Back to Journals » Infection and Drug Resistance » Volume 12
Molecular epidemiology and resistance profiles among healthcare- and community-associated Staphylococcus aureus keratitis isolates
Authors Peterson JC , Durkee H , Miller D , Maestre-Mesa J , Arboleda A, Aguilar MC , Relhan N, Flynn Jr HW , Amescua G, Parel JM , Alfonso E
Received 9 October 2018
Accepted for publication 18 January 2019
Published 11 April 2019 Volume 2019:12 Pages 831—843
DOI https://doi.org/10.2147/IDR.S190245
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Jeffrey C Peterson,1,2 Heather Durkee,1,2 Darlene Miller,3,4 Jorge Maestre-Mesa,3,4 Alejandro Arboleda,1 Mariela C Aguilar,1 Nidhi Relhan,1 Harry W Flynn Jr,3 Guillermo Amescua,3 Jean-Marie Parel,1–3 Eduardo Alfonso3
1Ophthalmic Biophysics Center, Department of Ophthalmology, Bascom Palmer Eye Institute, University of Miami Miller School of Medicine, Miami, FL, USA; 2Department of Biomedical Engineering, University of Miami, Coral Gables, FL, USA; 3Anne Bates Leach Eye Center, Department of Ophthalmology, Bascom Palmer Eye Institute, University of Miami Miller School of Medicine, Miami, FL, USA; 4Ocular Microbiology Laboratory, Department of Ophthalmology, Bascom Palmer Eye Institute, University of Miami Miller School of Medicine, Miami, FL, USA
Purpose: To characterize the molecular, epidemiological, and resistance profiles of methicillin-resistant (MRSA) and methicillin-susceptible (MSSA) keratitis isolates.
Patients and methods: We used a combination of standard microbiological techniques and DNA microarray analysis to characterize the molecular and antibiotic resistance profiles of 75 Staphylococcus aureus keratitis isolates collected over an 11-year period (2006–2016).
Results: Two major USA clonal complexes (CC), CC5 (n=30, 40%) and CC8 (n=28, 37.3%), accounted for 77.3% of the collected S. aureus isolates. USA100, traditionally healthcare associated (n=18/47, 38.3%), and USA300, traditionally community associated (n=12/47, 25.5%), were the dominant MRSA strains. Four (22.2%) of the USA100 MRSA isolates were recovered from patients with no prior healthcare exposure. Eleven (91.7%) of the USA300 isolates were recovered from patients with documented healthcare risk factors. MSSA isolates were polyclonal (n=13). Ninety-three percent of MSSA infections were of healthcare origin. Thirty-seven of 61 (60.6%) healthcare- and 11 of 14 (78.6%) community-associated strains were resistant to three or more antibiotic classes. Sixty-eight percent (n=51) of isolates harbored three of more resistance determinants (genes). The Panton-Valentine Leucocidin gene was detected in 11 (14.7%) of the study isolates. The majority (72.7%) of the strains were members of the USA300 MRSA clone.
Conclusion: Clonal complexes CC5 and CC8 were the most frequent clones detected among both the MSSA and the MRSA keratitis isolates. USA100 and USA300 clones were the dominant MRSA genotypes. The USA300 MRSA clone has become a leading cause of healthcare-associated keratitis in South Florida. The USA100 MRSA clone has emerged as an increasing cause of community-associated corneal infections in our outpatient population. This shifting epidemiology coupled with the increasing prevalence of multidrug resistance among both MSSA and MRSA keratitis is a cause of concern.
