Back to Journals » OncoTargets and Therapy » Volume 13
LOC441178 Overexpression Inhibits the Proliferation and Migration of Esophageal Carcinoma Cells via Methylation of miR-182
Authors Hu W, Chen Z, Chen J, Cai D, Chen C, Fang T
Received 10 July 2020
Accepted for publication 5 October 2020
Published 3 November 2020 Volume 2020:13 Pages 11253—11263
DOI https://doi.org/10.2147/OTT.S271711
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Arseniy Yuzhalin
Weitao Hu,* Zongchi Chen,* Jiangmu Chen, Daxing Cai, Congjie Chen, Taiyong Fang
Department of Gastroenterology, The Second Affiliated Hospital of Fujian Medical University, Quanzhou, Fujian 362000, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Taiyong Fang
Department of Gastroenterology, The Second Affiliated Hospital of Fujian Medical University, 34 North Zhongshan Road, Licheng District, Quanzhou, Fujian 362000, People’s Republic of China
Email [email protected]
Background: Long noncoding RNAs (lncRNAs) have been shown to play an important role in the development and progression of esophageal carcinoma (EC). Recently, lncRNA LOC441178 was shown to be dysregulated in many cancer types; however, the role of LOC441178 in EC remains unclear.
Materials and Methods: Flow cytometry, transwell and wound healing assays were used to measure the apoptosis and migration in esophageal squamous cell carcinoma (ESCC) cells. RT-qPCR was used to detect the level of miR-182 in LOC441178-overexpressed EC cells. In addition, DNA methylation status of miR-182 promoter in LOC441178-overexpressed ESCC cells was detected by methylation-specific PCR (MSP) and bisulfite sequencing PCR.
Results: In this study, we found that LOC441178 negatively regulated miR-182 expression in ESCC cells. In addition, overexpression of LOC441178 inhibited the proliferation and migration and induced apoptosis in ESCC cells via downregulation of miR-182. Moreover, overexpression of LOC441178 markedly inhibited the phosphorylation of Akt and phosphorylation FOXO3a and increased the expression of FOXO3a in ESCC cells via downregulation of miR-182. Mechanistically, LOC441178 overexpression epigenetically suppressed miR-182 expression via DNA methylation. In vivo experiments revealed that overexpression of LOC441178 inhibited ESCC tumor growth in mouse xenograft model.
Conclusion: Collectively, our data suggested that LOC441178 overexpression epigenetically inhibited tumorigenesis of ESCC via DNA methylation of miR-182. These data indicated that the LOC441178/miR-182 axis might represent a novel therapeutic option for the treatment of ESCC.
Keywords: esophageal carcinoma, lncRNA LOC441178, miR-182, DNA methylation
Introduction
Esophageal carcinoma (EC) is the eighth most common malignant tumor worldwide, which poses a great threat to public health.1,2 EC is characterized by aggressive invasion, high rate of metastasis and poor prognosis.3 It is reported that esophageal squamous cell carcinoma (ESCC) is the most common histological subtype of EC, representing nearly 80% of the incidence of all EC.4,5 The 5-year survival rate of patients with EC remains only about 15–25% following diagnosis.6 Currently, traditional therapies such as surgery, radiotherapy and chemotherapy are the effective treatments used for EC.7 However, the serious side effects and unexpected risks are not uncommon, including esophageal perforation, hemorrhage, dysphagia.8,9 Therefore, exploring the promising therapeutic molecular targets may help to understand the diagnosis and treatment of EC.
Long noncoding RNAs (lncRNAs) are a group of noncoding RNAs longer than 200 nucleotides.10 Evidences have been shown that several lncRNAs were dysregulated and participated in the progression of human cancers including ESCC.11,12 LncRNA may function as either oncogenes or tumor suppressors, affecting different cellular processes, such as proliferation, apoptosis, and metastasis.13 Xu et al found that LOC441178 was downregulated in oral squamous cell carcinoma (OSCC) tissues, and overexpression of LOC441178 could inhibit the invasion and migration of squamous carcinoma cells.14 However, the role of LOC441178 in the progression of ESCC remain largely unclear.
MiRNAs are a kind of non-coding RNA molecules, 18–22 nucleotides in length.15 Many microRNAs can function as tumor suppressors and oncogenes, which are associated with tumor carcinogenesis.16 MiR-182 plays an oncogenic role in the progression of human cancers, such as hepatocellular carcinoma, oral squamous cell carcinoma, melanoma, and EC.17–20 Zhu et al found that miR-182-5p was significantly upregulated in ESCC tissues, and overexpression of miR-182-5p could promote the growth and metastasis in ESCC cells.21 These findings support that miR-182 may functions as an oncogene in EC.
