Back to Journals » OncoTargets and Therapy » Volume 14
Emerging Mechanisms and Treatment Progress on Liver Metastasis of Colorectal Cancer
Authors Zheng W, Wu F, Fu K, Sun G, Sun G, Li X , Jiang W, Cao H , Wang H, Tang W
Received 10 January 2021
Accepted for publication 24 March 2021
Published 6 May 2021 Volume 2021:14 Pages 3013—3036
DOI https://doi.org/10.2147/OTT.S301371
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Jianmin Xu
Wubin Zheng,1,* Fan Wu,1,* Kai Fu,1,* Guangshun Sun,1 Guoqiang Sun,1 Xiao Li,1 Wei Jiang,1 Hongyong Cao,1 Hanjin Wang,1 Weiwei Tang2
1Department of General Surgery, Nanjing First Hospital, Nanjing Medical University, Nanjing, People’s Republic of China; 2Hepatobiliary/Liver Transplantation Center, The First Affiliated Hospital of Nanjing Medical University, Key Laboratory of Living Donor Transplantation, Chinese Academy of Medical Sciences, Nanjing, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Hongyong Cao; Hanjin Wang Email [email protected]; [email protected]
Abstract: Colorectal cancer is currently the third largest malignant tumor in the world, with high new cases and high mortality. Metastasis is one of the most common causes of death of colorectal cancer, of which liver metastasis is the most fatal. Since the beginning of the Human Genome Project in 2001, people have gradually recognized the 3 billion base pairs that make up the human genome, of which only about 1.5% of the nucleic acid sequences are used for protein coding, including proto-oncogenes and tumor suppressor genes. A large number of differences in the expression of proto-oncogenes and tumor suppressor genes have also been found in the study of colorectal cancer, which proves that they are also actively involved in the progression of colorectal cancer and promote the occurrence of liver metastasis. Except for 1.5% of the coding sequence, the rest of the nucleic acid sequence does not encode any protein, which is called non-coding RNA. With the deepening of research, genome sequences without protein coding potential that were originally considered “junk sequences” may have important biological functions. Many years of studies have found that a large number of abnormal expression of ncRNA in colorectal cancer liver metastasis, indicating that ncRNA plays an important role in it. To explore the role and mechanism of these coding sequences and non-coding RNA in liver metastasis of colorectal cancer is very important for the early diagnosis and treatment of liver metastasis of colorectal cancer. This article reviews the coding genes and ncRNA that have been found in the study of liver metastasis of colorectal cancer in recent years, as well as the mechanisms that have been identified or are still under study, as well as the clinical treatment of liver metastasis of colorectal cancer.
Keywords: colorectal cancer, liver metastasis, mechanisms, treatment
Introduction
Colorectal cancer has about 1.4 million new cases and 694,000 deaths worldwide every year, making it the third largest malignant tumor in the world.1,2 Metastasis is the most common cause of death from colorectal cancer, while liver metastasis is the most common.3–5 Some studies have shown that liver metastasis accounts for 75–83% of colorectal cancer metastasis.6,7 Therefore, the early diagnosis of liver metastasis of colorectal cancer is very important to reduce the mortality of patients. Due to the lack of sensitivity and specificity of serum markers such as CEA, in the diagnosis of colorectal cancer liver metastasis, the diagnosis of colorectal cancer liver metastasis still depends on imaging examination and focus biopsy.8–10 However, the sensitivity of imaging diagnosis of liver metastasis of colorectal cancer is still insufficient to achieve the effect of early diagnosis, so it is imperative to find a new biomarker that can better diagnose liver metastasis of colorectal cancer.
The mechanism of liver metastasis of colorectal cancer has not been fully elucidated. Current studies have shown that tumor cells spread from primary to distant organs and develop into metastatic foci, which is a continuous dynamic process with the participation of multiple genes and multiple links. It involves many links, such as tumor cell adhesion, invasion, extracellular matrix remodeling, neovascularization and lymphangiogenesis, body immunity and so on.11 In recent years, the relationship between abnormal expression of tumor genes and liver metastasis of colorectal cancer has become the focus of tumor molecular biology research, and made a breakthrough. Liver metastasis of colorectal cancer must have two prerequisites: first, colon cancer cells themselves have high metastatic characteristics; second, there must be target organs suitable for the growth of metastatic colon cancer cells. Thus, it can be seen that a large number of mutations of oncogenes or tumor suppressor genes make colon cancer cells easy to metastasize, and the liver is an organ suitable for the growth of metastatic colon cancer cells, both of which promote liver metastasis of colorectal cancer and lead to poor prognosis.
Over the past 10 years, with the development of next generation sequencing and bioinformatics, there has been a great variety. Thousands of Non-coding RNA has been discovered. In a sense, only the open reading frame (ORF) sequence on mRNA is the “coding” sequence, while other sequences on mRNA and a large number of non-mRNA transcripts that can not be used as translation templates are called non-coding RNA sequences.12–14 The non-coding RNA sequence on mRNA can regulate the translation, location and stability of mRNA.15–20 More and more evidence shows that ncRNA plays an important role in the occurrence, development and metastasis of colorectal cancer. It can be used not only as a tumor suppressor, but also as an oncogene to regulate the expression of its specific mRNA target.21–26
In this review, we summarize the research progress of coding genes and non-coding RNAs in colorectal liver metastasis, and briefly introduce the progress in clinical treatment in recent years. It is expected to contribute to the early detection and early treatment of liver metastasis of colorectal cancer, and to prolong the survival time and quality of life of patients.
Brief Overview of Coding Genes
Malignant tumor causes serious harm to human health, and its occurrence is the result of the joint action of multi-factors, multi-stages and multi-genes, which is closely related to the disturbance of cell cycle and apoptosis regulation. In the occurrence and development of tumor, when various environmental factors and genetic pathogenic factors cause DNA damage, thus activating proto-oncogenes and inactivating tumor suppressor genes, resulting in disturbance of cell cycle regulation and imbalance of apoptosis regulation, resulting in abnormal cell growth, is the most fundamental cause of tumorigenesis. We sorted out the coding genes and their products that have been found in the research of colorectal cancer liver metastasis (CRLM) in recent years, and briefly worked out the mechanism in Table 1.
 |  |  |  |
Table 1 Coding Genes in Liver Metastasis of Colorectal Cancer |
The Role of Tumor Suppressor Genes in Liver Metastasis of Colorectal Cancer
Tumor suppressor genes are a kind of genes that exist in normal cells which can inhibit cell growth and have potential anti-tumor effect. Tumor suppressor genes play a very important negative regulatory role in controlling cell growth, proliferation and differentiation. They restrict each other with proto-oncogenes and maintain the relative stability of positive and negative regulatory signals. When these genes are mutated, deleted or inactivated, they can cause malignant transformation of cells and lead to tumorigenesis. So far, scientists have isolated and identified about 100 kinds of tumor suppressor genes from cells, the most common of which are Rb, p53, APC, nm23 and so on. According to whether there is a mutation, it can be divided into two types: mutant type and wild type, and according to different acting organs, it can be divided into broad spectrum and specificity. Different tumor suppressor genes have different functions. Some of them can participate in cell signal transduction and epigenetic regulation, some can negatively regulate cell cycle, some can lead to DNA mismatch repair, some can regulate development and stem cell proliferation, and some can be used as negative transcriptional factors. These functions can participate in the occurrence and development of tumor. In CRLM, some tumor suppressor genes are also involved and play an irreplaceable role. The abnormally expressed tumor suppressor genes found in CRLM in recent years were sorted out in Table 1.
PTEN
PTEN gene (phosphatase and tensin homologue deleted on chromosome 10), also known as advanced tumor mutant gene, is the first tumor suppressor gene with dual specific phosphatase activity cloned from primary breast cancer, prostate cancer and glioblastoma cell lines in 1997.27 The anti-tumor effect of PTEN is mainly through MAPK pathway, PI3K/Akt pathway, FAK pathway and cell cycle pathway to participate in normal cell development and inhibit angiogenesis. The subcellular localization of PTEN is very important for its normal cellular function and its role as a tumor suppressor. In colorectal cancer, the mutation rate of PTEN gene is 6%, indicating that PTEN also has a high mutation rate in CRLM. In the cytoplasm, PTEN binds with its cytoplasmic targets to regulate cell proliferation, apoptosis, cell cycle progression, cell adhesion, migration and invasion. In the nucleus, PTEN plays a role in maintaining chromosome stability and DNA double-strand break repair, thus maintaining the integrity of the genome.28 The mislocalization of PTEN between nucleus and cytoplasm may lead to malignant growth.29 Therefore, the subcellular localization of PTEN is closely regulated and a variety of regulatory mechanisms have been found.