Keywords: MRSA, MSSA, DNA microarray, USA100, USA300, clones
Introduction
Staphylococcus aureus keratitis is a significant clinical and public health concern in the United States and worldwide.1–3 S. aureus is the second most frequently reported cause of bacterial keratitis with an estimated prevalence of 3%–49%. Methicillin-susceptible (MSSA) and methicillin-resistant (MRSA) S. aureus infections are more common following corneal transplants, contact lens wear, and cataract surgery. Rates differ by region, population sampled, and study period. Disease severity and patient outcomes are strain specific.4–6
In addition, increasing antibiotic resistance to fluoroquinolones and trimethoprim–sulfamethoxazole among both MSSA and MRSA keratitis isolates poses a growing therapeutic dilemma.20,21 S. aureus strains differ in molecular characteristics, virulence factors, and antibiotic resistance profiles. Several studies have linked specific S. aureus clonal groups and lineages with particular infections, patient outcomes, and antibiotic resistance profiles.7–10
S. aureus isolates have been grouped into ten major lineages based on a combination SCCmec, pulsed field gel electrophoresis (PFGE), multilocus sequencing (MLST), and sequence and spa types.11–14 Globally, S. aureus genotypes most frequently associated with invasive disease and postsurgical infections include clonal complex 5 (CC5)/USA100, CC30/USA200, CC8/USA300, CC1/USA400, and CC45/USA600.15,16 The most frequent healthcare-associated strains in the United States are USA100, USA200, USA500, USA600, and USA800.12,14,17,18 Healthcare-associated MRSA (HA-MRSA) strains historically contained the staphylococcal chromosomal cassette mecA (SCCmecA) genotypes I–III. The USA100 MRSA clone is the dominant healthcare-associated lineage and the leading cause of invasive infections in hospitalized patients. Community-associated MRSA (CA-MRSA) strains traditionally contained the SCCmecA IV or VII genotypes.12,18 The USA300 MRSA clone is the leading cause of severe community-associated soft tissue and skin infections. Genotype and resistance profiles differ for healthcare- and community-associated S. aureus strains but the distinctions between the two are beginning to blur.10,19
Few studies are available documenting the prevalence and diversity of clonal structure, virulence factors, and antibiotic resistance profiles among healthcare- or community-associated MRSA and MSSA keratitis isolates.5,21–26 A better understanding of the molecular epidemiology and resistance profiles of S. aureus keratitis infections is essential to help decipher the changing pathology and provide further insight into selection of optimal therapy. The purpose of this study was to investigate and provide data on the prevalence and diversity of S. aureus genotypes and antimicrobial resistance profiles for a convenient set of S. aureus keratitis isolates in South Florida.
Patients and methods
We used a combination of standard microbiological techniques and DNA microarray technology to characterize the epidemiology, molecular, and antibiotic resistance profiles for 47 MRSA and 28 MSSA keratitis isolates (n=75). This convenient subset of 75 isolates was selected from a group of 467 S. aureus corneal pathogens collected over an 11-year period, 2006–2016. Isolates were collected from corneal scrapings of patients presenting to our emergency room (n=45), outpatient clinics (n=27), or hospitalized patients (n=3). Standard microbiological procedures including beta hemolysis, coagulase tests, and cefoxitin screens were used to provide isolate identification and phenotype.2 Phenotypes were confirmed and antibiotic susceptibility patterns were determined using the Vitek 2 automated system (BioMerieux, Durham, NC, USA). Assignment of susceptible or resistance phenotype was based on standards published by the Clinical Laboratory Standards Institute.27
The DNA Microarray (Alere StaphyType, Alere, Technologies GmbH, Jena, Germany) was used to genotype and classify isolates according to matched probe hybridization profiles as previously described.13 Briefly, the isolates are enzymatically digested and then subjected to DNA purification. A linear primer elongation reaction then amplifies and labels target sequences. This creates ssDNA aplicons that are biotin labeled and hybridized to a microarray patterned with desired oligonucleotides. DNA hybridization is then visualized using a streptavidin–horseradish-peroxidase conjugate that causes a subsequent precipitation of dye. The microarray is scanned and compared to the background signal.13 The array is a unique system that uses 333 target sequences (probes) corresponding to 170 distinct genes and their allelic variants. This allows for simultaneous detection and identification of species markers, SCCmec elements, toxins, CCs, virulence factors, and antibiotic resistance determinants (genes).28,29
Keratitis isolates were defined as healthcare associated or community associated based on epidemiology and prior healthcare exposure.10,30 Healthcare-associated S. aureus isolates were identified as those recovered from patients with prior exposure to the healthcare system. Known risk factors among this group include ocular surface disease, prior ocular surgery, prior/prolonged topical antibiotic use, or residence in long-term care. Healthcare-associated isolates were further categorized as either healthcare-associated hospital onset (HAHO), ie, those originating in the hospital, or healthcare-associated community onset (HACO), ie, those originating from the community. Community-associated S. aureus keratitis isolates were those collected from patients with no recognizable healthcare exposure or known risk factors for microbial keratitis.31–33
Statistics
Chi-squared tests and independent two-sample t-tests were used for comparisons between groups (MRSA vs MSSA, healthcare vs community associated). The McNemar test was used to compare phenotypic vs genotypic resistance to antimicrobials. The calculations were performed using Excel 2013 (Redman, WA, USA) and SPSS version 24 (IBM Corporation, Chicago, IL, USA). P-values <0.05 were considered statistically significant.
Results
Patient and isolate characterization
Patient age, gender, and isolate characteristics are demonstrated in Table 1. Among the 75 nonconsecutive S. aureus corneal isolates included in the study, 47 (62.7%) were phenotyped as MRSA and the remaining 28 (37.3%) as MSSA.
Figure 1 highlights general trends in S. aureus keratitis over a 15-year period. There was a gradual increase in the percent of corneal ulcers caused by S. aureus for each 5-year period; rates of S. aureus keratitis ranged from 11.8% (2002–2006) to 17.4% (2012–2016).
