Back to Journals » OncoTargets and Therapy » Volume 13
Malic Enzyme 1 Indicates Worse Prognosis in Breast Cancer and Promotes Metastasis by Manipulating Reactive Oxygen Species
Authors Liu C, Cao J, Lin S, Zhao Y, Zhu M, Tao Z, Hu X
Received 5 April 2020
Accepted for publication 10 August 2020
Published 26 August 2020 Volume 2020:13 Pages 8735—8747
DOI https://doi.org/10.2147/OTT.S256970
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr William C. Cho
Chang Liu,1,2,* Jun Cao,1,2,* Shuchen Lin,1,2,* Yannan Zhao,1,2 Mingyu Zhu,1,2 Zhonghua Tao,1,2 Xichun Hu1,2
1Department of Medical Oncology, Fudan University Shanghai Cancer Center, Shanghai, People’s Republic of China; 2Department of Oncology, Shanghai Medical College, Fudan University, Shanghai, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Xichun Hu; Zhonghua Tao Email [email protected]; [email protected]
Purpose: Malic enzyme 1 (ME1) catalyzes malate to pyruvate and thus promotes glycolysis. Its function in breast cancer remains to be fully clarified. The aim of this work was to investigate the prognostic value of ME1 and its possible mechanism in breast cancer.
Methods: We evaluated ME1 expression in 220 early breast cancer patients with tissue microarray-based immunohistochemistry and explored the relationships between ME1 expression and clinicopathological features. Survival analyses were further performed to determine its prognostic value. The public database was used to confirm tissue microarray results. Further, cell proliferation, migration, invasion ability and reactive oxygen species (ROS) were examined in breast cancer cells.
Results: In breast cancer tissues, high ME1 expression was significantly associated with larger tumor size, higher incidence of lymph node metastasis and higher incidence of lymph-vascular invasion. High ME1 expression significantly correlated with worse recurrence-free survival (RFS), and was an independent prognostic factor for RFS, which was confirmed by mRNA results in the public database. In vitro, upregulation of ME1 by transfecting MCF-7 cells with virus vector remarkably enhanced viability, motility and epithelial–mesenchymal transition (EMT) and decreased ROS levels, whereas knockdown in MDA-MB-468 cells produced totally opposite effects as expected. When pretreated with oxidizing agent, MCF-7 cells overexpressing ME1 lost its motility, whereas MDA-MB-468 cells with knockdown of ME1 restored its motility when pretreated with antioxidant.
Conclusion: To our knowledge, these clinical and experiment works first suggested that ME1 may be a novel biomarker and potential therapeutic target for breast cancer metastasis, and its biological effect is mainly controlled by manipulating ROS.
Keywords: malic enzyme 1, breast cancer, prognosis, metastasis, reactive oxygen species
Plain Language Summary
Breast cancer is the most common malignancy in women worldwide. Tumor metastasis is the important cause for breast cancer patients’ deaths. It is also a hard problem for clinical treatment. In this article, Dr Hu’s team reported a novel risk factor and potential therapeutic target for breast cancer metastasis. Anaerobic glycolysis is a metabolic process in which glucose is broken down without the use of oxygen. However, cancer cells prefer anaerobic glycolysis even in the presence of oxygen. Various types of cancer including breast cancer express elevated levels of malic enzyme 1 (ME1), a metabolic enzyme which can promote anaerobic glycolysis. By evaluating the expression of ME1 in breast cancer tissues, Dr Hu’s team found high ME1 expression significantly correlated with risk factors for breast cancer recurrence, and indicated worse prognosis in breast cancer patients. In cellular experiments, Dr Hu’s team found ME1 could eliminate reactive oxygen species and thus promote the malignant biological behaviors of breast cancer cells. Future studies should address the strategies targeting ME1 for breast cancer treatment.
