Back to Journals » Journal of Inflammation Research » Volume 14
Identification of Prognostic Biomarkers and Molecular Targets Among JAK Family in Breast Cancer
Received 1 October 2020
Accepted for publication 8 December 2020
Published 14 January 2021 Volume 2021:14 Pages 97—114
DOI https://doi.org/10.2147/JIR.S284889
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Ning Quan
Fangteng Liu,1,2 Hengyu Wu1
1Department of Breast Surgery, The Third Hospital of Nanchang, Nanchang 330009, Jiangxi, People’s Republic of China; 2Faculty of Medicine, University of Munich, Munich 80336, Germany
Correspondence: Hengyu Wu Tel +86-15270280590
Email [email protected]
Background: Janus kinases (JAKs) are a family of non-receptor tyrosine kinases involved in multiple malignancies. However, clinical values of JAKs as prognostic markers and potential mechanism as molecular targets in breast invasive carcinoma (BC) are not completely clarified.
Methodology: TIMER, UALCAN and GEPIA were used to assess the expression and methylation levels of JAKs in BC. Kaplan–Meier Plotter, bc-GenExMiner, SurvExpress, TRGAted, MethSurv, and SurvivalMeth were used to assess the multilevel prognostic significance of JAKs in breast cancer patients. And cBioPortal, TIMER, STRING, GeneMANIA, NetworkAnalysis, LinkedOmics, DAVID 6.8, and Metascape were applied for multilayer networks and functional enrichment analyses. Correlations between immune cell infiltrates/their gene markers and JAKs were evaluated by TIMER.
Results: We first explored the expression and methylation level of JAKs in breast cancer and found significantly reduced JAK1 and JAK2 expression at mRNA and protein levels, significantly higher JAK3 protein expression, and significantly increased TYK2 expression at mRNA level but decreased at protein level. In addition, hypermethylation of JAK3 and TYK2 and hypomethylation of JAK1 were found in tumor samples. In terms of prognostic values of JAKs in BC patients, low transcriptional levels of JAK1, JAK2, JAK3, and TYK2 indicated worse OS/DMFS/PPS/RFS/DFS, inferior DFS, worse RFS, and shorter OS/DMFS/RFS, respectively. The mRNA signature analysis showed that high-risk group had unfavorable OS/RFS/MFS. Low JAK2 protein level indicated unfavorable DSS/PFS in BC patients. Five CpGs of JAK1, four CpGs of JAK2, 20 CpGs of JAK3, and 13 CpGs of TYK2 were significantly associated with prognosis in BC patients. The DNA methylation signature analysis also suggested worse prognosis in the high-risk group. For potential biological roles of JAKs, interaction analyses, functional enrichment analyses for biological process, cellular component, molecular function, and KEGG pathway analyses of JAKs and their neighbor genes in BC were conducted. Kinase targets, gene–miRNA interactions, and transcription factor–gene interactions of JAKs were also identified. Furthermore, JAKs were found to be significantly related to immune infiltrates as well as the expression levels of multiple immune markers in BC.
Conclusion: JAKs showed multilevel prognostic value and important biological roles in BC. They might serve as promising prognostic markers and possible targets in breast cancer.
Keywords: Janus kinase, breast cancer, prognosis, immunity, biomarker, target
Introduction
Breast cancer is the second most common cancer worldwide. It accounts for more than one-fifth of all female invasive tumors, and 16% of all cancers in women.1,2 Every year an estimated 1,700,000 new cases are diagnosed, this aggressive disease is the leading cause of cancer-related deaths under the age of 45.3 And according to the expression of estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER2), breast cancer is broadly divided into luminal ER positive (luminal A and luminal B), HER2 enriched, and basal-like molecular subtypes.4,5 Despite recent great progress in diagnostic methods and treatment technologies,6–8 long-term survival rate of these patients is still not satisfactory, especially in the metastatic setting (median survival less than 24 months). It would be beneficial to conduct more accurate risk assessment and optimize treatment for patients with breast cancer. Therefore, it is essential to develop and identify valuable prognostic markers and therapeutic targets for breast cancer.
JAK (Janus kinase) is a family of intracellular, non-receptor tyrosine kinases with the molecular weight between 120 kD and 140 kD. Structurally, they can be divided into two parts. At the C-terminal, there are two closely connected JAK active sample domains, and the closest to the C-terminal is the catalytic active reaction region of JAK kinase. At the N-terminal, there are five subdomains (A, B, C, D, and E), known as JH domain, which may be involved in molecular coupling of JAK kinase and cytokine receptor. JAKs can mediate the activation reaction of signal protein after binding of cytokines and receptors.9,10 The members of JAK family are JAK1, JAK2, JAK3, and TYK2. In the amino acid sequences of human JAK family proteins, JAK1, JAK2, and JAK3 have high homology with TYK2. JAKs are involved in autoimmune diseases and malignancies, so the targeting of specific JAK members can be used to treat related diseases.11,12 The activated JAK protein phosphorylates the STAT protein to facilitate the formation of STAT dimer, eventually activating the JAK/STAT pathway.13 Inhibition of the JAK1/STAT3 pathway can effectively inhibit ovarian tumor progression.14 JAK2 also promotes invasiveness and drug resistance in colorectal cancer cells through the JAK2/STAT3 signaling pathway.15 The JAK3 mutation is of great significance as a response marker for targeted therapy of JAK kinase or anti-PD1 immunotherapy.16 TYK2 is involved in tumor immune-surveillance in human cancers,17 and it is necessary for the survival of anaplastic large cell lymphoma cells by activating MCL1 expression.18
Although the JAK family members have been reported to be involved in the progression of various types of cancers, the clinical value of the entire JAK family remains poorly investigated in breast cancer. This study investigated the clinical value of JAK family members as prognostic markers and therapeutic targets in breast cancer, based on multiple public resources and reliable integrative bioinformatics analysis. The flow diagram of this study is shown in Supplementary Figure 1.
Materials and Methods
Expression and Methylation Levels of JAKs
The JAK expression and methylation levels were compared between tumor tissues of breast invasive carcinoma (BC) and normal tissues using the DiffExp module of TIMER (https://cistrome.shinyapps.io/timer/)19 and UALCAN (http://ualcan.path.uab.edu/) web resource.20 GEPIA (http://gepia.cancer-pku.cn/)21 was used for the multiple gene comparison and pathological stage plot of JAK family in BC.
