Back to Journals » OncoTargets and Therapy » Volume 13
Metformin Combined with 4SC-202 Inhibited the Migration and Invasion of OSCC via STAT3/TWIST1
Authors He Y, Fan Z, He L, Zhang C, Ping F, Deng M, Liu S, Wang Y, Cheng B, Xia J
Received 22 June 2020
Accepted for publication 11 September 2020
Published 29 October 2020 Volume 2020:13 Pages 11019—11029
DOI https://doi.org/10.2147/OTT.S268851
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Leo Jen-Liang Su
Yuan He,1,2,* Zhaona Fan,1,2,* Lihong He,1,2 Chi Zhang,1,2 Fan Ping,1,2 Miao Deng,1,2 Suyang Liu,1,2 Yanting Wang,1,2 Bin Cheng,1,2 Juan Xia1,2
1Hospital of Stomatology, Sun Yat-Sen University, Guangzhou, People’s Republic of China; 2Guangdong Provincial Key Laboratory of Stomatology, Guanghua School of Stomatology, Sun Yat-Sen University, Guangzhou, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Bin Cheng; Juan Xia
Guangdong Provincial Key Laboratory of Stomatology, Guanghua School of Stomatology, Sun Yat-Sen University, No. 55 Linyuan Xi Road, Guangzhou, Guangdong 510055, People’s Republic of China
Tel +862083741891
Fax +86 20 83822807
Email [email protected]; [email protected]
Background: Oral squamous cell carcinoma (OSCC), the most common epithelial malignant neoplasm in the head and neck, characterizes with local infiltration and metastasis of lymph nodes. The five-year survival rate of OSCC remains low despite the advances in clinical methods. Thus, it is necessary to develop a new effective therapeutic scheme for OSCC. Our previous results showed that metformin and 4SC-202 synergistically promoted the intrinsic apoptosis of OSCC in vitro and in vivo, but the effects on invasion and migration remained unclear.
Methods: Human OSCC cell lines HSC6 and CAL33 were cultured with metformin (16 mM) or/and 4SC-202 (0.4 μM) for 72 h. STAT3 inhibitor S31-201 was applied at concentration of 60 μM for 48 h. Wound-healing assays and transwell assays were used to determine the invasion and migration ability of OSCC. qRT-PCR and Western blot were performed to detect mRNA levels and protein levels.
Results: Metformin or/and 4SC-202 suppressed the migration and invasion of OSCC cells. Importantly, the expression of TWIST1 was suppressed by metformin and 4SC-202, while the invasion and migration inhibitory effects of metformin and 4SC-202 were countered by the overexpression of TWIST1. In addition, the phosphorylation level of STAT3 decreased after the administration of metformin or/and 4SC-202. Furthermore, inhibition of STAT3 by S31-201 suppressed the expression of TWIST1 and led to a decline in migration and invasion of OSCC, while overexpression of TWIST1 attenuated these effects.
Conclusion: Metformin and 4SC-202 suppressed the invasion and migration of OSCC through inhibition of STAT3/TWIST1, and this scheme can serve as a novel therapeutic strategy for OSCC.
Keywords: oral squamous cell carcinoma, metformin, 4SC-202, invasion, migration
Introduction
Oral squamous cell carcinoma (OSCC) is the most common oral cancer, and accounts for more than 90% of all oral tumors.1 Due to the delayed clinical detection and the high incidence of local invasion and metastasis, the prognosis of OSCC is poor.2 Despite advances in therapeutic strategies, five-year survival rate of OSCC has not improved significantly in recent years. Therefore, development of new therapeutic methods for OSCC becomes particularly important.
Protein acetylation modification plays a vital role in the tumorigenesis. In OSCC, high expression of HDACs such as HDAC1, HDAC2, HDAC6 were associated with poor prognosis, advanced stage, larger tumor size, and lymph node metastasis,3–5 suggesting that HDACs were involved in OSCC progress and could serve as potential drug targets. 4SC-202 is a novel selective class I histone deacetylase inhibitor (HDACi). In vitro, 4SC-202 was observed to inhibit survival and proliferation of several types of cancer cells including hepatocellular carcinoma cell, colorectal cancer cell, medulloblastoma cell, and urothelial carcinoma cell,6–9 and 4SC-202 was shown to inhibit TGF-β-induced epithelial-to-mesenchymal transition (EMT) of pancreatic cancer cells.10 Our previous findings revealed that 4SC-202 promoted the apoptosis of OSCC,11 however, the effects of 4SC-202 on migration and invasion in OSCC remained unclear.
Metformin, a low cost agent widely used for diabetes, received considerable attention for its antitumor effect. It was found that metformin suppressed invasion and migration of various types of cancers including esophageal squamous cell carcinoma, cholangiocarcinoma and prostate cancer.12–14 In addition, metformin could enhance the efficacy of several chemotherapeutics.15–17
Drug combination could achieve the therapeutic efficacy at reduced dosages, thus minimize the toxicity and other side effects of high doses of single drugs. Our previous results showed that metformin and 4SC-202 synergistically promoted the intrinsic apoptosis of OSCC in and in vivo,11 but the effects of this combination on invasion and migration remained elusive.