Epigenetic changes play an important role in the occurrence and development of cancers.22 Epigenetic mechanisms include DNA methylation, chromatin remodeling and histone modifications, which could regulate gene expression and cellular differentiation.23 DNA methylation plays an important role in the regulation of EC.24 It has been shown that lncRNAs can interact with miRNA and consequently mediate gene expression at the epigenetic, transcriptional, and post-transcriptional levels.25 In addition, lncRNAs can downregulate miRNAs via DNA methylation.26 However, the interaction of LOC441178 and miR-182 in EC was seldom explored.
In this study, we showed that overexpression of LOC441178 reduced the level of miR-182 in ESCC cells. In addition, LOC441178 overexpression was sufficient to suppress the tumorigenicity of ESCC cells through downregulating miR-182 expression. Mechanistically, LOC441178 may epigenetically inhibit miR-182 expression through DNA methylation. Our data suggested that LOC441178/miR-182 axis may serve as a promising therapeutic option for the treatment of ESCC.
Materials and Methods
Cell Culture
The human ESCC cell lines TE-9 and Eca-109 were purchased from Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). The cell lines were grown in DMEM medium supplemented with 10% fetal bovine serum (FBS; Thermo Fisher Scientific, Waltham, MA, USA) and 1% penicillin-streptomycin, and cultured in a humidified atmosphere of 37°C with 5% CO2.
Cell Transfection and Lentivirus Infection
MiR-182 agomir was purchased from GenePharma (Shanghai, China). Lipofectamine 2000 (Thermo Fisher Scientific) was used to transfect miR-182 agomir into TE-9 and Eca-109 cells.
The control and LOC441178-overexpressed lent viruses were obtained from Ribobio (Guangdong, China), named vector-control and LOC441178-OE. The plasmids expressing lentivirus, packaging plasmid (pAX2) and envelope plasmid (pMD2.G) were transfected into 293T cells for 72 h. After that, virus-containing supernatant was collected, and then added into TE-9 and Eca-109 cells in the presence of polybrene (Santa Cruz Biotechnology, CA, USA) for 48 h. Later on, infected cells were then cultured with 2.5 μg/mL puromycin (Thermo Fisher Scientific) to select stable LOC441178-overexpressing cells. Cells of the blank control group were not transfected.
Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
The total RNA from TE-9 and Eca-109 cells was isolated using TRIzol reagent (Thermo Fisher Scientific). Reverse transcription reactions were performed using the EntiLink™ 1st Strand cDNA Synthesis Kit (ELK Biotechnology). After that, real-time PCR was performed using SYBR Premix Ex TaqII Kit (Takara Bio Inc. Shiga, Japan) on the StepOnePlus System (Applied Biosystems, CA, USA). The relative expression of LOC441178 was calculated using the 2−ΔΔCt method, and β-actin was used as endogenous controls. The relative expressions of miR-21, miR-182, miR-141, miR-424 and miR-543 were calculated using the 2−ΔΔCt method, and U6 was used as endogenous controls. β-actin, Forward, 5ʹ-GTCCACCGCAAATGCTTCTA-3ʹ; Reverse, 5ʹ-TGCTGTCACCTTCACCGTTC-3ʹ. LOC441178, Forward, 5ʹ-TGGTGAAAACCACGTGGAAC-3ʹ; Reverse, 5ʹ-GAGTCCACCAAAGGGAGAAGTC-3ʹ. U6, Forward, 5ʹ-CTCGCTTCGGCAGCACAT-3ʹ; Reverse, 5ʹ-AACGCTTCACGAATTTGCGT-3ʹ. MiR-182, Forward, 5ʹ-CAACTACTCAACTGAATTGCCGA-3ʹ; Reverse, 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ. MiR-141, Forward, 5ʹ-GCATCTTCCAGTACAGTGTTGG-3ʹ; Reverse, 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ. MiR-21, Forward, 5ʹ-TGGGCTTATCAGACTGATGTTGA-3ʹ; Reverse, 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ. MiR-424, Forward, 5ʹ-ATAATCAAAACGTGAGGCGCT-3ʹ; Reverse, 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ. MiR-543, Forward, 5ʹ-AACATTCGCGGTGCACTTC-3ʹ; Reverse, 5ʹ-CTCAACTGGTGTCGTGGAGTC-3ʹ.
