Cooperation of Salmonella pathogenicity islands 1 and 4 is required to breach epithelial barriers. Cell Microbiol. 2008;10(11):2364–2376. doi:10.1111/j.1462-5822.2008.01218.x
15. Main-Hester KL, Colpitts KM, Thomas GA, Fang FC, Libby SJ. Coordinate regulation of Salmonella pathogenicity island 1 (SPI1) and SPI4 in Salmonella enterica serovar Typhimurium. Infect Immun. 2008;76(3):1024–1035. doi:10.1128/IAI.01224-07
16. Wagner C, Polke M, Gerlach RG, et al. Functional dissection of SiiE, a giant non-fimbrial adhesin of Salmonella enterica. Cell Microbiol. 2011;13(8):1286–1301. doi:10.1111/j.1462-5822.2011.01621.x
17. Latasa C, Roux A, Toledo-Arana A, et al. BapA, a large secreted protein required for biofilm formation and host colonization of Salmonella enterica serovar Enteritidis. Mol Microbiol. 2005;58(5):1322–1339. doi:10.1111/j.1365-2958.2005.04907.x
18. Romling U, Rohde M, Olsen A, Normark S, Reinkoster J. AgfD, the checkpoint of multicellular and aggregative behaviour in Salmonella typhimurium regulates at least two independent pathways. Mol Microbiol. 2000;36(1):10–23. doi:10.1046/j.1365-2958.2000.01822.x
19. González JF, Tucker L, Fitch J, Wetzel A, White P, Gunn JS. Human bile-mediated regulation of Salmonella curli fimbriae. J Bacteriol. 2019;
20. Rosselin M, Virlogeux-Payant I, Roy C, et al. Rck of Salmonella enterica, subspecies enterica serovar Enteritidis, mediates Zipper-like internalization. Cell Res. 2010;20(6):647. doi:10.1038/cr.2010.45
21. Heffernan EJ, Reed S, Hackett J, Fierer J, Roudier C, Guiney D. Mechanism of resistance to complement-mediated killing of bacteria encoded by the Salmonella typhimurium virulence plasmid gene rck. J Clin Invest. 1992;90(3):953–964. doi:10.1172/JCI115972
22. Wiedemann A, Mijouin L, Ayoub MA, et al. Identification of the epidermal growth factor receptor as the receptor for Salmonella Rck-dependent invasion. FASEB J. 2016;30(12):4180–4191. doi:10.1096/fj.201600701R
23. Wiedemann A, Rosselin M, Mijouin L, Bottreau E, Velge P. Involvement of c-Src tyrosine kinase upstream of class I phosphatidylinositol (PI) 3-kinases in Salmonella Enteritidis Rck protein-mediated invasion. J Biol Chem. 2012;287(37):31148–31154. doi:10.1074/jbc.M112.392134
24. Lambert MA, Smith SG. The PagN protein of Salmonella enterica serovar Typhimurium is an adhesin and invasin. BMC Microbiol. 2008;8(1):142. doi:10.1186/1471-2180-8-142
25. Lambert MA, Smith SG. The PagN protein mediates invasion via interaction with proteoglycan. FEMS Microbiol Lett. 2009;297(2):209–216. doi:10.1111/j.1574-6968.2009.01666.x
26. Lambert MA, Smith SGJ. The PagN protein mediates invasion via interaction with proteoglycan. FEMS Microbiol Lett. 2009;297(2):209–216. doi:10.1111/j.1574-6968.2009.01666.x
27. Prost LR, Miller SI. The Salmonellae PhoQ sensor: mechanisms of detection of phagosome signals. Cell Microbiol. 2008;10(3):576–582. doi:10.1111/cmi.2008.10.issue-3
28. Chowdhury R, Das S, Ta A, Das S. Epithelial invasion by Salmonella Typhi using STIV–met interaction. Cell Microbiol. 2019;21(3):e12982.
29. Chowdhury R, Mandal RS, Ta A, Das S. An AIL family protein promotes type three secretion system-1-independent invasion and pathogenesis of Salmonella enterica serovar Typhi. Cell Microbiol. 2015;17(4):486–503. doi:10.1111/cmi.2015.17.issue-4
30. Dorsey CW, Laarakker MC, Humphries AD, Weening EH, Bäumler AJ. Salmonella enterica serotype Typhimurium MisL is an intestinal colonization factor that binds fibronectin. Mol Microbiol. 2005;57(1):196–211. doi:10.1111/j.1365-2958.2005.04666.x
31. Kingsley RA, Santos RL, Keestra AM, Adams LG, Bäumler AJ. Salmonella enterica serotype Typhimurium ShdA is an outer membrane fibronectin-binding protein that is expressed in the intestine. Mol Microbiol. 2002;43(4):895–905. doi:10.1046/j.1365-2958.2002.02805.x
32. Cooley MB, Miller WG, Mandrell RE. Colonization of Arabidopsis thaliana with Salmonella enterica and enterohemorrhagic Escherichia coli O157: h7and competition by Enterobacter asburiae. Appl Environ Microbiol. 2003;69(8):4915–4926. doi:10.1128/AEM.69.8.4915-4926.2003
33. Tan MSF, White AP, Rahman S, Dykes GA. Role of fimbriae, flagella and cellulose on the attachment of Salmonella typhimurium ATCC 14028 to plant cell wall models. PLoS One. 2016;11(6):e0158311. doi:10.1371/journal.pone.0158311
34. Hara-Kaonga B, Pistole TG. OmpD but not OmpC is involved in adherence of Salmonella enterica serovar Typhimurium to human cells. Can J Microbiol. 2004;50(9):719–727. doi:10.1139/w04-056
35. Schmidt H, Hensel M. Pathogenicity islands in bacterial pathogenesis. Clin Microbiol Rev. 2004;17(1):14–56. doi:10.1128/CMR.17.1.14-56.2004
36. Ilyas B, Tsai CN, Coombes BK. Evolution of Salmonella-host cell interactions through a dynamic bacterial genome. Front Cell Infect Microbiol. 2017;7:428. doi:10.3389/fcimb.2017.00428
37. Wallis TS, Galyov EE. Molecular basis of Salmonella‐induced enteritis: microReview. Mol Microbiol. 2000;36(5):997–1005. doi:10.1046/j.1365-2958.2000.01892.x
38. Jantsch J, Chikkaballi D, Hensel M. Cellular aspects of immunity to intracellular Salmonella enterica. Immunol Rev. 2011;240(1):185–195. doi:10.1111/imr.2011.240.issue-1