In this study, we evaluated the effects and mechanisms of metformin and 4SC-202 on the invasion and migration of OSCC. Here, we found that metformin and 4SC-202 synergistically inhibited the migration and invasion of OSCC. Importantly, our results revealed that the combined treatment of metformin and 4SC-202 inhibited invasion and migration by suppressing STAT3/TWIST1 in OSCC. These findings further emphasized the combination of metformin and 4SC-202 as a promising potential therapeutic strategy for OSCC.
Materials and Methods
Cell Culture
Human OSCC cell lines HSC6 and CAL33 were kindly provided by J Silvio Gutkind (NIH, Bethesda, MD, USA), SCC25 (CRL-1628)was purchased from American Type Culture Collection (ATCC, Manassas, VA, USA), and HSC3 was obtained from Professor Qianming Chen (State Key Laboratory of Oral Diseases, Sichuan University, China). The use of these cell lines was approved by our institutional research ethics committee. The HSC3, HSC6 and CAL33 cells were maintained in DMEM (Gibco, Grand Island, NY, USA) supplemented with 10% FBS (Gibco). SCC25 was cultured in DMEM supplemented with 10% FBS and 400 ng/mL hydrocortisone. All cells were maintained at 37°C with 5% CO2. The cells were treated with metformin (16 mM) or/and 4SC-202 (0.4 μM) for 72 h, and STAT3 inhibitor (40 μM, 60 μM) was applied for 48 h.
Information regarding reagents and antibodies were listed in Supplementary Table 1.
Wound-healing Assays
The cells were seeded into six-well plates, and then cells were wounded in a line across the slides with a sterile 200 μL plastic pipette tip. All cell debris was removed by washing with PBS. Cell migration, indicating wound-healing effect, was demonstrated as percentage of wounded area, that was the distance migrated by OSCC cells at the indicated point in time relative to the initial length of the wound. Experiments were performed at least three times.
Transwell Assays
Cell migration and invasion capacities were examined with 24-well transwell chamber (pore size: 8 μm) (Corning Costar, Corning, NY, USA). The upper chambers were precoated with or without 0.1 mL (300 μg/mL) Matrigel matrix (Corning) for invasion or migration assay, respectively. The prepared cells were seeded in the upper chambers with serum-free DMEM, but the bottom chambers were filled with DMEM containing 10% FBS. Then, cells were incubated for 24 h. The noninvading cells were gently removed by a cotton swab, whereas the invading cells were fixed in 4% paraformaldehyde for 15 min, stained with 0.4% crystal violet for 10 min and photographed under an inverted light microscope (Olympus Corporation, Tokyo, Japan). Finally, the invasive cells in five randomly selected visual fields at 100× magnification were counted.
RT-PCR
Total RNA was isolated with TRIzol (Invitrogen, CA, USA) and cDNA was synthesized by reverse transcription according to the manufacturer’s procedure (Takara, Dalian, China). The real-time PCR was carried out utilizing the LightCycler 480 SYBR Green I Master system (Roche, Basel, Switzerland). Relative mRNA expression levels were normalized to GAPDH using the 2−ΔΔCt method. The primers used were listed in Table 1.
 |
Table 1 Primers Used for RT-PCR |
Western Blot (WB) Analysis
Cells were lysed with RIPA buffer supplemented with protease and phosphatase inhibitor cocktail (CW Biotech, Beijing, China). 40–60 μg denatured protein of each sample was separated on 10% SDS-PAGE gels and transferred to a PVDF membrane (Millipore, Billerica, MA, USA). Then the membrane was blocked in 5% non-fat milk for one hour at room temperature and incubated with primary antibodies overnight at 4°C. Subsequently, the membrane was washed by 0.1% TBST for three times and incubated with HRP-conjugated secondary antibody for one hour at room temperature. The immunoreactive bands were visualized by a highly sensitive chemiluminescence (ECL) detection system (Millipore) and the immunoreactive bands were analyzed by densitometry using Image J (Bethesda, MD, USA). Similar results were obtained from three independent experiments.
Cell Transfection
CAL33 and SCC25 cells were transfected with Twist1 siRNA (50 nM, RiboBio, Guangzhou, China) using the lipofectamine RNA iMAX transfection reagent (Invitrogen) following the manufacturer’s instructions. The OSCC cells were transfected with lentivirus plasmid sh-TWIST1 or lentivirus plasmid Flag-TWIST1 (Cyagen, CA, USA), then the stable transfected cells were screened with puromycin. The TWIST1 sequences were as follows: sh-TWIST1, 5ʹATGGCAAGCTGCAGCTATGTG3ʹ, si1-TWIST1, 5ʹ-TGAGCAACAGCGAGGAAGA-3ʹ, and si2-TWIST1, 5ʹ-GATGGCAAGCTGCAGCTAT-3ʹ.