Survival Outcomes and mRNA Levels of JAKs or Family Signature
Kaplan–Meier Plotter (https://kmplot.com/) 22 was used to obtain the associations between JAK mRNA levels and overall survival (OS)/distant metastasis-free survival (DMFS)/post-progression survival (PPS)/relapse-free survival (RFS) in breast cancer. The associations between JAK mRNA expression and prognosis in breast cancer cases experiencing different clinical conditions were also assessed using gene chips data of Kaplan–Meier Plotter. The prognostic value of JAK mRNA for OS and disease-free survival (DFS) was also analyzed using the RNA-seq data in bc-GenExMiner tool (http://bcgenex.ico.unicancer.fr).23 SurvExpress (http://bioinformatica.mty.itesm.mx:8080/Biomatec/SurvivaX.jsp)24 was applied for the prognostic analysis of JAK family signature in BC cases from Van De Vijver Nature 200225 cohort for the OS/recurrence-free survival (RFS)/metastasis-free survival (MFS) and TCGA cohort for OS.
Survival Outcomes and JAK Protein Levels
The survival analysis based on JAK protein levels was performed in TRGAted (https://nborcherding.shinyapps.io/TRGAted/).26 Only the JAK2 protein was available on this platform. We explored the relationships between JAK2 protein level and OS/disease-specific survival (DSS)/DFS/progression-free survival (PFS) in BC patients.
Survival Outcomes and DNA Methylation of JAK or Family Signature
A web tool named MethSurv (https://biit.cs.ut.ee/methsurv/)27 was used to assess the prognostic value of different CpG methylation patterns of JAK1/JAK2/JAK3/TYK2 in BC patients. The effect of DNA methylation of JAK family signature on BC prognosis was analyzed using SurvivalMeth (http://bio-bigdata.hrbmu.edu.cn/survivalmeth/).28
Genetic Alteration, Co-Expression, PPI Analyses, and Enrichment Network of JAKs
Genetic alteration of JAKs was available from cBioPortal for Cancer Genomics (https://www.cbioportal.org/).29,30 In addition, co-expression of JAKs was studied using the correlation module of TIMER (https://cistrome.shinyapps.io/timer/).19 Protein–protein interaction (PPI) network was obtained from STRING (https://string-db.org/)31 and GeneMANIA servers (https://genemania.org/).32 Visual analytics of enrichment network of the four members of JAK family was based on the NetworkAnalysis (https://www.networkanalyst.ca/).33
Functional Enrichment Analysis of JAKs and Their Most Frequently Altered Neighboring Genes
The top 50 genes with highest frequency associated with JAKs were available from cBioPortal for Cancer Genomics (https://www.cbioportal.org/).29,30 The integrated network data, including biological process (BP), cellular component (CC), and molecular function (MF) and KEGG pathway, were taken from DAVID 6.8 (https://david.ncifcrf.gov/).34,35 The functional enrichment analysis of JAKs and their neighboring genes was also conducted on Metascape website (https://metascape.org/gp/index.html).36
Kinase, miRNA, and Transcription Factor (TF) Targets of JAKs
The top 10 kinase targets of JAKs were also identified from LinkedOmics (http://www.linkedomics.org/),37 and the predicted miRNA or TFs and their connections with JAKs were available from NetworkAnalysis (https://www.networkanalyst.ca/).33
JAKs and Infiltrating Immune Cells
The “Gene” module of TIMER (https://cistrome.shinyapps.io/timer/)19 was used to visualize the correlation of JAK1, JAK2, JAK3, and TYK2 gene expression with immune infiltration levels in BC. The “SCNA” module provided the comparison of tumor infiltration levels among BC samples with different somatic copy number alterations (CNA) for JAK genes. The “Survival” module was used to explore the clinical relevance of tumor immune subsets in a multivariate Cox proportional hazard model.
Correlation Analysis Between JAK Expression and Immune Cell Markers
The correlation module of TIMER (https://cistrome.shinyapps.io/timer/)19 was used for the correlation analysis between JAK expression and immune cell markers together with the Spearman’s rho value and estimated statistical significance. The related gene markers of multiple immune cells were obtained from the related references38–40 and the CellMarker database (http://biocc.hrbmu.edu.cn/CellMarker/).41
Statistical Analysis
The survival curves related to OS, DMFS, PPS, RFS, DFS, DSS, PFS, and MFS of JAKs were drawn from Kaplan Meier plotter, bc-GenExMiner, SurvExpress, TRGAted, MethSurv, or SurvivalMeth, and the P-values with hazard ratios (HRs) were provided. In NetworkAnalysis, miRNA–gene interaction data were collected from TarBase, while transcription factor and gene ChIP-seq data were collected from ENCODE. Only peak intensity signal <500 and the predicted regulatory potential score <1 were used (using BETA Minus algorithm). In this study, TCGA_BC dataset was used in the “LinkInterpreter” module. Gene Set Enrichment Analysis (GSEA) tool was applied to explore the kinase target of JAKs in the BC, with a minimum number of genes (size) of 3 and a simulation of 500. Gene correlations were evaluated with Spearman correlation coefficients and P-values. A P-value lower than 0.05 was considered statistically significant.
Results
Expression of JAK mRNA and Protein, and Methylation Level in Breast Cancer
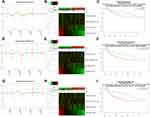
We first explored the expression of JAKs and methylation level in tumor and normal samples (Figure 1A–C). As shown in Figure 1A, JAK1 and JAK2 mRNA level was significantly decreased, while TYK2 mRNA level was significantly increased in tumor tissues of BC compared with the corresponding normal tissues. Higher JAK3 mRNA level was found in tumor tissues, but statistical significance was not reached. Protein levels of JAK1, JAK2, and TYK2 were significantly lower in tumor samples, while JAK3 protein level was significantly higher in tumor samples (Figure 1B). Significant hypermethylation of JAK3 and TYK2 as well as hypomethylation of JAK1 were found in BC tumor samples (Figure 1C).
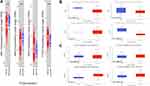 |
Figure 1 Expression of JAK mRNA (A) and protein (B), and methylation level (C) in the BC cases. |
We also compared the relative expression levels of JAK mRNA in BC and found that among the JAK family members the relative expression of JAK1 was the highest, while JAK3 had the lowest expression (Supplementary Figure 2). There was a significant correlation between JAK2 level and pathologic stage of BC (Supplementary Figure 3).
Prognostic Value of Individual JAK mRNA Expression Level in Breast Cancer Patients
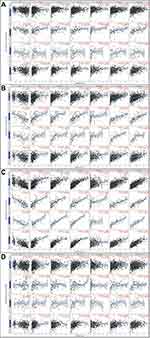
The associations between JAK mRNA expression and prognosis were analyzed using the data from gene chip in Kaplan–Meier Plotter (Figure 2A–P). Low expression level of JAK1 (P =1.4e-06) and TYK2 (P=0.0018) indicated significantly shorter OS in patients with breast cancer (Figure 2A and M), while low expression of JAK2 (P=0.096) and JAK3 (P=0.086) showed a modest association with a worse OS (Figure 2E and I).
