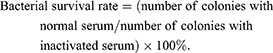Back to Journals » Drug Design, Development and Therapy » Volume 15
In vitro Antimicrobial Activity of Fosfomycin, Rifampin, Vancomycin, Daptomycin Alone and in Combination Against Vancomycin-Resistant Enterococci
Authors Tong J, Jiang Y, Xu H, Jin X, Zhang L, Ying S, Yu W, Qiu Y
Received 22 April 2021
Accepted for publication 27 June 2021
Published 12 July 2021 Volume 2021:15 Pages 3049—3055
DOI https://doi.org/10.2147/DDDT.S315061
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Jianbo Sun
Jiepeng Tong,1 Yiheng Jiang,2 Hao Xu,1 Xuehang Jin,1 Li Zhang,1 Shuaibing Ying,1 Wei Yu,1 Yunqing Qiu1
1State Key Laboratory for Diagnosis and Treatment of Infectious Diseases, National Clinical Research Center for Infectious Diseases, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, Zhejiang Provincial Key Laboratory for Drug Clinical Research and Evaluation, The First Affiliated Hospital, Zhejiang University School of Medicine, Hangzhou, People’s Republic of China; 2Clinical Medicine, Nanjing Medical University, Nanjing, People’s Republic of China
Correspondence: Wei Yu; Yunqing Qiu Email [email protected]; [email protected]
Purpose: The emergence of vancomycin resistant Enterococci (VRE) is shortening the choices for clinical anti-infective therapy. The aim of this study was to investigate the mechanism of vancomycin resistance and evaluate the effect of fosfomycin (FM), rifampin (RIF), vancomycin (VAN), linezolid (LNZ), daptomycin (DAP) alone or in combination against VRE.
Methods: Eight VRE isolates were collected. A total of 18 antibiotics susceptibility tests were further done for VRE. Whole genome sequencing and bioinformatics analysis were performed. The effect of FM, RIF, VNA, LNZ, DAP alone or in combination was determined using anti-biofilm testing and the time-kill assay.
Results: All isolates were susceptible to LNZ and DPA. The high-level resistance determinant of VAN in these strains was due to VanA-type cassette. MLST revealed two different STs for vancomycin-resistant Enterococcus faecium (VREm) and four different STs for vancomycin-resistant E. faecalis (VREs). Virulence genes in VREs were more than VREm, especially for 4942 isolated from blood. Gene acm and uppS were only identified in VREm, while virulence genes related to cytolysin were only found in E. faecalis. Further in vitro studies indicated FM (83 mg/L) combined with DAP (20.6 mg/L) and DAP monotherapy (47.1 mg/L) had bactericidal effect against VRE isolates at 24h.
Conclusion: High-level resistance determinant of VAN in tested isolates was due to VanA-type cassette. FM combined with DAP is a potential therapeutic option for VRE infections.
Keywords: vancomycin, daptomycin, combination therapy, biofilm
Introduction
Vancomycin resistant Enterococci (VRE) are increasingly becoming public health threat for hospitals worldwide. VRE infections caused significant mortality, ranging from 19% to 63%.1,2 In previous years, most VRE infections were caused by E. faecalis.3 However, since 2002 an increase in the prevalence of vancomycin-resistant Enterococcus faecium (VREm) has been observed, with reports of VREm being as common as vancomycin-resistant E. faecalis (VREs).4 This could be due to intrinsic and acquired resistance to many classes of antibiotics of E. faecium, making it better adapted to the hospital and environment where antibiotic use is common.5
The opportunistic invasive VRE infections are often broadly resistant to available antibiotics. Combination therapies, such as gentamicin and β-lactams, were reported to achieve reliable bactericidal effect for enterococcal endocarditis.6 However, isolates with high-level resistance of gentamicin do not show synergism during combination. Current effective treatments for VRE included quinupristin/dalfopristin, teicoplanin, telavancin, linezolid (LNZ), and daptomycin (DAP).7 There are relatively few studies in this important area of combination treatment with synergistic antibiotics. Thus, to elucidate the in vitro effectiveness, we compared the antibacterial effects tigecycline of fosfomycin (FM), rifampin (RIF), vancomycin (VAN), LNZ, DAP alone and in combination against VRE.