Statistics
All statistical analyses were performed using SPSS 20.0 software (SPSS, Chicago, IL, USA). All results were graphically depicted as mean ±SD from triplicate experiments performed in a parallel manner. Two-tailed Student’s t-test and one-way ANOVA were performed to detect statistically significant differences. P<0.05 was considered statistically significant.
Results
Metformin and 4SC-202 Inhibited Invasion and Migration of OSCC
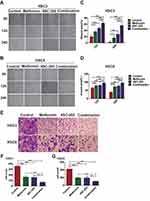
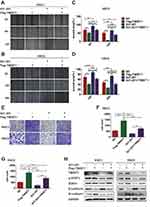
To investigate the effects of metformin and 4SC-202 on migration of OSCC, a wound-healing assay was performed. The results showed that metformin or/and 4SC-202 inhibited the migration of OSCC (Figure 1A and B), and the combination had the most significant inhibition. With metformin or/and 4SC-202 treatment, the wound area of HSC3 was increased from 20.53±5.33% to 36.74±3.79% (metformin), 47.68±5.49% (4SC-202) and 64.22±4.74% (combination) at 12 h and from 2.80±3.97% to 16.37±7.24% (metformin), 28.35±6.64% (4SC-202) and 56.39±8.86% (combination) at 24 h vs control (P<0.05), and the wound area of HSC6 was increased from 66.05±3.59% to 79.78±4.76% (metformin), 81.21±4.06% (4SC-202) and 95.731±2.80% (combination) at 12 h and from 39.92±3.88% to 73.10±6.58% (metformin), 77.52±4.86 (4SC-202) and 95.41±2.55% (combination) at 24 h vs control (P<0.05) (Figure 1C and D). Subsequently, transwell results revealed that metformin or/and 4SC-202 reduced the counts of the invading cells and the combination had the most dramatic reduction (Figure 1E). The counts of HSC3 cells invading to lower chamber were 164.33±19.67 with metformin alone, 157.33±19.29 with 4SC-202 alone, 71.00±13.37 with metformin and 4SC-202 treatment, and 358.33±11.32 in control group (P<0.01) (Figure 1F). Like HSC3, the counts of HSC6 cells invading the lower chamber were 196.67±9.98 with metformin, 192.67±11.6 with 4SC-202 and 104.67±7.04 with combination of metformin and 4SC-202, while the control group was 425.67±48.24 (P<0.01) (Figure 1G).
Metformin and 4SC-202 Suppressed the Expression of TWIST1
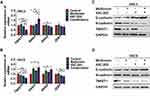
EMT plays vital roles in the invasion and migration of cancer cells,18 so we used qRT-PCR to determine the expression of EMT-related transcription factors including TWIST1, SNAI1, SNAI2, and ZEB1. The results showed that under the treatment of metformin and 4SC-202, the relative expression level of TWIST1 decreased to 0.51±0.01 (P<0.001) of control in HSC3 and 0.57±0.03 (P<0.001) of control in HSC6. However, the expression of SNAI1, SNAI2, and ZEB1 remained unperturbed or even increased (Figure 2A and B). Subsequently, the protein level of mesenchymal protein N-cadherin, epithelial protein E-cadherin and TWIST1 was evaluated by WB. These results revealed that metformin or/and 4SC-202 inhibited the expression of N-cadherin and TWIST1, but promoted the expression of E-cadherin in HSC3 and HSC6 cells, and the combination had the most significant effects (Figure 2C and D). Therefore, we speculated that TWIST1 may be the main target of metformin and 4SC-202.
Metformin and 4SC-202 Suppressed the Invasion and Migration by Inhibiting TWIST1
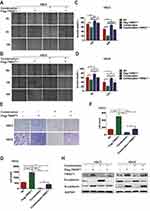
The efficiency of overexpression and knockdown of TWIST1 in OSCC was evaluated (Supplementary Figure 1). In line with previous reports, overexpression of TWIST1 promoted the invasion, migration and EMT of OSCC cells, and knockdown of TWIST1 had the reverse effects (Supplementary Figure 2). TWIST1 overexpression attenuated the invasion, migration and EMT-inhibiting effects of 4SC-202 and metformin, as the wound area was decreased from 57.74±3.68% to 38.49±9.50% (12 h) (P<0.05) in HSC3 cells (Figure 3A and C), and from 68.37±2.48% to 48.15±1.84% (24 h) (P<0.001) in HSC6 cells (Figure 3B and D), the counts of cells invading the lower chamber were increased from 66.00±8.84 to 161.00±33.22 (P<0.05) in HSC3 cells (Figure 3E and F), and from 46.00±38.68 to 404.00±56.70 in HSC6 cells (Figure 3G). Subsequently, the expression of E-cadherin and N-cadherin was determined by WB. The results revealed that TWIST1 overexpression partly rescued the expression of N-cadherin and impeded the downregulation of E-cadherin (Figure 3H). Overall, our results indicated that TWIST1 was the main target of metformin and 4SC-202.