PI3K/Akt signaling pathway is an important intracellular signal transduction pathway, which plays an important biological role in cell apoptosis, survival, proliferation and cytoskeleton changes. Its regulation disorder is closely related to the occurrence and development of tumor. Receptor tyrosine kinase (RTK) is the main upstream regulator of PI3K/Akt signal. PTEN is the main antagonist of PI3K and a negative regulator of this pathway. Inactivation of PTEN will lead to excessive activation of RTK/PI3K/Akt signal in many tumor types, thus driving tumorigenesis.30
Related studies found that in CRLM, the mutation rate of PTEN gene increased significantly, and suggested that the prognosis is obviously poor, PTEN gene may also be involved in liver metastasis through the above mechanism, but the related research has not yet been carried out, we can know the next research direction.
WTp53
P53 gene is a tumor suppressor gene, located on the short arm of chromosome 17p. 20 kb, encodes a tumor suppressor protein Tp53, with a molecular weight of 53 kD, which is involved in the control of a variety of cell events, such as cell cycle regulation, apoptosis, cell senescence, DNA repair, cell differentiation, gene amplification, DNA recombination, chromosome segregation and angiogenesis. P53 gene can be divided into wild type (WTp53) and mutant type (MTp53). The anti-tumor effect of WTp53 mainly prevents cells from entering S phase in G1/G0 phase, which inhibits cell proliferation and induces cell apoptosis.31 When the p53 gene is mutated, the stable protein can not inhibit the cell proliferation, promote the malignant transformation of the cell, enhance the invasiveness, and make the p53 gene change from tumor suppressor gene to proto-oncogene.
The occurrence of liver metastasis of colorectal cancer is mainly caused by reflux of portal vein and hepatic artery, in which the formation of tumor neovascularization is very important.32 Du T found that WTp53 has a synergistic effect with TK/GCV and CD/5-FC to inhibit tumor neovascularization and prevent liver metastasis.33
Claudin
In humans and animals, there is a tight junction between adjacent epithelial or endothelial cells. Tight junction is one of the main forms of cell adhesion. It is a macromolecular complex composed of cytoplasmic proteins and transmembrane proteins. It is the structural and functional basis of paracellular material transport pathways. It mainly has barrier and screening functions, which can maintain the stability of the internal environment of the body and maintain the polarity of epithelial cells. It plays an important role in the normal physiological process of cells. Claudins is a cytoskeleton protein of tight junction between cells and an important component of tight junction molecules. Its abnormal expression can lead to structural damage and functional damage of epithelial cells and endothelial cells, and then lead to tumor invasion and metastasis.
In 1998, in the study of transmembrane protein, Furuse found and identified a complete tight junction fragment in chicken liver for the first time, its relative molecular weight was 22,000, and named it Claudin.34 So far, 27 members of the Claudins protein family have been found in humans, with a sequence identity of 12.5%-69.7%, most of the relative molecular weight is 20,000–27,000. So far, 21 species have been identified to exist in the human body.35
Each molecular structure consists of four transmembrane domains and two extracellular annular domains. In the extracellular annular regions, the larger primary macrocycle is composed of 52 amino acid residues, while the secondary ring is composed of 16,033 residues. The same is that the terminal structures of amino and carboxyl groups are present in the cytoplasm, and there is a specific PDZ binding region on it. Among different Claudins protein molecules, the C-terminal sequence has different lengths of 210.63 residues, and the amino acid sequences of the first and fourth transmembrane fragments and the outer ring are highly conserved. These two annular structures of Claudins protein can contact the outside world through homologous or heterologous ways: (1) the longer extracellular ring mainly forms two different kinds of intercellular pores of paracellular open or closed, and these pores are the important structural basis for the formation of tight junctions, and pores play an important role in the formation of tight junction bands and ion-selective permeability. (1) the longer extracellular rings mainly form two different kinds of intercellular pores, which are open or closed, and these pores play an important role in the formation of tight junction bands and ion selective permeability. (2) the shorter extracellular loop is mainly responsible for the trans-binding between Claudins molecules and does not participate in the regulation of paracellular permeability, and the shorter extracellular loop is the binding site of enterotoxin from Cystitis perfringens, while the intracellular C-terminal is connected to tight junction protein (zonula occludens, ZO) molecules through different PDZ domains.36
The expression of Claudins protein is specific, and more than two kinds of Claudins protein molecules can be expressed in most tissues and organs. They are expressed not only in the tight junction but also on the side of some epithelial cells and on the basement membrane. It has been found that there are abnormal expressions of Claudin1 and Claudin4 in CRLM. Claudin1 can promote the activation of matrix metalloproteinase-2 precursor and enhance the ability of invasion and metastasis of cancer cells. At the same time, the increased expression of Claudin1 can inhibit the expression of E-cadherin, Claudin1 may be the target gene of β-catenin/Tcf (T cell factor) signal pathway, the up-regulation of β-catenin/Tcf signal pathway can promote the occurrence and development of tumor, while the decreased expression of E-cadherin can promote the up-regulation of β-catenin/Tcf signal pathway, so the overexpression of Claudin1 can promote tumorigenesis.37 Claudin4 was previously low or non-expressed in colon cancer cells and highly expressed in cholangiocarcinoma. Recently, it has been found that Claudin4 is highly expressed in CRLM, but the specific mechanism remains to be studied.
The Role of Oncogenes in Liver Metastasis of Colorectal Cancer
Proto-oncogenes exist in the genomes of normal biological cells. Like tumor suppressor genes, proto-oncogenes play an important role in the regulation of cell growth and proliferation. Under normal circumstances, the ability is that the proto-oncogenes in the genome are in a state of low expression or non-expression, but when they encounter certain conditions, such as viral infection, chemical carcinogens or radiation, the proto-oncogenes will be abnormally activated and converted to oncogenes to induce cell carcinogenesis. In Table 2, we sorted out the abnormally expressed oncogenes found in the study of CRLM in recent years.
 |  |  |
Table 2 Non-Coding RNAs in Liver Metastasis of Colorectal Cancer |
MACC1
Metastasis-associated in colon cancer (MACC1) is closely related to liver metastasis of colorectal cancer.38 It was discovered and named by Stein using differential display RT-PCR technology in 2009. There is a small amount of MACC1 expression in normal tissues and abnormally high expression in many kinds of malignant tumors, but the expression level is the highest in colon cancer cells. Studies have also shown that the increase of MACC1 expression is often accompanied by the occurrence of malignant tumors, and the 5-year survival rate of patients with high expression of MACC1 mRNA is significantly lower, and the risk of death is increased.39 MACC1 has a strong correlation with the clinicopathological features, development and prognosis of the tumor, and can be used as an independent evaluation index of metastasis and prognosis.
The MACC1 gene is located on human chromosome 7 (7p21. 1), there are about 76,762 base pairs, including 7 exons and 6 introns. Its cDNA has a sequence of 2559 nucleotides and encodes a protein of 852 amino acids. MACC1 protein is a protein complex composed of multiple domains, including ZU5, SH3 and two C-terminal death domains (death domain, DD). It has been found that transfection of MACC1 containing SH3 domain and proline-rich sequence into colon cancer cell lines that do not express MACC1 can enhance cell viability and proliferation, while MACC1 mutants without this domain lack these abilities, indicating that SH3 domain and SH3 binding motif are essential to maintain the biological function of MACC1.
HGF is a multiple growth factor, which is a growth stimulating factor secreted by hepatocytes and endothelial cells.40 MET protein is encoded by proto-oncogene c-met and is the membrane-specific receptor of HGF.41 HGF can bind to specific MET receptors through autocrine, paracrine and classical axonal reverse transport. The combination of the two can activate HGF/MET signal pathway, cause cell proliferation, mucosal epithelial transformation, promote angiogenesis, enhance cell movement, enhance tumor cell invasion and metastasis and so on.42 MACC1 is a key gene regulating HGF/MET pathway. It has been found that the expression of MACC1 mRNA and MET RNA is high or low in gastrointestinal tumors. As a transcription factor, MACC1 can bind to MET promoter and enhance the expression of MET.43 MACC1 siRNA can inhibit cell proliferation and movement, reduce metastasis in vivo, and greatly reduce the expression of MET RNA and its protein.