39. Mason D, Mallo GV, Terebiznik MR, et al. Alteration of epithelial structure and function associated with PtdIns (4, 5) P2 degradation by a bacterial phosphatase. J Gen Physiol. 2007;129(4):267–283. doi:10.1085/jgp.200609656
40. Knodler LA, Finlay BB, Steele-Mortimer O. The Salmonella effector protein SopB protects epithelial cells from apoptosis by sustained activation of Akt. J Biol Chem. 2005;280(10):9058–9064. doi:10.1074/jbc.M412588200
41. Garcia-Gil A, Galan-Enriquez CS, Perez-Lopez A, Nava P, Alpuche-Aranda C, Ortiz-Navarrete V. SopB activates the Akt-YAP pathway to promote Salmonella survival within B cells. Virulence. 2018;9(1):1390–1402. doi:10.1080/21505594.2018.1509664
42. Kum WWS, Lo BC, Yu HB, Finlay BB. Protective role of Akt2 in Salmonella enterica serovar typhimurium-induced gastroenterocolitis. Infect Immun. 2011;79(7):2554. doi:10.1128/IAI.01235-10
43. Bakowski MA, Braun V, Lam GY, et al. The phosphoinositide phosphatase SopB manipulates membrane surface charge and trafficking of the Salmonella-containing vacuole. Cell Host Microbe. 2010;7(6):453–462. doi:10.1016/j.chom.2010.05.011
44. Truong D, Boddy KC, Canadien V, et al. Salmonella exploits host Rho GTPase signalling pathways through the phosphatase activity of SopB. Cell Microbiol. 2018;20(10):e12938. doi:10.1111/cmi.v20.10
45. Hernandez LD, Hueffer K, Wenk MR, Galán JE. Salmonella modulates vesicular traffic by altering phosphoinositide metabolism. Science. 2004;304(5678):1805. doi:10.1126/science.1098188
46. Mallo GV, Espina M, Smith AC, et al. SopB promotes phosphatidylinositol 3-phosphate formation on Salmonella vacuoles by recruiting Rab5 and Vps34. J Cell Biol. 2008;182(4):741–752. doi:10.1083/jcb.200804131
47. Sana TG, Flaugnatti N, Lugo KA, et al. Salmonella Typhimurium utilizes a T6SS-mediated antibacterial weapon to establish in the host gut. Proc National Acad Sci. 2016;113(34):E5044–E5051. doi:10.1073/pnas.1608858113
48. Kaur J, Jain S. Role of antigens and virulence factors of Salmonella enterica serovar Typhi in its pathogenesis. Microbiol Res. 2012;167(4):199–210. doi:10.1016/j.micres.2011.08.001
49. Hart PJ, O’Shaughnessy CM, Siggins MK, et al. Differential killing of Salmonella enterica serovar typhi by antibodies targeting Vi and lipopolysaccharide o:9 antigen. PLoS One. 2016;11(1):e0145945. doi:10.1371/journal.pone.0145945
50. Reid RR, Prodeus AP, Khan W, Hsu T, Rosen FS, Carroll MC. Endotoxin shock in antibody-deficient mice: unraveling the role of natural antibody and complement in the clearance of lipopolysaccharide. J Immunol. 1997;159(2):970–975.
51. Pickard D, Wain J, Baker S, et al. Composition, acquisition, and distribution of the Vi exopolysaccharide-encoding Salmonella enterica pathogenicity island SPI-7. J Bacteriol. 2003;185(17):5055–5065. doi:10.1128/JB.185.17.5055-5065.2003
52. Seth-Smith HMB, Fookes MC, Okoro CK, et al. Structure, diversity, and mobility of the pathogenicity island 7 family of integrative and conjugative elements within enterobacteriaceae. J Bacteriol. 2012;194(6):1494.
53. Zhang X-L, Tsui ISM, Yip CMC, et al. Salmonella enterica serovar typhi uses type IVB pili to enter human intestinal epithelial cells. Infect Immun. 2000;68(6):3067. doi:10.1128/IAI.68.6.3067-3073.2000
54. Winter SE, Winter MG, Thiennimitr P, et al. The TviA auxiliary protein renders the Salmonella enterica serotype Typhi RcsB regulon responsive to changes in osmolarity. Mol Microbiol. 2009;74(1):175–193. doi:10.1111/mmi.2009.74.issue-1
55. Bajaj V, Hwang C, Lee CA. hilA is a novel ompR/toxR family member that activates the expression of Salmonella typhimurium invasion genes. Mol Microbiol. 1995;18(4):715–727. doi:10.1111/mmi.1995.18.issue-4
56. Hiyoshi H, Wangdi T, Lock G, et al. Mechanisms to Evade the Phagocyte Respiratory Burst Arose by Convergent Evolution in Typhoidal Salmonella Serovars. Cell Rep. 2018;22(7):1787–1797. doi:10.1016/j.celrep.2018.01.016
57. Sharma A, Qadri A. Vi polysaccharide of Salmonella typhi targets the prohibitin family of molecules in intestinal epithelial cells and suppresses early inflammatory responses. Proc Natl Acad Sci U S A. 2004;101(50):17492–17497. doi:10.1073/pnas.0407536101
58. Raffatellu M, Chessa D, Wilson RP, Dusold R, Rubino S, Bäumler AJ. The Vi capsular antigen of salmonella enterica serotype typhi reduces toll-like receptor-dependent interleukin-8 expression in the intestinal mucosa. Infect Immun. 2005;73(6):3367. doi:10.1128/IAI.73.6.3367-3374.2005
59. Hirose K, Ezaki T, Miyake M, et al. Survival of Vi-capsulated and Vi-deleted Salmonella typhi strains in cultured macrophage expressing different levels of CD14 antigen. FEMS Microbiol Lett. 1997;147(2):259–265. doi:10.1111/j.1574-6968.1997.tb10251.x
60. Kong Q, Yang J, Liu Q, Alamuri P, Roland KL, Curtiss R. Effect of deletion of genes involved in lipopolysaccharide core and O-antigen synthesis on virulence and immunogenicity of Salmonella enterica serovar Typhimurium. Infect Immun. 2011;79(10):4227. doi:10.1128/IAI.00040-11
61. Nevola JJ, Laux DC, Cohen PS. In vivo colonization of the mouse large intestine and in vitro penetration of intestinal mucus by an avirulent smooth strain of Salmonella typhimurium and its lipopolysaccharide-deficient mutant. Infect Immun. 1987;55(12):2884–2890.