Methods
Bacterial Strains and Antibiotic Susceptibility Test
A total of 8 VRE strains were isolated from clinical specimens for diagnosis and frozen at −80◦C in our laboratory. The pure cultures were put on identification plate and add 2 μL mixed liquid including trifluoroacetic acid, acetonitrile and distilled water. After drying, all isolates were definite identification using Matrix-assisted laser desorption ionisation-time of flight mass spectrometry (MALDI-TOF) (Bruker Diagnostics, Bremen, Germany).
The minimum inhibitory concentrations (MICs) for 18 antibiotics, including oxacillin, penicillin, meropenem, erythromycin, clindamycin, trimethoprim-sulfamethoxazole, amikacin, gentamicin, ciprofloxacin, levofloxacin, moxifloxacin, tetracycline, tigecycline, RIF, VAN, LNZ, FM, and DAP were determined by broth microdilution.8 The susceptibility to FM was tested by agar dilution. The media was supplemented with 50 mg/l Ca2+ for testing of DAP and 25 mg/l glucose-6-phosphate (G6P) for FM. E. faecalis ATCC 29,212 was used as quality control.
Whole-Genome Sequencing (WGS)
WGS was carried out for 8 VRE isolates with further analyses of gene-environment. Genomic DNA was extracted by FastDNA SPIN Kit for Soil (MP Biomedicals, United States) and sequenced using HiSeq 2000 (Illumina, SanDiego, CA, USA) with constructing 2x125-bp pair-end libraries. De novo assembly was done using the CLC Workbench v8.0 (QIAGEN, Hilden, Germany). The resistance genes, and virulence genes were identified by BLAST against the ResFinder 2.1 database (https://cge.cbs.dtu.dk/services/ResFinder/). The bioinformatics tools used in this study were available at the following web platforms: NCBI (National Center for Biotechnological Information), SMS (Sequence Manipulation Suite), and EBI (European Bioinformatics Institute).
This Whole Genome Shotgun BioProject for VRE has been deposited at GenBank under the accession PRJNA662846 and PRJNA662849 (Supplemental Table 1).
Anti-Biofilm Testing
All VRE isolates and ATCC 29,212 were inoculated into 96-well polystyrene microtiter plates with Mueller–Hinton II broth (MHB) and different RIF, LNZ, FM, and DAP concentration for 24h, 48h, and 72h to test the biofilms formation as a previous study.9 All experiments results were from three separate experiments.
Anti-Complement Killing Test
Mouse serum was purchased from Dalian Guangzhou Ruite Biotechnology (Guangzhou, China). This was placed in a water bath at 56°C for 30 min to inactivate complement, generating inactive serum. An overnight bacterial culture was diluted to a cell density of 2×106 CFU/mL, and normal and inactivated sera (180 μL) were separately mixed with 20 μL bacterial suspension and incubated at 37°C for 1 h. Samples were diluted 100-fold, spread onto plates, and incubated overnight, and colonies on plates were counted. The bacterial survival rate was calculated using the following formula:
ATCC 29,212 served as the control strain.
Time–Kill Assays
The bactericidal activities of FM, VAN, and DAP alone or in combination against four VRE (4942, 12,022, 19,372, 23,760) and ATCC 29,212 were investigated using the time–kill method.10 The antibiotic concentrations were calculated according to steady-state concentrations of drug in humans as described previously.11 The following concentrations were used: FM 83 mg/L;11 LNZ 10 mg/L;12 RIF 3 mg/L;13 VAN 13.3 mg/L;14 DAP 20.6 mg/L, 31.1 mg/L and 47.1 mg/L;15 FM 83 mg/L + DAP 20.6 mg/L; FM 83 mg/L + LNZ 10 mg/L; FM 83 mg/L + RIF 3 mg/L; FM 83 mg/L + VAN 13.3 mg/L; RIF 3 mg/L + DAP 20.6 mg/L; RIF 3 mg/L + LNZ 10 mg/L; RIF 3 mg/L + VAN 13.3 mg/L. The media was supplemented with 25 mg/l G6P for testing of FM and 50 mg/l Ca2+ for DAP. Each test had three replications.
