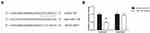Back to Journals » Cancer Management and Research » Volume 12
Long Non-Coding RNA TRPM2-AS Promotes Cell Migration and Invasion by Serving as a ceRNA of miR-138 and Inducing SOX4-Mediated EMT in Laryngeal Squamous Cell Carcinoma
Received 31 May 2020
Accepted for publication 27 July 2020
Published 25 August 2020 Volume 2020:12 Pages 7805—7812
DOI https://doi.org/10.2147/CMAR.S265412
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Harikrishna Nakshatri
Ning Wang,1,2,* Lei Wang,2,* Xinliang Pan1– 3
1Department of Otorhinolaryngology, Qilu Hospital, Cheeloo College of Medicine, Shandong University, Jinan, Shandong 250012, People’s Republic of China; 2Department of Otolaryngology, Qilu Hospital (Qingdao), Cheeloo College of Medicine, Shandong University, Qingdao, Shandong 266035, People’s Republic of China; 3NHC Key Laboratory of Otorhinolaryngology (Shandong University), Jinan, Shandong 250012, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Xinliang Pan Department of Otorhinolaryngology
Qilu Hospital, Cheeloo College of Medicine, Shandong University, Jinan, Shandong 250012, People’s Republic of China
Email [email protected]
Background: Laryngeal squamous cell carcinoma (LSCC) is a common type of malignant tumors of larynx, and in this study, we aimed to evaluate the functional role of long non-coding RNA TRPM2-AS in LSCC.
Methods: The expression levels of TRPM2-AS in LSCC tissues and cell lines were detected by RT-qPCR analysis. In vitro functional assays, including MTT assay and transwell assay, were performed to explore the biological effects of TRPM2-AS on LSCC cells. The expression levels of EMT-relevant proteins were detected by Western blot analysis. The interaction between TRPM2-AS and miR-138 in LSCC, predicted by bioinformatic method, was verified by dual-luciferase reporter assay.
Results: We observed that TRPM2-AS was highly expressed in human LSCC tissues and cell lines. LSCC patients with advanced clinical stage exhibited higher intratumoral TRPM2-AS expression. The results of functional assays demonstrated that TRPM2-AS knockdown remarkably inhibited the proliferation, migration and invasion of LSCC cells, whereas TRPM2-AS overexpression showed opposite effects. In mechanism, we further observed that TRPM2-AS directly bound to miR-138 and served as competing endogenous RNA (ceRNA), thereby increasing SOX4 expression and promoting EMT in LSCC. The oncogenic effects of TRPM2-AS in LSCC cells were partly diminished by miR-138 restoration.
Conclusion: In short, our findings provided first evidence that TRPM2-AS is highly expressed and exerts its oncogenic role in LSCC partly by miR-138/SOX4 axis.
Keywords: laryngeal squamous cell carcinoma, long non-coding RNA TRPM2-AS, miR-138, SOX4, EMT
Introduction
Laryngeal cancer is a common malignant neoplasm of head and neck, and laryngeal squamous cell carcinoma (LSCC) accounts for more than 90% of all laryngeal cancer cases.1 Although considerable progresses have been achieved in therapeutic methods, including surgery, chemotherapy and radiotherapy, due to frequent metastasis and recurrence, the long-term prognosis of LSCC patients remains largely dismal.2 Therefore, there is an urgent need to better understand the molecular mechanisms of LSCC, and identify potential therapeutic targets.
The human genome produces both coding and non-coding RNAs. Long non-coding RNAs (lncRNAs) are defined as a group of regulatory RNA transcripts longer than 200 nucleotides with limited protein-coding potential.3 Once regarded as part of the “dark matter” of genome, lncRNAs are aberrantly expressed in cancer cells and serve as oncogenes or tumor suppressors.4 TRPM2-AS is a newly discovered lncRNA generated from the antisense of TRPM2,5 and recently, it was reported to be upregulated and closely associated with tumor progression in non-small cell lung cancer6 and gastric cancer.7 In this study, we aimed to investigate the expression pattern and regulatory effects of TRPM2-AS in LSCC.