CCK-8 Assay
Cell Counting Kit-8 (CCK8, Dojindo Molecular Technologies, Inc., Kyushu, Japan) was used to analyze cell viability. TE-9 or Eca-109 cells were plated at 5 × 103 per well in 96-well plates, and incubated overnight at 37°C. After that, cells were infected with vector-control and LOC441178-OE for 12, 24 and 48 h, respectively. Then, each well was added with 10 μL CCK-8 reagent and incubated for another 2 h. Subsequently, optical density (OD) value was measured using a microplate reader (Thermo Fisher Scientific) at a wavelength of 450 nm.
5-Ethynyl-2′-Deoxyuridine (EdU) Assays
EdU DNA Proliferation Detection kit (RiboBio, China) was used to determine the cell proliferation. TE-9 and Eca-109 cells were infected with vector-control and LOC441178-OE for 48 h, and then cells were stained by EdU and DAPI. Later on, the images were captured with a fluorescence microscope (Leica, Buffalo Grove, IL, USA). ImageJ software was used to quantify the number of EdU-positive cells.
Methylation-Specific PCR (MSP) and Bisulfite Sequencing PCR (BSP)
Genomic DNA, extracted and purified by using the Genomic DNA Extraction Kit (TaKaRa, Dalian, China), was bisulfite-modified by the EZ-96 DNA Methylation Kit (Zymo Research, Irvine, CA, USA) according to the manufacturer’s protocol, and analyzed by MSP, collected the BSP product for sequencing. The unmethylated primers of miR-182 were Forward: 5ʹ-GGGGTTGTTTGGGTTAGTTG-3ʹ; Reverse: 5ʹ-AACAACCCCACCTACCTACAAT-3ʹ. The methylated primers of miR-182 were Forward: 5ʹ-GGGGTTGTTTGGGTTAGTCG-3ʹ; reverse: 5ʹ-ACAACCCCGCCTACCTACG-3ʹ. The PCR reaction conditions were as follows: 98°C for 4 min, 40 cycles of 98°C for 30 s, 56°C for 30 s, and 72°C for 30 s, and then 72°C for 10 min. Later on, PCR products were electrophoresed in 3% agarose gel. Subsequently, the images were captured with an Ultroscan XL Laser Densitometer (Pharmacia LKB, Uppsala, Sweden).
Flow Cytometry Assay
The TE-9 and Eca-109 cells were infected with LOC441178-OE in the presence or absence of miR-182 agomir for 72 h. Cell apoptosis was assessed using an FITC Annexin V/Dead cell apoptosis kit (Thermo Fisher Scientific), and was analyzed on the flow cytometry (BD FACSCanto II system; BD Biosciences, Franklin Lake, NJ, USA) using the CellQuest software (BD Biosciences).
Transwell Migration Assay
Cell migration assay was performed using 24‐well transwell chambers (0.8 μm; Corning Incorporated, Corning, NY, USA). TE-9 cells were suspended in 200 μL serum‐free medium and then added into the upper chamber. Later on, 600 μL of DMEM medium (containing 10% FBS) was added into the lower chamber. After that, cells that invaded the lower surface were stained with 0.2% crystal violet at 24 h. Then, cells were counted using a fluorescence microscope (Olympus CX23 Tokyo, Japan).
Wound Healing Assay
TE-9 cells (2×105 cells/well) were seeded into 12-well culture plate at 37°C overnight. When cells reach about 80% confluent, a wound area was carefully made in the cell monolayer with a sterile 20 μL pipette tip. After that, TE-9 cells were infected with LOC441178-OE in the presence or absence of miR-182 agomir. The width of the wound area was observed at 0 h and 48 h using a fluorescence microscope (Olympus CX23 Tokyo, Japan).
Western Blot Assay
Cells were lysed using a modified radioimmunoprecipitation assay (RIPA, (Beyotime, China) lysis buffer on ice, and the BCA Protein Assay Kit (Beyotime, China) was used to measure the protein concentration. Equivalent amounts of proteins (40 μg per lane) were separated by 10% SDS-PAGE, and then transferred onto a PVDF membrane (Millipore, Billerica, MA, USA). After that, the membrane was incubated with primary antibodies at 4°C overnight. Antibodies were diluted to 1:1000 for p-Akt (Abcam; ab81283), Akt (Abcam; ab179463), p-FOXO3a (Abcam; ab154786), FOXO3a (Abcam; ab23683), cleaved caspase 3 (Abcam; ab49822), β-actin (Abcam; ab6276). Later on, the membrane was incubated with secondary antibodies (Abcam; ab150077) for 1 h at room temperature. Subsequently, the bands were imaged by an enhanced chemiluminescent substrate kit (Thermo Fisher Scientific).