Metformin and 4SC-202 Suppressed STAT3 Signaling
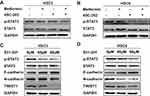
As the mRNA changed resembled to those of the protein level, TWIST1 could be controlled at the transcriptional level. STAT3 signal plays vital roles in the proliferation, invasion and migration of HNSCC,19–22 and STAT3 could regulate TWIST1 expression directly by binding to its promoter.23 The phosphorylation level of STAT3 was determined by WB, as the results showed, p-STAT3 was reduced after metformin or/and 4SC-202 treatment, and the combination had more remarkable inhibition compare to each alone (Figure 4A and B). Subsequently, one inhibitor of STAT3, S31-201, was applied at different concentrations to investigate regulatory relations between STAT3 and TWIST1. Our results showed that S31-201 inhibited the expression of TWIST1 and N-cadherin, but promoted the expression of E-cadherin (Figure 4C and D), indicating that TWIST1 and EMT was modulated by STAT3.
STAT3 Inhibited the Migration and Invasion of OSCC by Regulating TWIST1
Next, the effects of STAT3/TWIST1 on migration and invasion of OSCC cells were examined. The results showed that the migration was decreased significantly after treatment of S31-201, while overexpression of TWIST1 reversed this inhibitory effect (Figure 5A and B). With S31-201 administration, the wound area of HSC3 cells was increased from 51.15±2.90% to 67.34±7.30% (P<0.01) at 12 h, and the wound area of HSC6 cells was increased from 47.74±2.50% to 74.38±5.97% (P<0.01) at 24 h. While TWIST1 could reverse this inhibition, as with TWIST1 overexpression, the wound area was decreased to 39.84±3.97% (P<0.001) in HSC3 at 12 h and 41.32±3.75% (P<0.001) in HSC6 at 24 h (Figure 5C and D). Subsequently, transwell assays were performed to investigate the invasion of OSCC. The results demonstrated that S31-201 inhibited the invasion ability of HSC3 and HSC6, and overexpression of TWIST1 eliminated this inhibition (Figure 5E). The quantitative analysis demonstrated that S31-201 decreased the counts of HSC3 cells invading to lower chamber from 544.00±25.66 to 295.00±45.90 (P<0.05), while TWIST1 overexpression restored it to 734±58.70 (P<0.01), and a decrease of HSC6 cells from 533.00±19.38 to 259.00±65.01 was observed (P<0.01), while TWIST1 overexpression reinstated it to 883±58.35 (P<0.001) (Figure 5F and G). Furthermore, S31-201 was found to reduce the protein level of N-cadherin while it increased the protein level of E-cadherin in HSC3 and HSC6, and TWIST1 overexpression reversed these effects (Figure 5H). According to these data, we could infer that metformin and 4SC-202 inhibited the invasion and migration of OSCC by regulating STAT3/TWIST1.
Discussion
In present study, our results indicated that combination of metformin and 4SC-202 suppressed the invasion and migration of OSCC. Importantly, the expression of TWIST1 was inhibited by metformin and 4SC-202, and overexpression of TWIST1 attenuated the invasion and migration inhibiting effects of the drugs. In addition, phosphorylation of STAT3 was reduced by metformin and 4SC-202, and overexpression of TWIST1 reversed the invasion and migration inhibitory effects of STAT3 inhibitor S31-201. Our results revealed that metformin and 4SC-202 synergistically inhibited the invasion and migration of OSCC via STAT3/TWIST1, and could be a promising therapeutic scheme for OSCC.
As mentioned in the literature, HDACi could inhibit the growth of multiple cancer cells. However, the impacts of HDACi on cell invasion and migration seemed to be contradictory because the same agent had a contradictory effect on different cell types or the same cell type had a different response to different HDACi.24 In hepatocellular carcinoma, class II HDACi TSA administration was found to enhance migration and invasion,25 while pan HDACi panobinostat had an opposite effect.26 For head and neck cancer cells, prior studies have noted that SAHA enhanced migration and metastasis in human nasopharyngeal carcinoma cell line,26 while migration and invasion of nasopharyngeal carcinoma cells were inhibited by TSA.27 Different HDAC targets, different gene expression profile or side effects may account for these contradictions. In this study, we found that the invasion and migration of OSCC were inhibited by 4SC-202.