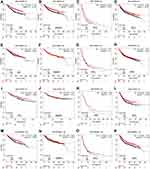 |
Figure 2 Survival curves of OS (A, E, I, M), DMFS (B, F, J, N), PPS (C, G, K, O), and RFS (D, H, L, P) for the expression of JAKs in patients with breast cancer. |
There were significant associations between DMFS and JAK1 (P=0.00014) and TYK2 (P=0.029) expression in breast cancer patients (Figure 2B and N). Only JAK1 (P=0.043) was significantly related to PPS in breast cancer patients (Figure 2C).
Low transcriptional levels of JAK1 (P=5e-05), JAK3 (P=0.0022), and TYK2 (P=1.9e-08) in breast cancer patients were significantly associated with worse RFS (Figure 2A, L and P), while a modest association was found between JAK2 and RFS (P=0.085) (Figure 2H).
In addition, we also assessed the prognostic value of JAKs using the data from RNA-seq in bc-GenExMiner. We found low mRNA level of JAK1 and JAK2, which indicated significantly shorter OS and worse DFS in patients with breast cancer (all P<0.001) (Supplementary Figure 4).
The Relationship Between JAK mRNA Expression and Clinical Variables in Breast Cancer
We further studied the association between JAK mRNA expression levels and clinical variables in breast cancer. We found that low expression of JAK1 was significantly related to reduced OS in patients with the following subtypes: ER-positive or -negative, PR-positive, TP53 wild-type, basal, luminal B, lymph node positive or negative, grade 2 or 3, and luminal androgen receptor subtype (Supplementary Table 1). Low expression of JAK2 was also significantly related to shorter OS in patients with basal, luminal B, lymph node positive or grade 3 subtype, and inferior DMFS in patients with ER-negative, HER2-negative, basal, luminal B, or luminal androgen receptor subtype (Supplementary Table 2). Low expression of JAK3 was also obviously related to worse OS in patients with basal, grade 3, and basal-like 1 subtype, and inferior DMFS in patients with basal subtype (Supplementary Table 3). Low expression of JAK3 or TYK2 was related to worse OS in patients with basal and luminal B subtype (Supplementary Table 4). The results of prognostic values of JAKs in clinicopathological subtypes regarding OS, DMFS, PPS, and RFS in breast cancer are detailed displayed in Supplementary Table 1–4.
Prognostic Value of JAK mRNA Signature in Breast Cancer
The JAK family signature was input for prognostic analysis in SurvExpress. All patients were assigned to the high/low-risk groups based on the score value with an optimal cutoff. The dataset from the cohort by Van De Vijver was applied. For OS (Figure 3A–C), distinct expression patterns of JAK1 and JAK2 were noticed between low- and high-risk groups. The high-risk group displayed a significantly shorter OS compared with the low-risk group (HR=1.91, 95% CI: 1.21–3.03, P=0.005469), which was also confirmed in the TCGA cohort (Supplementary Figure 5, HR=1.57, 95% CI: 1.11–2.21, P=0.01007). In addition, for RFS (Figure 3D–F), the expression levels of JAK1, JAK2, and TYK2 were higher in the low-risk group than in the high-risk group, and the high-risk group showed a significantly worse RFS than the low-risk group (HR=2.01, 95% CI: 1.36–2.99, P=0.0005106). For MFS (Figure 3G–I), significantly lower expression patterns of JAK1, JAK2, and TYK2 were noticed in the high-risk group, and the cases in the high-risk group had an inferior MFS compared with the low-risk group (HR=2.16, 95% CI: 1.45–3.21, P=0.0001424).
Prognostic Value of JAK2 Protein Expression in Breast Cancer
The prognostic values of JAK protein were investigated in the reverse-phase protein arrays data of The Cancer Proteome Atlas (TCPA) Portal. Of note, only JAK2 was available (Figure 4A–D). In fact, the BC patients with low protein expression of JAK2 had unfavorable DSS (HR=0.305, P=0.0068) and PFS (HR=0.48, P=0.014) compared with those with high protein expression.
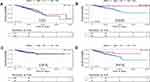 |
Figure 4 The prognostic value of JAK2 protein in BC. (A) The survival curve for OS. (B) The survival curve for DSS. (C) The survival curve for DFS. (D) The survival curve for PFS. |
Prognostic Value of Single CpG of JAKs in Breast Cancer
The prognostic value of DNA methylation of JAK family in breast cancer was analyzed by MethSurv. The heat map of DNA methylation results is displayed in Supplementary Figure 6. Specifically, cg18227442 of JAK1, cg13629201 of JAK2, cg11085762 of JAK3, and cg11369662 of TYK2 showed the highest DNA methylation level. Overall, we found that five CpGs of JAK1, four CpGs of JAK2, 20 CpGs of JAK3, and 13 CpGs of TYK2 were significantly associated with prognosis in breast cancer patients (Table 1).
 |
Table 1 The Prognostic Value of CpG in the JAK Family by MethSurv (P<0.05) |
Prognostic Value of the DNA Methylation of JAK Family Signature in Breast Cancer
The gene symbols of the JAK family members (JAK1, JAK2, JAK3, TYK2) were input for prognostic analysis in SurvivalMeth. Significant expression patterns of all JAKs were found between the low- and high-risk groups (Figure 5A). The heat map showed that DNA methylation level of JAK2 was the highest (Figure 5B), and the high-risk group displayed an unfavorable prognosis compared with the low-risk group (P=0.0060540538) (Figure 5C).
Genetic Alteration, Co-Expression, Physical Interaction Analyses, and Enrichment Network of JAKs
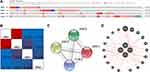
Comprehensive analyses of molecular characteristics of JAKs were further conducted. First, the genetic alterations of these members were analyzed. As displayed in Figure 6A, JAK1, JAK2, JAK3, and TYK2 were altered in 6%, 12%, 5%, and 9% of the 960 samples, respectively. Altered mRNA expression, amplification, and deep deletion of JAKs were also commonly found in these samples.
Then, the potential co-expressions of JAKs were explored. There was a strong correlation between JAK1 and JAK2, and low correlations were found among other JAK members (Figure 6B). Moreover, protein/protein interactions (PPI) of JAKs were explored using STRING. As expected, four nodes and six edges were obtained in the PPI network (Figure 6C). The physical interaction analyses of JAKs were also performed with GeneMANIA (Figure 6D). Moreover, the enrichment network of JAKs was done with NetworkAnalysis, and the integrated interactive visualization with the top 10 highly enriched items of JAKs (P<0.05) is displayed in Supplementary Figure 7. The results revealed that the functions of these four members were mainly related to immune response, immune system, binding and kinase activity in GO items (Supplementary Figure 7A–C) and Th1 and Th2 cell differentiation, Th17 cell differentiation, and JAK-STAT signaling path in KEGG analysis (Supplementary Figure 7D).