Our study population included 40 females (53.3%) and 35 males (46.7%). Patient age (n=60) ranged from 22 to 93 years with a mean of 65.1 years and a median of 67.5 years. Females (median age 74.5 years) were generally older than males (median age 56 years) and more likely to have MRSA (females: 31/40, 77.5% vs males: 16/35, 47.5%; P=0.0048, 95% CI: 9.79–50.13, chi-squared test). Patients with MSSA keratitis were younger (median age 60 years) and more likely to be males (males: 19/28, 67.9% vs females: 9/28, 32.1%, P=0.0079, 95% CI: 9.61–55.88, chi-squared test). The ages of 15 (20%) patients were not available; 7 of these 15 patients (46.7%) were males and 8 of 15 (53.3%) were females.
Patient setting
S. aureus isolates were significantly more likely to be recovered from patients presenting in the emergency room (n=45/75, 60%) than from patients visiting our outpatient clinics (n=27, 36%, P=0.0034, 95% CI: 8.01–38.34, chi-squared test). Twenty-two of the 27 (81.5%) isolates recovered from patients in our outpatient clinics were reported by the corneal specialty service. The remaining three (4%) isolates were isolated from hospitalized patients (Table 1).
Clinical presentation
Corneal ulcers were the most frequently presenting clinical diagnoses (n=30/75, 40%) followed by postsurgical infections (n=23/75, 30.7%). Twelve of 23 (52%) isolates from postsurgical infections were recovered following corneal transplants (Table 1).
S. aureus isolates recovered from corneal wounds/ulcers were more likely to be methicillin-resistant (n=23/30, 76.7%) than methicillin-susceptible (n=7/30, 23.3%, P<0.0001 95% CI: 28.52–69.71, chi-squared test). There was no significant difference between the proportion of MRSA isolates (n=12/23, 52.2%) and MSSA (n=11/23, 47.8%) isolates (P=0.9163, 95% CI: –19.3120–21.3182, chi-squared test) recovered from patients with postsurgical infections.
Healthcare- vs community-associated keratitis
Healthcare-associated strains constituted 61/75 (81.3%) of all S. aureus isolates. The majority of these (58/61, 95.1%) were from the community and classified as HACO. The remaining three (4.9%) isolates were recovered from hospitalized patients and classified as HAHO. Fourteen of the 75 (18.7%) isolates were recovered from outpatients with no known healthcare exposure or microbial risk factors. These strains were classified as community associated.10,34
The majority of the MSSA keratitis isolates (n=25/28, 89.3%) were more likely to be classified as HACO than were MRSA isolates (n=33/47, 70.2%, P=0.0577, 95% CI: –0.79–34.92, chi-squared test). Two of the three (66.7%) S. aureus isolates recovered from hospitalized patients were MRSA and both classified as HACO.
Molecular profile
The DNA microarray identified 13 CCs, 29 strains, and 4 USA PFGE types among this group of isolates (Table 2).
Two major USA CCs, CC5 (traditionally healthcare associated, n=30/75, 40%) and CC8 (traditionally community associated, n=28/75, 37.3%), accounted for 77.3% of the total keratitis isolates. The 47 MRSA strains clustered within the two major clones. Twenty-four or 51.1% of the isolates were identified as members of the CC5 clonal complex. The remaining 23 or 48.9% of the MRSA were classified as members of the CC8 clonal complex.
DNA microarray analysis confirmed the mecA genotype in 36/47 (76.6%) S. aureus isolates identified phenotypically as MRSA. Isolates designated as MRSA by DNA microarray (ie, mecA was detected) were split evenly between the CC5 clonal lineage (n=18/36, 50%) and the CC8 clonal lineage (n=18/36, 50%). Among the 18 CC5 isolates, 15/18 (83.3%) were confirmed as USA100 (traditionally HA-MRSA) and members of the ST5/225 New York–Japan clone. These possessed the classic molecular fingerprint of CC5/SCCmecII/ST5/t002 (Table 2). The remaining three CC5-MRSA isolates (2.1%) included two CC5-MRSA-[II + ACME] clones and a single CC5-MRSA-[II + ccrC] strain.
Among the 18 CC8-MRSA strains, 9/18 (50%) were confirmed as USA300 (traditionally CA-MRSA) with the classic molecular fingerprint of CC8/SCCmecIV/ST8/t008. Additional USA-confirmed genotypes/lineages in CC8 included 4/18 (22.2%) USA500/SCCmecIV/ST8/t064 and one (5.5%) USA700/SCCmec IV/ST72/t126 isolate. The remaining four CC8-MRSA strains were epidemic MRSA stains identified as CC8-MRSA-IV-UK-EMRSA14 (n=3) and CC8-MRSA-IV, (SEA+), Lyon Clone (n=1).