It has been found that the abnormal expression of MACC1 gene plays an important role in CRLM. Stein U experiments have confirmed that MACC1 has the ability to induce cell migration, infiltration and proliferation. They found similar results in vitro and in vivo animal experiments on SW480 cell line from primary focus of colon cancer and liver metastatic cell line SW620 from the same specimen. All these fully show that MACC1 has important research significance in colorectal liver metastasis. It can not only be used as a marker for early diagnosis but also guide clinical treatment and even drug research and development.
SNCG
γ-synaptophysin (SNCG) is a member of the neuronuclear protein family, which is located on chromosome 10q23. 2-q23.3. The transcriptional product mRNA is about 100bp, long and encodes a protein of less than 123 amino acid.44 Ji isolated human breast cancer cDNA library by differential cDNA sequence in 1997.45 Because of its high expression in advanced breast cancer, it was once called breast cancer-specific gene (BCSGI). However, recent studies have shown that SNCG is highly expressed in many organ tumors, especially in advanced malignant tumors, including breast cancer, ovarian cancer, pancreatic cancer, colorectal cancer and so on. It is almost undetectable or low expression in adjacent non-tumor tissues and is related to tumor invasion and metastasis.
Synaptophysin is a small molecular soluble protein family mainly expressed in nerve tissue, which is distributed among different species and genera, and contains 127–140 amino acid homologous sequences that are highly conserved in the process of evolution. SNCG is located on chromosome 10q23 and is about 5kb in length. It consists of five exons and is transcribed into a mRNA, polypeptide encoding 127AA of 1kb. The results of the SNCG sequence obtained from BAC clone 174P13 and the reported expression sequence tag (expressed sequence tags, ESTs) showed that there were four single nucleotide polymorphism (SNP) sites in the SNCG gene: the upstream-19 site of AUG, the 3 ‘untranslated region of AUG, the 122bp behind the stop codon UAG, and the third base of the 65th codon. In particular, the change of GTG glutamate GAG in exon 4 leads to the change of amino acid: valine (Val) 110 glutamate (Glu).
A great deal of evidence shows that the abnormal expression of SNCG protein in tumor cells is not due to gene mutation and amplification, but to the activation of transcription.46,47 There are four main factors that affect the transcription and expression of SNCG: (1) Activator protein 1 (AP1) can activate the AP1 site in the AP1,TPA binding intron through AP1 activators such as TPA to up-regulate the activity of the promoter and increase the expression of SNCG mRNA.48–50 (2) BubR1 is the main component of the cell division checkpoint, which can monitor the correct assembly of mitotic spindles, so as to ensure the correct separation of homologous chromosomes and prevent cells from escaping the checkpoint to make homologous chromosomes nonsynaptic. SNCG can inhibit the effect of BubR1 by binding to BubR1-microtubulin E (CENP-E) complex, which makes cells escape the control of mitotic checkpoints and form aneuploidy of chromosomes, thus increasing genetic instability and inducing tumorigenesis.51 (3) CpG island demethylation of SNCG can activate SNCG expression, which is to some extent consistent with the activation and expression mechanism of proto-oncogenes and growth-related factors in tissue cells.52 (4) SNCG can promote cell proliferation and inhibit the control of mitotic checkpoints by activating estrogen receptors, leading to tumorigenesis and progression.53
The latest study has found that there is a significant high expression of SNCG in CRLM.54 The 5-year survival rate and prognosis of patients with colorectal liver metastasis with SNCG positive are significantly lower than those with SNCG negative, indicating that the expression of SNCG can be used as an independent prognostic factor for colorectal cancer liver metastasis and can be used as an index for early diagnosis or to guide the next step of treatment.
Chemokine and Its Receptor
Chemokines are the largest known superfamily of cytokines, and more than 50 kinds of chemokine and more than 20 kinds of chemokine receptors have been found so far.55 By binding to receptors, chemokines participate in neovascularization,56 further affect tumor formation and metastasis,57 and play an important role in liver metastasis of colorectal cancer.
Chemokine is a small molecular cytokine with molecular weight of 8000–12,000, which has chemotactic effect on many kinds of cells, such as neutrophils, lymphocytes, monocytes and so on. According to the position of the conserved sequence of terminal hemisteminic acid, they can be divided into four categories: CXC, CC, and CX3C. Chemokines play a role by binding to the corresponding G-protein-coupled receptor superfamily. Chemokines affect tumor in many ways: some chemokines can inhibit tumor growth and development by activating immunoreactive cells or inhibiting tumor-related angiogenesis; other chemokines promote tumor development by enhancing tumor cell proliferation and migration, proteolytic enzyme secretion and inducing tumor angiogenesis, which show that chemokine network involved in tumor formation and metastasis is complex.58
There are four steps for cells to overcome vascular endothelial cell barrier between body fluid and tissue: cells flow with body fluid, cells are firmly attached to vascular endothelium, cells pass through endothelial intercellular space, and cells migrate to specific tissues. In this process, chemokines control the selection of exudative cells and the stable adhesion of the selected cells. The migration, invasion and metastasis of tumor cells are affected by chemokines and their receptors. It has been confirmed that chemokines regulate the metastasis of tumor cells and determine the target tissue of tumor metastasis. The interaction between chemokines specifically expressed in the target organs of tumor metastasis and the corresponding receptors expressed by the tumor can cause the polymerization of intracellular actin and regulate the movement and migration of cells. Therefore, the specific metastasis of tumor cells is closely related to the chemokine receptors expressed by tumor cells and the chemokines expressed by target organs.
There are also many chemokines involved in CRLM. There are also many chemokines involved in CRLM. CXCR4 is a chemokine receptor that plays an important role in CRLM and has a significantly high expression in CRLM, indicating that mediating tumor metastasis can become an important marker for the diagnosis of liver metastasis. The basic steps of the effect of CXCR4 on cancer cell metastasis may be: (1) cloning and expansion of cancer cells, infiltration of local tissue, angiogenesis, and expression of CXCR4; on the surface of tumor cells. (2) the cancer cells fall off from the primary tumor, pass through the lymphatic and vascular walls in the tumor, and enter the systemic circulation. (3) CXCR4 can make cancer cells invade normal tissue through CXCR4/TGF- β 1 axis, cell expansion and angiogenesis, which leads to liver metastasis.59 CXCL8 and its receptor CXCR1, CXCR2 can induce epithelial-mesenchymal transformation (EMT), of colorectal cancer cells to help them escape from host immune surveillance and enhance their resistance to anestrous apoptosis, thus promoting the formation of circulating tumor cells (CTCs) and their colonization in distant organs.60
With the further research, it has been found that chemokines and their receptors play a role in promoting distant metastasis in CRLM, but the mechanisms of a small number of chemokines have been declassified, but the mechanisms of most of the chemokines that have been found in CRLM cells are still unclear, which need to be further explored and expected to be transformed into early diagnostic indicators or drugs.
Brief Overview of Non-Coding RNAs
Over the past 10 years, with the development of next generation sequencing and bioinformatics, there has been a great variety. Thousands of Non-coding RNA has been discovered.19,20 MiRNA61–63 and piRNA64,65 were first discovered as a kind of small RNA. Then it was discovered that lncRNA. With the increase of species evolution, the number of small RNAs (especially miRNA) and lncRNA gradually increased. In recent years, it has been found that there are a large number of non-coding RNA, namely circRNA, in eukaryotic multicellular organisms.66–68 With the in-depth study of ncRNA, it has been found that ncRNA can participate in the occurrence and development of tumors through a variety of mechanisms. We summarized the role of ncRNA in liver metastasis of colorectal cancer. In Table 2, we summarize the progress in the field of non-coding RNAs research in recent years.