Introduction
Breast cancer is the most common malignancy in women and one of the three most common malignant tumors worldwide.1 In recent years, the curative effect of breast cancer has increased significantly. Nonetheless, even in patients with no lymph node metastasis, nearly 20% of them suffer from breast cancer recurrence.2,3 Therefore, it is necessary to further characterize patients with high recurrence risk and explore the molecular mechanism of breast cancer progression to help develop personalized treatment.
Reprogramming energy metabolism is one of the hallmarks of cancer, which is known as Warburg effect: cancer cells prefer glycolysis even in the presence of oxygen.4 It has been well accepted that cancer cells reprogram the metabolic patterns to satisfy their rapid proliferation and metastasis.5 In breast cancer, many glycolysis enzymes (hexokinase, phosphofructokinase, pyruvate kinase, etc.), glucose transporters, together with related signaling pathways (PI3K/AKT, mTOR, AMPK, etc.) and transcription factors (c-myc, p53, HIF-1, etc.) have been reported to be essential for the energy metabolism, and contribute to cell proliferation, metastasis, angiogenesis, and resistance to chemotherapy and radiation.6
Malic enzyme 1 (ME1, Enzyme Commission number: 1.1.1.40) is a cytosolic metabolic enzyme that catalyzes the citric acid cycle intermediate malate to pyruvate, the substrate of glycolysis, and converts Nicotinamide adenine dinucleotide phosphate (NADP) to Nicotinamide adenine dinucleotide phosphate hydrate (NADPH), the necessities of various biosynthetic reactions. Besides, ME1 can form physiological complexes with 6-Phosphogluconate dehydrogenase and further increase the pentose phosphate pathway flux and NADPH generation.7 Hence, ME1 promotes glycolysis and is involved in reactive oxygen species (ROS) homeostasis, glutamine metabolism and lipogenesis,8 playing an important role in Warburg effect. We performed data mining from the public Oncomine database and found that ME1 was upregulated in various cancer types including breast cancer. Recently, the potential function of ME1 has been revealed in tumor progression. However, there is a lacking of research on the role of ME1 in breast cancer. These findings prompted us to assess the value of ME1 in breast cancer.
In the present study, we evaluated the relationships between ME1 expression and clinicopathological features in breast cancer patients, and explored the potential role of ME1 in proliferation, migration and invasion of breast cancer cells. Our clinical and experimental work identified ME1 as a novel prognostic indicator, highlighted its potential role in the development of breast cancer and found its effect could be mainly blocked by manipulating ROS.
Materials and Methods
Patients, Tissue Specimens and Clinical Data
The study cohort consisted of 220 female breast cancer patients who underwent radical surgery between 11 August 2015 and 17 May 2016 in Fudan University Shanghai Cancer Center (FUSCC, Shanghai, China). Eligible patients were women who had histologically confirmed invasive breast cancer; had no evidence of distant metastasis; and provided sufficient tissues for further research. Patients were ineligible if they had received neoadjuvant chemotherapy or radiation therapy. Formalin-fixed paraffin-embedded breast cancer tissues were used as the study materials. Clinical data of the patients were retrieved from the Outcome unit. Molecular subtype was defined according to the St. Gallen International Expert Consensus Conference on the Primary Therapy of Early Breast Cancer 2013 (Supplementary Table S1).9 The medium follow-up period was 29.2 months (range 0.50–34.25). Recurrence-free survival (RFS) time was calculated from the date of radical surgery to the date of breast cancer recurrence (ipsilateral breast, local-regional, or distant), death or the last time of follow-up.
Immunohistochemistry (IHC)
Tissue microarray was constructed by Wei Ao biotech Company (Shanghai, China). Duplicate cores, which were selected morphologically as representative regions of tumor tissues, were taken from each specimen with a diameter of 2 mm. After deparaffinized and rehydrated, the sections were heated in an autoclave at 120°C in sodium citrate buffer (pH 6.0) for 10 min for antigen retrieval. The sections were then incubated with 3% hydrogen peroxide for 15 min. After blocking of nonspecific binding with QuickBlock™ Blocking Buffer (Beyotime, P0260, China) for 15 min, the sections were incubated at 4°C with ME1 antibody (Abcam, ab97445, 1:1000, USA) overnight. The IHC Kit (GTVision, GK500705, China) including second antibody and DAB substrate was used. After washing with PBS, the sections were incubated with the second antibody for 30 min at room temperature. Color was developed with DAB (1:100) for 2 min. The sections were counterstained with hematoxylin (Servicebio, G1004, China), and finally dehydrated. The TMA section omitting the incubation of primary antibody was used as negative control. We also verified that intercellular substance was not stained to exclude nonspecific background staining. ME1 staining was observed mainly in cytoplasm and a small part was in nucleus.