Among the eleven discordant MRSA results (methicillin-resistant with no mecA gene detected), six isolates (54.5%) were identified as USA100 (n=3) or USA300 (n=3) deletion mutants. The remaining five isolates were classified as MSSA (Table 2).
The 28 MSSA isolates were distributed across all 13 identified CCs. Five of these clonal complexes (CC1, CC5, CC8, CC15, and CC45) were associated with >1 MSSA isolate and accounted for 20/28 (70.4%) of the strains. Eleven of 28 (39.3%) of MSSA strains were identified as members of either CC5 (n=6, 21.4%) or CC8 (n=5, 18.5%).
Virulence factors
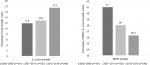
Figure 2 highlights select virulence factors from the 75 S. aureus isolates. The Panton-Valentine Leucocidin (PVL) toxin (n=11, 14.7%) and S. aureus enterotoxin A gene (SEA, n=7, 9.3%) were both detected in MSSA and MRSA isolates. The SEA-positive isolates were most frequently associated with USA500 MRSA isolates. The Arginine Catabolic Mobile Element (ACME) was found exclusively in CC8-MRSA isolates. The USA300 MRSA clone was associated with PVL and the presence of the ACME cluster. All three genes (PVL, ACME, and enterotoxin A) were absent in the USA100 strains.
Genotype: healthcare vs community associated
Five of the 24 (20.8%) CC5-MRSA isolates were recovered from community-associated infections while 15 of 23 (66.2%) of the CC8-MRSA isolates were recovered from infections of healthcare-associated origin. Four of the 18 (22.2%) MRSA isolates with the USA100 fingerprint were recovered from outpatients with no known healthcare exposure or risk factors for keratitis. Eleven of the 12 (91.7%) MRSA isolates characterized as USA300 were recovered from patients with known ocular healthcare risk factors. Ten of the 11 (90.9%) MRSA-USA300 isolates were classified as HACO, and one was classified as HAHO.
Resistance profiles
Tables 3–5 highlight phenotypic and genotypic antibiotic resistance prevalence and patterns for the 75 S. aureus keratitis isolates. Phenotypic susceptibility profiles were determined for 13 antibiotics representing 11 unique antimicrobial drug classes (Table 3). Phenotypic resistance to at least one antibiotic was detected in 63 (84%) of the 75 S. aureus isolates. Across all of these isolates, there were >90% in vitro susceptibility for gentamicin (92%), linezolid (100%), and vancomycin (100%). Susceptibility for moxifloxacin and trimethoprim-sulfamethoxazole were 33% and 89.3%, respectively.
Resistance to clindamycin (P<0.0001, 95% CI: 44.69–81.58, chi-squared test) and erythromycin (P=0.00175, 95% CI: 3.56–40.25, chi-squared test) were higher for CC5 isolates (90% and 96.7%, respectively) than were resistance for CC8 isolates (21.4% and 75%, respectively). All USA100 and USA300 strains were resistant to erythromycin (Table 3).
HACO strains were more likely to be resistant to erythromycin (43/58 or 74.1%) than community-associated isolates (9/14 or 64.2%). The rate of clindamycin resistance among USA100-MRSA isolates (18/18 or 100%) was approximately two times the rate documented for the USA300-MRSA strains (5/12 or 41.7%).
Multidrug resistance, defined as resistance to three or more classes of antibiotics, was documented for 48 or 64% of all isolates. MRSA strains (n=43/47, 91.5%) were significantly more likely to be multidrug resistant than MSSA strains (n=5/28, 17.8%, P<0.0001, 95% CI: 52.71–84.91, chi-squared test). The most common phenotypic resistance pattern included co-resistance to oxacillin, ciprofloxacin, clindamycin, erythromycin, and moxifloxacin (n=24/75 or 32%, Table 5).
The DNA microarray assay screened for the presence of 32 different resistance genes, representing 12 different antibiotic classes. Fourteen different resistance genes targeting at least eight antibiotic classes were detected among both MSSA and MRSA populations (Table 5).
At least one antibiotic resistance gene was detected in 72 of the 75 (96%) total isolates. Three or more antibiotic resistance genes were documented in 51 (68%) of the strains. Seventy-eight percent of the isolates carried the blaZ gene; there was a similar distribution of the blaZ gene among MSSA (75%) and MRSA (80%) isolates. A high rate of genotypic resistance (88%) to fosfomycin (fosB) was detected among all isolates.
Genes conferring resistance to the glycopeptides (vancomycin: vanA, vanB; teicoplanin: vanZ), chloramphenicol (cat, fexA), oxazolidinones (linezolid: cfr), and fusidic acid (far1) were not detected among the 75 S. aureus isolates. The microarray did not include probes for the detection of fluoroquinolone resistance.