62. Raetz CR, Whitfield C. Lipopolysaccharide endotoxins. Annu Rev Biochem. 2002;71(1):635–700. doi:10.1146/annurev.biochem.71.110601.135414
63. Murray GL, Attridge SR, Morona R. Altering the length of the lipopolysaccharide O antigen has an impact on the interaction of Salmonella enterica serovar Typhimurium with macrophages and complement. J Bacteriol. 2006;188(7):2735–2739. doi:10.1128/JB.188.7.2735-2739.2006
64. Vaara M. Antibiotic-supersusceptible mutants of Escherichia coli and Salmonella typhimurium. Antimicrob Agents Chemother. 1993;37(11):2255. doi:10.1128/AAC.37.11.2255
65. Pawlak A, Rybka J, Dudek B, et al. Salmonella O48 serum resistance is connected with the elongation of the lipopolysaccharide O-antigen containing sialic acid. Int J Mol Sci. 2017;18:10. doi:10.3390/ijms18102022
66. Delgado MA, Mouslim C, Groisman EA. The PmrA/PmrB and RcsC/YojN/RcsB systems control expression of the Salmonella O‐antigen chain length determinant. Mol Microbiol. 2006;60(1):39–50. doi:10.1111/mmi.2006.60.issue-1
67. McCONNELL M, Wright A. Variation in the structure and bacteriophage-inactivating capacity of Salmonella anatum lipopolysaccharide as a function of growth temperature. J Bacteriol. 1979;137(2):746–751.
68. McClelland M, Sanderson KE, Clifton SW, et al. Comparison of genome degradation in Paratyphi A and Typhi, human-restricted serovars of Salmonella enterica that cause typhoid. Nat Genet. 2004;36(12):1268. doi:10.1038/ng1470
69. Verma N, Reeves P. Identification and sequence of rfbS and rfbE, which determine antigenic specificity of group A and group D salmonellae. J Bacteriol. 1989;171(10):5694. doi:10.1128/jb.171.10.5694-5701.1989
70. Vázquez-Torres A. Less is best in the convergent evolution of typhoidal Salmonella. Cell Host Microbe. 2018;23(2):151–153. doi:10.1016/j.chom.2018.01.009
71. Knodler LA, Steele-Mortimer O. Taking possession: biogenesis of the Salmonella-containing vacuole. Traffic. 2003;4(9):587–599. doi:10.1034/j.1600-0854.2003.00118.x
72. Stebbins CE, Galan JE. Modulation of host signaling by a bacterial mimic: structure of the Salmonella effector SptP bound to Rac1. Mol Cell. 2000;6(6):1449–1460. doi:10.1016/S1097-2765(00)00141-6
73. Bonifacino JS, Glick BS. The mechanisms of vesicle budding and fusion. Cell. 2004;116(2):153–166. doi:10.1016/S0092-8674(03)01079-1
74. Semerdjieva S, Shortt B, Maxwell E, et al. Coordinated regulation of AP2 uncoating from clathrin-coated vesicles by rab5 and hRME-6. J Cell Biol. 2008;183(3):499–511. doi:10.1083/jcb.200806016
75. Steele-Mortimer O, Meresse S, Gorvel JP, Toh BH, Finlay BB. Biogenesis of Salmonella typhimurium-containing vacuoles in epithelial cells involves interactions with the early endocytic pathway. Cell Microbiol. 1999;1(1):33–49. doi:10.1046/j.1462-5822.1999.00003.x
76. Martinez-Lorenzo MJ, Meresse S, de Chastellier C, Gorvel JP. Unusual intracellular trafficking of Salmonella typhimurium in human melanoma cells. Cell Microbiol. 2001;3(6):407–416. doi:10.1046/j.1462-5822.2001.00123.x
77. Cai H, Reinisch K, Ferro-Novick S. Coats, tethers, Rabs, and SNAREs work together to mediate the intracellular destination of a transport vesicle. Dev Cell. 2007;12(5):671–682. doi:10.1016/j.devcel.2007.04.005
78. Scott CC, Cuellar-Mata P, Matsuo T, Davidson HW, Grinstein S. Role of 3-phosphoinositides in the maturation of Salmonella-containing vacuoles within host cells. J Biol Chem. 2002;277(15):12770–12776. doi:10.1074/jbc.M110399200
79. Sindhwani A, Arya SB, Kaur H, Jagga D, Tuli A, Sharma M. Salmonella exploits the host endolysosomal tethering factor HOPS complex to promote its intravacuolar replication. PLoS Pathog. 2017;13(10):e1006700.