Immunohistochemical Staining Score
The standards for IHC staining scoring were previously described (Supplementary Table S2).10 Herein the intensity range was 0 = negative; 1 = low; 2 = medium and 3 = high. The quantity 0 = no positivity; 1 = positivity in 0–10%; 2 = positivity in 11–50%; 3 = positivity in 51–80%; 4 = positivity in >80%. The final immunoreactive score (IRS, ranging from 0–12) was obtained by multiplying the intensity score and the quantity score. Two pathologists blinded to the patients’ information scored the immunohistochemical staining. In discrepant cases, they further reviewed the cases and reached a consensus. For ME1 low and high expression was defined as IRS ≤ 6 and IRS > 6, respectively.
Cell Lines and Cell Culture
Cell lines including MCF-7, MDA-MB-468, SKBR3, MDA-MB-231, ZR-75-1 and 293FT were obtained from the Cell Research Institute (Shanghai, China). Cells were routinely cultured in high-glucose DMEM (MCF-7, SKBR3, ZR-75-1 and 293FT) (Hyclone, USA) or L-15 (MDA-MB-468 and MDA-MB-231) (Hyclone, USA), supplemented with 10% (v/v) fetal bovine serum (Gibco, USA) at 37°C in a humidified 5% CO2 atmosphere (MCF-7, SKBR3, ZR-75-1 and 293FT) or 100% atmosphere (MDA-MB-468 and MDA-MB-231).
Protein Extraction and Western Blotting Analysis
Cells were washed twice with pre-chilled PBS, pelleted by centrifugation and lysed in RIPA (Beyotime, PC102, China). After incubation for 30 min on ice, lysates were centrifuged (12,000g, 10 min, 4°C). Supernatants were collected and the protein concentration was measured using a BCA protein assay reagent (Enzyme, ZJ101, China). Total protein (20ug) was separated on 10% SDS-PAGE and then transferred to nitrocellulose membranes for 2h at room temperature. The membranes were incubated with the appropriate primary antibodies (anti-ME1, Abcam, ab97445, 1:1000, USA; anti-β-actin, Proteintech, 60,008-1-Ig, 1:1000, USA) overnight at 4°C, washed three times with TBST, and then incubated with the corresponding secondary antibody (goat anti rabbit IgG, SA00001-2, 1:1000; goat anti mouse IgG, SA00001-1, 1:1000, Proteintech, USA) for 1h at room temperature. The bands were visualized by Immobilon Western Chemiluminescent HRP Substrate (Millipore, WBKLS0100, USA) and detected with a luminescent image analyzer (GE Image Quant LAS4000 mini, USA).
Plasmids, Constructs and Infection
ZsGreen-labelled pHBLV lentiviral vector was purchased from XY biotech Company (Shanghai, China). Full-length cDNA encoding ME1 was ligated to ZsGreen-labelled pHBLV vector according to the manufacturer’s instructions. Cherry-labelled GV298 lentiviral vector was purchased from Genechem (Shanghai, China). The siRNA target sequence (5-GCTGAGGTTATAGCTCAGCAA-3) for ME1 (Genebank ID, NM_002395) and two different single-stranded DNA oligonucleotides (5-CCGGGCTGAGGTTATAGCTCAGCAACTCGAGTTGCTGAGCTATAACCTCAGCTTTTTG-3 and 5-GATCCAAAAAGCTGAGGTTATAGCTCAGCAACTCGAGTTGCTGAGCTATAACCTCAGC-3) used to match to generate the ME1-siRNA constructs were designed by GeneChem. The ME1-siRNA constructs were inserted into Cherry-labelled GV298 vector to form ME1-shRNA. Lentivirus carrying ME1 cDNA or shRNA targeting ME1 was produced from 293FT cells. MCF-7 and MDA-MB-468 cells were infected with lentivirus and then selected with puromycin. The overexpression and knockdown efficacy were evaluated by Western blotting.