Patients and Methods
Patients and Tissue Samples
Paired tumor tissues and adjacent normal tissues were collected from 77 patients with LSCC who underwent surgery at Qilu Hospital (Jinan, China). None of the patients received any local or systemic treatment before surgery. All samples were immediately snap-frozen in liquid nitrogen and then stored at −80°C until further use. All the protocols in this study were approved by the Ethics Committee of Qilu Hospital. Written informed consent was obtained from all patients.
Cell Culture and Transfection
Human LSCC cell lines AMC-HN-8, Tu-177 and normal human bronchial epithelial cell line 16HBE, obtained from the Cell Bank of Chinese Academy of Sciences (Shanghai, China) and Shanghai Bioleaf Biotech Co., Ltd. (Shanghai, China), were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, Inc., Waltham, MA, USA) containing 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) and 1% penicillin/streptomycin in an incubator at 37°C with 5% CO2.
TRPM2-AS small interfering RNA (si-TRPM2-AS) and non-targeting small interfering RNA (si-NC), miR-138 mimics and mimics control were designed by Shanghai GenePharma Co., Ltd. (Shanghai, China). The full-length sequence of human TRPM2-AS cDNA was amplified by PCR, and then subcloned into the vector pcDNA3.1 (Invitrogen). Cells were transfected with the aforementioned plasmids and oligonucleotides using Lipofectamine 2000 (Invitrogen). After 48 h, the cells were harvested, and transfection efficiency was determined by RT-qPCR analysis.
RNA Extraction and RT-qPCR Analysis
Total RNA was extracted from tissues and cells using TRIzol reagent (Invitrogen). Cytoplasmic and nuclear RNAs of cells were extracted and purified using PARIS™ kit (Invitrogen). cDNA was synthesized using the PrimeScript™ RT reagent kit (TaKaRa, Dalian, China). PCR amplification was then conducted on an ABI PRISM 7300 Sequence Detection system (Applied Biosystems, Foster City, CA, USA) using a SYBR Premix Ex Taq II kit (TaKaRa). Relative gene expression was analyzed using 2−ΔΔCt method.8 GAPDH or U6 was the reference.
MTT Assay
Cells were plated into 96-well plates (5000 cells/well). Following incubation for indicated time points, 20 μL of MTT reagent (0.5 mg/mL; Merck KGaA, Darmstadt, Germany) was added to each well, and the cells were incubated at 37°C for additional 4 h. Then, the supernatant was discarded, and 200 μL of DMSO was added to dissolve the formazan. The absorbance at 570 nm was read using a microplate reader (MultiskanEX, Lab systems, Helsinki, Finland).
Transwell Assay
Cells were suspended in serum-free media and then plated into the upper chambers of transwell plates (8 μm pore size; Corning Inc., Corning, NY, USA). The lower chamber was supplemented with 600 μL medium containing 10% FBS. After 24 h, the cells that passed through the pores were fixed with methanol, and then stained with 0.1% crystal violet. Then, the stained cells were counted with five random fields.
Protein Extraction and Western Blot Analysis
Total protein was extracted from tissues and cells using radioimmunoprecipitation assay (RIPA) lysis buffer (Beyotime, Shanghai, China). Equal amounts of total protein were separated by SDS-PAGE, and then transferred to PVDF membranes (Millipore, Bedford, MA, USA). After blocking with 5% fat-free milk for 1 h, the membranes were incubated with primary antibodies at 4°C overnight, followed by the incubation with appropriate HRP-conjugated secondary antibody. Next, the membranes were exposed using an enhanced chemiluminescence kit (GE Healthcare, Little Chalfont, UK). GAPDH was used as the internal loading control.
Dual-Luciferase Reporter Assay
The wild-type (WT) or corresponding mutant (MUT) miR-138 binding sites in the TRPM2-AS or SOX4 mRNA fragment were cloned into the psiCHECK-2 luciferase reporter vector (Promega, Madison, WI, USA). By using Lipofectamine 2000, HEK293T cells were co-transfected with the luciferase reporter plasmid and miR-138 mimics or mimics control. After 48 h of transfection, the luciferase activity was detected using the Dual-Luciferase Reporter Assay System (Promega).