80. Mills IG, Jones AT, Clague MJ. Involvement of the endosomal autoantigen EEA1 in homotypic fusion of early endosomes. Curr Biol. 1998;8(15):881–884. doi:10.1016/S0960-9822(07)00351-X
81. Méresse S, Steele‐Mortimer O, Finlay BB, Gorvel JP. The rab7 GTPase controls the maturation of Salmonella typhimurium‐containing vacuoles in HeLa cells. EMBO J. 1999;18(16):4394–4403. doi:10.1093/emboj/18.16.4394
82. Kaniuk NA, Canadien V, Bagshaw RD, et al. Salmonella exploits Arl8B‐directed kinesin activity to promote endosome tubulation and cell‐to‐cell transfer. Cell Microbiol. 2011;13(11):1812–1823.
83. Harrison RE, Brumell JH, Khandani A, et al. Salmonella impairs RILP recruitment to Rab7 during maturation of invasion vacuoles. Mol Biol Cell. 2004;15(7):3146–3154. doi:10.1091/mbc.e04-02-0092
84. Bujny MV, Ewels PA, Humphrey S, Attar N, Jepson MA, Cullen PJ. Sorting nexin-1 defines an early phase of Salmonella-containing vacuole-remodeling during Salmonella infection. J Cell Sci. 2008;121(12):2027–2036.
85. McGourty K, Thurston TL, Matthews SA, Pinaud L, Mota LJ, Holden DW. Salmonella inhibits retrograde trafficking of mannose-6-phosphate receptors and lysosome function. Science. 2012;338(6109):963–967. doi:10.1126/science.1227037
86. Carlton J, Bujny M, Peter BJ, et al. Sorting nexin-1 mediates tubular endosome-to-TGN transport through coincidence sensing of high- curvature membranes and 3-phosphoinositides. Curr Biol. 2004;14(20):1791–1800. doi:10.1016/j.cub.2004.09.077
87. Chakraborty S, Mizusaki H, Kenney LJ. A FRET-based DNA biosensor tracks OmpR-dependent acidification of Salmonella during macrophage infection. PLoS Biol. 2015;13(4):e1002116. doi:10.1371/journal.pbio.1002116
88. Krieger V, Liebl D, Zhang Y, et al. Reorganization of the endosomal system in Salmonella-infected cells: the ultrastructure of Salmonella-induced tubular compartments. PLoS Pathog. 2014;10(9):e1004374. doi:10.1371/journal.ppat.1004374
89. Hannemann S, Gao B, Galán JE. Salmonella modulation of host cell gene expression promotes its intracellular growth. PLoS Pathog. 2013;9(10):e1003668. doi:10.1371/journal.ppat.1003668
90. Tuli A, Sharma M. How to do business with lysosomes: Salmonella leads the way. Curr Opin Microbiol. 2019;47:1–7. doi:10.1016/j.mib.2018.10.003
91. Drecktrah D, Levine‐Wilkinson S, Dam T, et al. Dynamic behavior of Salmonella‐induced membrane tubules in epithelial cells. Traffic. 2008;9(12):2117–2129. doi:10.1111/j.1600-0854.2008.00830.x
92. Perrett CA, Zhou D. Salmonella type III effector SopB modulates host cell exocytosis. Emerging Microbes Infect. 2013;2(1):1–6. doi:10.1038/emi.2013.37
93. Santos JC, Enninga J. At the crossroads: communication of bacteria-containing vacuoles with host organelles. Cell Microbiol. 2016;18(3):330–339. doi:10.1111/cmi.v18.3
94. Santos JC, Duchateau M, Fredlund J, et al. The COPII complex and lysosomal VAMP7 determine intracellular Salmonella localization and growth. Cell Microbiol. 2015;17(12):1699–1720. doi:10.1111/cmi.12475
95. D’Costa VM, Braun V, Landekic M, et al. Salmonella disrupts host endocytic trafficking by SopD2-mediated inhibition of Rab7. Cell Rep. 2015;12(9):1508–1518. doi:10.1016/j.celrep.2015.07.063
96. McEwan DG, Richter B, Claudi B, et al. PLEKHM1 regulates Salmonella-containing vacuole biogenesis and infection. Cell Host Microbe. 2015;17(1):58–71. doi:10.1016/j.chom.2014.11.011
97. Zhang Y, Higashide WM, McCormick BA, Chen J, Zhou D. The inflammation-associated Salmonella SopA is a HECT-like E3 ubiquitin ligase. Mol Microbiol. 2006;62(3):786–793. doi:10.1111/mmi.2006.62.issue-3
98. McGhie EJ, Brawn LC, Hume PJ, Humphreys D, Koronakis V. Salmonella takes control: effector-driven manipulation of the host. Curr Opin Microbiol. 2009;12(1):117–124. doi:10.1016/j.mib.2008.12.001
99. Kolodziejek AM, Altura MA, Fan J, et al. Salmonella translocated effectors recruit OSBP1 to the phagosome to promote vacuolar membrane integrity. Cell Rep 2019;27:2147–2156. doi:10.2139/ssrn.3188415
100. Abrahams GL, Muller P, Hensel M. Functional dissection of SseF, a type III effector protein involved in positioning the salmonella-containing vacuole. Traffic. 2006;7(8):950–965. doi:10.1111/j.1600-0854.2006.00454.x
101. Yu XJ, Liu M, Holden DW. Salmonella effectors SseF and SseG interact with mammalian protein ACBD3 (GCP60) to anchor salmonella-containing vacuoles at the golgi network. MBio. 2016;7:4. doi:10.1128/mBio.00474-16
102. Mesquita FS, Thomas M, Sachse M, Santos AJ, Figueira R, Holden DW. The Salmonella deubiquitinase SseL inhibits selective autophagy of cytosolic aggregates. PLoS Pathog. 2012;8(6):e1002743. doi:10.1371/journal.ppat.1002743
103. Domingues L, Ismail A, Charro N, et al. The Salmonella effector SteA binds phosphatidylinositol 4-phosphate for subcellular targeting within host cells. Cell Microbiol. 2016;18(7):949–969. doi:10.1111/cmi.12558
104. Bayer-Santos E, Durkin CH, Rigano LA, et al. The Salmonella effector SteD mediates MARCH8-dependent ubiquitination of MHC II molecules and inhibits T cell activation. Cell Host Microbe. 2016;20(5):584–595. doi:10.1016/j.chom.2016.10.007