Animal Study
BALB/c nude mice (4–6 weeks old) were purchased from the Shanghai SLAC Animal Center (Shanghai, China). All animal experiments were approved by the Institutional Ethical Committee of the Second Affiliated Hospital of Fujian Medical University, and animals were maintained following the guidelines of the Institutional Animal Care and Use Committee. Animals were randomized into two groups: control and LOC441178-OE groups. 1 x 106 TE-9 or TE-9 cells stably expressing LOC441178-OE cells in 100 μL PBS were injected subcutaneously into the left flank of nude mice. Tumor volume was monitored every week with a caliper and tumor size was calculated as V: volume = length × width2/2. After 28 days of treatment, mice were sacrificed, and the entire tumors were weighed.
Statistical Analysis
All statistical analyses were performed using GraphPad Prism software (version 7.0, La Jolla, CA, USA). Student’s t-test was performed for two group comparisons. One-way analysis of variance (ANOVA) and Tukey’s tests were carried out for multiple group comparisons. All experiments were repeated three times. Data were expressed as the mean ± standard deviation (S.D.). *P<0.05 were considered statistically significant.
Results
Overexpression of LOC441178 Inhibited the Proliferation of ESCC Cells
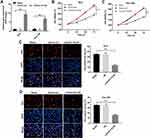
To investigate the role of LOC441178 in ESCC, we established ESCC cell lines (TE-9 and Eca-109) with LOC441178 stable overexpression. As shown in Figure 1A, the level of LOC441178 was significantly upregulated in TE-9 and Eca-109 cells following transfection with LOC441178-OE plasmids. In addition, CCK-8 assay indicated that overexpression of LOC441178 markedly suppressed the viability of TE-9 and Eca-109 cells, compared with the vector-ctrl group (Figure 1B and C). Moreover, upregulation of LOC441178 notably inhibited proliferation of TE-9 and Eca-109 cells, as determined using EdU assays (Figure 1D and E). These data suggested that overexpression of LOC441178 could inhibit the proliferation of ESCC cells.
LOC441178 Regulated DNA Methylation of miR-182 Promoter Region
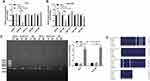
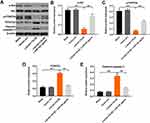
Next, Starbase (http://starbase.sysu.edu.cn) was used to determine the potential targets of LOC441178. The data indicated that miR-21, miR-182, miR-141, miR-424 and miR-543 were closely associated with LOC441178. Meanwhile, these five miRNAs have been found to play important roles in the progression of EC.20,27–30 As shown in Figure 2A and B, overexpression of LOC441178 notably downregulated the level of miR-182 in TE-9 and Eca-109 cells, while no significant changes were observed in the levels of miR-21, miR-141, miR-424 and miR-543 in these conditions in TE-9 and Eca-109 cells.
Evidence has been shown that lncRNAs can regulate miRNA level at the epigenetic level.25 To investigate the effect of LOC441178 on DNA methylation level of miR-182 promoter region, MSP was performed. As shown in Figure 2C, overexpression of LOC441178 in TE-9 and Eca-109 cells significantly decreased DNA methylation levels in the promoter region of miR-182. This data was confirmed by bisulfite sequencing PCR (Figure 2D). These data indicated that LOC441178 may epigenetically reduce miR-182 expression via DNA methylation.
Overexpression of LOC441178 Induced Apoptosis of ESCC Cells Through Downregulating miR-182 Expression
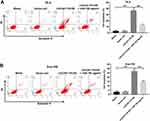
To investigate whether LOC441178 could induce apoptosis via suppressing miR-182 in ESCC, flow cytometry assay was applied. As shown in Figure 3A and B, overexpression of LOC441178 markedly induced apoptosis of TE-9 and Eca-109 cells, and that effect was reversed in the presence of miR-182 agomir. However, miR-182 agomir control had no effect on cell apoptosis (Supplementary Figure 1A). These data indicated that overexpression of LOC441178 could induce apoptosis of ESCC cells through inhibition of miR-182.
