Back to Journals » OncoTargets and Therapy » Volume 14
lncRNA GSEC Promotes the Progression of Triple Negative Breast Cancer (TNBC) by Targeting the miR-202-5p/AXL Axis
Authors Zhang J, Du C, Zhang L, Wang Y, Zhang Y, Li J
Received 24 November 2020
Accepted for publication 19 March 2021
Published 20 April 2021 Volume 2021:14 Pages 2747—2759
DOI https://doi.org/10.2147/OTT.S293832
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Jianhua Zhang, Chuang Du, Linfeng Zhang, Yan Wang, Yingying Zhang, Jingruo Li
Department of Breast Surgery, The First Affiliated Hospital of Zhengzhou University, Zhengzhou City, Henan Province, 450000, People’s Republic of China
Correspondence: Jingruo Li
Department of Breast Surgery, The First Affiliated Hospital of Zhengzhou University, No. 1 Jianshe East Road, Erqi District, Zhengzhou City, Henan Province, 450000, People’s Republic of China
Email [email protected]
Background: This study aimed to explore the biological functions of G-quadruplex-forming sequence containing lncRNA (GSEC) in triple negative breast cancer (TNBC).
Methods: The expression of GSEC in TNBC tissues was evaluated by qRT-PCR. Cell viability was evaluated by Cell Counting Kit-8 assay. Cell proliferation was evaluated by 5-ethynyl-20-deoxyuridine (EdU) staining assay. Cell invasion and migration were evaluated by Transwell assay. Gain- and loss-function assays were performed to assess the biological functions of GSEC in TNBC. The interactions between GSEC, miR-202-5p and AXL were determined by luciferase report assay and RNA immunoprecipitation (RIP) assay. In addition, a nude mouse xenograft model was used to confirm the oncogenic role of GSEC in TNBC.
Results: GSEC was significantly upregulated in TNBC tissues and cancer cell lines, and high level of GSEC was associated with advanced tumor stage, positive lymph-node metastasis and the poor prognosis of TNBC patients. Knockdown of GSEC effectively inhibited TNBC cell proliferation, invasion and migration in vitro. GSEC regulated the expression of AXL by directly sponging miR-202-5p. Downregulation of miR-202-5p attenuated GSEC knockdown-induced inhibition on TNBC cell proliferation, invasion and migration in vitro. Meanwhile, overexpression of AXL obviously reversed the inhibitory effects of miR-202-5p mimics in TNBC progression in vitro.
Conclusion: GSEC functioned as a potential oncogene and promoted AXL-mediated TNBC progression by sponging miR-202-5p, which might be a novel diagnostic and therapeutic target for TNBC.
Keywords: TNBC, GSEC, miR-202-5p, AXL
Introduction
Triple negative breast cancer (TNBC) is an aggressive subtype of breast cancer and accounts for approximately 15-20% of breast cancer cases worldwide.1,2 TNBC has a higher incidence of local recurrence and metastasis, which brings a huge burden to human health.3 In the last decades, the main treatments for TNBC are mainly the combination of surgery, conventional chemotherapy and radiotherapy.3 Recent studies suggest that immunotherapy agents in combination with traditional systemic chemotherapy is a promising selection for TNBC treatment.4 However, the prognosis of TNBC patients is still poor mainly due to its recurrence and metastasis.5 Therefore, better understanding of the underlying mechanisms involved in TNBC progression would contribute to the development of effective therapeutic strategies.
Long non-coding RNAs (lncRNAs) are a group of non-coding RNA molecules with more than 200 nucleotides in length and lack the protein encoding function.6 It has been demonstrated that lncRNAs play crucial roles in various biological processes involved in cancer development such as cell proliferation, differentiation, and apoptosis.7 For example, high expression levels of TUC338 were correlated with the poor prognosis of patients with lung cancer, and overexpression of TUC338 enhances cell activity and invasion ability in lung cancer cells.8 PVT1 is highly expressed in prostate cancer, and knockdown of PVT1 significantly suppresses prostate cancer development both in vivo and in vitro.9 Abnormal expression of lncRNAs is frequently observed in breast cancer including TNBC. LINC02273 is significantly upregulated in metastatic lymph nodes of breast cancer, and overexpression of LINC02273 promotes the metastasis of breast cancer by upregulating AGR2 transcription.10 Low expression levels of GAS5 lead to a lower percentage of apoptosis in TNBC cells, and overexpression of GAS5 promotes cell apoptosis in TNBC through targeting the miR-378a-5p/SUFU signaling axis.11 In addition, various lncRNAs have been reported to be dysregulated in TNBC including DANCR,12 MIR100HG,13 NRON,14 and AWPPH.15 An upregulation of G-quadruplex-forming sequence containing lncRNA GSEC has only been observed in osteosarcoma16 and colon cancer.17 However, the function and underlying mechanism of GSEC in TNBC remain unclear.
In this study, we investigated the role of GSEC in TNBC and found that GSEC was significantly upregulated in TNBC tissues and cell lines. High expression levels of GSEC were correlated to advanced tumor stage, positive lymph-node metastasis and poor survival time of TNBC patients. Knockdown of GSEC effectively inhibited cell viability and proliferation of TNBC cells in vitro, and also suppressed tumor growth in vivo. Our results demonstrated that knockdown of GSEC prevented against TNBC progression partially through regulating the miR-202-5p/AXL axis, suggesting that GSEC/miR-202-5p/AXL might be potential diagnostic and therapeutic targets for TNBC.