Correlation between phenotype and genotype
Overall agreement between phenotypic and genotypic antibiotic resistance was 94.6% with a sensitivity of 82.6% and a specificity of 99.3% (Table 6).
Significant disagreement (P=0.0492, 95% CI: 0.14–30.77, McNemar’s test) was observed only for the oxacillin/mecA pair. Forty-seven or 62.7% of all 75 S. aureus isolates were resistant to methicillin by phenotype while presence of the mecA gene was confirmed in only 36 (48%) of isolates (P=0.003, 95% CI: 2.51 to Infinity, McNemar’s test). Agreement between the remaining antibiotic/gene pairs ranged from 89.3% to 97.3% (Table 6).
Discussion
Few studies are available documenting the genomic, phenotypic, demographic, and antibiotic profiles of healthcare- and community-associated MSSA and MRSA keratitis. This information is important because of the increasing recovery of MSSA and MRSA from surgical and nonsurgical infections. Additionally, this information is essential to help document the evolution and clonality of ocular S. aureus and their resistant profiles.3,35
To our knowledge, this is the first report confirming the presence of USA100-MRSA and USA300-MRSA clones among S. aureus keratitis isolates in the United States. These two strains were the predominant clones circulating among keratitis isolates in South Florida.
Seventy-seven percent of the 18 USA100-MRSA strains were healthcare-associated and constituted 24% of all S. aureus keratitis isolates. Ninety percent of the 12 USA300-MRSA isolates were also healthcare-associated and accounted for only one (7%) of the community-associated isolates.
Hesje et al used a combination of molecular techniques including PFGE, MLST, and spa typing to characterize 40 MRSA and 16 MSSA ocular isolates collected during a 5-year period (2006–2008). Study isolates were recovered from the conjunctiva, cornea, and intraocular fluids of patients representing 24 US hospitals in 14 states.24 The number of keratitis isolates in this collection was not identified.
Similar to our data, two major SCCmec element types, II and IV, were detected among the 38 typeable MRSA isolates: twenty-two (57%) were classified as SCCmecII/t002 consistent with the USA100 healthcare-associated genotype and 16 (42.1%) were classified as SCCmecIV/t008 consistent with the USA300 genotype. All CC5/SCCmecII isolates were more likely to be multidrug resistant in both studies. MSSA isolates were polyclonal and belonged to 12 or more clones. Similarly, our MSSA isolates were polyclonal and distributed across 13 clonal groups.
In our study, DNA microarray confirmed the presence of the PVL toxin in 14.7% of the isolates with >80% associated with USA300 isolates. PVL is a cytotoxic pore-forming toxin associated with USA300, skin and soft tissue infections.36 The role of the PVL toxin in keratitis is unclear. Zaidi et al demonstrated in general, the presence and expression of the PVL gene resulted in increased toxicity, clinical morbidity, and disease in a mouse model of keratitis.36 However, the overall outcome was strain and clone specific; the team reported experimental differences between the USA300 community strain compared with the USA400 strain in a mouse keratitis model. In another study, Sueke et al found that patients with PVL-positive S. aureus keratitis isolates were more likely to trend toward worse clinical outcomes and required more surgical interventions.37
In this study, the Enterotoxin A toxin (SEA gene) was detected in seven (9.3%) of the isolates. Greater than 50% of these isolates were associated with the USA500 MRSA clone, the progenitor of the current USA300 epidemic MRSA strain.38,39 The USA500 MRSA clone is frequently associated with invasive disease in the USA. This is the first report of SEA-positive MRSA keratitis isolates.
Fujishima et al demonstrated a link between S. aureus enterotoxins and corneal ulceration in 11 patients with atopic keratoconjunctivitis.40 S. aureus enterotoxin B (SEB) was detected in two (18.2%) S. aureus isolates, enterotoxin G (SEG) in eight (72.7%) isolates, and enterotoxin I (SEI) in eight (72.7%) isolates. No SEA strains were detected among the 11 S. aureus isolates.40
The ACME, a virulent factor contributing to enhanced growth and invasiveness of USA300 MRSA, was present in 13 of our isolates. All of the ACME-positive isolates were associated with the USA300 MRSA clone.
Our in vitro susceptibility data differed from those reported in a 20-year study of S. aureus keratitis isolates by Chang et al.41 Specifically, there was a difference in the rates of resistance among healthcare- and community-associated MSSA and MRSA keratitis isolates to the fourth generation fluoroquinolones and gentamicin. In our study, 85% of our community-associated S. aureus isolates and 61% of our healthcare-associated isolates were resistant to moxifloxacin. Ninety-one percent of MRSA isolates and 18% of MSSA isolates were resistant to moxifloxacin. Only vancomycin (100%) and gentamicin (92%) provided in vitro susceptibilities of ≥90%.