Overexpression of LOC441178 Inhibited the Migration Abilities of ESCC Cells Through Downregulating miR-182 Expression
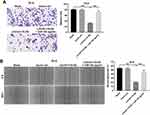
To investigate whether LOC441178 could the migration of ESCC cells via inhibition of miR-182, transwell migration and wound healing assays were used. As indicated in Figure 4A and B, overexpression of LOC441178 notably inhibited the migration ability of TE-9 cells, and this phenomenon was reversed following transfection with miR-182 agomir. However, miR-182 agomir control had no effect on cell migration (Supplementary Figure 1B). These results suggested that overexpression of LOC441178 could inhibit the migration abilities of ESCC cells through inhibition of miR-182.
Overexpression of LOC441178 Inhibited the Growth of ESCC Cells via Regulating the miR-182/Akt/FOXO3a Axis
Next, we further explored the mechanisms underlying the role of LOC441178 in the progression of ESCC. Akt/FOXO3 signaling has been known as the important regulators of cell growth in ESCC.31 As shown in Figure 5A–E, overexpression of LOC441178 decreased phosphorylation of Akt (p-Akt) and phosphorylation of FOXO3a (p-FOXO3a) and increased expressions of FOXO3a and cleaved caspase 3 in TE-9 cells; however, these changes were reversed by miR-182 overexpression. These results illustrated that overexpression of LOC441178 could inhibit the growth of ESCC cells via regulating the miR-182/AktFOXO3a axis.
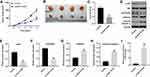
Overexpression of LOC441178 Inhibited the Tumorigenesis of ESCC Subcutaneous Xenografts in vivo
To investigate the role of LOC441178 in regulating the tumor growth of ESCC in vivo, TE-9 and Eca-109 subcutaneous xenograft models were established. As shown in Figure 6A–C, overexpression of LOC441178 significantly inhibited the tumor volume and tumor weight of TE-9 subcutaneous xenografts, compared with control group. In addition, overexpression of LOC441178 obviously decreased p-Akt and p-FOXO3a and increased the expressions of FOXO3a and cleaved caspase 3 in tumor tissues, compared with control group (Figure 6D–H). Moreover, LOC441178-OE markedly upregulated the level of LOC441178 in tumor tissues (Figure 6I). Consistently, the anti-tumor effect of LOC441178 OE on Eca-109 subcutaneous xenografts in vivo was reserved by miR-182 agomir (Supplementary Figure 2A–C). These data revealed that overexpression of LOC441178 could inhibit tumorigenesis of ESCC subcutaneous xenografts in vivo.
Discussion
It has been shown that lncRNAs are aberrantly expressed in almost all cancers and emerging as important diagnostic and prognostic biomarkers in human cancers.32 In addition, dysregulation of lncRNAs has been shown to play important roles in the occurrence and development of EC.33 Li et al indicated that exosomal lncRNA ZFAS1 could promote the proliferation, invasion and migration in ESCC cells.11 Yao et al found that lncRNA-ECM could promote the invasion and metastasis in ESCC cells.34 In this study, we found that overexpression of LOC441178 significantly inhibited the proliferation and migration via inducing apoptosis in ESCC cells. These data indicate that LOC441178 may serve as a tumor-suppressive role in ESCC.
The data in Starbase dataset indicated that miR-21, miR-182, miR-141, miR-424 and miR-543 were closely associated with LOC441178. Considering these five miRNAs are dysregulated in EC,20,27–30 whether LOC441178 is involved in the regulation of miRNAs expression in EC remains unclear. In this study, we identified that LOC441178 negatively regulated miR-182 expression in which LOC441178 overexpression downregulated miR-182 level. In addition, our data showed that overexpression of LOC441178 significantly inhibited the proliferation and induced apoptosis in ESCC cells; however, these changes were reversed by miR-182 agomir. Bai et al found that overexpression of miR-182 promoted the growth and metastasis of ESCC cells, whereas downregulation of miR-182 inhibited the growth and metastasis in cells, indicating that miR-182 serves as an oncogenic role in EC.20 These data indicated that LOC441178 might inhibit the growth of ESCC cells via suppressing miR-182 expression.