Materials and Methods
Tissue Specimens
A total of 45 pairs of cancer tissues and matched adjacent normal tissues were obtained by surgical excision from TNBC patients admitted at the First Affiliated Hospital of Zhengzhou University from 2017 to 2019. All patients signed the written informed consent. This study was approved by the Ethics Committee of the First Affiliated Hospital of Zhengzhou University in accordance with the Declaration of Helsinki. The clinicopathological features of 45 TNBC patients are shown in Supplementary Table 1.
Cell Culture
Five TNBC cell lines (MDA-MB-231, BT-549, MDA-MB-468, HCCl937, BT-549 and SK-BR-3) and human breast epithelial cell line MCF-10A were purchased from American Tissue Culture Collection (ATCC). All cells were cultured in DMEM/F12 medium supplementing with 10% FBS (FBS, Hyclone, Life Technologies, CA) at 37°C with 5% CO2.
Cell Transfection
Short hairpin RNA targeting GSEC (sh-GSEC) and scrambled control sh-NC were constructed by GenePharma (Shanghai, China). MiR-202-5p mimics, miR-202-5p inhibitor and corresponding negative controls (miR-NC and inhibitor NC) were synthesized by RiboBio (Guangzhou, China). To overexpress GSEC and AXL, the full length of GSEC and AXL were synthesized by RiboBio (Guangzhou, China) and cloned into pcDNA3.1 expression vector. Empty vector pcDNA3.1 was used as the negative control. The sh-RNAs, overexpression vectors or miR-202-5p mimic/inhibitor were transfected into BT-549 and MDA-MB-468 cells using Lipofectamine 2000 reagent (Invitrogen). Cells were harvested at 48 h after transfection to detect the transfection efficiency and used for the following experiments. The sequence of sh-GSEC was as follows: 5ʹ-ATTGGGAGCTGTGGTCAATTA-3ʹ.
RNA Isolation and qRT-PCR
Total RNAs of tissue samples and cultured cells were extracted using Ribozol RNA Extraction Reagent (Thomas Scientific). Reverse transcription (RT) assay was performed using High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, U.S.A.). Subsequently, qRT-PCR reactions were performed on the 7500 real‑time PCR machine using SYBR Green PCR Master Mix (Applied Biosystems). The relative expression levels were calculated based on the 2−ΔΔCt method. The expression levels of GSEC and AXL were normalized to GADPH, and the expression levels of miR-202-5p were normalized to U6. The primers used in this study were listed as follows: GSEC forward: 5′-TTCCAATTAACCTGGCCGGAG-3′, reverse: 5′-GTCAGCCAACCCATTGCAAC-3′; AXL forward: 5′-TGCCAGAGGACCATTCTCCC-3′, reverse: 5′-GGTGCATTTGTACGGCCCTT-3′; GAPDH forward: 5′-GGAAGGACTCATGACCACAGTCC-3′, reverse: 5′-TCGCTGTTGAAGTCAGAGGAGACC-3′; miR-202-5p forward: 5′-TTCTCCCAAGGAAAGCACTTTCTG-3′, reverse: 5′-TGGTGTTGTTGAGTTG-3′; U6 forward: 5′-CGCTTCACGAATTTGCGTGTCAT-3′, reverse: 5′-GCTTCGGCAGCACATATACTAAAAT-3′.
Western Blot
Total proteins were extracted from tissue samples and cultured cells using a RIPA lysis buffer kit (Santa Cruz Biotechnology, Inc., USA). Approximately equal amounts of protein samples were separated by SDS-PAGE and transferred onto PVDF membranes. The membranes were incubated with specific primary antibodies including anti-AXL (1:500, Abcam, USA), anti-N-cadherin (1:500, Abcam, USA), anti-E-cadherin (1:500, Abcam, USA), anti-Vimentin (1:500, Abcam, USA) and anti-GAPDH (1:500, Abcam, Cambridge, Britain) at 4°C overnight. On the following day, the membranes were incubated with appropriate HRP-linked secondary antibodies (1:10,000, Sigma) at room temperature for 2 h. The protein bands were visualized by enhanced chemiluminescence (ECL) kit (Takara), and the intensity of targets were analyzed using the Quantity One software.
CCK-8 Assay
Cell viability was evaluated using a Cell Counting Kit-8 (CCK8, Dojindo, Japan). In brief, approximately 5×105 transfected BT-549 and MDA-MB-231 cells were seeded into 96-well plates. To detect cell proliferation, 10 μL CCK-8 solution was added into each well every 24 h until 96 h and incubated for another 3 h. Finally, the absorbance at 450 nm was evaluated using a microplate reader.
5-Ethynyl-20-deoxyuridine (EdU) Staining Assay
EdU staining assay was performed using EdU Staining Proliferation Kit (Abcam) as previously described.18 Briefly, approximately 5 × 104 transfected BT-549 and MDA-MB-231 cells were seeded into 96-well plates, followed by EdU reaction mix (provided by the kit) for 30 min. Then, DAPI dye was added and incubated for 15 min to counterstain the cell nucleus. The cells were then observed under a fluorescence microscope (200× magnification) and the number of EdU positive cells was counted in five random selected fields.