105. Odendall C, Rolhion N, Forster A, et al. The Salmonella kinase SteC targets the MAP kinase MEK to regulate the host actin cytoskeleton. Cell Host Microbe. 2012;12(5):657–668. doi:10.1016/j.chom.2012.09.011
106. Freeman JA, Rappl C, Kuhle V, Hensel M, Miller SI. SpiC is required for translocation of Salmonella pathogenicity island 2 effectors and secretion of translocon proteins SseB and SseC. J Bacteriol. 2002;184(18):4971–4980. doi:10.1128/JB.184.18.4971-4980.2002
107. Bakowski MA, Braun V, Brumell JH. Salmonella-containing vacuoles: directing traffic and nesting to grow. Traffic. 2008;9(12):2022–2031. doi:10.1111/tra.2008.9.issue-12
108. Jackson LK, Nawabi P, Hentea C, Roark EA, Haldar K. The Salmonella virulence protein SifA is a G protein antagonist. Proc National Acad Sci. 2008;105(37):14141–14146. doi:10.1073/pnas.0801872105
109. Brawn LC, Hayward RD, Koronakis V. Salmonella SPI1 effector SipA persists after entry and cooperates with a SPI2 effector to regulate phagosome maturation and intracellular replication. Cell Host Microbe. 2007;1(1):63–75. doi:10.1016/j.chom.2007.02.001
110. Freeman JA, Ohl ME, Miller SI. The Salmonella enterica serovar typhimurium translocated effectors SseJ and SifB are targeted to the Salmonella-containing vacuole. Infect Immun. 2003;71(1):418–427. doi:10.1128/IAI.71.1.418-427.2003
111. Núñez-Hernández C, Alonso A, Pucciarelli MG, Casadesús J, García-del Portillo F. Dormant intracellular Salmonella enterica serovar Typhimurium discriminates among Salmonella pathogenicity island 2 effectors to persist inside fibroblasts. Infect Immun. 2014;82(1):221–232. doi:10.1128/IAI.01304-13
112. Abrahams GL, Hensel M. Manipulating cellular transport and immune responses: dynamic interactions between intracellular Salmonella enterica and its host cells. Cell Microbiol. 2006;8(5):728–737. doi:10.1111/cmi.2006.8.issue-5
113. Henry T, Couillault C, Rockenfeller P, et al. The Salmonella effector protein PipB2 is a linker for kinesin-1. Proc National Acad Sci. 2006;103(36):13497–13502. doi:10.1073/pnas.0605443103
114. Knodler LA, Celli J, Hardt W-D, Vallance BA, Yip C, Finlay BB. Salmonella effectors within a single pathogenicity island are differentially expressed and translocated by separate type III secretion systems. Mol Microbiol. 2002;43(5):1089–1103. doi:10.1046/j.1365-2958.2002.02820.x
115. Miao EA, Mao DP, Yudkovsky N, et al. Innate immune detection of the type III secretion apparatus through the NLRC4 inflammasome. Proc Natl Acad Sci U S A. 2010;107(7):3076–3080. doi:10.1073/pnas.0913087107
116. Reyes Ruiz VM, Ramirez J, Naseer N, et al. Broad detection of bacterial type III secretion system and flagellin proteins by the human NAIP/NLRC4 inflammasome. Proc Natl Acad Sci U S A. 2017;114(50):13242–13247. doi:10.1073/pnas.1710433114
117. Karki R, Lee E, Place D, et al. IRF8 regulates transcription of naips for NLRC4 inflammasome activation. Cell. 2018;173(4):920–933.e913. doi:10.1016/j.cell.2018.02.055
118. Cummings LA, Barrett SLR, Wilkerson WD, Fellnerova I, Cookson BT. FliC-specific CD4+ T cell responses are restricted by bacterial regulation of antigen expression. J Immunol. 2005;174(12):7929–7938. doi:10.4049/jimmunol.174.12.7929
119. Miao EA, Leaf IA, Treuting PM, et al. Caspase-1-induced pyroptosis is an innate immune effector mechanism against intracellular bacteria. Nat Immunol. 2010;11:1136. doi:10.1038/ni.1960
120. Humphries F, Yang S, Wang B, Moynagh P. RIP kinases: key decision makers in cell death and innate immunity. Cell Death Differ. 2015;22(2):225. doi:10.1038/cdd.2014.126
121. Behnsen J, Perez-Lopez A, Nuccio S-P, Raffatellu M. Exploiting host immunity: the Salmonella paradigm. Trends Immunol. 2015;36(2):112–120. doi:10.1016/j.it.2014.12.003
122. McCoy MW, Moreland SM, Detweiler CS. Hemophagocytic macrophages in murine typhoid fever have an anti-inflammatory phenotype. Infect Immun. 2012;80(10):3642–3649. doi:10.1128/IAI.00656-12
123. Libby SJ, Lesnick M, Hasegawa P, Weidenhammer E, Guiney DG. The Salmonella virulence plasmid spv genes are required for cytopathology in human monocyte-derived macrophages. Cell Microbiol. 2000;2(1):49–58. doi:10.1046/j.1462-5822.2000.00030.x
124. Browne SH, Lesnick ML, Guiney DG. Genetic requirements for salmonella-induced cytopathology in human monocyte-derived macrophages. Infect Immun. 2002;70(12):7126–7135. doi:10.1128/IAI.70.12.7126-7135.2002
125. Haneda T, Ishii Y, Shimizu H, et al. Salmonella type III effector SpvC, a phosphothreonine lyase, contributes to reduction in inflammatory response during intestinal phase of infection. Cell Microbiol. 2012;14(4):485–499. doi:10.1111/j.1462-5822.2011.01733.x
126. Mazurkiewicz P, Thomas J, Thompson JA, et al. SpvC is a Salmonella effector with phosphothreonine lyase activity on host mitogen-activated protein kinases. Mol Microbiol. 2008;67(6):1371–1383. doi:10.1111/mmi.2008.67.issue-6
127. Singh Y, Saxena A, Kumar R, Saxena MK. Virulence system of salmonella with special reference to Salmonella enterica. Salmonella. 2018;41.