Recently, metformin was shown to impede the invasion and migration of several types of cancers including prostate adenocarcinoma, colorectal cancer, gastric cancer, cervical cancer.28 Several studies have shown that metformin could increase oral cancer cell sensitivity to chemotherapeutic drugs (such as 5-FU, gefitinib), improve the therapeutic efficacy, and reduce the dose and toxicity.29,30 Therefore, combination of metformin with other chemotherapeutics could be a potential candidate for the development of new treatment strategies for OSCC. In line with previous reports, our finding demonstrated that metformin inhibited invasion and migration of OSCC. Importantly, combination of metformin and 4SC-202 synergistically suppressed the migration and invasion of OSCC.
EMT plays a key role in tumor progression and is associated with stemness, invasiveness, drug-resistance, and the ability to form metastases in distant organs.22 HDACi exerted different effects on EMT in head and neck cancer cells, the EMT progression was induced by TSA,31 but class I, II HDACi valproic acid suppressed EMT.32 Pervious study of pancreatic cancer indicated that 4SC-202 had negative effects on EMT,10 and metformin suppressed EMT in various cancer types.12,14,33,34 In addition, metformin sensitized HNSCC to gefitinib and inhibited EMT.30 In our study, both 4SC-202 and metformin had negative impact on EMT of OSCC. Interestingly, metformin combined with 4SC-202 resulted in a more pronounced inhibition of tumor EMT compared with either drug alone.
EMT is modulated by TWIST1, a conserved helix-loop-helix (bHLH) transcription factor, which plays pivotal roles in multiple stages of embryonic development35 and significantly contributes to tumor metastasis.36 In OSCC, there is positive association between TWIST1 and poor prognosis, and TWIST1 promotes invasion and migration.37 Our results showed that metformin and 4SC-202 induced the decrease of TWIST1, and when applied together, a more dramatic suppression was observed, while the expression of other EMT transcription factors such as SNAI2, SNAI1, ZEB1 remained unperturbed or even increased. Overexpression of TWIST1 attenuated the invasion, migration and EMT inhibitory effects of metformin and 4SC-202, indicating that TWIST1 was the main target of metformin and 4SC-202.
TWIST1 is regulated by multiple signaling pathways including ERK, STAT3 and NF-κB.38 As a novel prognostic factor in HNSCC, STAT3 contributes to tumor growth,19 and induces invasion and migration.20–22 STAT3 was clarified to activate TWIST1 expression by directly binding to its promoter,23 and STAT3/TWIST1 regulatory axis was shown to be a critical regulator that drove the EMT progress in multiple types of cancer.39–44 Therefore, STAT3 could be the upstream regulator of TWIST1. In the present study, the inhibition of STAT3 led to a dramatic decrease of TWIST1 level in OSCC, supporting that TWIST1 was a downstream gene of STAT3. We speculated that the expression of TWIST1 was induced by STAT3 via direct transcriptional activation.
Metformin was reported to suppress invasion and migration of several cancers including bladder cancer, prostate cancer and pancreatic cancer though STAT3 inhibition.12,45,46 HDACi also exerts its tumor inhibitory impact through STAT3, as the class II HDAC inhibitor mercaptoacetamide suppresses migration and invasion in human glioma cells by inhibiting FAK/STAT3,47 and class I HDAC inhibitor chidamide and BG45 inhibits the viability of MDS and AML cells by suppressing JAK2/STAT3 signaling.48,49 Thus, STAT3 could be the target of anticancer drugs. Our results revealed that phosphorylation of STAT3 decreased under metformin or/and 4SC-202 treatment, suggesting metformin or/and 4SC-202 may suppress TWIST1 through STAT3, and intriguingly, combination of metformin and 4SC-202 had a more dramatic inhibition that either alone. Metformin activates AMPK, which in turn deactivates mTOR,28 and mTOR could phosphorylate STAT3 directly to activate it.50–52 Hence, we speculated that metformin suppressed STAT3 via AMPK activation in OSCC. Class I HDAC HDAC3 forms a complex with STAT3, and HDAC3 inhibition by HDACi represses STAT3 phosphorylation.53,54 Therefore, a possible explanation for STAT3 inhibition by 4SC-202 was that as selective class I HDAC inhibitor, 4SC-202 suppressed the enzymatic activity of HDAC3, and inhibition of HDAC3 deactivated STAT3. Thus, it could be assumed that metformin and 4SC-202 exerts its STAT3-suppressing effects via a different approach, and further work is required to determine the exact mechanisms.
A more dramatic decrease of TWIST1 was observed under metformin and 4SC-202 treatment, together with STAT3 inhibitor administration, and overexpression of TWIST1 rescued the invasion, migration, and the expression of EMT genes repressed by STAT3 inhibitor. These results demonstrated STAT3 was the key component that linked the anticancer effects of the drugs and TWIST1.
In summary, this study showed that metformin and 4SC-202 synergistically inhibited the invasion and migration of OSCC through STAT3/TWIST1. Our results complemented our previous studies, corroborating metformin and 4SC-202 as a promising therapeutic scheme for OSCC.