The Role of miRNAs in Liver Metastasis of Colorectal Cancer
MiRNA is a class of highly conserved, 18–25 nucleotides long endogenous single-stranded non-coding small molecule RNA.69 MiRNA exists widely in eukaryotes, and it does not have an open reading frame and does not encode proteins, but miRNA has the characteristics of clustering and can be co-expressed with related genes.70 In tumors, miRNA usually plays the role of tumor genes or tumor suppressor genes in the occurrence and development of tumors.71 Onco-miRNA such as miR-21 and miR-10b have oncogene-like functions and are positively correlated with tumorigenesis. Their overexpression or continuous activation will directly lead to tumorigenesis and development.72,73 Suppressor-miRNA such as miR-15/16 and Let-7 have the function similar to tumor suppressor gene. It has a negative correlation with tumorigenesis, and its down-regulation or inactivation will directly lead to tumorigenesis and development.74,75
MiRNA participates in tumorigenesis, development and metastasis in many ways. Drosha and Dicer are members of the RNase III family, mediating the splicing of the double-stranded complex from primiRNA to pre-miRNA and from pre-miRNA to miRNA, respectively. Many studies have shown that the expression and activity of Drosha and Dicer participate in the occurrence and development of tumor by affecting the synthesis of mature miRNA.76,77 Some scholars have proposed the “ceRNA hypothesis”, that is, ceRNA (competing endogenous RNA) can compete with the MRE elements of the 3ʹ-UTR region of tumor genes, tumor suppressor genes and tumor-related signaling pathways to combine with miRNA, antagonize or regulate the function of tumor suppressor or tumorigenic miRNA, so it plays an important role in tumorigenesis, development, invasion, metastasis and drug efficacy.78
The Main Mechanism of miRNAs Regulation in CRLM
In the cells of colorectal cancer liver metastasis (CRLM), researchers found a variety of abnormal expression of miRNA, indicating that miRNA is closely related to colorectal liver metastasis. Among them, epithelial–mesenchymal transition (EMT) is the main mechanism, and most miRNA can directly or indirectly affect EMT to promote the occurrence of liver metastasis. EMT specifically refers to the transformation of cell phenotypes between epithelium and mesenchyme, which was first discovered in the process of embryonic development. In recent years, researchers have gradually paid more and more attention to its role in tumorigenesis and other diseases.79,80 EMT makes epithelial cells remove differentiated phenotypes, including cell-cell adhesion, top-bottom polarity, low motor ability, and obtain the phenotypes of some mesenchymal cells, including migration, invasion, anti-apoptosis and so on.81 MiRNAs can regulate EMT through a variety of gene loci in the process of liver metastasis of colorectal cancer. The miRNA-200 family is the hub for the regulation of EMT found by the latest research, which mainly includes miR-200a, miR-200b, miR-200c, miR-200s, miR-141, miR-429 and so on. The study of Shelton Pm found that the expression of miR-200s was significantly increased in CRLM. It promotes metastasis and regulates EMT through the PKC ζ/ADAR2 axis.82 KRAS is an important targeted tumor suppressor gene. In tumors, the miRNAs, cascade of some targeted oncogenes KRAS activates KRAS, activated RAS response original binding protein (RREB1), which directly inhibits the transcription of miRNA clusters including miR-384, miR-487b and down-regulates the expression of KRAS in subcutaneous or xenotransplanted tumors by transcriptional up-regulation of these miRNA or liposome delivery systems.83 At the same time, a variety of miRNAs can pass through various signals such as mTOR, PTEN, PKCζ/ADAR2, etc, affect the proliferation of blood vessels around the tumor, reduce tumor cell adhesion, and promote the progress of CRLM.
Thus, many miRNAs have been found in CRLM, and the related mechanisms of their regulation of CRLM are gradually becoming clear. It is believed that miRNA will play an important role in the diagnosis and treatment of CRLM.
The Role of lncRNAs in Liver Metastasis of Colorectal Cancer
In 2002, Japanese researchers first put forward the concept of lncRNA, which refers to non-coding RNA with a length of more than 200 nt.84 Until 2007, a highly expressed functional lncRNA gene of 2.2 kb was found for the first time in breast cancer, proving the role of lncRNA in tumorigenesis and development.85
LncRNA has a 5ʹ- cap structure and a 3ʹ- terminal polyadenylate tail, and its gene structure is similar to that of mRNA.86 LncRNA can act as the sponge of microRNA and adsorb some specific microRNA, in the way of bait so as to regulate the expression of some microRNA target genes. The lncRNA with this function is also known as ceRNA (Competing endogenous RNA).87 At the same time, because the expression of most long non-coding genes is transcribed by RNA polymerase II (RNAP II), the specific expression of these genes in tissues and developmental stages indicates that the transcription of these genes is strictly regulated, so people can infer the state of chromatin according to the expression of lncRNA. In addition, lncRNA can also regulate tumor suppression or oncogene expression. Many studies have shown that lncRNA has a variety of other effect mechanisms, it can be used as signal molecules, bait molecules and guiding, functional scaffolds, but also can interact with other biomolecules to form a certain module structure to exert its biological function.88,89 It can be seen that LncRNA is closely related to the occurrence and development of tumors.90
The Main Mechanism of lncRNAs Regulation in CRLM
ZEB2 can directly down-regulate the expression of cell junction-related genes, including adhesive junctions, tight junctions, desmosomes and gap junctions. ZWB2 can increase the expression of interstitial proteins (such as vimentin and N-cadherin), which is common in the cell cord of hepatocellular carcinoma.91 The study of Chen Dl found that the expression of lncRNA-UICLM in CRLM was significantly increased, while the expression of tumor suppressor gene ZEB2 was significantly decreased. Further study of the mechanism found that lncRNA-UICLM is a competitive endogenous RNA, of miR-215, that is, lncRNA-UICLM can be used as a “sponge” of miR-215, that is, it absorbs miR-215, to inhibit the expression of ZEB2 and accelerate the occurrence of liver metastasis of colorectal cancer.92
PDI2 (Peptidyl arginine deiminase 2) was first confirmed in animal experiments to have a high expression in HER2/ERBB2 (+) breast cancer, and when PDI2 is inhibited, the tumor cells will be significantly suppressed. Several recent studies have reported that PDI2 is involved in tumorigenesis and plays an important role in the development of many malignant tumors such as spontaneous skin tumors, multiple myeloma, breast cancer and colorectal cancer.93–95 MiR-125a-5p is a classic tumor suppressor gene. At present, many studies at home and abroad have found that it can effectively inhibit the invasion and metastasis of common tumors, including hepatocellular carcinoma, gastric cancer, breast cancer, lung cancer, glioma, melanoma and so on. The regulatory targets of its inhibitory effect in different tumors are SIRT7, PI3K, E2F3, ERBB2, TSTA3, HDAC4, HDAC5, EGFR, Gab2, Lin28B and other genes.96–100 Chen et al found that lncRNA HOXA11-AS can act as a “sponge” of miR-125a-5p, promoting the expression of PADI2, which leads to liver metastasis of colorectal cancer.101
LncRNA can affect the occurrence and development of CRLM through a variety of mechanisms, but it is still necessary to further explore the different mechanisms and pathways of different lncRNA, which is very important for the early diagnosis and even treatment of CRLM in the future.
The Role of circRNAs in Liver Metastasis of Colorectal Cancer
The role and mechanism of circRNA in tumor is one of the recent research hotspots. In 2013, Jeck detected more than 25,000 circRNAs in human fibroblasts.68 They realized that circRNA is widespread in the human body and may play an important role in the spectrum of human diseases. CircRNA is a special type of endogenous non-coding RNA whose 3ʹend and 5ʹend are connected to form a closed covalent ring structure. This structure makes circRNA more stable and highly conserved. Since circRNA has one or more miRNA binding sites, it can reduce the activity of miRNA by binding to miRNA, weaken the inhibitory effect of miRNA on target genes, thereby regulating the expression of related target genes, and ultimately leading to increased production of messenger RNA. The biological effect is strengthened, this is the competitive-binding mechanism, and this effect is called “miRNA sponge”. At the same time, circRNA can combine with specific proteins to form RNA-protein complexes, thereby affecting protein translation and regulating protein functions.