Transwell Assay
Cell invasion and migration abilities were assessed using Transwell assay as previously described.19 In brief, 200 μL serum-free DMEM/F12 medium containing 2 × 104 transfected BT-549 and MDA-MB-231 cell were seeded into the upper Transwell chambers (Corning, NY, USA) containing 8 μm pores for invasion. Migration assay was performed without pores. The low chamber was filled with normal medium containing 10% FBS. After incubation at 37°C for 48 h, cells on the lower surface were stained with 0.1% crystal violet for 15 min. Invasive and migrated cells were photographed and counted using a light microscopy (Nikon, Tokyo, Japan) (magnification: × 200) in 10 random selected fields.
Luciferase Reporter Assay
Starbase was used to predict the putative binding site between GSEC and miR-202-5p, and between miR-202-5p and AXL. The wild-type (WT) or mutant (MUT) GSEC and AXL containing the putative miR-202-5p binding site were synthesized by GenePharma (Shanghai, China) and cloned into pGL3 Basic vector (Promega). These luciferase reporter vectors (10 μg) were co-transfected with miR-202-5p mimics or miR-NC into BT-549 and MDA-MB-231 cells. After transfection for 48 h, cells were lysed and relative luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega).
RNA Immunoprecipitation (RIP) Assay
RIP assay was performed using the Magna RIP™ RNA-Binding Protein Immunoprecipitation Kit (Millipore, Bedford, MA). In brief, total cell lysates of BT-549 and MDA-MB-231 cells were incubated with magnetic beads containing human anti-Ago2 antibody (Millipore) and the negative control anti-IgG antibody (Millipore) at 4°C overnight. Then the enriched RNA was purified by using Trizol reagent and the Input group is the RNA of total cell lysate. The relative enrichment of GSEC and miR-202-5p was detected using qRT-PCR.
Xenograft Model
BALB/c nude mice (female, five-weeks-old) were kept under standard conditions. All animal procedures were approved by the animal Ethics Committee of the First Affiliated Hospital of Zhengzhou University and performed following the guidelines of National Institutes of Health (USA). For construction of the xenograft tumor model, approximately 5×106 BT-549 cells transfected with sh-GSEC, sh-NC, miR-202-5p inhibitor, or co-transfected with sh-GSEC and miR-202-5p inhibitor were subcutaneously inoculated into the dorsal of the nude mice (Six mice in each group). Tumor volume was evaluated every 7 d for 5 weeks based on the formula of (length × width2)/2. At the 35th day, mice were sacrificed by cervical dislocation, and tumors were excised, weighted and photographed. Then, tumors were used for immunohistochemistry analysis.
Immunohistochemistry (IHC) Analysis
The tumors excised from mice of different groups were fixed in 4% paraformaldehyde, dehydrated, embedded in paraffin and cut into 4-μm sections using a microtome. The sections were incubated with the anti-ki-67 antibody (1:500, Abcam) to detect the proliferation index in vivo. The sections were observed using an ordinary fluorescence microscope (Leica, Germany) at 200× magnification.
Statistical Analysis
All experiments were repeated in triplicate and data were presented as means ± standard deviation (SD) using the SPSS v.19.0 software (IBM Corp., Armonk, NY). Difference between two groups was tested by Student’s t test. The difference among multiple groups was tested by one-way ANOVA. The overall survival of TNBC patients was evaluated using Kaplan-Meier survival curve. Pearson’s correlation analysis was used to evaluate the correlation between the expression levels of GSEC, miR-202-5p and AXL in TNBC tissues. P < 0.05 was considered as the significant threshold.
Results
GSEC Was Highly Expressed in TNBC
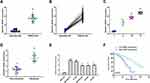
To explore the role of GSEC in TNBC, the expression levels of GSEC in TNBC tissues were measured by qRT-PCR and it showed that GSEC was significantly upregulated in TNBC tissues compared with that in adjacent normal tissues (n = 45, p < 0.01, Figure 1A). Meanwhile, the expression levels of GSEC were also obviously elevated in all TNBC tissues than that in matched adjacent normal tissues (n = 45, p < 0.01, Figure 1B). In addition, high expression levels of GSEC were positively associated with advanced tumor stage (p < 0.01, Figure 1C) and lymph-node metastasis (p < 0.01, Figure 1D) in TNBC patients. qRT-PCR results showed that GSEC was significantly upregulated in five TNBC cell lines including MDA-MB-231 (p < 0.001), BT-549 (p < 0.01), MDA-MB-468 (p < 0.01), HCCl937 (p < 0.01), BT-549 and SK-BR-3 (p < 0.01) compared with that in human breast epithelial cell line MCF-10A (Figure 1E). Kaplan-Meier curve analysis showed that high level of GSEC predicted a poor prognosis of TNBC patients (p < 0.01, Figure 1F). These results indicated that GSEC might play an oncogenic role in TNBC progression.