Epigenetic alterations, such as DNA methylation, are now recognized as essential players in the initiation and progression of human cancers.35 Xun et al found that lncRNA miR143HG epigenetically suppresses miR-21 expression in EC cells via DNA methylation.36 Cheng et al found that lncRNA HOTAIR mediated miR-122 expression via DNMTs-mediated DNA methylation.37 In this study, overexpression of LOC441178 decreased the level of miR-182 in ESCC cells. Meanwhile, overexpression of LOC441178 in ESCC cells significantly decreased DNA methylation levels in the promoter region of miR-182, suggesting that miR-182 was epigenetically suppressed via DNA methylation in ESCC cells. Evidence has been shown that lncRNAs could regulate the maintenance of DNA methylation and then inhibit the expression of their target genes in cancers.38 These data indicated that LOC441178 could inhibit the expression of miR-182 in ESCC cells via DNA methylation, thus inhibiting the tumorigenesis of EC. It is possible that LOC441178 regulates the DNA methylation of miR-182 promoter region via mediating DNA methyltransferases; however, more investigations are needed and no conclusion was reached so far.
Wang et al indicated that miR-182 could activate the AKT signaling pathway via increasing AKT phosphorylation.39 Liu et al found that overexpression of miR-182 could promote the chemoresistance of colorectal cancer cells through activation of PI3K/AKT signaling pathway.40 Cao et al revealed that miR-182 promote the proliferation and invasion of hepatocellular carcinoma cells via activating AKT/FOXO3a pathway.17 In addition, phosphorylation of Akt could lead to decreased expression of FOXO3a.41 In this study, our data indicated that overexpression of LOC441178 led to the decreased phosphorylation of Akt and increased expressions of FOXO3a and cleaved caspase3 in TE-9 cells; however, these changes were reversed by miR-182 agomir. Thus, these data indicated that overexpression of LOC441178 induced apoptosis and inhibited migration in ESCC cells via inhibiting miR-182/Akt/FOXO3a pathway.
Conclusion
Collectively, our data showed for the first time that overexpression of LOC441178 inhibited the progression of ESCC through downregulation of miR-182. LOC441178 epigenetically suppressed miR-182 expression via DNA methylation. Thus, our results provide a novel mechanistic link between two non-coding RNAs in ESCC: LOC441178 and miR-182 and suggest that the LOC441178/miR-182 axis might represent a novel therapeutic option for the treatment of ESCC. However, further studies are needed to determine the correlation between LOC441178 and miR-182 in ESCC tissues.
Ethical Approval
All of the experiments were approved by the Institutional Ethical Committee of the Second Affiliated Hospital of Fujian Medical University.
Funding
This study was supported by Quanzhou Science and Technology Bureau, Fujian Province, China (Grant No. 2017Z013).
Disclosure
The authors declare no conflicts of interest in this work and no competing financial interests.
These authors contributed equally to this work and should be considered as co-first authors: Weitao Hu and Zongchi Chen.
References
1. Wu G, Chen G, Zhou J, et al. Liriodenine enhances radiosensitivity in esophageal cancer ECA-109 cells by inducing apoptosis and G(2)/M arrest. Oncol Lett. 2018;16(4):5020–5026.
2. Zhang Y, Li S, He H, et al. Influence of Tanshinone IIA on apoptosis of human esophageal carcinoma Eca-109 cells and its molecular mechanism. Thorac Cancer. 2017;8(4):296–303. doi:10.1111/1759-7714.12441
3. Luo HL, Huang MD, Guo JN, et al. AFAP1-AS1 is upregulated and promotes esophageal squamous cell carcinoma cell proliferation and inhibits cell apoptosis. Cancer Med. 2016;5(10):2879–2885. doi:10.1002/cam4.848