128. Kim Y-G, Kim J-H, Kim K-J. Crystal structure of the Salmonella enterica serovar typhimurium virulence factor SrfJ, a glycoside hydrolase family enzyme. J Bacteriol. 2009;191(21):6550–6554. doi:10.1128/JB.00641-09
129. Bishop AL, Hall A. Rho GTPases and their effector proteins. Biochem J. 2000;348(Pt 2):241–255. doi:10.1042/bj3480241
130. Friebel A, Ilchmann H, Aepfelbacher M, Ehrbar K, Machleidt W, Hardt W-D. SopE and SopE2 from Salmonella typhimurium activate different sets of RhoGTPases of the host cell. J Biol Chem. 2001;276(36):34035–34040. doi:10.1074/jbc.M100609200
131. Hardt WD, Chen LM, Schuebel KE, Bustelo XR, Galan JE. S. typhimurium encodes an activator of Rho GTPases that induces membrane ruffling and nuclear responses in host cells. Cell. 1998;93(5):815–826. doi:10.1016/S0092-8674(00)81442-7
132. Keestra AM, Winter MG, Auburger JJ, et al. Manipulation of small Rho GTPases is a pathogen-induced process detected by NOD1. Nature. 2013;496(7444):233. doi:10.1038/nature12025
133. Johnson R, Byrne A, Berger CN, et al. The type III secretion system effector SptP of Salmonella enterica serovar Typhi. J Bacteriol. 2017;199(4):e00647–e00616. doi:10.1128/JB.00647-16
134. Pearson G, Robinson F, Beers Gibson T, et al. Mitogen-activated protein (MAP) kinase pathways: regulation and physiological functions. Endocr Rev. 2001;22(2):153–183. doi:10.1210/edrv.22.2.0428
135. Lin SL, Le TX, Cowen DS. SptP, a Salmonella typhimurium type III-secreted protein, inhibits the mitogen-activated protein kinase pathway by inhibiting Raf activation. Cell Microbiol. 2003;5(4):267–275. doi:10.1046/j.1462-5822.2003.t01-1-00274.x
136. Arricau N, Hermant D, Waxin H, Popoff M. Molecular characterization of the Salmonella typhi StpA protein that is related to both Yersinia YopE cytotoxin and YopH tyrosine phosphatase. Res Microbiol. 1997;148(1):21–26. doi:10.1016/S0923-2508(97)81896-7
137. Sabbagh SC, Forest CG, Lepage C, Leclerc J-M, Daigle F. So similar, yet so different: uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi. FEMS Microbiol Lett. 2010;305(1):1–13. doi:10.1111/fml.2010.305.issue-1
138. McIntosh A, Meikle LM, Ormsby MJ, et al. SipA activation of caspase-3 is a decisive mediator of host cell survival at early stages of Salmonella enterica serovar Typhimurium infection. Infect Immun. 2017;85(9):e00393–e00317. doi:10.1128/IAI.00393-17
139. Singh PK, Kapoor A, Lomash RM, et al. Salmonella SipA mimics a cognate SNARE for host Syntaxin8 to promote fusion with early endosomes. J Cell Biol. 2018;217(12):4199. doi:10.1083/jcb.201802155
140. Xu J, Luo F, Zhang Z, et al. SNARE proteins synaptobrevin, SNAP-25, and syntaxin are involved in rapid and slow endocytosis at synapses. Cell Rep. 2013;3(5):1414–1421. doi:10.1016/j.celrep.2013.03.010
141. Hallstrom KN, McCormick BA. The type three secreted effector SipC regulates the trafficking of PERP during Salmonella infection. Gut Microbes. 2016;7(2):136–145. doi:10.1080/19490976.2015.1128626
142. Hallstrom KN, Srikanth C, Agbor TA, et al. PERP, a host tetraspanning membrane protein, is required for S almonella‐induced inflammation. Cell Microbiol. 2015;17(6):843–859. doi:10.1111/cmi.2015.17.issue-6
143. Lara-Tejero M, Galán JE. Salmonella enterica serovar typhimurium pathogenicity island 1-encoded type III secretion system translocases mediate intimate attachment to nonphagocytic cells. Infect Immun. 2009;77(7):2635–2642. doi:10.1128/IAI.00077-09
144. Eswarappa SM, Janice J, Nagarajan AG, et al. Differentially evolved genes of Salmonella pathogenicity islands: insights into the mechanism of host specificity in Salmonella. PLoS One. 2008;3(12):e3829. doi:10.1371/journal.pone.0003829
145. Shivcharan S, Yadav J, Qadri A. Host lipid sensing promotes invasion of cells with pathogenic Salmonella. Sci Rep. 2018;8(1):15501. doi:10.1038/s41598-018-33319-9
146. Knodler LA, Nair V, Steele-Mortimer O. Quantitative assessment of cytosolic Salmonella in epithelial cells. PLoS One. 2014;9(1):e84681. doi:10.1371/journal.pone.0084681
147. Jones RM, Wu H, Wentworth C, Luo L, Collier-Hyams L, Neish AS. Salmonella AvrA coordinates suppression of host immune and apoptotic defenses via JNK pathway blockade. Cell Host Microbe. 2008;3(4):233–244. doi:10.1016/j.chom.2008.02.016
148. Fedor Y, Vignard J, Nicolau-Travers ML, et al. From single-strand breaks to double-strand breaks during S-phase: a new mode of action of the Escherichia coli cytolethal distending toxin. Cell Microbiol. 2013;15(1):1–15. doi:10.1111/cmi.12028
149. Guidi R, Guerra L, Levi L, et al. Chronic exposure to the cytolethal distending toxins of Gram-negative bacteria promotes genomic instability and altered DNA damage response. Cell Microbiol. 2013;15(1):98–113. doi:10.1111/cmi.12034
150. Del Bel Belluz L, Guidi R, Pateras IS, et al. The typhoid toxin promotes host survival and the establishment of a persistent asymptomatic infection. PLoS Pathog. 2016;12(4):e1005528–e1005528. doi:10.1371/journal.ppat.1005528
151. Chong A, Lee S, Yang Y-A, Song J. The role of typhoid toxin in Salmonella typhi virulence. Yale J Biol Med. 2017;90(2):283–290.