Downregulation of GSEC Inhibited the Proliferation, Invasion and Migration of TNBC Cells in vitro
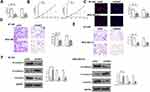
To determine the oncogenic role of GSEC in TNBC, BT-549 and MDA-MB-231 cells were transfected with sh-GSEC or sh-NC, and the transfection efficiency was confirmed by qRT-PCR (p < 0.01, Figure 2A). The results of CCK-8 assay and EdU staining assay both showed that sh-GSEC significantly inhibited cell proliferation compared with sh-NC in two cell lines (p < 0.01, Figure 2B and C). Transwell assay showed that sh-GSEC effectively inhibited cell invasion and migration ability compared with sh-NC in two cell lines (p < 0.01, Figure 2D and E). The effects of GSEC on epithelial–mesenchymal transition (EMT) related genes were also explored, and it showed that knockdown of GSEC markedly upregulated the expression of E-cadherin, while downregulated the expression of N-cadherin and Vimentin in both BT-549 and MDA-MB-231 cells (p < 0.01, Figure 2F). These results demonstrated that knockdown of GSEC could effectively inhibit cell proliferation, invasion and migration while promoted EMT process in TNBC.
GSEC Served as a Sponge of miR-202-5p
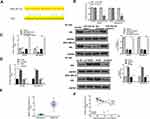
To explore the potential mechanism of GSEC in TNBC, BT-549 and MDA-MB-231 cells were transfected with miR-202-5p mimics/inhibitor and negative controls (miR-NC and inhibitor NC), and the transfection efficiency was confirmed by qRT-PCR (p < 0.01, Figure 3A). Then, Starbase v2.0 was used to predict the potential targets of GSEC and the results showed that GSEC might serve as a sponge of miR-202-5p (Figure 3B). To verify the interaction, luciferase reporter assay was performed, and the results showed that miR-202-5p mimics significantly reduced the relative luciferase activity of Luc-GSEC-WT compared with miR-NC in two cell lines (p < 0.01), and it exhibited no obvious change on luciferase activity of Luc-GSEC-MUT (Figure 3C). Meanwhile, RIP assay revealed that both GSEC and miR-202-5p were significantly enriched in miRNPs containing anti-Ago2 compared with that in anti-IgG group in two cell lines (p < 0.01, Figure 3D). In both BT-549 and MDA-MB-231 cells, sh-GSEC significantly increased the expression levels of miR-202-5p compared with sh-NC (p < 0.01), and overexpression of GSEC obviously reduced the expression levels of miR-202-5p compared with pc-NC (p < 0.01, Figure 3E). Next, we found that the expression of miR-202-5p was significantly downregulated in TNBC tissues compared with that in adjacent normal tissues (n = 45, p < 0.01, Figure 3F). Meanwhile, low expression levels of miR-202-5p was related to advanced tumor stage (p < 0.01, Figure 3G) and positive lymph-node metastasis (p < 0.01, Figure 3H) in TNBC patients. In addition, Pearson’s correlation analysis showed that the expression levels of GSEC were negatively correlated to the expression levels of miR-202-5p in TNBC tissues (n = 45, p < 0.01, r = −0.681, Figure 3I). These results suggested that miR-202-5p was a target of GSEC in TNBC.
Downregulation of miR-202-5p Effectively Reversed Knockdown of GSEC Induced Inhibition on TNBC Progression in vitro
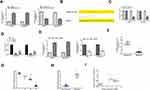
Next, whether GSEC regulated TNBC development through miR-202-5p was explored. BT-549 and MDA-MB-231 cells were transfected with sh-NC, sh-GSEC, miR-202-5p inhibitor, or co-transfected with sh-GSEC and miR-202-5p inhibitor. The results of CCK-8 assay and EdU staining assay showed that compared with sh-NC group, sh-GSEC significantly inhibited cell proliferation (p < 0.05), and miR-202-5p inhibitor markedly enhanced cell proliferation (p < 0.05), while co-transfection of sh-GSEC with miR-202-5p inhibitor obviously eliminated the inhibitory effect of sh-GSEC on cell proliferation (p < 0.05, Figure 4A and B). Transwell assay showed that sh-GSEC significantly inhibited cell invasion and migration ability compared with sh-NC group (p < 0.05), and miR-202-5p inhibitor significantly enhanced cell invasion and migration ability compared with sh-NC group (p < 0.05), while co-transfection of sh-GSEC with miR-202-5p inhibitor obviously eliminated the inhibitory effect of sh-GSEC on cell invasion and migration (p < 0.05, Figure 4C and D). In addition, compared with sh-NC group, sh-GSEC significantly upregulated E-cadherin, while downregulated N-cadherin and Vimentin; and miR-202-5p inhibitor downregulated E-cadherin, while upregulated N-cadherin and Vimentin; meanwhile, the effects of sh-GSEC on the expression of these EMT-related proteins were obviously reversed by co-transfection of sh-GSEC with miR-202-5p inhibitor (p < 0.01, Figure 4E). These results demonstrated that inhibition of miR-202-5p could effectively attenuate knockdown of GSEC induced inhibition of TNBC progression in vitro.