Functional Enrichment Analysis of JAKs and Their Most Frequently Altered Neighboring Genes
Moreover, the top 50 genes with the highest frequency associated with JAKs were available from cBioPortal. The data suggested that TP53, TTN, PIK3CA, SYNE2, CDH1, COL4A2, HMGB3, ACACA, HMGXB4, LY75, PRKDC, CUBN, MYO9A, ATM, ADGRG4, SDK1, FRMPD4, MUC12, GATA3, NEB, MUC16, RNF213, NHS, CTCF, DNAH11, OBSCN, MDN1, MAP3K1, ABCC1, ANKRD36, APLP1, BCAS2, BEND5, CACNA1S, CATSPERB, CCDC138, CCDC34, CCT4, CRX, CWC27, CYP4V2, DENND2D, DGCR8, DNAJB7, FADS1, FAM49A, FGFR2, GAS2, GOSR1, and GRIN2B were primarily connected with the modulation and function of JAKs in breast cancer. The integrated network using DAVID 6.8 with these 54 genes is shown in Supplementary Figure 8.
In the BP category, the most highly enriched GO items, including MAPK cascade, innate immune response, phosphatidylinositol-3-phosphate biosynthetic process, and response to antibiotic, might be associated with the tumorigenesis and progression of BC (Supplementary Figure 8A). In the CC category, the most significantly highly enriched items included extrinsic component of cytoplasmic side of plasma membrane, nucleolus, and cytoskeleton (Supplementary Figure 8A). In the MF category, the JAKs and their neighboring genes were mainly enriched in binding and kinase activities (Supplementary Figure 8A). As expected in KEGG pathway analysis (Supplementary Figure 8B), hsa04151 (PI3K-Akt signaling pathway), hsa04550 (signaling pathways regulating pluripotency of stem cells), hsa04630 (JAK/STAT signaling pathway), and hsa05200 (pathways in cancer) were significantly related to the tumorigenesis and development of breast cancer.
In addition, we also performed the functional enrichment analysis in Metascape website. Supplementary Figures 9 and 10 show that the functions of JAKs and their neighboring genes were mainly enriched in ST interleukin 4 pathway, PID IFNG pathway (IFN-gamma pathway), and regulation of innate immune response. To better understand the correlation between JAKs and BC, the PPI network and mCODE components were also analyzed. We found that biological function was mainly associated with ST interleukin 4 pathway, interleukin receptor SHC signaling, interleukin-2 family signaling, and interleukin-20 family signaling (Supplementary Figure 11).
Kinase Targets, Gene–miRNA Interaction, and Transcription Factor (TF)-Gene Interaction of JAKs
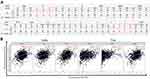
In addition, the top 10 kinase targets of JAKs were also identified from LinkedOmics (Table 2). We also further explored potential miRNAs and TFs of JAKs as well as their interactions using NetworkAnalysis (Figure 7). The predicted miRNAs or TFs are shown in Figure 7A and B. Namely, there were 75 miRNA targets predicted in JAK1, 25 miRNAs in JAK2, 10 miRNAs in JAK3, and 23 miRNAs in TYK2. There were 15 TF targets predicted in JAK1, 22 TFs in JAK2, 6 TFs in JAK3, and 27 TFs in TYK2. miRNAs or TFs were also found to synchronously correlate with all JAK members or at least with three of them.
 |
Table 2 The Top 10 Kinase Targets of JAKs in BC (P<0.05) |
Infiltrating Immune Cell and JAK Expression in Breast Cancer
Given the close relationship between immunological features and prognosis in cancer, we further investigated the correlation between immune infiltrates and JAK expression in BC (Figure 8). Overall, JAK1, JAK2, and JAK3 expression was highly positively associated with all six immune infiltrates (B cells, CD4+ T cells, CD8+ T cells, macrophages, neutrophils, and dendritic cells) (all P<0.05). A significant positive correlation was found between TYK2 expression and four kinds of immune cell infiltration, including B cells, CD4+ T cells, neutrophils, and dendritic cells (P< 0.05). In different breast cancer subtypes, including basal-like, HER2+, and luminal, the correlations between JAK expression and the six types of infiltrating immune cells were not all the same (Figure 8).
We also compared the tumor infiltration levels among BC with different somatic copy number alterations for JAKs (Supplementary Figure 12). Moreover, the multivariate Cox proportional hazard model was applied for JAKs and six tumor-infiltrating immune cells in BC and the subtypes. Overall, macrophages (P=0.059) and JAK2 (P=0.075) were marginally associated with the clinical outcome of all BC patients (Supplementary Table 5). In addition, macrophages (P=0.010), B cells (P=0.028), and dendritic cells (P=0.042) were significantly related to the clinical outcome in basal, HER2, and luminal subtype, respectively (Supplementary Tables 6–8).
Correlation Analysis Between JAK Expression and Various Markers of Immune Cells
To investigate the effects of JAK expression on tumor-infiltrating immune cells, we analyzed the correlations between JAK1/JAK2/JAK3/TYK2 expression and various markers of immune cells via public databases. The innate and adaptive immune cells investigated included monocytes, tumor-associated macrophages (TAMs), M1 and M2 macrophages, neutrophils, natural killer cells, dendritic cells, CD8+ T cells, B cells, and T cells (general), as well as the subsets of T cells (Th1, Th2, Tfh, Th17, Tregs, and exhausted T cells). The correlation was also adjusted for tumor purity due to its influences on the immune infiltration analysis.
We found that the JAK1 expression significantly correlated with the expression of gene markers in TAMs and Treg in BC patients after adjusting for tumor purity (Figure 9). The correlations between the expression level of JAK2 and the expression of gene marker sets of TAMs, natural killer cells, dendritic cells, CD8+ T cells, T cells (general), Th1, Tfh, Th17, Treg, and T cell exhaustion were still significant after adjustment for purity (Supplementary Figure 13). JAK3 expression was significantly associated with TAMs, M2, natural killer cells, dendritic cells, CD8+ T cells, T cells (general), and Th1 and T cell exhaustion in BC after adjustment for purity (Supplementary Figure 14). Regarding the correlation between TYK2 and immune cells in BC cases, significant associations were found for dendritic cells, CD8+ T cells, T cells (general), Th2, Treg, and T cell exhaustion (Supplementary Figure 15). The results strongly indicated that JAKs might influence prognosis of BC through regulating immune response.