The Main Mechanism of circRNAs Regulation in CRLM
In CRLM, most of the circRNA found at present acts as the “sponge” of miRNA to play its role. GRF-H1 is a kind of regulatory protein that regulates the activity of Rho GTP by regulating its GDP/GTP exchange. As an important regulator, it plays an important role in the biological process of mammals, especially in the proliferation and migration of tumor cells.102 The study of Yang H found that circ-133 can up-regulate the expression of RhoA by competitively binding to miR-133a in colorectal cancer cells; the overexpression of RhoA can reverse the inhibitory effect of circ-133 on the progression of colorectal cancer, indicating that circ-133 can promote the progression of colorectal cancer through the miR-133a/GEF-H1/RhoA axis.103
The HMGA2 gene is located in 12q13-15. There are 5 exons in 200 Kb, and each exon is independently encoded by HMGA2 gene.104 In normal human tissues, HMGA2 encodes a complete protein product with a relative molecular weight of 12,000. The protein is mainly located in the nucleus and belongs to non-histone chromosome protein.105 Widely expressed in eukaryotes, it does not have the ability of direct transcriptional regulation, but it can bind to the AT enrichment region of the regulated gene DNA through the unique AT hook structure, causing DNA to bend, stretch, ring or unchain, thus changing the structure of chromatin and enhancing its transcriptional activity, also known as structural transcription factors.106 Transcripts of HMGA2 gene can hardly be detected in normal adult tissues. In recent years, studies have shown that HMGA2 is abnormally expressed in a variety of cancer tissues, and participates in the regulation of a variety of biological behaviors, and plays a role in promoting the occurrence and development of cancer. The degree of expression is closely related to the clinicopathological features and the survival and prognosis of patients.107 Chen RX’s study found that CircNSUN2 and HMGA2 were highly expressed in colorectal liver metastatic cancer cells. Further studies showed that N6-methyladenosine modification of CircNSUN2 increased cytoplasmic output, thus through the formation of CircNSUN2/IGF2BP2/HMGA2RNA–protein ternary complex in the cytoplasm to enhance the stability of HMGA2 mRNA and promote colorectal cancer metastasis.108
According to the current research, there are still few studies on CRLM in circRNA, but it has been found that circRNA can affect the liver metastasis of colorectal cancer through different mechanisms. For example, as a sponge of miRNA, or regulating the expression of coding genes, it is proved that circRNA plays an important role in the occurrence of CRLM, which points out the direction for further research.
Clinical Treatment of Liver Metastasis of Colorectal Cancer
The liver is one of the most common organs with distant metastasis of colorectal cancer. Correct and effective treatment of liver metastasis can not only delay the progress of the disease and prolong the life span of patients but also provide a good guarantee for the quality of prognosis of patients. At present, surgical resection is still the first choice for liver metastasis of colorectal cancer. With the continuous development of related technologies and the in-depth study of liver metastasis of colorectal cancer, new treatment methods such as radiotherapy, cryotherapy and radiofrequency ablation are also being put into practice. Multidisciplinary comprehensive treatment will provide a new research direction for the clinical treatment of liver metastasis of colorectal cancer.
Surgical Resection
Surgical resection is the first choice for the treatment of liver metastasis of colorectal cancer at present. Standardized surgical treatment is generally advocated for CRC patients with surgical indications.109 The progress of medicine is changing with each passing day. With the improvement of surgery and multidisciplinary comprehensive treatment, the concept of resectability of liver metastatic cancer is constantly updated. The latest concept of surgical resection is: (1) all metastatic lesions of the liver can be removed; (2) the residual liver volume after resection is more than 30% of the original volume; and (3) extrahepatic metastatic lesions can be removed. At present, experts agree that as long as the residual liver can be compensated, the scope of resection should be expanded as far as possible, so that 20% of patients have the opportunity to operate.110
If the primary focus of colorectal cancer is found to have liver metastasis at the same time, or if liver metastasis occurs within 6 months after radical operation, it is called synchronous liver metastasis; if the primary focus is cured for more than 6 months, the recurrent liver metastasis is called metachronous liver metastasis. For patients with simultaneous liver metastasis, surgeons still have difficulties in the choice of simultaneous resection or staging resection. Simultaneous resection avoids the pain of the second operation and correspondingly reduces the incidence of postoperative complications. The survival benefit of staging resection depends on the biological behavior of the tumor, but the operative complications and mortality are higher, but it can make the occult liver metastasis more obvious and reduce the postoperative recurrence rate of simultaneous resection.109
To sum up, surgical resection is still the first choice at present, but the formulation of treatment plan needs to be based on the actual situation of patients, according to multi-disciplinary comprehensive evaluation, and have a real-time understanding of the status of liver metastasis of patients.
Radiofrequency Ablation Therapy
Radiofrequency ablation is to make the electrode needle directly contact the metastatic tumor tissue under the guidance of ultrasound, CT and laparoscopy, connect the radio frequency current to the electrode needle, and produce heat through the ion shock produced by the current around the electrode needle, so as to achieve the therapeutic purpose of promoting coagulative necrosis of tumor tissue around the electrode needle.111 In general, the heat production temperature of the target area of radiofrequency ablation electrode needle can be as high as 100 °C, which can cause irreversible thermal coagulation and necrosis of local tissue proteins, apoptosis and microcirculatory vascular embolism in surrounding tissues, inhibit the formation of blood vessels around the tumor, and prevent the blood dissemination of tumor cells, which has a definite effect on the treatment of liver metastatic lesions of colorectal cancer.112,113
The size of liver metastases, the number of liver metastases and the presence or absence of extrahepatic metastasis in patients with colorectal cancer are significantly related to the prognosis, efficacy and survival rate. Studies have shown that patients with liver metastases less than 3 cm in diameter, less than three and without extrahepatic metastasis can be treated with radiofrequency ablation with good accuracy, high repeatability, greater benefit and good prognosis. Most of the patients with liver metastasis of colorectal cancer who can not be operated are due to the lack of liver reserve and complications. In view of this situation, radiofrequency ablation can be used.
Chemotherapy
As a patient with unresectable colon cancer with liver metastasis, systemic chemotherapy can effectively prolong the survival time. According to the relevant information, the use of oxaliplatin combined with calcium folinate and fluorouracil has achieved considerable results. Using this kind of chemotherapy and continuing to maintain chemotherapy can increase the mPFS of patients with liver metastasis of colorectal cancer (median progression-free survival). However, this kind of method has severe adverse reactions to its own body, which seriously damages the life and health of patients. However, due to the frequency of chemotherapy, patients with liver metastasis of colorectal cancer are not as tolerant as they used to be.
Targeted Therapy
The application of molecular targeted therapy strategy to patients with liver metastasis of colorectal cancer also has a certain effect on the improvement of survival time of patients. At present, the selected molecular targeting drugs are mostly used for epidermal growth factor receptor or vascular endothelial growth factor receptor, and the common drugs are panizumab, bevacizumab, cetuximab and so on. Among them, panizumab and cetuximab are mostly used as epidermal growth factor receptor inhibitors, but after more in-depth studies, it is found that the applicable range of the population is gradually shrinking. In addition, cetuximab can be used as a first-line regimen in patients with liver metastasis of colorectal cancer. The obvious effect is the improvement of PFS and OS as well as the efficiency, but it is worth mentioning that if the opposite is a patient with BRAF gene mutation, the effect is very poor. In addition, when bevacizumab and cetuximab were used in patients with liver metastasis of colorectal cancer, the objective effective rate of bevacizumab and cetuximab was not significant. Therefore, in the selection of the above molecular targeted drugs, it is effective for patients with liver metastasis of colorectal cancer, but for patients to have a clear understanding of KRIS, BRAF gene status.
In the selection of molecular targeting drugs mentioned above, it is effective for patients with liver metastasis of colorectal cancer, but it is necessary to have a clear understanding of KRIS and BRAF gene status in patients. As for vascular endothelial growth factor inhibitor, it has not been found that gene status can affect receptor inhibitors. As a result, bevacizumab, which is led by bevacizumab, has gradually become a more respected treatment.
Immunotherapy
Tumor microenvironment (TME) is a local growth environment composed of tumor cells, tumor stem cells and surrounding infiltrating mesenchymal cells, immune cells, extracellular matrix, interstitial tissue near tumor cells, microvessels, and cytokines.114 It can be divided into a microenvironment dominated by immune cells and a non-immune microenvironment dominated by fibroblasts.115 When the tumor cells break through the immune defense line, the immune cells play an anti-tumor effect at the beginning of tumor invasion, but they also show the role of assisting tumor immune escape and distant metastasis during tumor progression. Immune cell infiltration can secrete inflammatory mediators, creating an inflammatory environment with a high degree of specificity.116 A variety of immune cells can mediate immune suppression and promote tumor progression in the tumor microenvironment.
With the continuous study of tumor immune mechanisms, immunotherapy has made major breakthroughs in the treatment of malignant tumors. Immunotherapy drugs have achieved many results in clinical trials, and some immunotherapy drugs have been approved for marketing. For CRLM patients, a variety of immunotherapy options can significantly prolong survival, such as tumor vaccines, immunosuppressive agents, and small molecule therapy.
Tumor vaccine therapy uses tumor antigen genes and antigen peptide vaccines to improve the activity of the immune system, and induces the body to produce humoral immunity and cellular immunity through tumor antigen substances and tumor cells to enhance the ability of anti-cancer. Autologous transplantation of dendritic cells (DC) is a classic method of tumor vaccination. DCs can be modified to present tumor antigens and stimulate the adaptive immune system.117 DC is a powerful TAA presenting cell with MHC (major histocompatibility complex)-I and MHC-II, which activates T cell immunity, has good safety for the treatment of advanced malignant tumors, and can prolong the survival period of CRLM patients.