Cell Proliferation Assays
Cells were seeded in 96-well plates (2000 cells/well) in triplicate and cell viability was examined by Cell Counting Kit-8 (CCK-8) assay (Dojindo Laboratories, Japan). Cells were seeded in 6-well plates (1000 cells/well) in triplicate and cultured for two weeks in colony-formation assay. Colonies were washed three times in PBS, fixed with 4% formaldehyde and stained with Crystal violet for 5 min.
Cell Migration and Invasion Assays
The migration and invasion assays were done in a 24-well Chemotaxis chamber with 8-μm pores (Corning, USA). For migration assays, 5×104 cells were seeded into the Matrigel-uncoated upper insert at 24-well in medium without serum. Medium containing 20% serum was added to the well as a chemoattractant. Following a culture of 36 h, non-invading cells were removed from the upper surface. For invasion assays, 1×106 cells were seeded into the Matrigel-coated upper insert at 24 wells in medium without serum. Medium containing 20% serum was added to the well as a chemoattractant. Following a culture of 48 h, non-invading cells were removed. Then, the membranes were fixed with 4% formaldehyde for 15 min. The invading cells were stained with Crystal violet for 5 min.
Detection of ROS Level
Cells in 6-well plates were washed with PBS three times and incubated with Dihydroethidium (DHE, Beyotime, S0063, China) or 2ʹ,7ʹ-Dichlorodihydrofluorescein diacetate (DCFH-DA, Beyotime, S0033, China) for 30 min at room temperature. Afterwards, cells were collected, washed with PBS three times and resuspended in PBS. DHE or DCF fluorescence was measured by FACScan Flow Cytometer (Beckman-Coulter, USA) within 30 min.
Statistical Analyses
We evaluated the correlations between ME1 expression and clinicopathological parameters by Pearson Chi-square test or Fisher’s exact test. Recurrence-free survival was plotted and calculated using Kaplan–Meier (KM) curve while differences between groups were compared by Log rank test. Univariate Cox proportional hazard model was used to estimate the influence of each variable on RFS. Variables with p values <0.1 in univariate Cox proportional hazard models were further included in multivariate Cox proportional hazard models. In laboratory experiments, quantitative variables were illustrated as means ± SD and analyzed with the Student’s t-test. Two-sided p values <0.05 were considered statistically significant. All analyses were performed by SPSS 22.0.
Results
High ME1 Expression Was Associated with Risk Factors of Breast Cancer Recurrence
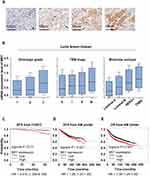
To evaluate the role of ME1 in breast cancer, we first performed IHC staining of ME1 in a tissue microarray containing 220 early-stage breast cancer patients. Several representative cases are shown in Figure 1A. ME1 mainly located in cytoplasm and a small part was observed in nucleus. As defined above, 51.8% (114/220) and 48.2% (106/220) of the patients were categorized as ME1-high and ME1-low cases, respectively.