Discussion
Janus kinases are a family of intracellular, nonreceptor tyrosine kinases, which comprise four members (JAK1, JAK2, JAK3 and TYK2). JAKs were initially identified using polymerase chain reaction and verified by the human genome project.10,42–45 JAKs possess two near-identical phosphate-transferring domains and mainly exertive functions through transducing cytokine-mediated signals via the JAK-STAT pathway.46 JAKs have been reported to be implicated in the pathogenesis of inflammatory and immune disorders as well as malignant tumors.47–49 And the clinical significance of JAKs in breast cancer was also reported. For example, JAK1 expression inversely correlates with tumor size, lymph node status, and TNM of breast cancer patients.50 And somatic mutations of JAKs (including JAK1, JAK2 and JAK3) have been commonly found in breast cancer and have potential implications for clinical management.51–53 TYK2 plays important roles in growth and metastasis of breast cancer.54,55 JAK inhibitors, such as CP-690,550 (JAK3 target), Lestaurtinib (multikinase target), and AZ-01/AZ-60 (JAK2 target) are currently under development for the treatment of autoimmune diseases and hematological malignancy.46,56,57 JAKs are closely associated with cytokine receptors, such as IFNγ, IL-2, IL-4 and IL-10,58,59 and have important immunologic and inflammation functions.60 In addition, JAK expression levels were also associated with the therapeutic effect in cases treated with anti-PD1 immunotherapy. The anti-PD1 therapy could improve the survival time of mice with NK/T-cell neoplasms and the level of JAK2 was greatly increased in mice treated with anti-PD1.61 Another study found that in patients with non-small cell lung cancer (NSCLC), JAK3 mutations upregulated PD-L1 expression, which explains their benefit from anti-PD1 treatment.16 However, the prognostic significance and biological functions of JAKs in breast cancer have not been comprehensively analyzed.
First, we examined the expression levels of JAKs in the breast cancer. Compared to normal tissues, tumor samples showed significantly decreased JAK1 and JAK2 expression at both mRNA and protein levels, significantly higher JAK3 expression at protein level, and significantly increased TYK2 expression at mRNA level but decreased at protein level. Besides, JAK3 and TYK2 showed hypermethylation, while JAK1 showed hypo-methylation in BC tumor samples. Moreover, in BC tissues, the relative transcriptional level was the highest for JAK1 and the lowest for JAK3, and there was a significant correlation between JAK2 level and BC pathologic stage. These data suggested that JAKs might play multiple important roles in the breast carcinoma tumorigenesis and progression.
Then, we provided systematic prognostic landscape of the expression level of JAK mRNA, protein, and DNA methylation status in breast cancer via multiple databases. First, we found that, through the gene chips data from K-M plotter, low transcriptional level of JAK1 was significantly related to worse OS/DMFS/PPS/RFS, and a modest association was found between worse OS/RFS and decreased JAK2 mRNA level. JAK3 mRNA level displayed a modest association with shorter OS, and a significant relationship with a worse RFS. Low TYK2 level indicated worse OS/DMFS/RFS. Then, we further assessed the association between JAK mRNA expression levels and OS/DMFS/PPS/RFS in breast cancer cases experiencing different clinical conditions. The results using the RNA-seq data of bc-GenExMiner tools showed that breast cancer patients with low mRNA level of JAK1 or JAK2 had significantly shorter OS and worse DFS. Second, we found low JAK2 protein expression group had unfavorable DSS and PFS. Third, we found that five CpGs of JAK1, four CpGs of JAK2, 20 CpGs of JAK3, and 13 CpGs of TYK2 were significantly related to prognosis in breast cancer patients. The mRNA signature analysis showed unfavorable OS/RFS/MFS in the high-risk group of BC patients, and the DNA methylation signature analysis indicated that the high-risk group had worse survival than the low-risk group. These results revealed strong evidence for powerful predictive abilities of JAKs as promising prognostic factors in BC patients.
As breast cancer emerges and progresses by a multistep and multifactorial process involving progressively accumulated genetic, epigenetic, and microenvironmental alterations,62–65 we further performed comprehensive analyses of the molecular characteristics, interaction networks, and the immune microenvironment.
Available from cBioPortal for Cancer Genomics, alterations of JAK1, JAK2, JAK3, and TYK2 were found in 6%, 12%, 5%, and 9% of the BC cases, respectively. Altered mRNA expression, amplification, and deep deletion were also commonly observed. In addition, a strong correlation was found between JAK1 and JAK2, and low correlations were found among other JAK members. These data confirmed that accumulated genetic alterations are involved in the occurrence and progression of breast tumors.
The related molecular biology functions of JAKs were also analyzed. The PPI and the enrichment network of JAKs showed that JAK family was mainly related to immune response, differentiation of immune cells (Th1, Th2, and Th17 cells) and JAK-STAT signaling path. Th1, Th2 and Th17 cells, the subsets of CD4+ T-cell, execute diverse important immune functions.66 It is known that CD4⁺T cells are essential to achieve an effective immune response to pathogens, and the differentiation of naive CD4⁺T cells mainly depends on the cytokine microenvironment, the extracellular matrix and transcription factors.66–71 In breast cancer, Th1 produces IFN-γ and IL-12, exerting anti-tumor activity. Th2 produces IL-4 and IL-10 to promote tumor activity.72 The Th1/Th2 balance is closely related to tumor immunity in breast tumor.72,73 Th17 cells, as the most important factor involved in the inflammation, play significant roles in multiple diseases by production of IL17 cytokine, including breast cancer.74,75 And the JAK-STAT signaling pathway is involved in various fundamental processes such as cell division and apoptosis, stem cell niche maintenance, inflammation and immunity, as well as tumor formation and cancer drug resistance.76–80 The functional enrichment analysis of JAKs and their most frequently altered neighboring genes was also conducted. The mainly highly enriched items included MAPK cascade, innate immune response, binding and kinase activities in BP/CC/MF categories, and hsa04151 (PI3K-Akt signaling pathway), hsa04630 (JAK-STAT signaling pathway), and hsa05200 (pathways in cancer) in KEGG pathway analysis. They play critical roles in diverse cellular functions, including cell growth, proliferation, and survival, and have also been implicated in therapeutic resistance, angiogenesis, tumor migration, invasion, and metastasis.81–85 Moreover, we also sought to characterize the kinase targets, potential miRNAs, and TFs of JAKs as well as their interactions. The top 10 kinase targets of JAKs were identified, and there were 75 miRNA targets predicted in JAK1, 25 miRNAs in JAK2, 10 miRNAs in JAK3, and 23 miRNAs in TYK2. Fifteen TF targets were predicted in JAK1, 22 TFs in JAK2, 6 TFs in JAK3, and 27 TFs in TYK2. The interactions of JAK-miRNA and JAK-TF and the potential kinase targets might be involved in the formation and development of breast tumors.