152. Haghjoo E, Galan JE. Salmonella typhi encodes a functional cytolethal distending toxin that is delivered into host cells by a bacterial-internalization pathway. Proc Natl Acad Sci U S A. 2004;101(13):4614–4619. doi:10.1073/pnas.0400932101
153. Spanò S, Ugalde JE, Galán JE. Delivery of a Salmonella Typhi exotoxin from a host intracellular compartment. Cell Host Microbe. 2008;3(1):30–38.
154. Eichelberg K, Galán JE. Differential regulation of Salmonella typhimurium type III secreted proteins by pathogenicity island 1 (SPI-1)-encoded transcriptional activators InvF and HilA. Infect Immun. 1999;67(8):4099.
155. Ellermeier JR, Slauch JM. Adaptation to the host environment: regulation of the SPI1 type III secretion system in Salmonella enterica serovar Typhimurium. Curr Opin Microbiol. 2007;10(1):24–29. doi:10.1016/j.mib.2006.12.002
156. Olekhnovich IN, Kadner RJ. Crucial roles of both flanking sequences in silencing of the hilA promoter in Salmonella enterica. J Mol Biol. 2006;357(2):373–386. doi:10.1016/j.jmb.2006.01.007
157. Fabrega A, Vila J. Salmonella enterica serovar Typhimurium skills to succeed in the host: virulence and regulation. Clin Microbiol Rev. 2013;26(2):308–341.
158. Ellermeier CD, Ellermeier JR, Slauch JM. HilD, HilC and RtsA constitute a feed forward loop that controls expression of the SPI1 type three secretion system regulator hilA in Salmonella enterica serovar Typhimurium. Mol Microbiol. 2005;57(3):691–705. doi:10.1111/mmi.2005.57.issue-3
159. Kenney LJ. The role of acid stress in Salmonella pathogenesis. Curr Opin Microbiol. 2019;47:45–51. doi:10.1016/j.mib.2018.11.006
160. Cheeseman G, Fuller R. Changes in the pH activity profile of the lysine decarboxylase during incubation of Escherichia coli. J Appl Bacteriol. 1968;31(2):253–258. doi:10.1111/jam.1968.31.issue-2
161. Crawford RW, Rosales-Reyes R, de la Luz Ramírez-aguilar M, Chapa-Azuela O, Alpuche-Aranda C, Gunn JS. Gallstones play a significant role in Salmonella spp. gallbladder colonization and carriage. Proc National Acad Sci. 2010;107(9):4353–4358. doi:10.1073/pnas.1000862107
162. Quinn HJ, Cameron AD, Dorman CJ. Bacterial regulon evolution: distinct responses and roles for the identical OmpR proteins of Salmonella Typhimurium and Escherichia coli in the acid stress response. PLoS Genet. 2014;10(3):e1004215. doi:10.1371/journal.pgen.1004215
163. Bustamante VH, Martínez LC, Santana FJ, Knodler LA, Steele-Mortimer O, Puente JL. HilD-mediated transcriptional cross-talk between SPI-1 and SPI-2. Proc National Acad Sci. 2008;105(38):14591–14596. doi:10.1073/pnas.0801205105
164. Pérez-Morales D, Banda MM, Chau NYE, et al. The transcriptional regulator SsrB is involved in a molecular switch controlling virulence lifestyles of Salmonella. PLoS Pathog. 2017;13(7):e1006497. doi:10.1371/journal.ppat.1006497
165. Gantois I, Ducatelle R, Pasmans F, et al. Butyrate specifically down-regulates salmonella pathogenicity island 1 gene expression. Appl Environ Microbiol. 2006;72(1):946. doi:10.1128/AEM.72.1.946-949.2006
166. Bronner DN, Faber F, Olsan EE, et al. Genetic ablation of butyrate utilization attenuates gastrointestinal Salmonella disease. Cell Host Microbe. 2018;23(2):266–273.e264. doi:10.1016/j.chom.2018.01.004
167. Wick MJ. The role of dendritic cells during Salmonella infection. Curr Opin Immunol. 2002;14(4):437–443. doi:10.1016/S0952-7915(02)00364-3
168. Cheminay C, Schoen M, Hensel M, et al. Migration of Salmonella typhimurium –harboring bone marrow–derived dendritic cells towards the chemokines CCL19 and CCL21. Microb Pathog. 2002;32(5):207–218. doi:10.1006/mpat.2002.0497
169. Carden SE, Walker GT, Honeycutt J, et al. Pseudogenization of the secreted effector gene sseI confers rapid systemic dissemination of S. Typhimurium ST313 within migratory dendritic cells. Cell Host Microbe. 2017;21(2):182–194. doi:10.1016/j.chom.2017.01.009
170. Reyes M, Fisher S, McLaughlin LM, et al. A microfluidic-based genetic screen to identify microbial virulence factors that inhibit dendritic cell migration. Integr Biol. 2014;6(4):438–449. doi:10.1039/C3IB40177D
171. Riol-Blanco L, Sanchez-Sanchez N, Torres A, et al. The chemokine receptor CCR7 activates in dendritic cells two signaling modules that independently regulate chemotaxis and migratory speed. J Immunol. 2005;174(7):4070–4080. doi:10.4049/jimmunol.174.7.4070