Results
Antimicrobial Susceptibility and Multi-Locus Sequence Typing (MLST)
There were two VREs and six VREm isolates included in this study. Eight VRE isolates obtained from culture including urine (n = 3), bile (n = 1), and blood (n = 4) (Table 1). All isolates showed high-level resistance to VAN, whereas they were susceptible to tigecycline, LNZ, DPA (Table 1). Only two E. faecalis isolates (4942 and 12,022) were susceptible to RIF.
 |
Table 1 Minimum Inhibitory Concentrations of 18 Antimicrobial Agents Against 8 VRE Isolates |
MLST revealed two different STs for VREs isolates 4942 (ST4) and 12,022 (ST179). There were four STs (ST412, ST564, ST78, ST17) for VREm.
Resistance Genes and Virulence Genes
Isolate 23,760 has the least number of resistance genes. Three genes vanRA, vanSA, vanYA were found in all isolates. VanA, VanXA were not found in 12,022 and 23,760. In addition, vanHA was not identified in 12,022 and vanZA was not found in 23,760 (Supplemental Table 2). The vanA operon was carried on Tn1546 transposon in six isolates (4942, 5057, 5173, 5734, 9604, 12,022). Isolates 4942, 5734, and 9604 had genomic island including vanZ, vanY, vanX, vanA, vanH, vanS, vanR (Supplemental Figure 1), while 5057, 5173, and 12,022 had genetic rearrangements in Tn1546 transposon.
Virulence genes bopD and efaA were found in all six isolates. Notably, virulence genes in VREs were more than VREm, especially for 4942 isolated from blood (Supplemental Table 3). Gene acm and uppS were only identified in VREm, while virulence genes related to cytolysin were only found in VREs.
Anti-Complement Killing Test and Anti-Biofilm Formation Test
The survival rate of 5057 was less than 30%, while the rate of 5173 and 9604 were above 90%. (Supplemental Figure 2).
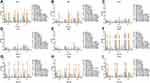
As shown in Figure 1 and Supplemental Table 4, the biofilm formations of isolate 5057, 5173, 5743, 9604 were less than other four VRE isolates. Except for isolate 19,372, FM monotherapy could not efficiently inhibit the formation of biofilms. LNZ and DAP monotherapy showed effective anti-biofilm formation during 72 hours. The biofilm inhibitory effect against planktonic VRE isolates for FM combined with LNZ or VAN was better than monotherapy. In addition, the effect of FM (83 mg/L) combined with DAP (20.6 mg/L) anti-biofilm formation was similar to DAP monotherapy (47.1 mg/L).
Time–Kill Assays
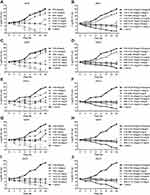
Four VRE isolates (4942, 12,022, 19,372, 23,760) with stronger ability of biofilm formations and ATCC 29,212 were treated with antibiotics at average steady-state serum concentrations. The results were shown in Figure 2. Among monotherapy time–kill studies, DAP showed bactericidal activity against four VRE isolates at 24 h. The bactericidal activity of DAP was concentration-dependent. It is noteworthy that FM (83 mg/L) combined with DAP (20.6 mg/L) and DAP monotherapy (47.1 mg/L) reduced the population of four VRE isolates to zero without re-growth at 24h.
Discussion
The increasing prevalence of VRE is posing a constraint on therapeutic options. Furthermore, the effects of antibiotics for VRE were reported limit in vitro and in vivo studies. In the present study, the resistance determinant of VAN in these strains was due to VanA-type cassette. Virulence genes in VREs were more than VREm. In addition, FM (83 mg/L) combined with DAP (20.6 mg/L) showed pronounced biofilm elimination effects and bactericidal activity.