In contrast, the Chang et al study reported only 35% of MRSA and 6% of MSSA isolates resistant to moxifloxacin.41 Thirteen percent of MRSA isolates and 2.2% of MSSA isolates in the Chang et al study were resistant to gentamicin, while our study reported less than 10% gentamicin resistance for both MRSA (8%) and MSSA (7%). Similar to our study, resistance rates were higher for MRSA than for MSSA isolates.
Agreement between susceptible/resistant phenotype as determined by the Vitek 2 and detection of the corresponding resistance gene determinants was >90% for erythromycin, clindamycin, gentamicin, linezolid, tetracycline, trimethoprim–sulfamethoxazole, and vancomycin for the 75 S. aureus isolates. The mecA gene was present in only three-quarters (76.6%) of the 47 isolates identified phenotypically as MRSA. The mecC gene was negative in all isolates. These genes (mecA, mecC) reside on the mobile SCCmec genetic element and code for an alternative penicillin binding protein (PBP2a) with low affinity for broad-spectrum beta-lactam antibiotics. Expression is heterogeneous and variable. Several reports have identified mecA-negative, methicillin-resistant isolates, as the resistance mechanism can be independent of the mecA gene. Methicillin-resistant, mecA-negative strains may confer beta-lactam resistance via intrinsic chromosomal mutations, beta-lactamase hyper production, production of methicillinases, and presence of small colony variants.10,17,42–44
The blaZ gene, which confers resistance to penicillin, was detected in >70% of the MSSA and MRSA keratitis isolates. Rates of detection of resistance determinants for the aminoglycosides tobramycin (aadD), kanamycin (aphA3), and neomycin (aphA3) were three to four times higher than the rate of detection for the gentamicin resistance determinant (aacA-aphD). Fifteen percent of the study isolates possessed the mupR gene conferring resistance to mupirocin; the rate was twice as high for MRSA isolates compared with detection rate for the MSSA isolates. Interestingly, the most prevalent resistance determinant among the study group was fosB, conferring resistance to the fosfomycin. The gene was detected in 100% of MRSA isolates and two-thirds of the MSSA isolates. Taken together, our data confirm a high prevalence of resistance gene determinants among S. aureus and support the use of molecular methods such as DNA microarray technology for their simultaneous detection.28
Conclusion
USA100 and USA300 MRSA clones were common among South Florida keratitis isolates. The shifting and increasing rates of antibiotic resistance among these two clones, as well as among MSSA isolates, are a cause for concern. Understanding the molecular characteristics of S. aureus keratitis may offer insights into this changing epidemiology and support for the development of new treatment strategies.
Limitations
Our study has several limitations. First, it is a retrospective study and contains a small subset of the total isolates collected during the 11-year period. Analysis of all S. aureus isolates from this time period could offer different results. Second, these results are reported from a single institution and may not be generalizable to other parts of the United States or the world. Nevertheless, we offer the first documentation of the intermixing of healthcare- and community-associated MRSA and MSSA among S. aureus keratitis isolates in South Florida. We also confirm the presence of high levels of genetic resistance to common ocular antibiotics, which has implications for patient management and institutional infection control strategies. The cumulative data suggest that vancomycin and gentamicin may provide broader in vitro susceptibility for the management of S. aureus keratitis in South Florida. Taken together, we provide an early “snapshot” of the evolving epidemiology, molecular characteristics, and antibiotic resistance of S. aureus keratitis isolates in South Florida.
Acknowledgments
This study was supported in part by the University of Miami Scientific Awards Committee (SAC) Interdisciplinary Team Science Pilot Award (UM SAC 2016-26), the Edward D and Janet K. Robson Foundation, Florida Lions Eye Bank and Beauty of Sight Foundation, Drs. KR Olsen and ME Hildebrandt, Drs. Raksha Urs and Aaron Furtado, Australian Federal Government Cooperative Research Centre Scheme through the Vision Cooperative Research Centre, the Brien Holden Vision Institute, NIH Center Grant P30EY14801, an unrestricted grant from Research to Prevent Blindness to the Department of Ophthalmology, and the Henri and Flore Lesieur Foundation (JMP).
Disclosure
The authors report no conflicts of interest in this work.