172. Miao EA, Brittnacher M, Haraga A, Jeng RL, Welch MD, Miller SI. Salmonella effectors translocated across the vacuolar membrane interact with the actin cytoskeleton. Mol Microbiol. 2003;48(2):401–415. doi:10.1046/j.1365-2958.2003.t01-1-03456.x
173. McLaughlin LM, Govoni GR, Gerke C, et al. The Salmonella SPI2 effector SseI mediates long-term systemic infection by modulating host cell migration. PLoS Pathog. 2009;5(11):e1000671. doi:10.1371/journal.ppat.1000671
174. Arpaia N, Godec J, Lau L, et al. TLR signaling is required for Salmonella typhimurium virulence. Cell. 2011;144(5):675–688. doi:10.1016/j.cell.2011.01.031
175. Iwasaki A, Medzhitov R. Toll-like receptor control of the adaptive immune responses. Nat Immunol. 2004;5(10):987. doi:10.1038/ni1112
176. Hsu LC, Park JM, Zhang K, et al. The protein kinase PKR is required for macrophage apoptosis after activation of Toll-like receptor 4. Nature. 2004;428(6980):341–345. doi:10.1038/nature02405
177. Park JM, Greten FR, Li ZW, Karin M. Macrophage apoptosis by anthrax lethal factor through p38 MAP kinase inhibition. Science. 2002;297(5589):2048–2051. doi:10.1126/science.1073163
178. Forstner J, Oliver M, Sylvester F. Production, structure and biologic relevance of gastrointestinal mucins. Infect Gastrointest Tract. 1995;1995:71–88.
179. Vimal DB, Khullar M, Gupta S, Ganguly NK. Intestinal mucins: the binding sites for Salmonella typhimurium. Mol Cell Biochem. 2000;204(1–2):107–117. doi:10.1023/A:1007015312036
180. Arabyan N, Weis AM, Huang BC, Weimer BC. Implication of sialidases in Salmonella infection: genome release of sialidase knockout strains from Salmonella enterica serovar typhimurium LT2. Genome Announc. 2017;5(19):e00341–e00317. doi:10.1128/genomeA.00341-17
181. Songhet P, Barthel M, Stecher B, et al. Stromal IFN-γR-signaling modulates goblet cell function during Salmonella typhimurium infection. PLoS One. 2011;6(7):e22459. doi:10.1371/journal.pone.0022459
182. Johansson C, Ingman M, Wick MJ. Elevated neutrophil, macrophage and dendritic cell numbers characterize immune cell populations in mice chronically infected with Salmonella. Microb Pathog. 2006;41(2–3):49–58. doi:10.1016/j.micpath.2006.03.004
183. Broz P, Newton K, Lamkanfi M, Mariathasan S, Dixit VM, Monack DM. Redundant roles for inflammasome receptors NLRP3 and NLRC4 in host defense against Salmonella. J Exp Med. 2010;207(8):1745. doi:10.1084/jem.20100257
184. De Groote MA, Ochsner UA, Shiloh MU, et al. Periplasmic superoxide dismutase protects Salmonella from products of phagocyte NADPH-oxidase and nitric oxide synthase. Proc Natl Acad Sci U S A. 1997;94(25):13997–14001. doi:10.1073/pnas.94.25.13997
185. Godinez I, Raffatellu M, Chu H, et al. Interleukin-23 orchestrates mucosal responses to Salmonella enterica serotype typhimurium in the intestine. Infect Immun. 2009;77(1):387.
186. Zindl CL, Lai J-F, Lee YK, et al. IL-22–producing neutrophils contribute to antimicrobial defense and restitution of colonic epithelial integrity during colitis. Proc National Acad Sci. 2013;110(31):12768–12773. doi:10.1073/pnas.1300318110
187. Sutton CE, Lalor SJ, Sweeney CM, Brereton CF, Lavelle EC, Mills KH. Interleukin-1 and IL-23 induce innate IL-17 production from γδ T cells, amplifying Th17 responses and autoimmunity. Immunity. 2009;31(2):331–341. doi:10.1016/j.immuni.2009.08.001
188. Mooney JP, Galloway LJ, Riley EM. Malaria, anemia, and invasive bacterial disease: a neutrophil problem? J Leukoc Biol. 2019;105(4):645–655. doi:10.1002/jlb.2019.105.issue-4
189. Corbin BD, Seeley EH, Raab A, et al. Metal chelation and inhibition of bacterial growth in tissue abscesses. Science. 2008;319(5865):962. doi:10.1126/science.1152449
190. Diaz-Ochoa Vladimir E, Lam D, Lee Carlin S, et al. Salmonella mitigates oxidative stress and thrives in the inflamed gut by evading calprotectin-mediated manganese sequestration. Cell Host Microbe. 2016;19(6):814–825. doi:10.1016/j.chom.2016.05.005
191. Lin D, Kim B, Slauch JM. DsbL and DsbI contribute to periplasmic disulfide bond formation in Salmonella enterica serovar typhimurium. Microbiology. 2009;155(Pt 12):4014–4024. doi:10.1099/mic.0.032904-0
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