AXL Was a Target of miR-202-5p
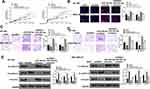
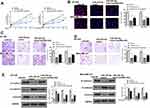
Then, Starbase v2.0 was also used to predict the potential targets of miR-202-5p and it showed that there was a putative binding site between miR-202-5p and AXL (Figure 5A). Luciferase reporter assay results showed that miR-202-5p mimics significantly reduced the relative luciferase activity of Luc-AXL-WT compared with miR-NC in two cell lines (p < 0.01), and it exhibited no obvious change in luciferase activity of Luc-AXL-MUT (Figure 5B). Meanwhile, miR-202-5p mimics significantly reduced the expression levels of AXL compared with miR-NC in two cell lines (p < 0.01), and downregulation of miR-202-5p significantly elevated the expression levels of AXL compared with inhibitor NC in two cell lines (p < 0.01, Figure 5C). In addition, overexpression of GSEC significantly increased the expression levels of AXL compared with the negative control (pc-NC) in two cell lines (p < 0.01), and sh-GSEC significantly decreased the expression levels of AXL compared with sh-NC in two cells (p < 0.01, Figure 5D). Moreover, the expression levels of AXL in TNBC tissues were markedly increased compared with that in adjacent normal tissues (n = 45, p < 0.01, Figure 5E). Pearson’s correlation analysis showed that the expression levels of miR-202-5p was negatively correlated to the expression levels AXL in TNBC tissues (n = 45, p < 0.01, r = −0.741, Figure 5F). These results indicated that AXL was a target of miR-202-5p in TNBC.
Overexpression of AXL Effectively Reversed miR-202-5p Mimics-Induced Inhibition on TNBC Progression in vitro
To determine whether miR-202-5p was mediated by AXL in TNBC, BT-549 and MDA-MB-231 cells were transfected with miR-NC, miR-202-5p mimics, or co-transfected with miR-202-5p mimics and pc-AXL. CCK-8 assay and EdU staining assay showed that miR-202-5p mimics significantly inhibited the proliferation of two cell lines compared with miR-NC (p < 0.01), while co-transfection of miR-202-5p mimics and pc-AXL effectively attenuated the inhibition of miR-202-5p mimics on cell proliferation (p < 0.05, Figure 6A and B). Transwell assay revealed that miR-202-5p mimics significantly inhibited the invasion and migration ability of two cell lines compared with miR-NC (p < 0.01), while co-transfection of miR-202-5p mimics and pc-AXL effectively reversed the inhibitory effect of miR-202-5p mimics on cell invasion and migration (p < 0.05, Figure 6C and D). Meanwhile, similar to knockdown of GSEC, overexpression of miR-202-5p significantly upregulated E-cadherin, while downregulated N-cadherin and Vimentin compared with miR-NC in two cell lines (p < 0.01), while co-transfection of miR-202-5p mimics and pc-AXL effectively reversed the effects of miR-202-5p mimics on the expression of EMT-related proteins (p < 0.05, Figure 6E). These data demonstrated that overexpression of AXL could effectively attenuate miR-202-5p mimics-induced inhibition on TNBC progression in vitro.
Knockdown of GSEC Inhibited Tumor Growth by Regulating the miR-202-5p/AXL Axis in vivo
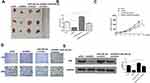
To further confirm the oncogenic role of GSEC in TNBC, a xenograft tumor model was used. The representative images of tumors showed that the sizes of the xenograft tumors in sh-GSEC group were obviously smaller than that in sh-NC group, the tumor sizes in miR-202-5p inhibitor group were bigger than that in sh-NC group, while the tumor sizes in sh-GSEC + miR-202-5p inhibitor group were between sh-GSEC and miR-202-5p inhibitor group (Figure 7A). Meanwhile, compared with sh-NC group, sh-GSEC significantly reduced tumor volume and weight (p < 0.01), and miR-202-5p inhibitor increased tumor volume and weight (p < 0.05), while co-transfection of sh-GSEC with miR-202-5p inhibitor obviously reversed the inhibitory effect of sh-GSEC on tumor growth (p < 0.05, Figure 7B and C). Moreover, Ki-67 IHC staining in tumor tissues showed that sh-GSEC reduced cell proliferation, miR-202-5p inhibitor exacerbated cell proliferation, while co-transfection of sh-GSEC with miR-202-5p inhibitor effectively reversed the inhibitory effect of sh-GSEC on cell proliferation in vivo (Figure 7D). Western blot results showed that the expression levels of AXL were significantly decreased in sh-GSEC group (p < 0.01), increased in miR-202-5p inhibitor group (p < 0.05), while co-transfection of sh-GSEC with miR-202-5p inhibitor significantly attenuated the inhibitory effect of sh-GSEC on the expression of AXL (p < 0.05, Figure 7E). These results indicated that GSEC played its oncogenic role in TNBC through regulating the miR-202-5p/AXL axis.
Discussion
As an aggressive subtype of breast cancer, TNBC results in a significant morbidity and mortality worldwide.20 Recent studies have identified several lnRNAs, including HOTAIR, SPRY4-IT1, GAS5 and PANDAR to play essential roles in tumor progression in breast cancer, and might be considered as potential molecular therapeutic targets.21 However, the roles and underlying molecular mechanisms of most lncRNAs in TNBC have not been well elucidated. In the present study, we explored the role and specific mechanism of GSEC in TNBC and demonstrated that GSEC was highly upregulated in TNBC. GSEC promoted TNBC development partially through miR-202-5p mediated upregulation of AXL. Our results suggested that GSEC might be a novel biomarker for the treatment of TNBC.