Eight isolates showed high-level resistance to VAN, relating to VanA-type cassette. So far, one of the most relevant VAN resistance traits is the acquisition of van genes.16 vanA (80–90%) and vanB (10–20%) are the most predominant among 9 van genotypes.17 The vanA operon usually consists of five genes (vanHAXYZ) for glycopeptide resistance, two regulatory genes (vanRS), a transposase (orf1)/resolvase (orf2) region.17 Tn1546 transposon was contributed to the increase of VRE infections.18 Genetic variations, such as deletions and/or addition of some insertion sequences, have been reported in Tn1546.19 Consistent with previous studies, three VRE isolates (5057, 5173, and 12,022) were found polymorphisms in upstream of vanR among Tn1546 as well.20 Due to horizontal transfer of plasmids, it is necessary to reinforce managements to prevent spreading of VRE.
Several studies have investigated the importance of putative virulence genes in VRE, however, there are no clear conclusions as to what constituents decisively contribute to the pathogenicity of VRE yet.21 We found virulence genes in E. faecalis were more than E. faecium. The only virulence genes confirmed to be associated with VRE infection are the enterococcal surface protein gene (esp) and the hyaluronidase gene (hyl).22,23 The putative virulence factors include proteins that attack several different constituents of cells, such as cytolysin that targets cell membranes, as well as gelatinase and serine protease that attack various proteins such as collagen, fibrinogen, and insulin.24
Notably, a few effective therapies are available for VRE infections.25 Other options, such as pump inhibitors and essential oils, have been proved to have antibacterial activity against multidrug-resistant Enterococci in vitro.26,27 Although previous studies showed 0.3% to 20% VRE were resistance to DAP, all isolates in our study were sensitive to DAP.28,29 LNZ and DAP monotherapy showed effective anti-biofilm formation during 72 hours. It is of note that LNZ treatment for VRE bloodstream infection was associated with higher mortality and microbiologic failure in comparison to DAP.30 DAP showed bactericidal activity against four VRE isolates at 24 h in the time-kill assay. There is a concern for the toxicity of higher doses of DAP due to increase of creatine kinase levels and muscle toxicity.31 Fortunately, similar to DAP monotherapy (47.1 mg/L), FM (83 mg/L) combined with DAP (20.6 mg/L) reduced the population of four VRE isolates to zero without re-growth at 24h. Previous reports demonstrated FM combined with DAP had synergistic effects as well.15,32 Therefore, the combination of FM and DAP for patients with VRE infections, especially for patients with renal impairment, is of great significance for further clinical trials.
In conclusion, the high-level resistance determinant of sporadic VRE in the present study was due to VanA-type cassette. The most efficient regimen of bactericidal effect against VRE and biofilm inhabitation was the combination of FM and DAP. Further in vivo investigation and clinical trials are needed to define the effect of different drug combinations.
Data Sharing Statement
The BioProject for VRE has been deposited at GenBank under the accession PRJNA662846 and PRJNA662849.
Ethics Approval
In our study, we did not perform any experiments with animals or higher invertebrates, neither performed experiments on humans nor the use of human tissue samples. Our data have been originated from bacteria which were frozen for antimicrobial resistance monitoring, not linked to clinical information. Therefore, our research was exempt from ethics approval.
Consent for Publication
All authors have seen and approved the content and fulfil the journal’s requirements for authorship.
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Funding
This work was supported by Department of Health of Zhejiang province [2019RC003].
Disclosure
The authors report no conflicts of interest in this work.
References
1. Akhtar N, Sultan F, Nizamuddin S, Zafar W. Risk factors and clinical outcomes for vancomycin-resistant enterococcus bacteraemia in hospitalised cancer patients in Pakistan: a case-control study. J Pak Med Assoc. 2016;66(7):829–836.