References
Mah FS, Davidson R, Holland EJ, et al. Current knowledge about and recommendations for ocular methicillin-resistant Staphylococcus aureus. J Cataract Refract Surg. 2014;40(11):1894–1908. | ||
Miller D. Ocular microbiology. In: Mahon M, editor. Diagnostic Microbiology. 4th ed. Philadelphia: WB Saunders; 2011:1104. | ||
Ritterband D. Methicillin-resistant Staphylococcus aureus and the eye: current concepts and management strategies. Curr Ophthalmol Rep. 2013;1(4):151–160. | ||
Chuang C-C, Hsiao C-H, Tan H-Y, et al. Staphylococcus aureus ocular infection: methicillin-resistance, clinical features, and antibiotic susceptibilities. PLoS One. 2012;8(8):e42437. | ||
Ong SJ, Huang Y-C, Tan H-Y, et al. Staphylococcus aureus keratitis: a review of hospital cases. PLoS One. 2013;8(11):e80119. | ||
Wong ES, Chow CWY, Luk WK, Fung KSC, Li KKW. A 10-year review of ocular methicillin-resistant Staphylococcus aureus infections: epidemiology, clinical features, and treatment. Cornea. 2017;36(1):92–97. | ||
McNicholas S, Shore AC, Coleman DC, Humphreys H, Hughes DF. DNA microarray genotyping and virulence and antimicrobial resistance gene profiling of methicillin-resistant Staphylococcus aureus bloodstream isolates from renal patients. J Clin Microbiol. 2011;49(12):4349–4351. | ||
O’Callaghan RJ. The pathogenesis of Staphylococcus aureus eye infections. Pathogens. 2018;7(1). | ||
Tenover FC, Mcdougal LK, Goering RV, et al. Characterization of a strain of community-associated methicillin-resistant Staphylococcus aureus widely disseminated in the United States. J Clin Microbiol. 2006;44(1):108–118. | ||
Uhlemann A-C, Otto M, Lowy FD, Deleo FR. Evolution of community- and healthcare-associated methicillin-resistant Staphylococcus aureus. Infect Genet Evol. 2014;21:563–574. | ||
Deleo FR, Chambers HF. Reemergence of antibiotic-resistant Staphylococcus aureus in the genomics era. J Clin Invest. 2009;119(9):2464–2474. | ||
King JM, Kulhankova K, Stach CS, Vu BG, Salgado-Pabón W. Phenotypes and virulence among Staphylococcus aureus USA100, USA200, USA300, USA400, and USA600 clonal lineages. mSphere. 2016;1(3):pii: e00071-16. | ||
Monecke S, Coombs G, Shore AC, et al. A field guide to pandemic, epidemic and sporadic clones of methicillin-resistant Staphylococcus aureus. PLoS One. 2011;6(4):e17936. | ||
O’Hara FP, Amrine-Madsen H, Mera RM, et al. Molecular characterization of Staphylococcus aureus in the United States 2004-2008 reveals the rapid expansion of USA300 among inpatients and outpatients. Microb Drug Resist. 2012;18(6):555–561. | ||
McDouglas LK, Steward CD, Killgore GE, Chaitram JM, Mcallister SK, Tenover FC. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: establishing a national database. J Clin Microbiol. 2003;41(11):5113–5120. | ||
Tenover FC, Mcallister S, Fosheim G, et al. Characterization of Staphylococcus aureus isolates from nasal cultures collected from individuals in the United States in 2001 to 2004. J Clin Microbiol. 2008;46(9):2837–2841. | ||
Deleo FR, Otto M, Kreiswirth BN, Chambers HF. Community-associated methicillin-resistant Staphylococcus aureus. Lancet. 2010;375(9725):1557–1568. | ||
Goering RV, Shawar RM, Scangarella NE, et al. Molecular epidemiology of methicillin-resistant and methicillin-susceptible Staphylococcus aureus isolates from global clinical trials. J Clin Microbiol. 2008;46(9):2842–2847. | ||
Klevens RM, Morrison MA, Fridkin SK, et al. Community-associated methicillin-resistant Staphylococcus aureus and healthcare risk factors. Emerg Infect Dis. 2006;12(12):1991–1993. | ||
Asbell PA, Sanfilippo CM, Pillar CM, DeCory HH, Sahm DF, Morris TW. Antibiotic resistance among ocular pathogens in the United States: five-year results from the Antibiotic Resistance Monitoring in Ocular Microorganisms (ARMOR) Surveillance Study. JAMA Ophthalmol. 2015;133(12):1445–1454. | ||
Hsiao C-H, Chuang C-C, Tan H-Y, et al. Methicillin-resistant Staphylococcus aureus ocular infection: a 10-year hospital-based study. Ophthalmology. 2012;119(3):522–527. | ||
Hsiao CH, Ong SJ, Chuang CC, Ma DH, Huang YC. A comparison of clinical features between community-associated and healthcare-associated methicillin-resistant Staphylococcus aureus keratitis. J Ophthalmol. 2015;2015:923941. | ||
Solomon R, Donnenfeld ED, Perry HD, et al. Methicillin-resistant Staphylococcus aureus infectious keratitis following refractive surgery. Am J Ophthalmol. 2007;143(4):629–634. | ||