Moreover, the correlation between JAKs and immune cell infiltration in BC was also assessed. JAK1, JAK2 and JAK3 expression were found to be highly positively associated with six immune infiltrates (B cells, CD4+ T cells, CD8+ T cells, macrophages, neutrophils and dendritic cells), and TYK2 expression was highly positively related to infiltrations of B cells, CD4+ T cells, Neutrophils, and Dendritic cells. In different breast cancer subtypes, including basal-like, HER2+ and luminal, the correlations between JAKs expression and immune cells infiltrates were also analyzed. We also explored the correlation between somatic CNA of JAKs and abundance of immune infiltrates. In addition, Macrophage, Dendritic cells, and B_cell infiltrates could be an independent factor for clinical survival in basal, luminal, and Her2 cases, respectively. We also performed correlation analysis between JAKs expression and various markers of immune cells. After adjusting for tumor purity, we found that JAK1 expression was significantly associated with the expression of gene markers of TAMs and Treg in BC patients. JAK2 expression correlated with the expression of gene marker sets of TAMs, natural killer cells, dendritic cells, CD8+ T cells, T cells (general), Th1, Tfh, Th17, Treg, and T cell exhaustion. JAK3 expression was significantly related to TAMs, M2, natural killer cells, dendritic cells, CD8+ T cells, T cells (general), Th1, and T cell exhaustion. Significant correlation was found between TYK2 and some of the immune cells (dendritic cells, CD8+ T cells, T cells [general], Th2, Treg, and T cell exhaustion) in BC. These results above indicated JAKs might have an impact on survival of BC patients through regulating immune response and tumor-infiltrating lymphocytes (TILs).
In previous studies, TILs were demonstrated to be closely related to clinical outcomes of BC patients, a systematic evaluation of these cell infiltrations may be able to guide the prognosis and appropriate treatment of breast cancer.86–88 In this work, our findings further showed the expression of JAKs was associated to the infiltration of immune/inflammatory cells. JAKs could be used as prognostic factors and immunotherapy targets in breast cancer.
Our results provided new insights into the identification of JAKs as promising drug targets and powerful prognostic markers in breast cancer. However, some limitations to our study should be acknowledged. First, there might be potential biases caused by the confounders from bioinformatics databases. Second, the findings in our work are based on the data from public databases, and the exact mechanisms need to be validated in future in vitro or in vivo experiments.
In conclusion, the observations of this work suggest a potential role of JAKs as prognostic factors and possible targets in breast cancer.
Data Sharing Statement
All datasets used and/or analyzed during the current study are available from the public database.
Acknowledgments
We would like to extend our sincere gratitude to the editors and anonymous reviewers for their professional reviews and valuable suggestions. We also thank LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Disclosure
All authors declare no conflicts of interest in this study.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Harbeck N, Penault-Llorca F, Cortes J, et al. Breast cancer. Nature Reviews Disease Primers. 2019;5(1):66. doi:10.1038/s41572-019-0111-2
3. Anastasiadi Z, Lianos GD, Ignatiadou E, Harissis HV, Mitsis M. Breast cancer in young women: an overview. Updates Surg. 2017;69(3):313–317. doi:10.1007/s13304-017-0424-1
4. Fragomeni SM, Sciallis A, Jeruss JS. Molecular subtypes and local-regional control of breast cancer. Surg Oncol Clin N Am. 2018;27(1):95–120. doi:10.1016/j.soc.2017.08.005
5. Turashvili G, Brogi E. Tumor heterogeneity in breast cancer. Front Med. 2017;4:227. doi:10.3389/fmed.2017.00227
6. Nieto C, Vega MA, Martín Del Valle EM. Trastuzumab: more than a guide in her2-positive cancer nanomedicine. Nanomaterials. 2020;10(9). doi:10.3390/nano10091674
7. Şahin S, Caglayan MO, Üstündağ Z. Recent advances in aptamer-based sensors for breast cancer diagnosis: special cases for nanomaterial-based VEGF, HER2, and MUC1 aptasensors. Mikrochim Acta. 2020;187(10):549. doi:10.1007/s00604-020-04526-x
8. de Melo Gagliato D, Buzaid AC, Perez-Garcia J, Cortes J. Immunotherapy in breast cancer: current practice and clinical challenges. BioDrugs. 2020. doi:10.1007/s40259-020-00436-9
9. Ladyman SR, Fieldwick DM, Grattan DR. Suppression of leptin-induced hypothalamic JAK/STAT signalling and feeding response during pregnancy in the mouse. Reproduction. 2012;144(1):83–90. doi:10.1530/rep-12-0112
10. Wilks AF. Two putative protein-tyrosine kinases identified by application of the polymerase chain reaction. Proc Natl Acad Sci U S A. 1989;86(5):1603–1607. doi:10.1073/pnas.86.5.1603
11. Schwartz DM, Bonelli M, Gadina M, O’Shea JJ. Type I/II cytokines, JAKs, and new strategies for treating autoimmune diseases. Nat Rev Rheumatol. 2016;12(1):25–36. doi:10.1038/nrrheum.2015.167
12. Kleppe M, Kwak M, Koppikar P, et al. JAK-STAT pathway activation in malignant and nonmalignant cells contributes to MPN pathogenesis and therapeutic response. Cancer Discov. 2015;5(3):316–331. doi:10.1158/2159-8290.Cd-14-0736
13. Yeh YT, Ou-Yang F, Chen IF, et al. Altered p-JAK1 expression is associated with estrogen receptor status in breast infiltrating ductal carcinoma. Oncol Rep. 2007;17(1):35–39. doi:10.3892/or.17.1.35
14. Wen W, Liang W, Wu J, et al. Targeting JAK1/STAT3 signaling suppresses tumor progression and metastasis in a peritoneal model of human ovarian cancer. Mol Cancer Ther. 2014;13(12):3037–3048. doi:10.1158/1535-7163.Mct-14-0077
15. Park SY, Lee CJ, Choi JH, et al. The JAK2/STAT3/CCND2 Axis promotes colorectal Cancer stem cell persistence and radioresistance. J Experimental Clin Cancer Res. 2019;38(1):399. doi:10.1186/s13046-019-1405-7
16. Li SD, Ma M, Li H, et al. Cancer gene profiling in non-small cell lung cancers reveals activating mutations in JAK2 and JAK3 with therapeutic implications. Genome Med. 2017;9(1):89. doi:10.1186/s13073-017-0478-1
17. Wöss K, Simonović N, Strobl B, Macho-Maschler S, Müller M. TYK2: an upstream kinase of STATs in cancer. Cancers. 2019;11(11). doi:10.3390/cancers11111728.