GSEC is a newly identified lncRNA and its function has only been investigated in a few human cancers. For example, the expression of GSEC is significantly upregulated in osteosarcoma cell lines, and overexpression of GSEC enhances the proliferating and migratory ability, while reduces the apoptotic rate of osteosarcoma cells in vitro.16 GSEC is also upregulated in colorectal cancer, and knockdown of GSEC reduces the motility of colon cancer cells by targeting DEAH box polypeptide 36 (DHX36) RNA helicase.17 In addition, differential expression analysis between lung squamous cell carcinoma and lung adenocarcinoma showed that GSEC had more co-repressed relationships with differentially expressed genes, indicating that GSEC might participate in the development of lung cancer.22 The role of GSEC in most human cancers including TNBC remains unclear. Here, we found that GSEC was upregulated in TNBC and high expression levels of GSEC were associated with advanced tumor stage, positive lymph-node metastasis and the poor survival time of TNBC patients. Moreover, knockdown of GSEC effectively inhibited cell proliferation, invasion and migration of TNBC cell lines. In addition, knockdown of GSEC downregulated N-cadherin and Vimentin, and upregulated E-cadherin involved in EMT process in TNBC cell lines. These results demonstrated that GSEC might exert as an oncogenic role in TNBC development.
Increasing studies have reported that lncRNAs exert their roles by sponging endogenous miRNAs in cancer including TNBC.23 For example, HCP5 promotes TNBC progression as a competing endogenous RNA (ceRNA) to upregulate BIRC3 by directly sponging miR-219a-5p.24 Overexpression of HEIH enhances cell proliferation and inhibits apoptosis of TNBC cells through targeting the miR-4458/SOCS1 axis.25 To explore the mechanism of GSEC in TNBC, we predicted the potential targets of GSEC using Starbase v2.0 and miR-202-5p was identified as a putative target of GSEC. MiR-202-5p is often downregulated and has been identified to act as a tumor suppressor in different types of human cancer such as osteosarcoma,26 colorectal carcinoma,27 thyroid carcinoma28 and ovarian cancer.29 Recent studies revealed that low expression levels of miR-202-5p reduces doxorubicin resistance and proliferation, while induces the apoptosis of TNBC cells in vitro.30 In this study, luciferase reporter assay and RIP assay confirmed the interaction between GSEC and miR-202-5p. Meanwhile, we found that miR-202-5p was downregulated in TNBC tissues and cell lines. Overexpression of miR-202-5p exhibited a similar inhibitory effect as knockdown of GSEC on TNBC progression in vitro, while downregulation of miR-202-5p obviously enhanced the aggressive phenotypes including proliferation, invasion, migration and EMT process. Moreover, downregulation of miR-202-5p attenuated the protective effects of GSEC knockdown on TNBC development both in vitro and in vivo. Our study revealed a tumor suppressive role of miR-202-5p and mediated the function of miR-202-5p in TNBC development. In one previous study, miR-388 was identified as a target of GSEC and played a crucial role in osteosarcoma.16 However, whether miR-388 mediated the role of GSCE in TNBC needs to be explored in the future.
Previous studies have revealed that miRNAs play essential roles through directly binding to the 3ʹ-untranslated regions (3ʹ-UTR) of target mRNAs.31 In TNBC, more miRNAs play oncogenic or tumor suppressive roles through directly targeting and downregulating the transcription of target mRNAs. For example, miR-211-5p inhibits tumor cell proliferation, invasion, migration and metastasis in TNBC by directly targeting SETBP1.32 MiR-25-3p promotes cell proliferation of TNBC cells through directly targeting BTG2.33 In this study, we identified AXL receptor tyrosine kinase as the target of miR-202-5p. AXL has been studied in different types of human cancer including lung cancer,34 pancreatic cancer,35 colorectal cancer,36 and ovarian cancer.37 In TNBC, AXL was found to be highly expressed in TNBC, and it could contribute to the migration of TNBC cells.38 One study performed a microarray and pattern miner analysis and found that AXL was upregulated in the TNBC cell line MDA-MB-231.39 Moreover, AXL inhibitor and small molecules targeting AXL have been identified to be a promising therapeutic agent for human cancer including TNBC.40,41 Here, we observed that AXL was significantly upregulated in TNBC tissues, which is consistent with previous reports.42,43 Moreover, overexpression of AXL effectively reversed the inhibitory effect of miR-202-5p mimics on TNBC cell proliferation, invasion and migration. These results demonstrated that GSEC/miR-202-5p participate in TNBC development by modulating AXL.
Conclusion
Our study revealed that GSEC was highly upregulated in TNBC and regulated the expression of AXL by directly sponging miR-202-5p. Meanwhile, we revealed the specific mechanism of GSEC/miR-202-5p/AXL in TNBC progression, suggesting that this axis may be a potential therapeutic target for TNBC.
Abbreviations
GSEC, G-quadruplex-forming sequence containing lncRNA; qRT-PCR, quantitative Real-time PCR; RIP, RNA immunoprecipitation; EdU, Ethynyl-20-deoxyuridine.