Abbreviations
OSCC, oral squamous cell carcinoma; HDAC, histone deacetylase; HDACi, histone deacetylase inhibitor; EMT, epithelial-to-mesenchymal transition; qRT-PCR, quantitative reverse transcription-polymerase chain reaction; WB, Western blot; ECL, chemiluminescence.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (no. 81671000, 81870769) and the Science and Technology Program of Guangzhou, China (no. 201803010019, 201607010354).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Chen JH, Chen WL, Sider KL, Yip CY, Simmons CA. β-catenin mediates mechanically regulated, transforming growth factor-β1-induced myofibroblast differentiation of aortic valve interstitial cells. Arterioscler Thromb Vasc Biol. 2011;31(3):590. doi:10.1161/ATVBAHA.110.220061
2. Rivera C. Essentials of oral cancer. Int J Clin Exp Pathol. 2015;8(9):11884–11894.
3. Chang HH, Chiang CPHung HC, Hung H-C, Lin C-Y, Deng Y-T, Kuo MY-P. Histone deacetylase 2 expression predicts poorer prognosis in oral cancer patients. Oral Oncol. 2009;45(7):610–614. doi:10.1016/j.oraloncology.2008.08.011
4. Sakuma T, Uzawa K, Onda T, et al. Aberrant expression of histone deacetylase 6 in oral squamous cell carcinoma. Int J Oncol. 2006;29(1):117–124.
5. Stamatios T, Jerzy K, Constantinos G, et al. Histone deacetylase-1 and −2 expression in mobile tongue squamous cell carcinoma: associations with clinicopathological parameters and patients survival. J Oral Pathol Med. 2011;40(9):706–714.
6. Zhang F, Wan F, Li Z, Fu M. 4SC-202 activates ASK1-dependent mitochondrial apoptosis pathway to inhibit hepatocellular carcinoma cells. Biochem Biophys Res Commun. 2016;471(2):267–273.
7. Messerli SM, Hoffman MM, Gnimpieba EZ, Kohlhof H, Bhardwaj RD. 4SC-202 as a Potential Treatment for the Pediatric Brain Tumor Medulloblastoma. Brain Sci. 2017;7(11):147.
8. Pinkerneil M, Hoffmann MJ, Kohlhof H, Schulz WA, Niegisch G. Evaluation of the Therapeutic Potential of the Novel Isotype Specific HDAC Inhibitor 4SC-202 in Urothelial Carcinoma Cell Lines. Target Oncol. 2016;11(6):783–798.
9. Zhijun H, Shusheng W, Han M, Jianping L, Lisen Q, Dechun L. Pre-clinical characterization of 4SC-202, a novel class I HDAC inhibitor, against colorectal cancer cells. Tumour Biol. 2016;37(8):10257–10267.
10. Mishra VK, Wegwitz F, Kosinsky RL, et al. Histone deacetylase class-I inhibition promotes epithelial gene expression in pancreatic cancer cells in a BRD4- and MYC-dependent manner. Nucleic Acids Res. 2017;45(11):6334–6349. doi:10.1093/nar/gkx212
11. He Y, Tai S, Deng M, et al. Metformin and 4SC-202 synergistically promote intrinsic cell apoptosis by accelerating DeltaNp63 ubiquitination and degradation in oral squamous cell carcinoma. Cancer Med. 2019;8(7):3479–3490. doi:10.1002/cam4.2206
12. Tong D, Liu Q, Liu G, et al. Metformin inhibits castration-induced EMT in prostate cancer by repressing COX2/PGE2/STAT3 axis. Cancer Lett. 2017;389:23–32. doi:10.1016/j.canlet.2016.12.031
13. Trinh SX, Nguyen HT, Saimuang K, Prachayasittikul V, On W C. Metformin Inhibits Migration and Invasion of Cholangiocarcinoma Cells. Asian Pac J Cancer Prev. 2017;18(2):473–477.
14. Nakayama A, Ninomiya I, Harada S, et al. Metformin inhibits the radiation-induced invasive phenotype of esophageal squamous cell carcinoma. Int J Oncol. 2016;49(5):1890–1898. doi:10.3892/ijo.2016.3676
15. Peng M, Darko KO, Tao T, et al. Combination of metformin with chemotherapeutic drugs via different molecular mechanisms. Cancer Treat Rev. 2017;54:24–33. doi:10.1016/j.ctrv.2017.01.005
16. Zhang HH, Guo XL. Combinational strategies of metformin and chemotherapy in cancers. Cancer Chemother Pharmacol. 2016;78(1):13–26. doi:10.1007/s00280-016-3037-3
17. Duo J, Ma Y, Wang G, Han X, Zhang C. Metformin synergistically enhances antitumor activity of histone deacetylase inhibitor trichostatin a against osteosarcoma cell line. DNA Cell Biol. 2013;32(4):156–164. doi:10.1089/dna.2012.1926
18. Zhang Y, Weinberg RA. Epithelial-to-mesenchymal transition in cancer: complexity and opportunities. Front Med. 2018;12(4):361–373. doi:10.1007/s11684-018-0656-6
19. Masuda M, Suzui M, Yasumatu R, et al. Constitutive activation of signal transducers and activators of transcription 3 correlates with cyclin D1 overexpression and may provide a novel prognostic marker in head and neck squamous cell carcinoma. Cancer Res. 2002;62(12):3351–3355.