4. Kamangar F, Dores GM, Anderson WF. Patterns of cancer incidence, mortality, and prevalence across five continents: defining priorities to reduce cancer disparities in different geographic regions of the world. J Clin Oncol. 2006;24(14):2137–2150. doi:10.1200/JCO.2005.05.2308
5. Napier KJ, Scheerer M, Misra S. Esophageal cancer: a review of epidemiology, pathogenesis, staging workup and treatment modalities. World J Gastrointest Oncol. 2014;6(5):112–120. doi:10.4251/wjgo.v6.i5.112
6. Pennathur A, Farkas A, Krasinskas AM, et al. Esophagectomy for T1 esophageal cancer: outcomes in 100 patients and implications for endoscopic therapy. Ann Thorac Surg. 2009;87(4):1048–1054. doi:10.1016/j.athoracsur.2008.12.060
7. Tantai JC, Zhang Y, Zhao H. Heterophyllin B inhibits the adhesion and invasion of ECA-109 human esophageal carcinoma cells by targeting PI3K/AKT/β-catenin signaling. Mol Med Rep. 2016;13(2):1097–1104. doi:10.3892/mmr.2015.4659
8. Li J, Zhao Z, Du G, et al. Safety and efficacy of pulsed low-dose rate radiotherapy for local recurrent esophageal squamous cell carcinoma after radiotherapy: study protocol for a prospective multi-center Phase II trial. Medicine (Baltimore). 2019;98(26):e16176. doi:10.1097/MD.0000000000016176
9. Bi Y, Li J, Chen H, et al. Fluoroscopic guidance biopsy for severe anastomotic stricture after esophagogastrostomy of esophageal carcinoma: a STROBE-compliant article. Medicine (Baltimore). 2018;97(37):e12316. doi:10.1097/MD.0000000000012316
10. Cabianca DS, Casa V, Bodega B, et al. A long ncRNA links copy number variation to a polycomb/trithorax epigenetic switch in FSHD muscular dystrophy. Cell. 2012;149(4):819–831. doi:10.1016/j.cell.2012.03.035
11. Li Z, Qin X, Bian W, et al. Exosomal lncRNA ZFAS1 regulates esophageal squamous cell carcinoma cell proliferation, invasion, migration and apoptosis via microRNA-124/STAT3 axis. J Exp Clin Cancer Res. 2019;38(1):477. doi:10.1186/s13046-019-1473-8
12. Li Y, Chen D, Gao X, Li X, Shi G. LncRNA NEAT1 regulates cell viability and invasion in esophageal squamous cell carcinoma through the miR-129/CTBP2 axis. Dis Markers. 2017;2017:5314649. doi:10.1155/2017/5314649
13. Li J, Li Z, Zheng W, et al. LncRNA-ATB: an indispensable cancer-related long noncoding RNA. Cell Prolif. 2017;50:6. doi:10.1111/cpr.12381
14. Xu K, Tian H, Zhao S, et al. Long noncoding RNA LOC441178 reduces the invasion and migration of squamous carcinoma cells by targeting ROCK1. Biomed Res Int. 2018;2018:4357647. doi:10.1155/2018/4357647
15. He X, Zhang Z, Li M, et al. Expression and role of oncogenic miRNA-224 in esophageal squamous cell carcinoma. BMC Cancer. 2015;15:575. doi:10.1186/s12885-015-1581-6
16. Esquela-Kerscher A, Slack FJ. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer. 2006;6(4):259–269. doi:10.1038/nrc1840
17. Cao MQ, You AB, Zhu XD, et al. miR-182-5p promotes hepatocellular carcinoma progression by repressing FOXO3a. J Hematol Oncol. 2018;11(1):12. doi:10.1186/s13045-018-0555-y
18. Li N, Nan CC, Zhong XY, et al. miR-182-5p promotes growth in oral squamous cell carcinoma by inhibiting CAMK2N1. Cell Physiol Biochem. 2018;49(4):1329–1341. doi:10.1159/000493411
19. Liu X, Li H, Wu G, Cui S. miR-182 promotes cell proliferation and invasion by inhibiting APC in melanoma. Int J Clin Exp Pathol. 2018;11(4):1900–1908.
20. Bai Q, Yu J, Li Y, Ma J, Gou Y. MicroRNA-182 promoted esophageal squamous cell carcinoma cell growth and metastasis via targeting YWHAG. J BUON. 2018;23(5):1439–1447.
21. Zhu ZQ, Hu HF, Zheng XY, Wang F. [Effects of microRNA-182-5p on cell proliferation and invasion of esophageal squamous cell carcinoma and related molecular mechanisms]. Zhonghua Zhong Liu Za Zhi. 2020;42(8):635–643.