Hesje CK, Sanfilippo CM, Haas W, Morris TW. Molecular epidemiology of methicillin-resistant and methicillin-susceptible Staphylococcus aureus isolated from the eye. Curr Eye Res. 2011;36(2):94–102. | ||
Khan MA, Ahmad S, Banu N. Molecular characterisation of methicillin-resistant Staphylococcus aureus (MRSA) from keratitis patients: a microbiological analysis. Br J Ophthalmol. 2010;94(8):994–998. | ||
Nadig S, Velusamy N, Lalitha P, Kar S, Sharma S, Arakere G. Staphylococcus aureus eye infections in two Indian hospitals: emergence of ST772 as a major clone. Clin Ophthalmol. 2012;6:165–173. | ||
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 26th edn. Vol 26. Wayne, Pennsylvania: Clinical and Laboratory Standards Institute; 2016. | ||
Monecke S, Müller E, Dorneanu OS, Vremeră T, Ehricht R. Molecular typing of MRSA and of clinical Staphylococcus aureus isolates from Iaşi, Romania. PLoS One. 2014;9(5):e97833. | ||
Shore AC, Deasy EC, Slickers P, et al. Detection of staphylococcal cassette chromosome mec type XI carrying highly divergent mecA, mecI, MecR1, blaZ, and CCR genes in human clinical isolates of clonal complex 130 methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2011;55(8):3765–3773. | ||
Baron EJ, Tenover FC. Methicillin-resistant Staphylococcus aureus diagnostics: state of the art. Expert Opin Med Diagn. 2012;6(6):585–592. | ||
David MZ, Glikman D, Crawford SE, et al. What is community-associated methicillin-resistant Staphylococcus aureus? J Infect Dis. 2008;197(9):1235–1243. | ||
McCarthy NL, Sullivan PS, Gaynes R, Rimland D. Health care-associated and community-associated methicillin-resistant Staphylococcus aureus infections: a comparison of definitions. Am J Infect Control. 2010;38(8):600–606. | ||
Naimi TS, Ledell KH, Como-Sabetti K, et al. Comparison of community- and health care-associated methicillin-resistant Staphylococcus aureus infection. JAMA. 2003;290(22):2976–2984. | ||
Roberts JC. Community-associated methicillin-resistant Staphylococcus aureus epidemic clone USA100; more than a nosocomial pathogen. Springerplus. 2013;2(1):133. | ||
O’Hara FP, Guex N, Word JM, et al. A geographic variant of the Staphylococcus aureus Panton-Valentine leukocidin toxin and the origin of community-associated methicillin-resistant S. aureus USA300. J Infect Dis. 2008;197(2):187–194. | ||
Zaidi T, Zaidi T, Yoong P, Pier GB. Staphylococcus aureus corneal infections: effect of the Panton-Valentine leukocidin (PVL) and antibody to PVL on virulence and pathology. Invest Ophthalmol Vis Sci. 2013;54(7):4430–4438. | ||
Sueke H, Shankar J, Neal T, et al. lukSF-PV in Staphylococcus aureus keratitis isolates and association with clinical outcome. Invest Ophthalmol Vis Sci. 2013;54(5):3410–3416. | ||
Diep BA, Chambers HF, Graber CJ, et al. Emergence of multidrug-resistant, community-associated, methicillin-resistant Staphylococcus aureus clone USA300 in men who have sex with men. Ann Intern Med. 2008;148(4):249–257. | ||
Frisch MB, Castillo-Ramírez S, Petit RA, et al. Invasive methicillin-resistant Staphylococcus aureus USA500 strains from the U.S. Emerging Infections Program constitute three geographically distinct lineages. mSphere. 2018;3(3): pii: e00571-17. | ||
Fujishima H, Okada N, Dogru M, et al. The role of staphylococcal enterotoxin in atopic keratoconjunctivitis and corneal ulceration. Allergy. 2012;67(6):799–803. | ||
Chang VS, Dhaliwal DK, Raju L, Kowalski RP. Antibiotic resistance in the treatment of Staphylococcus aureus keratitis: a 20-year review. Cornea. 2015;34(6):698–703. | ||
Elhassan MM, Ozbak HA, Hemeg HA, Elmekki MA, Ahmed LM. Absence of the mecA gene in methicillin resistant Staphylococcus aureus isolated from different clinical specimens in Shendi City, Sudan. Biomed Res Int. 2015;2015:895860. | ||
Ba X, Harrison EM, Edwards GF, et al. Novel mutations in penicillin-binding protein genes in clinical Staphylococcus aureus isolates that are methicillin resistant on susceptibility testing, but lack the mec gene. J Antimicrob Chemother. 2014;69(3):594–597. | ||
Banerjee R, Gretes M, Harlem C, Basuino L, Chambers HF. A mecA-negative strain of methicillin-resistant Staphylococcus aureus with high-level β-lactam resistance contains mutations in three genes. Antimicrob Agents Chemother. 2010;54(11):4900–4902. |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.