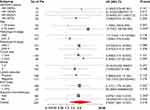
Next, we explored the correlations between ME1 expression by IHC and clinicopathological parameters in our cohort (Table 1). Comparing to ME1-low cases, ME1-high cases were significantly associated with larger tumor size (p = 0.036), higher incidence of lymph node metastasis (p < 0.001) and higher incidence of lymph-vascular invasion (p = 0.001). However, we did not find any specific relations of ME1 expression with age, histopathologic type, histologic grade, ki67 index or molecular subtype, partially due to the limited sample size of our cohort. Moreover, we investigated the correlation between ME1 mRNA level and clinicopathological parameters in Curtis Breast dataset via Oncomine database.11 High ME1 mRNA level correlated to higher histologic grade, later TNM stage, TNBC and HER2 positive breast cancer (Figure 1B). In short, high expression of ME1 in both the mRNA and protein levels was associated with risk factors of breast cancer recurrence.
 |
Table 1 Clinicopathological Parameters of Patients and ME1 Protein Expression |
High ME1 Expression Indicated Worse Prognosis of Breast Cancer Patients
Since ME1 expression correlated with risk factors of breast cancer recurrence, we further evaluated the association between ME1 expression and RFS in our cohort. Survival analysis by the Log rank test showed high ME1 expression was significantly correlated with worse RFS (p < 0.01) (Figure 1C). In addition, univariable cox regression showed pathologic N stage ≥2, lymph-vascular invasion and ME1 high expression were associated with worse RFS (p < 0.1). Further multivariable cox regression including the above three variables identified ME1 high expression as an independent negative prognostic factor for RFS (HR = 5.343, 95% CI = 1.191–23.971, p = 0.029) (Table 2). Moreover, we conducted stratified analysis which revealed high ME1 expression was related to worse RFS among cases more than 45 years old (HR = 10.725, 95% CI = 1.394–82.482, p = 0.023) and among those with Ki67 index ≥20% (HR = 4.127, 95% CI = 1.176–14.485, p = 0.027) (Figure 2).
 |
Table 2 Survival Analysis of RFS by Cox Regression |
Finally, we investigated the relationships between ME1 mRNA expression and survival in breast cancer via an online tool KM plotter.12 Consistently, high ME1 mRNA expression related to both worse RFS (HR = 1.29, 95% CI = 1.04–1.59, p < 0.05) (Figure 1D) and worse overall survival (HR = 1.39, 95% CI = 1.24–1.55, p < 0.05) (Figure 1E). Collectively, high expression of ME1 indicated worse prognosis of breast cancer patients.
Breast Cancer Cell Lines with High Malignancy Showed High ME1 Expression
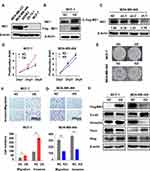
The correlations between ME1 expression with clinicopathological features and outcomes suggest ME1 could participate in breast cancer progression. To further evaluate the role of ME1 in breast cancer progression in the experimental context, we first investigate the endogenous ME1 expression in several representative breast cancer cell lines with diverse malignancy by Western blotting assays.13 ME1 showed higher expression in the more aggressive cell lines including TNBC cell lines (MDA-MB-468 and MDA-MB-231) and HER2 positive cell line (SKBR3), while lower expression in luminal cell lines (ZR-75-1 and MCF-7) (Figure 3A).
ME1 Promoted Proliferation, Migration, Invasion, and EMT in Breast Cancer Cells
To determine the effects of ME1 in breast cancer cells, we constructed MCF-7 cell line stably overexpressing ME1 and MDA-MB-468 cell line stably expressing shRNA against ME1. The overexpressing and silencing effect were verified via Western blotting assays (Figure 3B and C). We first conducted cell proliferation assays. Ectopic expression of ME1 in MCF-7 cells significantly accelerated cell proliferation in CCK-8 assays and increased colony formation in colony growth assays (p < 0.001). On the contrary, cell proliferation and clonogenicity were suppressed with knockdown of ME1 in MDA-MB-468 cells (p < 0.01) (Figure 3D and E). Next, we examined whether ME1 could influence the ability of cell migration and invasion, which is another distinct feature of malignant tumor. Overexpression of ME1 in MCF-7 cells enhanced cell motility in both transwell migration and invasion assays (p < 0.05) (Figure 3F), accompanied by the development of epithelial–mesenchymal transition (EMT) (Figure 3H). In contrast, knockdown of ME1 in MDA-MB-468 cells produced totally opposite effects as expected (p < 0.05) (Figure 3G and H).