Cellular immunotherapy refers to the separation of tumor-infiltrating lymphocytes, natural killer cells and other lymphocytes from tumor tissues and blood, which are transformed, activated, expanded and then re-infused into the body to achieve the killing effect on tumor cells. Cellular immunotherapy includes cytotoxic T cell therapy (CTL), genetically modified T cell receptor therapy (TCR-T), lymphokine activated killer cell therapy (LAK) and other treatment options.118 A recent study showed that cellular immunotherapy combined with oxaliplatin, fluorouracil, and folic acid combined with chemotherapy can appropriately prolong the overall survival and progression-free survival of patients with metastatic colon cancer.119 TCR-T is a type of cellular immunotherapy. The treatment process is to chimerize tumor-specific antigen receptors into T cells, and then genetically modify T cells back into the body to stimulate T cells to specifically recognize and kill tumor cells. Katz SC’s research confirmed that the percutaneous hepatic artery injection method can significantly reduce the CEA level in patients with liver metastases.120
Immunosuppressant therapy is one of the current research hotspots. The target of this treatment is the activation and inhibitory receptors distributed on the surface of T lymphocytes. The activated receptors include CD28, GITR, CD27, etc, and the inhibitory receptors include programmed death 1 (PD-1), cytotoxic T-lymphocyte antigen-4 (CTLA-4), etc. These receptors are also called immune checkpoints. PD-1 is expressed on the surface of immune cells, and programmed cell death ligand 1 (PD-L1) is a tumor cell surface ligand. The surface of solid tumor invading lymphocytes shows PD-1 positive, and PD-1 binds to PD-L1 Inhibit the function of T lymphocytes and induce immune escape of tumor cells. Therefore, blocking PD-1 binding to PD-L1 can reduce tumor cell immune escape.121 Immunosuppressants against PD-1/PD-L1 that have been marketed include nivolumab, pembrolizumab, and atezolizumab. Relevant studies have shown that immunosuppressive therapy can significantly prolong the median progression-free survival of patients and the advantage of progression-free survival is more prominent.122 The reason may be that chemotherapy drugs have a fast onset of cytotoxicity, and PD-1 inhibitors are induced by regulating the immune microenvironment. T cells kill tumor cells, and checkpoint inhibitors work slower than chemotherapy drugs.123
Immunotherapy has become a hot spot in current tumor treatment research, and obvious curative effects can also be seen in CRLM. However, there are still some problems: on the one hand, immunotherapy has narrow indications and is not suitable for any cancer patients. It cannot be used by breastfeeding women, organ transplant patients or patients with advanced cachexia; on the other hand, after the use of immunotherapy. Some patients can get obvious relief in a short time, but there are also cases of drug resistance in a short time. These issues still need follow-up research.
Conclusion and Future Perspectives
Colorectal cancer is still the third largest malignant tumor in the world, and metastasis is the most common cause of death of colorectal cancer, of which liver metastasis is more common. Early diagnosis and treatment of liver metastasis of colorectal cancer is a necessary measure to reduce mortality and improve quality of life and survival time of patients. This paper introduces the research progress of coding genes and ncRNA in the research field of CRLM in recent years, there is an urgent need to understand their mechanism of action in CRLM, looking forward to finding new early diagnostic indicators, and even guiding clinical treatment. Secondly, multidisciplinary comprehensive therapy is one of the main directions in the clinical treatment of liver metastasis of colorectal cancer in recent years. The comprehensive application of surgical treatment, cryotherapy, radiofrequency ablation, absolute ethanol injection, hepatic artery chemoembolization, portal vein chemoembolization, radiotherapy, laser therapy, liver perfusion therapy and immunotherapy will provide a new opportunity for the prognosis of patients with liver metastasis of colorectal cancer (Figure 1).
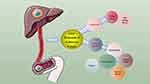 |
Figure 1 A summary of mechanism and treatment on liver metastasis of colorectal cancer. |
Acknowledgments
Wubin Zheng, Fan Wu and Kai Fu are co-first authors.
Funding
This research was supported by Jiangsu Natural Science Foundation (SBK2019021253) and by the Medical Science and Technology Development Foundation (ZKX18027).
Disclosure
The authors declare that they have no competing interests.
References
1. Favoriti P, Carbone G, Greco M, Pirozzi F, Pirozzi RE, Corcione F. Worldwide burden of colorectal cancer: a review. Updates Surg. 2016;68(1):7–11. doi:10.1007/s13304-016-0359-y
2. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
3. Smith KJ, Williams J, Skelton H. Metastatic adenocarcinoma of the esophagus to the skin: new patterns of tumor recurrence and alternate treatments for palliation. J Cutan Pathol. 2001;28(8):425–431. doi:10.1034/j.1600-0560.2001.028008425.x
4. Petri A, Hohn J, Balogh A, Kovách K, Andrási L, Lázár G. [Surgical treatment of liver metastasis in colorectal cancer with simultaneous liver resection]. Magy Onkol. 2010;54(2):125–128. Hungarian. doi:10.1556/MOnkol.54.2010.2.6
5. Nordlinger B, Sorbye H, Glimelius B, et al. Perioperative chemotherapy with FOLFOX4 and surgery versus surgery alone for resectable liver metastases from colorectal cancer (EORTC Intergroup trial 40983): a randomised controlled trial. Lancet. 2008;371(9617):1007–1016. doi:10.1016/S0140-6736(08)60455-9
6. Scholefield JH, Steele RJ; British Society For Gastroenterology, Association of Coloproctology for Great Britain and Ireland. Guidelines for follow up after resection of colorectal cancer. Gut. 2002;51(Suppl 5):V3–5. doi:10.1136/gut.51.suppl_5.v3
7. Weidle UH, Birzele F, Krüger A. Molecular targets and pathways involved in liver metastasis of colorectal cancer. Clin Exp Metastasis. 2015;32(6):623–635. doi:10.1007/s10585-015-9732-3
8. Fang C, Fan C, Wang C, et al. CD133+CD54+CD44+ circulating tumor cells as a biomarker of treatment selection and liver metastasis in patients with colorectal cancer. Oncotarget. 2016;7(47):77389–77403. doi:10.18632/oncotarget.12675
9. McAuliffe JC, Qadan M, D’Angelica MI. Hepatic resection, hepatic arterial infusion pump therapy, and genetic biomarkers in the management of hepatic metastases from colorectal cancer. J Gastrointest Oncol. 2015;6(6):699–708. doi:10.3978/j.issn.2078-6891.2015.081
10. Duffy MJ, van Dalen A, Haglund C, et al. Tumour markers in colorectal cancer: European Group on Tumour Markers (EGTM) guidelines for clinical use. Eur J Cancer. 2007;43(9):1348–1360. doi:10.1016/j.ejca.2007.03.021
11. Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003;3(6):453–458. doi:10.1038/nrc1098
12. Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482(7385):339–346. doi:10.1038/nature10887
13. Cech TR, Steitz JA. The noncoding RNA revolution-trashing old rules to forge new ones. Cell. 2014;157(1):77–94. doi:10.1016/j.cell.2014.03.008
14. Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2016;17(1):47–62. doi:10.1038/nrg.2015.10
15. Fu XD. Non-coding RNA: a new frontier in regulatory biology. Natl Sci Rev. 2014;1(2):190–204. doi:10.1093/nsr/nwu008
16. Liu J, Valencia-Sanchez MA, Hannon GJ, Parker R. MicroRNA-dependent localization of targeted mRNAs to mammalian P-bodies. Nat Cell Biol. 2005;7(7):719–723. doi:10.1038/ncb1274
17. Hu S, Wu J, Chen L, Shan G. Signals from noncoding RNAs: unconventional roles for conventional pol III transcripts. Int J Biochem Cell Biol. 2012;44(11):1847–1851. doi:10.1016/j.biocel.2012.07.013
18. Ulitsky I, Bartel DP. lincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154(1):26–46. doi:10.1016/j.cell.2013.06.020
19. Morris KV, Mattick JS. The rise of regulatory RNA. Nat Rev Genet. 2014;15(6):423–437.
20. Necsulea A, Soumillon M, Warnefors M, et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature. 2014;505(7485):635–640. doi:10.1038/nature12943
21. Mitchell PS, Parkin RK, Kroh EM, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105(30):10513–10518. doi:10.1073/pnas.0804549105
22. Tsai MC, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71(1):3–7. doi:10.1158/0008-5472.CAN-10-2483
23. DiStefano JK. The emerging role of long noncoding RNAs in human disease. Methods Mol Biol. 2018;1706:91–110.
24. Xue M, Zhuo Y, Shan B. MicroRNAs, long noncoding RNAs, and their functions in human disease. Methods Mol Biol. 2017;1617:1–25.
25. Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends Cell Biol. 2011;21(6):354–361. doi:10.1016/j.tcb.2011.04.001
26. Lalevée S, Feil R. Long noncoding RNAs in human disease: emerging mechanisms and therapeutic strategies. Epigenomics. 2015;7(6):877–879. doi:10.2217/epi.15.55
27. Planchon SM, Waite KA, Eng C. The nuclear affairs of PTEN. J Cell Sci. 2008;121(3):249–253. doi:10.1242/jcs.022459
28. Ming M, He YY. PTEN in DNA damage repair. Cancer Lett. 2012;319(2):125–129. doi:10.1016/j.canlet.2012.01.003
29. Milella M, Falcone I, Conciatori F, et al. PTEN: multiple functions in human malignant tumors. Front Oncol. 2015;5:24. doi:10.3389/fonc.2015.00024
30. Chen M, Zhang L, Cui X, et al. Wt1 directs the lineage specification of sertoli and granulosa cells by repressing Sf1 expression. Development. 2017;144(1):44–53. doi:10.1242/dev.144105
31. Vikhanskaya F. p73 competes with p53 and attenuates its response in a human ovarian cancer cell line. Nucleic Acids Res. 2000;28(2):513–519. doi:10.1093/nar/28.2.513
32. Porter PL, Gown AM, Kramp SG, Coltrera MD. Widespread p53 overexpression in human malignant tumors. An immunohistochemical study using methacarn-fixed, embedded tissue. Am J Pathol. 1992;140(1):145–153.
33. Du T, Niu H. Inhibitory effect of gene combination in a mouse model of colon cancer with liver metastasis. Exp Ther Med. 2014;8(3):913–918. doi:10.3892/etm.2014.1809
34. Furuse M, Fujita K, Hiiragi T, Fujimoto K, Tsukita S. Claudin-1 and −2: novel integral membrane proteins localizing at tight junctions with no sequence similarity to occludin. J Cell Biol. 1998;141(7):1539–1550. doi:10.1083/jcb.141.7.1539
35. Turksen K, Troy TC. Barriers built on claudins. J Cell Sci. 2004;117(12):2435–2447. doi:10.1242/jcs.01235
36. Kasapkara CS, Tumer L, Okur I, Hasanoglu A. A novel mutation of the claudin 16 gene in familial hypomagnesemia with hypercalciuria and nephrocalcinosis mimicking rickets. Genet Couns. 2011;22(2):187–192.
37. Li J, Chigurupati S, Agarwal R, et al. Possible angiogenic roles for claudin-4 in ovarian cancer. Cancer Biol Ther. 2009;8(19):1806–1814. doi:10.4161/cbt.8.19.9427
38. Stein U, Walther W, Arlt F, et al. MACC1, a newly identified key regulator of HGF-MET signaling, predicts colon cancer metastasis. Nat Med. 2009;15(1):59–67. doi:10.1038/nm.1889
39. Shimokawa H, Uramoto H, Onitsuka T, et al. Overexpression of MACC1 mRNA in lung adenocarcinoma is associated with postoperative recurrence. J Thorac Cardiovasc Surg. 2011;141(4):895–898. doi:10.1016/j.jtcvs.2010.09.044
40. Sattler M, Salgia R. c-Met and hepatocyte growth factor: potential as novel targets in cancer therapy. Curr Oncol Rep. 2007;9(2):102–108. doi:10.1007/s11912-007-0005-4
41. Dubocovich ML, Langer SZ. Negative feed-back regulation of noradrenaline release by nerve stimulation in the perfused cat’s spleen: differences in potency of phenoxybenzamine in blocking the pre- and post-synaptic adrenergic receptors. J Physiol. 1974;237(3):505–519. doi:10.1113/jphysiol.1974.sp010495
42. Birchmeier C, Birchmeier W, Gherardi E, Vande Woude GF. Met, metastasis, motility and more. Nat Rev Mol Cell Biol. 2003;4(12):915–925. doi:10.1038/nrm1261
43. Ge SH, Wu XJ, Wang XH, et al. Over-expression of Metastasis-associated in Colon Cancer-1 (MACC1) associates with better prognosis of gastric cancer patients. Chin J Cancer Res. 2011;23(2):153–159. doi:10.1007/s11670-011-0153-9
44. Frandsen PM, Madsen LB, Bendixen C, Larsen K. Porcine gamma-synuclein: molecular cloning, expression analysis, chromosomal localization and functional expression. Mol Biol Rep. 2009;36(5):971–979. doi:10.1007/s11033-008-9270-z
45. Ji H, Liu YE, Jia T, et al. Identification of a breast cancer-specific gene, BCSG1, by direct differential cDNA sequencing. Cancer Res. 1997;57(4):759–764.
46. Ninkina NN, Alimova-Kost MV, Paterson JW, et al. Organization, expression and polymorphism of the human persyn gene. Hum Mol Genet. 1998;7(9):1417–1424. doi:10.1093/hmg/7.9.1417
47. Bruening W, Giasson BI, Klein-Szanto AJ, Lee VM, Trojanowski JQ, Godwin AK. Synucleins are expressed in the majority of breast and ovarian carcinomas and in preneoplastic lesions of the ovary. Cancer. 2000;88(9):2154–2163. doi:10.1002/(SICI)1097-0142(20000501)88:9<2154::AID-CNCR23>3.0.CO;2-9
48. Lu A, Zhang F, Gupta A, Liu J. Blockade of AP1 transactivation abrogates the abnormal expression of breast cancer-specific gene 1 in breast cancer cells. J Biol Chem. 2002;277(35):31364–31372. doi:10.1074/jbc.M201060200
49. Steinmüller L, Cibelli G, Moll JR, Vinson C, Thiel G. Regulation and composition of activator protein 1 (AP-1) transcription factors controlling collagenase and c-Jun promoter activities. Biochem J. 2001;360(3):599–607. doi:10.1042/bj3600599
50. Przybyszewski J, Yaktine AL, Duysen E, et al. Inhibition of phorbol ester-induced AP-1-DNA binding, c-Jun protein and c-jun mRNA by dietary energy restriction is reversed by adrenalectomy in SENCAR mouse epidermis. Carcinogenesis. 2001;22(9):1421–1427. doi:10.1093/carcin/22.9.1421
51. Inaba S, Li C, Shi YE, Song DQ, Jiang JD, Liu J. Synuclein gamma inhibits the mitotic checkpoint function and promotes chromosomal instability of breast cancer cells. Breast Cancer Res Treat. 2005;94(1):25–35. doi:10.1007/s10549-005-6938-0
52. Liu H, Zhou Y, Boggs SE, Belinsky SA, Liu J. Cigarette smoke induces demethylation of prometastatic oncogene synuclein-gamma in lung cancer cells by downregulation of DNMT3B. Oncogene. 2007;26(40):5900–5910. doi:10.1038/sj.onc.1210400
53. Jiang Y, Liu YE, Lu A, et al. Stimulation of estrogen receptor signaling by gamma synuclein. Cancer Res. 2003;63(14):3899–3903.
54. Guo CG, Sun LC, Liu Q, Xie YB, Wang X. [SNCG expression and clinical significance in colorectal cancer liver metastasis]. Zhonghua Wei Chang Wai Ke Za Zhi. 2012;15(6):625–628. Chinese.
55. Bonecchi R, Galliera E, Borroni EM, Corsi MM, Locati M, Mantovani A. Chemokines and chemokine receptors: an overview. Front Biosci (Landmark Ed). 2009;14:540–551. doi:10.2741/3261
56. Bruserud Ø, Kittang AO. The chemokine system in experimental and clinical hematology. Curr Top Microbiol Immunol. 2010;341:3–12. doi:10.1007/82_2010_18
57. Singh S, Sadanandam A, Singh RK. Chemokines in tumor angiogenesis and metastasis. Cancer Metastasis Rev. 2007;26(3–4):453–467.