20. Wheeler SE, Suzuki S, Thomas SM, et al. Epidermal growth factor receptor variant III mediates head and neck cancer cell invasion via STAT3 activation. Oncogene. 2010;29(37):5135–5145. doi:10.1038/onc.2009.279
21. Sun SS, Zhou X, Huang YY, et al. Targeting STAT3/miR-21 axis inhibits epithelial-mesenchymal transition via regulating CDK5 in head and neck squamous cell carcinoma. Mol Cancer. 2015;14(1):213. doi:10.1186/s12943-015-0487-x
22. Wang Y, Wu C, Zhang C, et al. TGF-beta-induced STAT3 overexpression promotes human head and neck squamous cell carcinoma invasion and metastasis through malat1/miR-30a interactions. Cancer Lett. 2018;436:52–62. doi:10.1016/j.canlet.2018.08.009
23. Cheng GZ, Zhang WZ, Sun M, et al. Twist is transcriptionally induced by activation of STAT3 and mediates STAT3 oncogenic function. J Biol Chem. 2008;283(21):14665–14673. doi:10.1074/jbc.M707429200
24. Wawruszak A, Kalafut J, Okon E, et al. Histone Deacetylase Inhibitors and Phenotypical Transformation of Cancer Cells. Cancers. 2019;11(2):148. doi:10.3390/cancers11020148
25. Yang L, Chang Y, Cao P. CCR7 preservation via histone deacetylase inhibition promotes epithelial-mesenchymal transition of hepatocellular carcinoma cells. Exp Cell Res. 2018;371(1):231–237. doi:10.1016/j.yexcr.2018.08.015
26. Song X, Wang J, Zheng T, et al. LBH589 Inhibits proliferation and metastasis of hepatocellular carcinoma via inhibition of gankyrin/STAT3/Akt pathway. Mol Cancer. 2013;12(1):114.
27. Shen Z, Liao X, Shao Z, et al. Short-term stimulation with histone deacetylase inhibitor trichostatin a induces epithelial-mesenchymal transition in nasopharyngeal carcinoma cells without increasing cell invasion ability. BMC Cancer. 2019;19(1):262. doi:10.1186/s12885-019-5482-y
28. Sui X, Xu Y, Wang X, Han W, Pan H, Xiao M. Metformin: A Novel but Controversial Drug in Cancer Prevention and Treatment. Mol Pharm. 2015;12(11):3783–3791. doi:10.1021/acs.molpharmaceut.5b00577
29. Harada K, Ferdous T, Harada T, Ueyama Y. Metformin in combination with 5-fluorouracil suppresses tumor growth by inhibiting the Warburg effect in human oral squamous cell carcinoma. Int J Oncol. 2016;49(1):276–284. doi:10.3892/ijo.2016.3523
30. Yin X, Wei Z, Song C, et al. Metformin sensitizes hypoxia-induced gefitinib treatment resistance of HNSCC via cell cycle regulation and EMT reversal. Cancer Manag Res. 2018;10:5785–5798. doi:10.2147/CMAR.S177473
31. Giudice FS, Pinto DS, Nor JE, Squarize CH, Castilho RM. Inhibition of histone deacetylase impacts cancer stem cells and induces epithelial-mesenchyme transition of head and neck cancer. PLoS One. 2013;8(3):e58672. doi:10.1371/journal.pone.0058672
32. Kanamoto A, Ninomiya I, Harada S, et al. Valproic acid inhibits irradiation-induced epithelial-mesenchymal transition and stem cell-like characteristics in esophageal squamous cell carcinoma. Int J Oncol. 2016;49(5):1859–1869.