Regulating ROS Inhibited the Effect of ME1 on Cell Migration
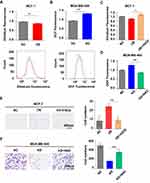
It is acknowledged that ME1 catalyzes malate to pyruvate, accompanied by NADP+ converting to NADPH, which plays a crucial role on ROS homeostasis. Thus, we assessed ROS level to explore the possible mechanism by which ME1 could promote breast cancer progression. In accordance with our speculation, DHE fluorescence revealed that ME1 overexpression significantly decreased ROS level in MCF-7 cells (p < 0.01) (Figure 4A). Oppositely, ME1 knockdown apparently increased ROS level in MDA-MB-468 cells by DCF fluorescence (p < 0.001) (Figure 4B). When pretreated with hydrogen peroxide, an oxidizing agent, MCF-7 cells overexpressing ME1 showed higher ROS level (p < 0.05) and lost its motility in transwell migration assays (p < 0.001) (Figure 4C and E). Moreover, MDA-MB-468 cells with knockdown of ME1 showed lower ROS level (p < 0.001) and restored its motility when pretreated with N-acetyl cysteine, an antioxidant (p < 0.001) (Figure 4D and F).
Discussion
In the present study, we demonstrated that ME1 high expression was associated with risk factors of breast cancer recurrence and indicated worse prognosis. Thus, we found out the important role of ME1 in breast cancer. Further work showed ME1 promoted cell proliferation, migration, invasion ability, and EMT in breast cancer cells, probably by decreasing intracellular ROS level, which probably reflects ROS level in the tumor micro-environment.
Previous study of ME1 in breast cancer only focusing on TNBC subtype drew the same conclusion as ours that ME1 was highly expressed in TNBC cells,14 but its upstream up-regulators have not been reported yet. We also found the relationship between ME1 expression and HER2 positive in Curtis Breast Dataset. This phenomenon was supported by the evidence that NR1D1, the positive transcription factor of ME1, resides on ERBB2-containing 17q12-21 amplicons and is part of the ERBB2 expression signature.15 p53 can inhibit ME1, and downregulation of ME1 reciprocally activates p53 through distinct MDM2- and AMP-activated protein kinase-mediated mechanisms in a feed-forward manner.8 ME2 is frequently hemizygously codeleted with SMAD4, and ME1, as its isoenzyme, is transcriptionally upregulated in an ETV4-dependent manner.16 ME1 upregulation occurs via direct transcriptional activation mediated by β-catenin/TCF1, in which way oncogenic WNT/β-catenin signaling facilitates tumor growth.17 These findings show the complex regulation mechanisms of ME1 and further exploration is needed.
The role of ME1 in breast cancer has not been fully elucidated. Although ME1 is reported to enhance cell proliferation and metastasis capacity in multiple cancer types,14,18-20 the underlying mechanism of which still remains inexplicit. On one hand, ME1 catalyzes malate to pyruvate, inducing cell glucose uptake and lactate production and thus promote Warburg effect, which is favorable for tumor.14 On the other hand, NADPH generated from ME1-catalyzed reaction reduces ROS accumulation, which is the common byproduct during tumor progression. Otherwise, excessive ROS can result in macromolecules damage including lipids, proteins, and nucleic acids, which is unfavorable for tumor.21 Moreover, ROS released to micro-environment may lead to ROS activation in other cell types including fibroblasts, vascular endothelial cells, immune cells, etc., which can in turn have a crosstalk with tumor cells. Our study implied that ME1 could help maintain the homeostasis of ROS and partially explained the function of ME1, in accordance with the results in gastric cancer cells.16
As far as we know, there are still no anti-tumor drugs targeting ME1. Considering the important role of ME1 in tumor progression, we also reflect on whether ME1 is a potential therapeutic target. Since ME1 is widely expressed in human tissues and it proves essential for the survival of normal fibroblasts,22 ME1 inhibition might do harm to normal tissues severely and may not be a suitable therapeutic strategy. As demonstrated above, ME1 is highly expressed in HER2 positive cancer; hence, ME1 inhibition conjugated to trastuzumab could be a possible treatment for HER2 positive patients. Alternatively, targeting glycolysis, NADPH-producing enzymes, redox-regulating enzymes23 or potential downstream molecules of ME1 may be a better approach.