18. Prutsch N, Gurnhofer E, Suske T, et al. Dependency on the TYK2/STAT1/MCL1 axis in anaplastic large cell lymphoma. Leukemia. 2019;33(3):696–709. doi:10.1038/s41375-018-0239-1
19. Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–e110. doi:10.1158/0008-5472.Can-17-0307
20. Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19(8):649–658. doi:10.1016/j.neo.2017.05.002
21. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–W102. doi:10.1093/nar/gkx247
22. Györffy B, Lanczky A, Eklund AC, et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1809 patients. Breast Cancer Res Treat. 2010;123(3):725–731. doi:10.1007/s10549-009-0674-9
23. Jézéquel P, Campone M, Gouraud W, et al. bc-GenExMiner: an easy-to-use online platform for gene prognostic analyses in breast cancer. Breast Cancer Res Treat. 2012;131(3):765–775. doi:10.1007/s10549-011-1457-7
24. Aguirre-Gamboa R, Gomez-Rueda H, Martínez-Ledesma E, et al. SurvExpress: an online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS One. 2013;8(9):e74250. doi:10.1371/journal.pone.0074250
25. van ‘T Veer LJ, Dai H, van de Vijver MJ, et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature. 2002;415(6871):530–536. doi:10.1038/415530a
26. Borcherding N, Bormann NL, Voigt AP, Zhang W. TRGAted: a web tool for survival analysis using protein data in the cancer genome atlas. F1000Research. 2018;7:1235. doi:10.12688/f1000research.15789.2
27. Modhukur V, Iljasenko T, Metsalu T, Lokk K, Laisk-Podar T, Vilo J. MethSurv: a web tool to perform multivariable survival analysis using DNA methylation data. Epigenomics. 2018;10(3):277–288. doi:10.2217/epi-2017-0118
28. Zhang C, Zhao N, Zhang X, et al. SurvivalMeth: a web server to investigate the effect of DNA methylation-related functional elements on prognosis. Brief Bioinform. 2020. doi:10.1093/bib/bbaa162
29. Cerami E, Gao J, Dogrusoz U, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–404. doi:10.1158/2159-8290.Cd-12-0095
30. Gao J, Aksoy BA, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1. doi:10.1126/scisignal.2004088
31. Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–d613. doi:10.1093/nar/gky1131
32. Warde-Farley D, Donaldson SL, Comes O, et al. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010;38(Web Server issue):W214–20. doi:10.1093/nar/gkq537
33. Zhou G, Soufan O, Ewald J, Hancock REW, Basu N, Xia J. NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 2019;47(W1):W234–w241. doi:10.1093/nar/gkz240
34. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. doi:10.1038/nprot.2008.211
35. Huang da W, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37(1):1–13. doi:10.1093/nar/gkn923
36. Zhou Y, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10(1):1523. doi:10.1038/s41467-019-09234-6
37. Vasaikar SV, Straub P, Wang J, Zhang B. LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 2017;46(D1):D956–D963. doi:10.1093/nar/gkx1090
38. Song D, Wang Y, Zhu K, et al. DCK is a promising prognostic biomarker and correlated with immune infiltrates in hepatocellular carcinoma. World J Surg Oncol. 2020;18(1):176. doi:10.1186/s12957-020-01953-1
39. Dai D, Chen B, Feng Y, et al. Prognostic value of prostaglandin I2 synthase and its correlation with tumor-infiltrating immune cells in lung cancer, ovarian cancer, and gastric cancer. Aging. 2020;12(10):9658–9685. doi:10.18632/aging.103235
40. Liu F, Wu H. Prognostic value of gastrokine-2 (gkn2) and its correlation with tumor-infiltrating immune cells in lung cancer and gastric cancers. J Inflamm Res. 2020;13:933–944. doi:10.2147/JIR.S277353
41. Zhang X, Lan Y, Xu J, et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 2018;47(D1):D721–D728. doi:10.1093/nar/gky900
42. Wilks AF. Cloning members of protein-tyrosine kinase family using polymerase chain reaction. Methods Enzymol. 1991;200:533–546. doi:10.1016/0076-6879(91)00169-w
43. Harpur AG, Andres AC, Ziemiecki A, Aston RR, Wilks AF. JAK2, a third member of the JAK family of protein tyrosine kinases. Oncogene. 1992;7(7):1347–1353.
44. Firmbach-Kraft I, Byers M, Shows T, Dalla-Favera R, Krolewski JJ. tyk2, prototype of a novel class of non-receptor tyrosine kinase genes. Oncogene. 1990;5(9):1329–1336.
45. Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002;298(5600):1912–1934. doi:10.1126/science.1075762
46. Pesu M, Laurence A, Kishore N, Zwillich SH, Chan G, O’Shea JJ. Therapeutic targeting of Janus kinases. Immunol Rev. 2008;223:132–142. doi:10.1111/j.1600-065X.2008.00644.x
47. Vainchenker W, Dusa A, Constantinescu SN. JAKs in pathology: role of Janus kinases in hematopoietic malignancies and immunodeficiencies. Semin Cell Dev Biol. 2008;19(4):385–393. doi:10.1016/j.semcdb.2008.07.002
48. Cance WG, Liu ET. Protein kinases in human breast cancer. Breast Cancer Res Treat. 1995;35(1):105–114. doi:10.1007/bf00694751
49. Ciobanu DA, Poenariu IS, Crînguș LI, et al. JAK/STAT pathway in pathology of rheumatoid arthritis (Review). Exp Ther Med. 2020;20(4):3498–3503. doi:10.3892/etm.2020.8982
50. Chen B, Lai J, Dai D, Chen R, Li X, Liao N. JAK1 as a prognostic marker and its correlation with immune infiltrates in breast cancer. Aging. 2019;11(23):11124–11135. doi:10.18632/aging.102514
51. Balko JM, Schwarz LJ, Luo N, et al. Triple-negative breast cancers with amplification of JAK2 at the 9p24 locus demonstrate JAK2-specific dependence. Sci Transl Med. 2016;8(334):334ra53. doi:10.1126/scitranslmed.aad3001
52. Naeem MA, Shah TH, Zafar N, Khan M, Bhutto AA, Rabbani S. Detection of JAK2 gene mutation in Pakistani women with triple-negative breast cancer. Breast J. 2020;26(4):829–830. doi:10.1111/tbj.13609
53. Jeong EG, Kim MS, Nam HK, et al. Somatic mutations of JAK1 and JAK3 in acute leukemias and solid cancers. Clinical Cancer Res. 2008;14(12):3716–3721. doi:10.1158/1078-0432.Ccr-07-4839
54. Zhang Q, Sturgill JL, Kmieciak M, et al. The role of Tyk2 in regulation of breast cancer growth. J Interferon Cytokine Res. 2011;31(9):671–677. doi:10.1089/jir.2011.0023
55. Sang QX, Man YG, Sung YM, et al. Non-receptor tyrosine kinase 2 reaches its lowest expression levels in human breast cancer during regional nodal metastasis. Clin Exp Metastasis. 2012;29(2):143–153. doi:10.1007/s10585-011-9437-1
56. Furumoto Y, Gadina M. The arrival of JAK inhibitors: advancing the treatment of immune and hematologic disorders. BioDrugs. 2013;27(5):431–438. doi:10.1007/s40259-013-0040-7
57. Harrington R, Al Nokhatha SA, Conway R. JAK inhibitors in rheumatoid arthritis: an evidence-based review on the emerging clinical data. J Inflamm Res. 2020;13:519–531. doi:10.2147/jir.S219586
58. Smith GA, Uchida K, Weiss A, Taunton J. Essential biphasic role for JAK3 catalytic activity in IL-2 receptor signaling. Nat Chem Biol. 2016;12(5):373–379. doi:10.1038/nchembio.2056
59. Rodig SJ, Meraz MA, White JM, et al. Disruption of the Jak1 gene demonstrates obligatory and nonredundant roles of the Jaks in cytokine-induced biologic responses. Cell. 1998;93(3):373–383. doi:10.1016/s0092-8674(00)81166-6
60. Buchert M, Burns CJ, Ernst M. Targeting JAK kinase in solid tumors: emerging opportunities and challenges. Oncogene. 2016;35(8):939–951. doi:10.1038/onc.2015.150
61. Xue W, Li W, Zhang T, et al. Anti-PD1 up-regulates PD-L1 expression and inhibits T-cell lymphoma progression: possible involvement of an IFN-γ-associated JAK-STAT pathway. Onco Targets Ther. 2019;12:2079–2088. doi:10.2147/ott.S187280
62. Yu D, Lu J. Breast cancer multistep development. In: Schwab M, editor. Encyclopedia of Cancer. Berlin Heidelberg: Springer; 2011:522–526.