33. Valaee S, Yaghoobi MM, Shamsara M. Metformin inhibits gastric cancer cells metastatic traits through suppression of epithelial-mesenchymal transition in a glucose-independent manner. PLoS One. 2017;12(3):e0174486. doi:10.1371/journal.pone.0174486
34. Cheng K, Hao M. Metformin Inhibits TGF-beta1-Induced Epithelial-to-Mesenchymal Transition via PKM2 Relative-mTOR/p70s6k Signaling Pathway in Cervical Carcinoma Cells. Int J Mol Sci. 2016;17(12):12. doi:10.3390/ijms17122000
35. Bildsoe H, Loebel DA, Jones VJ, et al. The mesenchymal architecture of the cranial mesoderm of mouse embryos is disrupted by the loss of Twist1 function. Dev Biol. 2013;374(2):295–307. doi:10.1016/j.ydbio.2012.12.004
36. Kang Y, Massague J. Epithelial-mesenchymal transitions: twist in development and metastasis. Cell. 2004;118(3):277–279. doi:10.1016/j.cell.2004.07.011
37. da Silva SD, Alaoui-Jamali MA, Soares FA, et al. TWIST1 is a molecular marker for a poor prognosis in oral cancer and represents a potential therapeutic target. Cancer. 2014;120(3):352–362. doi:10.1002/cncr.28404
38. Qin Q, Xu Y, He T, Qin C, Xu J. Normal and disease-related biological functions of Twist1 and underlying molecular mechanisms. Cell Res. 2012;22(1):90–106.
39. Hsu KW, Hsieh RH, Huang KH, et al. Activation of the Notch1/STAT3/Twist signaling axis promotes gastric cancer progression. Carcinogenesis. 2012;33(8):1459–1467.
40. Wu K, Wang B, Chen Y, et al. DAB2IP regulates the chemoresistance to pirarubicin and tumor recurrence of non-muscle invasive bladder cancer through STAT3/Twist1/P-glycoprotein signaling. Cell Signal. 2015;27(12):2515–2523.
41. Cho KH, Choi MJ, Jeong KJ, et al. A ROS/STAT3/HIF-1alpha signaling cascade mediates EGF-induced TWIST1 expression and prostate cancer cell invasion. Prostate. 2014;74(5):528–536.
42. Zhang C, Guo F, Xu G, Ma J, Shao F. STAT3 cooperates with Twist to mediate epithelial-mesenchymal transition in human hepatocellular carcinoma cells. Oncol Rep. 2015;33(4):1872–1882.
43. Cho KH, Jeong KJ, Shin SC, Kang J, Park CG, Lee HY. STAT3 mediates TGF-beta1-induced TWIST1 expression and prostate cancer invasion. Cancer Lett. 2013;336(1):167–173.
44. Li Y, Bai M, Xu Y, Zhao W, Liu N, Yu J. TPPP3 Promotes Cell Proliferation, Invasion and Tumor Metastasis via STAT3/Twist1 Pathway in Non-Small-Cell Lung Carcinoma. Cell Physiol Biochem. 2018;50(5):2004–2016.
45. Pan Q, Yang GL, Yang JH, et al. Metformin can block precancerous progression to invasive tumors of bladder through inhibiting STAT3-mediated signaling pathways. J Exp Clin Cancer Res. 2015;34:77.
46. Liu Q, Yuan W, Tong D, et al. Metformin represses bladder cancer progression by inhibiting stem cell repopulation via COX2/PGE2/STAT3 axis. Oncotarget. 2016;7(19):28235–28246.
47. Nam JH, Cho HJ, Kang H, et al. II Histone Deacetylase Inhibitor Suppresses Cell Migration and Invasion in Monomorphic Malignant Human Glioma Cells by Inhibiting FAK/STAT3 Signaling. J Cell Biochem. 2017;118(12):4672–4685.
48. Zhao S, Guo J, Zhao Y, et al. Chidamide, a novel histone deacetylase inhibitor, inhibits the viability of MDS and AML cells by suppressing JAK2/STAT3 signaling. Am J Transl Res. 2016;8(7):3169–3178.
49. Tang S, Cheng B, Zhe N, et al. Histone deacetylase inhibitor BG45-mediated HO-1 expression induces apoptosis of multiple myeloma cells by the JAK2/STAT3 pathway. Anticancer Drugs. 2018;29(1):61–74.
50. Rajan P, Panchision DM, Newell LF, McKay RD. BMPs signal alternately through a SMAD or FRAP-STAT pathway to regulate fate choice in CNS stem cells. J Cell Biol. 2003;161(5):911–921.
51. Kim JH, Kim JE, Liu HY, Cao W, Chen J. Regulation of interleukin-6-induced hepatic insulin resistance by mammalian target of rapamycin through the STAT3-SOCS3 pathway. J Biol Chem. 2008;283(2):708–715.
52. Yokogami K, Wakisaka S, Avruch J, Reeves SA. Serine phosphorylation and maximal activation of STAT3 during CNTF signaling is mediated by the rapamycin target mTOR. Curr Biol. 2000;10(1):47–50.
53. Ni L, Wang L, Yao C, et al. The histone deacetylase inhibitor valproic acid inhibits NKG2D expression in natural killer cells through suppression of STAT3 and HDAC3. Sci Rep. 2017;7:45266.
54. Gupta M, Han JJ, Stenson M, Wellik L, Witzig TE. Regulation of STAT3 by histone deacetylase-3 in diffuse large B-cell lymphoma: implications for therapy. Leukemia. 2012;26(6):1356–1364.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.