There are some limitations in our study. The follow-up period of our cohort was still short and the recurrence rate was only 7.7% (17/220) by the end-up point of our study, but our results were supported by data from KM plotter with a longer follow-up period. In order to identify whether ME1 influence the efficacy of treatment, information of adjuvant therapy including chemotherapy, anti-HER2 therapy and radiotherapy requires to be updated, which is ongoing in our center. Besides, we did not find the exact molecular mechanisms by which ME1 promoted tumor progression, which is now in active research in our laboratory. Despite these limitations, this study can clearly indicate the important role of ME1 in breast cancer and give emphasis on the relationship between Warburg effect and breast cancer.
Conclusions
In breast cancer tissues, high ME1 expression determined by IHC was significantly associated with risk factors of breast cancer recurrence and indicated worse prognosis. In vitro, ME1 enhanced viability, motility, and EMT. The influence of ME1 on cell motility can be reversed by manipulating ROS. These clinical and experiment works first suggested that ME1 may be a novel biomarker and potential therapeutic target for breast cancer metastasis, and its biological effect is mainly controlled by manipulating ROS.
Abbreviations
CCK-8, Cell counting kit-8; CI, Confidence interval; DCFH-DA, 2ʹ,7ʹ-Dichlorodihydrofluorescein diacetate; DHE, Dihydroethidium; ER, Estrogen receptor; HER2, Human epidermal growth factor receptor-2; HR, Hazard ratio; IHC, Immunohistochemistry; IRS, Immunoreactive score; KM, Kaplan–Meier; ME1, Malic enzyme 1; NADP, Nicotinamide adenine dinucleotide phosphate; NADPH, Nicotinamide adenine dinucleotide phosphate hydrate; RFS, Recurrence-free survival; ROS, Reactive oxygen species; shRNA, Short hairpin RNA; TNBC, Triple negative breast cancer.
Data Sharing Statement
The datasets used and/or analysed during the current study are available from the corresponding author Zhonghua Tao (Email: [email protected]) on reasonable request.
Ethics Approval and Consent to Participate
This study was approved by ethics committee of Fudan University Shanghai Cancer Center. Written informed consent was obtained from all patients to utilize tissues for research. The study was performed in accordance with the Declaration of Helsinki.
Disclosure
The authors declare that they have no competing interests for this work.
References
1. Harbeck N, Gnant M. Breast cancer. Lancet. 2017;389(10074):1134–1150. doi:10.1016/S0140-6736(16)31891-8
2. Pondé NF, Zardavas D, Piccart M. Progress in adjuvant systemic therapy for breast cancer. Nat Rev Clin Oncol. 2019;16(1):27–44. doi:10.1038/s41571-018-0089-9
3. Krag DN, Anderson SJ, Julian TB, et al. Sentinel-lymph-node resection compared with conventional axillary-lymph-node dissection in clinically node-negative patients with breast cancer: overall survival findings from the NSABP B-32 randomised Phase 3 trial. Lancet Oncol. 2010;11(10):927–933. doi:10.1016/S1470-2045(10)70207-2
4. Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–674. doi:10.1016/j.cell.2011.02.013
5. Vander Heiden MG, Cantley LC, Thompson CB. Understanding the warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324(5930):1029–1033. doi:10.1126/science.1160809
6. Wu Z, Wu J, Zhao Q, Fu S, Jin J. Emerging roles of aerobic glycolysis in breast cancer. Clin Transl Oncol. 2020;22(5):631–646.