Funding
This work was supported by the Key Scientific Research Project of Colleges and Universities in Henan Province (Grant No. 19A320074).
Disclosure
The authors confirmed there are no conflicts of interest in this work.
References
1. Kim C, Gao R, Sei E, et al. Chemoresistance evolution in triple-negative breast cancer delineated by single-cell sequencing. Cell. 2018;173:879–893.e813. doi:10.1016/j.cell.2018.03.041
2. Foulkes WD, Smith IE, Reis-Filho JS. Triple-negative breast cancer. N Engl J Med. 2010;363(1938–1948):1938–1948. doi:10.1056/NEJMra1001389
3. Al-Mahmood S, Sapiezynski J, Garbuzenko OB, Minko T. Metastatic and triple-negative breast cancer: challenges and treatment options. Drug Deliv Transl Res. 2018;8:1483–1507. doi:10.1007/s13346-018-0551-3
4. Bergin ART, Loi S. Triple-negative breast cancer: recent treatment advances. F1000Research2019;8. doi:10.12688/f1000research.18888.1
5. Bianchini G, Balko JM, Mayer IA, Sanders ME, Gianni L. Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat Rev Clin Oncol. 2016;13:674–690. doi:10.1038/nrclinonc.2016.66
6. Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 2009;23:1494–1504. doi:10.1101/gad.1800909
7. Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15(7–21). doi:10.1038/nrg3606
8. Zhang YX, Yuan J, Gao ZM, Zhang ZG. LncRNA TUC338 promotes invasion of lung cancer by activating MAPK pathway. Eur Rev Med Pharmacol Sci. 2018;22:443–449. doi:10.26355/eurrev_201801_14193
9. Yang J, Li C, Mudd A, Gu X. LncRNA PVT1 predicts prognosis and regulates tumor growth in prostate cancer. Biosci Biotechnol Biochem. 2017;81:2301–2306. doi:10.1080/09168451.2017.1387048
10. Xiu B, Chi Y, Liu L, et al. LINC02273 drives breast cancer metastasis by epigenetically increasing AGR2 transcription. MolCancer. 2019;18(1):187. doi:10.1186/s12943-019-1115-y
11. Zheng S, Li M, Miao K, Xu H. lncRNA GAS5-promoted apoptosis in triple-negative breast cancer by targeting miR-378a-5p/SUFU signaling. J Cell Biochem. 2020;121(3):2225–2235. doi:10.1002/jcb.29445
12. Tang J, Zhong G, Zhang H, et al. LncRNA DANCR upregulates PI3K/AKT signaling through activating serine phosphorylation of RXRA. Cell Death Dis. 2018;9(1167). doi:10.1038/s41419-018-1220-7
13. Wang S, Ke H, Zhang H, et al. LncRNA MIR100HG promotes cell proliferation in triple-negative breast cancer through triplex formation with p27 loci. Cell Death Dis. 2018;9:805. doi:10.1038/s41419-018-0869-2
14. Niu L, Fan Q, Yan M, Wang L. LncRNA NRON down-regulates lncRNA snaR and inhibits cancer cell proliferation in TNBC. Biosci Rep. 2019;39. doi:10.1042/bsr20190468
15. Liu AN, Qu H-J, Gong W-J, et al. LncRNA AWPPH and miRNA-21 regulates cancer cell proliferation and chemosensitivity in triple-negative breast cancer by interacting with each other. J Cell Biochem. 2019;120(9):14860–14866. doi:10.1002/jcb.28747
16. Liu R, Ju C, Zhang F, et al. LncRNA GSEC promotes the proliferation, migration and invasion by sponging miR-588/EIF5A2 axis in osteosarcoma. Biochem Biophys Res Commun. 2020;532(300–307):300–307. doi:10.1016/j.bbrc.2020.08.056
17. Matsumura K, Kawasaki Y, Miyamoto M, et al. The novel G-quadruplex-containing long non-coding RNA GSEC antagonizes DHX36 and modulates colon cancer cell migration. Oncogene. 2017;36(1191–1199):1191–1199. doi:10.1038/onc.2016.282
18. Dong F, Ruan S, Wang J, et al. M2 macrophage-induced lncRNA PCAT6 facilitates tumorigenesis and angiogenesis of triple-negative breast cancer through modulation of VEGFR2. Cell Death Dis. 2020;11(728). doi:10.1038/s41419-020-02926-8
19. Marshall J. Transwell(®) invasion assays. Methods Mol Biol. 2011;769(97–110). doi:10.1007/978-1-61779-207-6_8
20. O’Sullivan CC, Davarpanah NN, Abraham J, Bates SE. Current challenges in the management of breast cancer brain metastases. Semin Oncol. 2017;44(2):85–100. doi:10.1053/j.seminoncol.2017.06.006
21. Nagini S. Breast cancer: current molecular therapeutic targets and new players. Anticancer Agents Med Chem. 2017;17(152–163). doi:10.2174/1871520616666160502122724
22. Tian Y, Yu M, Sun L, et al. Distinct patterns of mRNA and lncRNA expression differences between lung squamous cell carcinoma and adenocarcinoma. J Computational Biol. 2020;27(1067–1078):1067–1078. doi:10.1089/cmb.2019.0164
23. Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17(272–283):272–283. doi:10.1038/nrg.2016.20
24. Wang L, Luan T, Zhou S, et al. LncRNA HCP5 promotes triple negative breast cancer progression as a ceRNA to regulate BIRC3 by sponging miR-219a-5p. Cancer Med. 2019;8(4389–4403):4389–4403. doi:10.1002/cam4.2335
25. Li P, Zhou B, Lv Y, Qian Q. LncRNA HEIH regulates cell proliferation and apoptosis through miR-4458/SOCS1 axis in triple-negative breast cancer. Hum Cell. 2019;32(522–528):522–528. doi:10.1007/s13577-019-00273-1
26. Li C, Ma D, Yang J, Lin X, Chen B. miR-202-5p inhibits the migration and invasion of osteosarcoma cells by targeting ROCK1. Oncol Lett. 2018;16(829–834). doi:10.3892/ol.2018.8694
27. Ke SB, Qiu H, Chen JM, Shi W, Chen YS. MicroRNA-202-5p functions as a tumor suppressor in colorectal carcinoma by directly targeting SMARCC1. Gene. 2018;676(329–335):329–335. doi:10.1016/j.gene.2018.08.064
28. He H, Yang H, Liu D, Pei R. LncRNA NORAD promotes thyroid carcinoma progression through targeting miR-202-5p. Am J Transl Res. 2019;11:290–299.