Statistical Analysis
All statistical analyses were conducted using GraphPad Prism 6.0 software (GraphPad Software, Inc., La Jolla, CA, USA) and SPSS 18.0 software (SPSS Inc., Chicago, IL, USA). Data were expressed as mean ± standard deviation (SD) of three repeated experiments. Comparisons between groups were conducted using Student’s t-test or one-way analysis of variance (ANOVA) test. Survival curves were plotted using Kaplan–Meier method and analyzed by Log rank test. A value of P<0.05 was considered to indicate a statistically significant difference.
Results
TRPM2-AS is Upregulated in LSCC
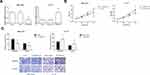
The relative expression of TRPM2-AS detected by RT-qPCR analysis was notably higher in the LSCC tissues than that in the adjacent normal tissues (Figure 1A). In addition, as exhibited in Figure 1B, LSCC cell lines (AMC-HN-8, Tu-177) had a higher TRPM2-AS expression level compared with normal 16HBE cell line. Next, according to the median expression level of intratumoral TRPM2-AS, these LSCC patients were allocated into high expression group (n=40) and low expression group (n=37). As listed in Table 1, high TRPM2-AS expression was closely associated with advanced clinical stage (P=0.023) of LSCC patients.
 |
Table 1 Association Between TRPM2-AS Expression and Clinicopathological Characteristics of LSCC Patients |
TRPM2-AS Promotes LSCC Cell Proliferation, Migration and Invasion
To further understand the regulatory functions of TRPM2-AS on LSCC cells, we used si-TRPM2-AS to decrease the endogenous level of TRPM2-AS in AMC-HN-8 cells (Figure 2A). In parallel, TRPM2-AS was markedly overexpressed in Tu-177 cells by transfection with pcDNA3.1-TRPM2-AS. MTT assay indicated that the proliferation ability of AMC-HN-8 cells was notably impaired by TRPM2-AS knockdown (Figure 2B). In contrast, TRPM2-AS overexpression caused a significant promotion in Tu-177 cell proliferation. Moreover, as shown in Figure 2C, the migration and invasion of AMC-HN-8 cells were notably inhibited when TRPM2-AS was silenced, while Tu-177 cells with TRPM2-AS overexpression showed enhanced migratory and invasive abilities.
TRPM2-AS Binds with miR-138 in LSCC
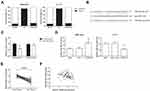
We noticed that TRPM2-AS was mainly located in the cytoplasm of AMC-HN-8 and Tu-177 cells (Figure 3A), indicating its potential role as ceRNAs to sequester miRNAs. We therefore screened miRNAs that have complementary base pairing with TRPM2-AS by the Starbase database (http://starbase.sysu.edu.cn/index.php). Among the results, miR-138, a tumor suppressor widely identified in many cancers, was chosen for follow-up investigation (Figure 3B). The direct binding relation was then validated by dual-luciferase reporter assay, and the results showed that the luciferase activity of HEK293T cells transfected with TRPM2-AS-WT plasmid was markedly reduced by miR-138 mimics (Figure 3C). Besides, the miR-138 expression level was increased or decreased by TRPM2-AS knockdown or overexpression, respectively, in AMC-HN-8 and Tu-177 cells (Figure 3D). As shown in Figure 3E, the expression of miR-138 was notably reduced in LSCC tissues, compared with adjacent normal tissues. We also identified a closely negative correlation between the expression of TRPM2-AS and miR-138 in LSCC tissues (r=−0.269, P=0.018; Figure 3F).
SOX4 is a Direct Target of miR-138 in LSCC
We further searched the online database TargetScan (http://www.targetscan.org/vert_71/) and found that SOX4 mRNA had miR-138 binding sites in its 3ʹ-UTR (Figure 4A). Dual-luciferase reporter assay showed that miR-138 mimics could markedly reduce the luciferase activity of SOX4-WT plasmid in HEK293T cells (Figure 4B).