7. Yao P, Sun H, Xu C, et al. Evidence for a direct cross-talk between malic enzyme and the pentose phosphate pathway via structural interactions. J Biol Chem. 2017;292(41):17113–17120. doi:10.1074/jbc.M117.810309
8. Jiang P, Du W, Mancuso A, Wellen KE, Yang X. Reciprocal regulation of p53 and malic enzymes modulates metabolism and senescence. Nature. 2013;493(7434):689–693. doi:10.1038/nature11776
9. Goldhirsch A, Winer EP, Coates AS, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen international expert consensus on the primary therapy of early breast cancer 2013. Ann Oncol. 2013;24(9):2206–2223. doi:10.1093/annonc/mdt303
10. Gdynia G, Sauer SW, Kopitz J, et al. The HMGB1 protein induces a metabolic type of tumour cell death by blocking aerobic respiration. Nat Commun. 2016;7(1):10764. doi:10.1038/ncomms10764
11. Curtis C, Shah SP, Chin S-F, et al. The genomic and transcriptomic architecture of 2000 breast tumours reveals novel subgroups. Nature. 2012;486(7403):346–352. doi:10.1038/nature10983
12. Lánczky A, Nagy Á, Bottai G, et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res Treat. 2016;160(3):439–446. doi:10.1007/s10549-016-4013-7
13. Neve RM, Chin K, Fridlyand J, et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell. 2006;10(6):515–527. doi:10.1016/j.ccr.2006.10.008
14. Liao R, Ren G, Liu H, et al. ME1 promotes basal-like breast cancer progression and associates with poor prognosis. Sci Rep. 2018;8(1):16743. doi:10.1038/s41598-018-35106-y
15. Kourtidis A, Jain R, Carkner RD, Eifert C, Brosnan MJ, Conklin DS. An RNA interference screen identifies metabolic regulators NR1D1 and PBP as novel survival factors for breast cancer cells with the ERBB2 signature. Cancer Res. 2010;70(5):1783–1792. doi:10.1158/0008-5472.CAN-09-1550
16. Lu YX, Ju HQ, Liu ZX, et al. ME1 regulates NADPH homeostasis to promote gastric cancer growth and metastasis. Cancer Res. 2018;78(8):1972–1985. doi:10.1158/0008-5472.CAN-17-3155
17. Zhu Y, Gu L, Lin X, et al. Dynamic regulation of ME1 phosphorylation and acetylation affects lipid metabolism and colorectal tumorigenesis. Mol Cell. 2020;77(1):138–149.e5. doi:10.1016/j.molcel.2019.10.015
18. Wen D, Liu D, Tang J, et al. Malic enzyme 1 induces epithelial-mesenchymal transition and indicates poor prognosis in hepatocellular carcinoma. Tumour Biol. 2015;36(8):6211–6221. doi:10.1007/s13277-015-3306-5
19. Zheng FJ, Ye HB, Wu MS, Lian YF, Qian CN, Zeng YX. Repressing malic enzyme 1 redirects glucose metabolism, unbalances the redox state, and attenuates migratory and invasive abilities in nasopharyngeal carcinoma cell lines. Chin J Cancer. 2012;11:519–531.
20. Nakashima C, Yamamoto K, Fujiwara-Tani R, et al. Expression of cytosolic malic enzyme (ME1) is associated with disease progression in human oral squamous cell carcinoma. Cancer Sci. 2018;109(6):2036–2045. doi:10.1111/cas.13594
21. Tsang CK, Liu Y, Thomas J, Zhang Y, Zheng XF. Superoxide dismutase 1 acts as a nuclear transcription factor to regulate oxidative stress resistance. Nat Commun. 2014;5:3446. doi:10.1038/ncomms4446
22. Dey P, Baddour J, Muller F, et al. Genomic deletion of malic enzyme 2 confers collateral lethality in pancreatic cancer. Nature. 2017;542(7639):119–123. doi:10.1038/nature21052
23. Murai S, Ando A, Ebara S, Hirayama M, Satomi Y, Hara T. Inhibition of malic enzyme 1 disrupts cellular metabolism and leads to vulnerability in cancer cells in glucose-restricted conditions. Oncogenesis. 2017;6(5):e329. doi:10.1038/oncsis.2017.34
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.