29. Yu HY, Pan SS. MiR-202-5p suppressed cell proliferation, migration and invasion in ovarian cancer via regulating HOXB2. Eur Rev Med Pharmacol Sci. 2020;24(2256–2263). doi:10.26355/eurrev_202003_20491
30. Liu T, Guo J, Zhang X. MiR-202-5p/ PTEN mediates doxorubicin-resistance of breast cancer cells via PI3K/Akt signaling pathway. Cancer Biol Ther. 2019;20(989–998):989–998. doi:10.1080/15384047.2019.1591674
31. Qi X, Zhang D-H, Wu N, et al. ceRNA in cancer: possible functions and clinical implications. J Med Genet. 2015;52(710–718):710–718. doi:10.1136/jmedgenet-2015-103334
32. Chen LL, Zhang ZJ, Yi ZB, Li JJ. MicroRNA-211-5p suppresses tumour cell proliferation, invasion, migration and metastasis in triple-negative breast cancer by directly targeting SETBP1. Br J Cancer. 2017;117:78–88. doi:10.1038/bjc.2017.150
33. Chen H, Pan H, Qian Y, Zhou W, Liu X. MiR-25-3p promotes the proliferation of triple negative breast cancer by targeting BTG2. Mol Cancer. 2018;17:4. doi:10.1186/s12943-017-0754-0
34. Zhang Z, Lee JC, Lin L, et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat Genet. 2012;44:852–860. doi:10.1038/ng.2330
35. Ludwig KF, Du W, Sorrelle NB, et al. Small-molecule inhibition of axl targets tumor immune suppression and enhances chemotherapy in pancreatic cancer. Cancer Res. 2018;78(246–255):246–255. doi:10.1158/0008-5472.can-17-1973
36. Uribe DJ, et al. The receptor tyrosine kinase AXL promotes migration and invasion in colorectal cancer. 2017;12:e0179979. doi:10.1371/journal.pone.0179979
37. Quinn JM, Greenwade MM, Palisoul ML, et al. Therapeutic inhibition of the receptor tyrosine kinase AXL improves sensitivity to platinum and taxane in ovarian cancer. Mol Cancer Ther. 2019;18(389–398):389–398. doi:10.1158/1535-7163.mct-18-0537
38. Zajac O, Leclere R, Nicolas A, et al. AXL controls directed migration of mesenchymal triple-negative breast cancer cells. Cells. 2020;9(1). doi:10.3390/cells9010247
39. Natarajan S, Sumantran VN, Ranganathan M, Madheswaran S. Microarray and pattern miner analysis of AXL and VIM gene networks in MDA‑MB‑231 cells. Mol Med Rep. 2018;18(4147–4155):4147–4155. doi:10.3892/mmr.2018.9404
40. Shen Y, Chen X, He J, Liao D, Zu X. Axl inhibitors as novel cancer therapeutic agents. Life Sci. 2018;198(99–111):99–111. doi:10.1016/j.lfs.2018.02.033
41. Leconet W, Chentouf M, Du Manoir S, et al. Therapeutic activity of Anti-AXL antibody against triple-negative breast cancer patient-derived xenografts and metastasis. Clin Cancer Res. 2017;23(2806–2816):2806–2816. doi:10.1158/1078-0432.ccr-16-1316
42. Wei J, Sun H, Zhang A, et al. A novel AXL chimeric antigen receptor endows T cells with anti-tumor effects against triple negative breast cancers. Cell Immunol. 2018;331(49–58):49–58. doi:10.1016/j.cellimm.2018.05.004
43. Yue CH, Liu L-C, Kao E-S, et al. Protein kinase C α is involved in the regulation of AXL receptor tyrosine kinase expression in triple-negative breast cancer cells. Mol Med Rep. 2016;14(1636–1642). doi:10.3892/mmr.2016.5424
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.