2. Vergis EN, Hayden MK, Chow JW, et al. Determinants of vancomycin resistance and mortality rates in enterococcal bacteremia. a prospective multicenter study. Ann Intern Med. 2001;135(7):484–492. doi:10.7326/0003-4819-135-7-200110020-00007
3. Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10(4):266–278. doi:10.1038/nrmicro2761
4. Treitman AN, Yarnold PR, Warren J, Noskin GA. Emerging incidence of Enterococcus faecium among hospital isolates (1993 to 2002). J Clin Microbiol. 2005;43(1):462–463. doi:10.1128/JCM.43.1.462-463.2005
5. Cattoir V, Giard JC. Antibiotic resistance in Enterococcus faecium clinical isolates. Expert Rev Anti Infect Ther. 2014;12(2):239–248. doi:10.1586/14787210.2014.870886
6. Miller WR, Murray BE, Rice LB, Arias CA. Resistance in vancomycin-resistant enterococci. Infect Dis Clin North Am. 2020;34(4):751–771. doi:10.1016/j.idc.2020.08.004
7. Prematunge C, MacDougall C, Johnstone J, et al. VRE and VSE bacteremia outcomes in the era of effective VRE therapy: a systematic review and meta-analysis. Infect Control Hosp Epidemiol. 2016;37(1):26–35. doi:10.1017/ice.2015.228
8. Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. 24th informational supplement. 2014. Available from: http://www.clsi.org/.
9. Yu W, Zhang J, Tong J, et al. In vitro antimicrobial activity of fosfomycin, vancomycin and daptomycin alone, and in combination, against linezolid-resistant Enterococcus faecalis. Infect Dis Ther. 2020;9(4):927–934. doi:10.1007/s40121-020-00342-1
10. Yu W, Shen P, Bao Z, et al. In vitro antibacterial activity of fosfomycin combined with other antimicrobials against KPC-producing Klebsiella pneumoniae. Int J Antimicrob Agents. 2017;50(2):237–241. doi:10.1016/j.ijantimicag.2017.03.011
11. Tängdén T, Hickman RA, Forsberg P, Lagerbäck P, Giske CG, Cars O. Evaluation of double- and triple-antibiotic combinations for VIM- and NDM-producing Klebsiella pneumoniae by in vitro time–kill experiments. Antimicrob Agents Chemother. 2014;58:1757–1762. doi:10.1128/AAC.00741-13
12. Mokline A, Gharsallah L, Rahmani I, Gaies E, Tabelsi S, Messadi AA. Pharmacokinetics and pharmacodynamics of linezolid in burn patients. Ann Burns Fire Disasters. 2018;31(2):118–121.
13. Medellín-Garibay SE, Milán-Segovia Rdel C, Magaña-Aquino M, Portales-Pérez DP, Romano-Moreno S. Pharmacokinetics of rifampicin in Mexican patients with tuberculosis and healthy volunteers. J Pharm Pharmacol. 2014;66(10):1421–1428. doi:10.1111/jphp.12275
14. Shokouhi S, Alavi Darazam I. Determination of vancomycin trough level in serum and cerebrospinal fluid of patients with acute community-acquired meningitis: a prospective study. J Infect. 2014;39(5):424–429. doi:10.1016/j.jinf.2014.06.010
15. D’Avolio A, Pensi D, Baietto L, Pacini G, Di Perri G, De Rosa FG. Daptomycin pharmacokinetics and pharmacodynamics in septic and critically ill patients. Drugs. 2016;76(12):1161–1174. doi:10.1007/s40265-016-0610-3
16. Courvalin P. Vancomycin resistance in gram-positive cocci. Clin Infect Dis. 2006;42(Suppl 1):S25–S34. doi:10.1086/491711
17. Cattoir V, Leclercq R. Twenty-five years of shared life with vancomycin-resistant enterococci: is it time to divorce? J Antimicrob Chemother. 2013;68(4):731–742. doi:10.1093/jac/dks469
18. Kuo AJ, Shu JC, Liu TP, et al. Vancomycin-resistant Enterococcus faecium at a university hospital in Taiwan, 2002–2015: fluctuation of genetic populations and emergence of a new structure type of the Tn1546-like element. J Microbiol Immunol Infect. 2018;51(6):821–828. doi:10.1016/j.jmii.2018.08.008