63. Beckmann MW, Niederacher D, Schnürch H-G, Gusterson BA, Bender HG. Multistep carcinogenesis of breast cancer and tumour heterogeneity. J Mol Med. 1997;75(6):429–439. doi:10.1007/s001090050128
64. Brenner AJ, Aldaz CM. The genetics of sporadic breast cancer. Prog Clin Biol Res. 1997;396:63–82.
65. Ingvarsson S. Molecular genetics of breast cancer progression. Semin Cancer Biol. 1999;9(4):277–288. doi:10.1006/scbi.1999.0124
66. Wan YY, Flavell RA. How diverse–CD4 effector T cells and their functions. J Mol Cell Biol. 2009;1(1):20–36. doi:10.1093/jmcb/mjp001
67. Tesmer LA, Lundy SK, Sarkar S, Fox DA. Th17 cells in human disease. Immunol Rev. 2008;223:87–113. doi:10.1111/j.1600-065X.2008.00628.x
68. Romagnani S. Lymphokine production by human T cells in disease states. Annu Rev Immunol. 1994;12:227–257. doi:10.1146/annurev.iy.12.040194.001303
69. Zhu J, Yamane H, Paul WE. Differentiation of effector CD4 T cell populations (*). Annu Rev Immunol. 2010;28:445–489. doi:10.1146/annurev-immunol-030409-101212
70. Murphy KM, Ouyang W, Farrar JD, et al. Signaling and transcription in T helper development. Annu Rev Immunol. 2000;18:451–494. doi:10.1146/annurev.immunol.18.1.451
71. Luckheeram RV, Zhou R, Verma AD, Xia B. CD4⁺T cells: differentiation and functions. Clin Dev Immunol. 2012;2012:925135. doi:10.1155/2012/925135
72. Zhao X, Liu J, Ge S, et al. Saikosaponin A inhibits breast cancer by regulating th1/th2 balance. Front Pharmacol. 2019;10:624. doi:10.3389/fphar.2019.00624
73. Gonda K, Shibata M, Ohtake T, et al. Myeloid-derived suppressor cells are increased and correlated with type 2 immune responses, malnutrition, inflammation, and poor prognosis in patients with breast cancer. Oncol Lett. 2017;14(2):1766–1774. doi:10.3892/ol.2017.6305
74. Alinejad V, Dolati S, Motallebnezhad M, Yousefi M. The role of IL17B-IL17RB signaling pathway in breast cancer. Biomedicine Pharmacotherapy. 2017;88:795–803. doi:10.1016/j.biopha.2017.01.120
75. Wang J, Cai D, Ma B, Wu G, Wu J. Skewing the balance of regulatory T-cells and T-helper 17 cells in breast cancer patients. J Int Med Res. 2011;39(3):691–701. doi:10.1177/147323001103900301
76. Tabassum S, Abbasi R, Ahmad N, Farooqi AA. Targeting of JAK-STAT signaling in breast cancer: therapeutic strategies to overcome drug resistance. Adv Exp Med Biol. 2019;1152:271–281. doi:10.1007/978-3-030-20301-6_14
77. Villarino AV, Kanno Y, O’Shea JJ. Mechanisms and consequences of Jak-STAT signaling in the immune system. Nat Immunol. 2017;18(4):374–384. doi:10.1038/ni.3691
78. Banerjee S, Biehl A, Gadina M, Hasni S, Schwartz DM. JAK-STAT signaling as a target for inflammatory and autoimmune diseases: current and future prospects. Drugs. 2017;77(5):521–546. doi:10.1007/s40265-017-0701-9
79. Kiu H, Nicholson SE. Biology and significance of the JAK/STAT signalling pathways. Growth Factors. 2012;30(2):88–106. doi:10.3109/08977194.2012.660936
80. Stine RR, Matunis EL. JAK-STAT signaling in stem cells. Adv Exp Med Biol. 2013;786:247–267. doi:10.1007/978-94-007-6621-1_14
81. Carnero A, Blanco-Aparicio C, Renner O, Link W, Leal JF. The PTEN/PI3K/AKT signalling pathway in cancer, therapeutic implications. Curr Cancer Drug Targets. 2008;8(3):187–198. doi:10.2174/156800908784293659
82. Hers I, Vincent EE, Tavaré JM. Akt signalling in health and disease. Cell Signal. 2011;23(10):1515–1527. doi:10.1016/j.cellsig.2011.05.004
83. Reddy D, Kumavath R, Tan TZ, Ampasala DR, Kumar AP. Peruvoside targets apoptosis and autophagy through MAPK Wnt/β-catenin and PI3K/AKT/mTOR signaling pathways in human cancers. Life Sci. 2020;241:117147. doi:10.1016/j.lfs.2019.117147
84. Qazi AK, Hussain A, Hamid A, et al. Recent development in targeting PI3K-Akt-mTOR signaling for anticancer therapeutic strategies. Anticancer Agents Med Chem. 2013;13(10):1552–1564. doi:10.2174/1871520613666131125123241
85. Hosford SR, Miller TW. Clinical potential of novel therapeutic targets in breast cancer: CDK4/6, Src, JAK/ STAT, PARP, HDAC, and PI3K/AKT/mTOR pathways. Pharmgenomics Pers Med. 2014;7:203–215. doi:10.2147/pgpm.S52762
86. Gu-Trantien C, Loi S, Garaud S, et al. CD4⁺ follicular helper T cell infiltration predicts breast cancer survival. J Clin Invest. 2013;123(7):2873–2892. doi:10.1172/jci67428
87. Jagtap SV. Evaluation of CD4+ T-cells and CD8+ T-cells in triple-negative invasive breast cancer. Indian J Pathol Microbiol. 2018;61(4):477–478. doi:10.4103/ijpm.Ijpm_201_18
88. Stanton SE, Disis ML. Clinical significance of tumor-infiltrating lymphocytes in breast cancer. J Immunotherapy Cancer. 2016;4:59. doi:10.1186/s40425-016-0165-6
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.