TRPM2-AS Promotes EMT in LSCC Cells
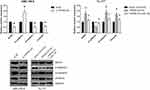
We then evaluated the effects of TRPM2-AS on EMT of LSCC cells by Western blot analysis. The results showed that TRPM2-AS knockdown caused the reduction of SOX4 protein expression and impairment of EMT in AMC-HN-8 cells, as indicated by the increased expression of epithelial markers (E-cadherin) and decreased expression of mesenchymal markers (N-cadherin, Vimentin) (Figure 5). Besides, the increased SOX4 protein level and enhanced EMT in TRPM2-AS-overexpressing Tu-177 cells were effectively diminished by co-transfection with miR-138 mimics.
miR-138 Restoration Blocks the Role of TRPM2-AS in LSCC Cells
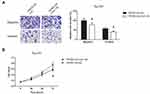
Rescue experiments were then carried out, and as indicated by transwell assay, ectopic expression of miR-138 reversed the promotion effect of TRPM2-AS on Tu-177 cell migration and invasion (Figure 6A). By MTT assay, we also noticed that miR-138 restoration partly reversed the enhanced proliferation of TRPM2-AS-overexpressing Tu-177 cells (Figure 6B).
Discussion
Tumor progression is a multistep process that involves the genetic dysregulation of oncogenes and tumor suppressor genes. In-depth study on the mechanisms underlying LSCC occurrence and development is of critical importance. Lots of lncRNAs are confirmed to be LSCC-associated. For example, overexpression of lncRNA snaR is correlated with progression and predicts poor survival of LSCC,9 while lncRNA NEF may inhibit proliferation and promote apoptosis of LSCC cells.10
In this study, for the first time, the expression level of TRPM2-AS was identified to be markedly higher in LSCC samples. Abnormal proliferation, migration and invasion are important characteristics of tumor cells. A series of in vitro functional experiments were then performed, and the results showed that these malignant behaviors of LSCC cells were impaired following TRPM2-AS knockdown, but TRPM2-AS overexpression showed the opposite effects, indicating the oncogenic role of TRPM2-AS in LSCC. Epithelial–mesenchymal transition (EMT) is a highly conserved cellular program by which tumor cells obtain migratory and invasive abilities to disseminate at distant organs.11 EMT increases the risk of metastasis and reduces overall survival time in LSCC patients.12 Increasing evidence highlights a relevant role of lncRNAs on EMT regulation,13 and this study also verified the effect of TRPM2-AS in inducing EMT of LSCC cells.
microRNAs (miRNAs), another type of non-coding RNAs, are also major players in cancer biology.14 As elucidated by the competing endogenous RNA (ceRNA) hypothesis, lncRNAs can function as ceRNAs to sequester miRNAs, thereby blocking the regulation of their target protein-coding genes.15 This ceRNA model has been identified in many cancer types, including LSCC.16,17 This study selected miR-138 as a candidate, and the direct binding relation between TRPM2-AS and miR-138 was validated in LSCC. We also found the downregulation of miR-138 in LSCC, in agreement with the previous literature.18 SOX4, a master regulator of EMT,19 is further proven as a direct target of miR-138 in LSCC. Rescue experiments indicated that the oncogenic role of TRPM2-AS in LSCC cells was blocked by miR-138 restoration. Based on these findings, we speculated that TRPM2-AS could sponge miR-138 to eliminate its repression on SOX4 function in LSCC cells.
Collectively, our research, for the first time, came into a conclusion that TRPM2-AS is highly expressed and exerts its oncogenic role in LSCC partly by miR-138/SOX4 axis, shedding new light on TRPM2-AS-directed diagnostics and therapeutics in LSCC.
Disclosure
The authors report no conflicts of interest for this work.