19. Willems RJ, Top J, van den Braak N, et al. Molecular diversity and evolutionary relationships of Tn1546-like elements in enterococci from humans and animals. Antimicrob Agents Chemother. 1999;43(3):483–491. doi:10.1128/AAC.43.3.483
20. Novais C, Freitas AR, Sousa JC, Baquero F, Coque TM, Peixe LV. Diversity of Tn1546 and its role in the dissemination of vancomycin-resistant enterococci in Portugal. Antimicrob Agents Chemother. 2008;52(3):1001–1008. doi:10.1128/AAC.00999-07
21. Reinseth IS, Ovchinnikov KV, Tønnesen HH, Carlsen H, Diep DB. The increasing issue of vancomycin-resistant Enterococci and the Bacteriocin solution. Probiotics Antimicrob Proteins. 2020;12(3):1203–1217. doi:10.1007/s12602-019-09618-6
22. Leavis HL, Willems RJ, van Wamel WJ, Schuren FH, Caspers MP, Bonten MJ. Insertion sequence-driven diversification creates a globally dispersed emerging multiresistant subspecies of E. faecium. PLoS Pathog. 2007;3(1):e7. doi:10.1371/journal.ppat.0030007
23. Leendertse M, Heikens E, Wijnands LM, et al. Enterococcal surface protein transiently aggravates Enterococcus faecium-induced urinary tract infection in mice. J Infect Dis. 2009;200(7):1162–1165. doi:10.1086/605609
24. Ispirli H, Demirbaş F, Dertli E. Characterization of functional properties of Enterococcus faecium strains isolated from human gut. Can J Microbiol. 2015;61(11):861–870. doi:10.1139/cjm-2015-0446
25. Millette M, Cornut G, Dupont C, Shareck F, Archambault D, Lacroix M. Capacity of human nisin- and pediocin-producing lactic acid bacteria to reduce intestinal colonization by vancomycin-resistant enterococci. Appl Environ Microbiol. 2008;74(7):1997–2003. doi:10.1128/AEM.02150-07
26. Usai D, Donadu M, Bua A, et al. Enhancement of antimicrobial activity of pump inhibitors associating drugs. J Infect Dev Ctries. 2019;13(2):162–164. doi:10.3855/jidc.11102
27. Quendera AP, Barreto AS, Semedo-Lemsaddek T. Antimicrobial activity of essential oils against foodborne multidrug-resistant enterococci and aeromonads in planktonic and biofilm state. Food Sci Technol Int. 2019;25(2):101–108. doi:10.1177/1082013218799027
28. Kamboj M, Cohen N, Gilhuley K, Babady NE, Seo SK, Sepkowitz KA. Emergence of daptomycin-resistant VRE: experience of a single institution. Infect Control Hosp Epidemiol. 2011;32(4):391–394. doi:10.1086/659152
29. Sader HS, Jones RN. Antimicrobial susceptibility of gram-positive bacteria isolated from US medical centers: results of the Daptomycin Surveillance Program (2007–2008). Diagn Microbiol Infect Dis. 2009;65(2):158–162. doi:10.1016/j.diagmicrobio.2009.06.016
30. Britt NS, Potter EM, Patel N, Steed ME. Comparison of the effectiveness and safety of linezolid and daptomycin in vancomycin-resistant Enterococcal bloodstream infection: a National Cohort Study of veterans affairs patients. Clin Infect Dis. 2015;61(6):871–878. doi:10.1093/cid/civ444
31. Moise PA, Hershberger E, Amodio-Groton MI, Lamp KC. Safety and clinical outcomes when utilizing high-dose (> or =8 mg/kg) daptomycin therapy. Ann Pharmacother. 2009;43(7):1211–1219. doi:10.1345/aph.1M085
32. Zheng JX, Sun X, Lin ZW, et al. In vitro activities of daptomycin combined with fosfomycin or rifampin on planktonic and adherent linezolid-resistant isolates of Enterococcus faecalis. J Med Microbiol. 2019;68(3):493–502. doi:10.1099/jmm.0.000945
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.