22. Toh TB, Lim JJ, Chow EK. Epigenetics in cancer stem cells. Mol Cancer. 2017;16(1):29.
23. Hwang JY, Aromolaran KA, Zukin RS. The emerging field of epigenetics in neurodegeneration and neuroprotection. Nat Rev Neurosci. 2017;18(6):347–361. doi:10.1038/nrn.2017.46
24. Ma K, Cao B, Guo M. The detective, prognostic, and predictive value of DNA methylation in human esophageal squamous cell carcinoma. Clin Epigenetics. 2016;8:43. doi:10.1186/s13148-016-0210-9
25. Zhang X, Wang W, Zhu W, et al. Mechanisms and functions of long non-coding RNAs at multiple regulatory levels. Int J Mol Sci. 2019;20(22):5573. doi:10.3390/ijms20225573
26. Liu HT, Fang L, Cheng YX, Sun Q. LncRNA PVT1 regulates prostate cancer cell growth by inducing the methylation of miR-146a. Cancer Med. 2016;5(12):3512–3519. doi:10.1002/cam4.900
27. Wen SW, Zhang YF, Li Y, et al. Characterization and effects of miR-21 expression in esophageal cancer. Genet Mol Res. 2015;14(3):8810–8818. doi:10.4238/2015.August.3.4
28. Xue L, Yu X, Jiang X, et al. TM4SF1 promotes the self-renewal of esophageal cancer stem-like cells and is regulated by miR-141. Oncotarget. 2017;8(12):19274–19284. doi:10.18632/oncotarget.13866
29. Wang F, Wang J, Yang X, Chen D, Wang L. MiR-424-5p participates in esophageal squamous cell carcinoma invasion and metastasis via SMAD7 pathway mediated EMT. Diagn Pathol. 2016;11(1):88. doi:10.1186/s13000-016-0536-9
30. Zhao H, Diao C, Wang X, et al. MiR-543 promotes migration, invasion and epithelial-mesenchymal transition of esophageal cancer cells by targeting phospholipase A2 group IVA. Cell Physiol Biochem. 2018;48(4):1595–1604. doi:10.1159/000492281
31. Lu Z, Lu C, Li C, et al. Dracorhodin perchlorate induces apoptosis and G2/M cell cycle arrest in human esophageal squamous cell carcinoma through inhibition of the JAK2/STAT3 and AKT/FOXO3a pathways. Mol Med Rep. 2019;20(3):2091–2100.
32. Ma W, Zhang CQ, Li HL, et al. LncRNA FER1L4 suppressed cancer cell growth and invasion in esophageal squamous cell carcinoma. Eur Rev Med Pharmacol Sci. 2018;22(9):2638–2645.
33. Mao Y, Fu Z, Zhang Y, et al. A seven-lncRNA signature predicts overall survival in esophageal squamous cell carcinoma. Sci Rep. 2018;8(1):8823. doi:10.1038/s41598-018-27307-2
34. Yao J, Shen X, Li H, et al. LncRNA-ECM is overexpressed in esophageal squamous cell carcinoma and promotes tumor metastasis. Oncol Lett. 2018;16(3):3935–3942.
35. Toiyama Y, Okugawa Y, Goel A. DNA methylation and microRNA biomarkers for noninvasive detection of gastric and colorectal cancer. Biochem Biophys Res Commun. 2014;455(1–2):43–57. doi:10.1016/j.bbrc.2014.08.001
36. Xun W, Cen W, Dahai Y, et al. LncRNA miR143HG suppresses miR-21 through methylation to inhibit cell invasion and migration. Laryngoscope. 2019. doi:10.1002/lary.28474
37. Cheng D, Deng J, Zhang B, et al. LncRNA HOTAIR epigenetically suppresses miR-122 expression in hepatocellular carcinoma via DNA methylation. EBioMedicine. 2018;36:159–170. doi:10.1016/j.ebiom.2018.08.055
38. Li W, Zheng J, Deng J, et al. Increased levels of the long intergenic non-protein coding RNA POU3F3 promote DNA methylation in esophageal squamous cell carcinoma cells. Gastroenterology. 2014;146(7):1714–1726.e1715. doi:10.1053/j.gastro.2014.03.002
39. Wang WM, Lu G, Su XW, Lyu H, Poon WS. MicroRNA-182 regulates neurite outgrowth involving the PTEN/AKT pathway. Front Cell Neurosci. 2017;11:96.
40. Liu B, Liu Y, Zhao L, et al. Upregulation of microRNA-135b and microRNA-182 promotes chemoresistance of colorectal cancer by targeting ST6GALNAC2 via PI3K/AKT pathway. Mol Carcinog. 2017;56(12):2669–2680. doi:10.1002/mc.22710
41. Cho HJ, Baek MO, Khaliq SA, et al. Microgravity inhibits decidualization via decreasing Akt activity and FOXO3a expression in human endometrial stromal cells. Sci Rep. 2019;9(1):12094. doi:10.1038/s41598-019-48580-9
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.