References
1. Hunter KD, Parkinson EK, Harrison PR. Profiling early head and neck cancer. Nat Rev Cancer. 2005;5(2):127–135. doi:10.1038/nrc1549
2. Shah JP, Karnell LH, Hoffman HT, et al. Patterns of care for cancer of the larynx in the United States. Arch Otolaryngol Head Neck Surg. 1997;123(5):475–483. doi:10.1001/archotol.1997.01900050021002
3. Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12(12):861–874. doi:10.1038/nrg3074
4. Lavorgna G, Vago R, Sarmini M, Montorsi F, Salonia A, Bellone M. Long non-coding RNAs as novel therapeutic targets in cancer. Pharmacol Res. 2016;110:131–138. doi:10.1016/j.phrs.2016.05.018
5. Orfanelli U, Wenke AK, Doglioni C, Russo V, Bosserhoff AK, Lavorgna G. Identification of novel sense and antisense transcription at the TRPM2 locus in cancer. Cell Res. 2008;18(11):1128–1140. doi:10.1038/cr.2008.296
6. Huang C, Qin Y, Liu H, et al. Downregulation of a novel long noncoding RNA TRPM2-AS promotes apoptosis in non-small cell lung cancer. Tumour Biol. 2017;39(2):1010428317691191.
7. Xiao J, Lin L, Luo D, et al. Long noncoding RNA TRPM2-AS acts as a microRNA sponge of miR-612 to promote gastric cancer progression and radioresistance. Oncogenesis. 2020;9(3):29. doi:10.1038/s41389-020-0215-2
8. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
9. Liang K, Yang Y, Zha D, Yue B, Qiu J, Zhang C. Overexpression of lncRNA snaR is correlated with progression and predicts poor survival of laryngeal squamous cell carcinoma. J Cell Biochem. 2019;120(5):8492–8498.
10. Cui X, Fang N, Cui Y, Xiao D, Wang X. Long non-coding RNA NEF inhibits proliferation and promotes apoptosis of laryngeal squamous cell carcinoma cells by inhibiting Wnt/beta-catenin signaling. Oncol Lett. 2019;17(6):4928–4934. doi:10.3892/ol.2019.10150
11. Ombrato L, Malanchi I. The EMT universe: space between cancer cell dissemination and metastasis initiation. Crit Rev Oncog. 2014;19(5):349–361. doi:10.1615/CritRevOncog.2014011802
12. Zhu GJ, Song PP, Zhou H, et al. Role of epithelial-mesenchymal transition markers E-cadherin, N-cadherin, beta-catenin and ZEB2 in laryngeal squamous cell carcinoma. Oncol Lett. 2018;15(3):3472–3481. doi:10.3892/ol.2018.7751
13. Gugnoni M, Ciarrocchi A. Long noncoding RNA and epithelial–mesenchymal transition in cancer. Int J Mol Sci. 2019;20(8). doi:10.3390/ijms20081924
14. Munker R, Calin GA. MicroRNA profiling in cancer. Clin Sci. 2011;121(4):141–158. doi:10.1042/CS20110005
15. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the rosetta stone of a hidden RNA language? Cell. 2011;146(3):353–358. doi:10.1016/j.cell.2011.07.014
16. Sanchez-Mejias A, Tay Y. Competing endogenous RNA networks: tying the essential knots for cancer biology and therapeutics. J Hematol Oncol. 2015;8:30. doi:10.1186/s13045-015-0129-1
17. Kong X, Qi J, Yan Y, et al. Comprehensive analysis of differentially expressed profiles of lncRNAs, mRNAs, and miRNAs in laryngeal squamous cell carcinoma in order to construct a ceRNA network and identify potential biomarkers. J Cell Biochem. 2019;120(10):17963–17974. doi:10.1002/jcb.29063
18. Si F, Sun J, Wang C. MicroRNA-138 suppresses cell proliferation in laryngeal squamous cell carcinoma via inhibiting EZH2 and PI3K/AKT signaling. Exp Ther Med. 2017;14(3):1967–1974. doi:10.3892/etm.2017.4733
19. Tiwari N, Tiwari VK, Waldmeier L, et al. Sox4 is a master regulator of epithelial-mesenchymal transition by controlling Ezh2 expression and epigenetic reprogramming. Cancer Cell. 2013;23(6):768–783. doi:10.1016/j.ccr.2013.04.020
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.