58. Balkwill F. Chemokine biology in cancer. Semin Immunol. 2003;15(1):49–55. doi:10.1016/S1044-5323(02)00127-6
59. Tan HX, Gong WZ, Zhou K, et al. CXCR4/TGF-β1 mediated hepatic stellate cells differentiation into carcinoma-associated fibroblasts and promoted liver metastasis of colon cancer. Cancer Biol Ther. 2020;21(3):258–268. doi:10.1080/15384047.2019.1685157
60. Bie Y, Ge W, Yang Z, et al. The crucial role of CXCL8 and its receptors in colorectal liver metastasis. Dis Markers. 2019;2019:8023460. doi:10.1155/2019/8023460
61. Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75(5):855–862. doi:10.1016/0092-8674(93)90530-4
62. Ambros V. The functions of animal microRNAs. Nature. 2004;431(7006):350–355. doi:10.1038/nature02871
63. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. doi:10.1016/S0092-8674(04)00045-5
64. Seto AG, Kingston RE, Lau NC. The coming of age for Piwi proteins. Mol Cell. 2007;26(5):603–609. doi:10.1016/j.molcel.2007.05.021
65. Siomi MC, Sato K, Pezic D, Aravin AA. PIWI-interacting small RNAs: the vanguard of genome defence. Nat Rev Mol Cell Biol. 2011;12(4):246–258. doi:10.1038/nrm3089
66. Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7(2):e30733. doi:10.1371/journal.pone.0030733
67. Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci U S A. 1976;73(11):3852–3856. doi:10.1073/pnas.73.11.3852
68. Jeck WR, Sorrentino JA, Wang K, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19(2):141–157. doi:10.1261/rna.035667.112
69. Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ. Processing of primary microRNAs by the microprocessor complex. Nature. 2004;432(7014):231–235. doi:10.1038/nature03049
70. Ruvkun G. Glimpses of a tiny RNA world. Science. 2001;294(5543):797–799. doi:10.1126/science.1066315
71. Søkilde R, Vincent M, Møller AK, et al. Efficient identification of miRNAs for classification of tumor origin. J Mol Diagn. 2014;16(1):106–115. doi:10.1016/j.jmoldx.2013.10.001
72. Pan X, Wang ZX, Wang R. MicroRNA-21: a novel therapeutic target in human cancer. Cancer Biol Ther. 2010;10(12):1224–1232. doi:10.4161/cbt.10.12.14252
73. Ibrahim SA, Yip GW, Stock C, et al. Targeting of syndecan-1 by microRNA miR-10b promotes breast cancer cell motility and invasiveness via a Rho-GTPase- and E-cadherin-dependent mechanism. Int J Cancer. 2012;131(6):E884–96. doi:10.1002/ijc.27629
74. Xue G, Yan HL, Zhang Y, et al. c-Myc-mediated repression of miR-15-16 in hypoxia is induced by increased HIF-2α and promotes tumor angiogenesis and metastasis by upregulating FGF2. Oncogene. 2015;34(11):1393–1406. doi:10.1038/onc.2014.82
75. Barh D, Malhotra R, Ravi B, Sindhurani P. MicroRNA let-7: an emerging next-generation cancer therapeutic. Curr Oncol. 2010;17(1):70–80. doi:10.3747/co.v17i1.356
76. Jaskiewicz L, Filipowicz W. Role of Dicer in posttranscriptional RNA silencing. Curr Top Microbiol Immunol. 2008;320:77–97. doi:10.1007/978-3-540-75157-1_4
77. Zhang X, Zeng Y. The terminal loop region controls microRNA processing by Drosha and Dicer. Nucleic Acids Res. 2010;38(21):7689–7697. doi:10.1093/nar/gkq645
78. Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language. Cell. 2011;146(3):353–358. doi:10.1016/j.cell.2011.07.014
79. Demirkan B. The roles of Epithelial-to-Mesenchymal Transition (EMT) and Mesenchymal-to-Epithelial Transition (MET) in breast cancer bone metastasis: potential targets for prevention and treatment. J Clin Med. 2013;2(4):264–282. doi:10.3390/jcm2040264
80. Brabletz T. EMT and MET in metastasis: where are the cancer stem cells. Cancer Cell. 2012;22(6):699–701. doi:10.1016/j.ccr.2012.11.009
81. De Craene B, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer. 2013;13(2):97–110. doi:10.1038/nrc3447
82. Shelton PM, Duran A, Nakanishi Y, et al. The secretion of miR-200s by a PKCζ/ADAR2 signaling axis promotes liver metastasis in colorectal cancer. Cell Rep. 2018;23(4):1178–1191. doi:10.1016/j.celrep.2018.03.118
83. Hata T, Mokutani Y, Takahashi H, et al. Identification of microRNA-487b as a negative regulator of liver metastasis by regulation of KRAS in colorectal cancer. Int J Oncol. 2017;50(2):487–496. doi:10.3892/ijo.2016.3813
84. Okazaki Y, Furuno M, Kasukawa T, et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature. 2002;420(6915):563–573.
85. Rinn JL, Kertesz M, Wang JK, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007;129(7):1311–1323. doi:10.1016/j.cell.2007.05.022
86. Guttman M, Amit I, Garber M, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458(7235):223–227. doi:10.1038/nature07672
87. Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465(7301):1033–1038. doi:10.1038/nature09144
88. Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43(6):904–914. doi:10.1016/j.molcel.2011.08.018
89. Zhang F, Zhang L, Zhang C. Long noncoding RNAs and tumorigenesis: genetic associations, molecular mechanisms, and therapeutic strategies. Tumour Biol. 2016;37(1):163–175. doi:10.1007/s13277-015-4445-4
90. Kwok ZH, Tay Y. Long noncoding RNAs: lincs between human health and disease. Biochem Soc Trans. 2017;45(3):805–812. doi:10.1042/BST20160376
91. Miyoshi A, Kitajima Y, Sumi K, et al. Snail and SIP1 increase cancer invasion by upregulating MMP family in hepatocellular carcinoma cells. Br J Cancer. 2004;90(6):1265–1273. doi:10.1038/sj.bjc.6601685
92. Chen DL, Lu YX, Zhang JX, et al. Long non-coding RNA UICLM promotes colorectal cancer liver metastasis by acting as a ceRNA for microRNA-215 to regulate ZEB2 expression. Theranostics. 2017;7(19):4836–4849. doi:10.7150/thno.20942
93. McElwee JL, Mohanan S, Horibata S, et al. PAD2 overexpression in transgenic mice promotes spontaneous skin neoplasia. Cancer Res. 2014;74(21):6306–6317. doi:10.1158/0008-5472.CAN-14-0749
94. Tanday S. Targeting PADI2 could stop the progression of myeloma. Lancet Oncol. 2016;17(8):e325. doi:10.1016/S1470-2045(16)30314-X
95. Wang H, Xu B, Zhang X, Zheng Y, Zhao Y, Chang X. PADI2 gene confers susceptibility to breast cancer and plays tumorigenic role via ACSL4, BINC3 and CA9 signaling. Cancer Cell Int. 2016;16:61. doi:10.1186/s12935-016-0335-0
96. Wang G, Mao W, Zheng S, Ye J. Epidermal growth factor receptor-regulated miR-125a-5p–a metastatic inhibitor of lung cancer. FEBS J. 2009;276(19):5571–5578. doi:10.1111/j.1742-4658.2009.07238.x
97. Sun Y, Liu X, Zhang Q, et al. Oncogenic potential of TSTA3 in breast cancer and its regulation by the tumor suppressors miR-125a-5p and miR-125b. Tumour Biol. 2016;37(4):4963–4972. doi:10.1007/s13277-015-4178-4
98. Xu Y, Huang Z, Liu Y. Reduced miR-125a-5p expression is associated with gastric carcinogenesis through the targeting of E2F3. Mol Med Rep. 2014;10(5):2601–2608. doi:10.3892/mmr.2014.2567
99. Kim JK, Noh JH, Jung KH, et al. Sirtuin7 oncogenic potential in human hepatocellular carcinoma and its regulation by the tumor suppressors MiR-125a-5p and MiR-125b. Hepatology. 2013;57(3):1055–1067. doi:10.1002/hep.26101
100. Nishida N, Mimori K, Fabbri M, et al. MicroRNA-125a-5p is an independent prognostic factor in gastric cancer and inhibits the proliferation of human gastric cancer cells in combination with trastuzumab. Clin Cancer Res. 2011;17(9):2725–2733. doi:10.1158/1078-0432.CCR-10-2132
101. Chen D, Sun Q, Zhang L, et al. The lncRNA HOXA11-AS functions as a competing endogenous RNA to regulate PADI2 expression by sponging miR-125a-5p in liver metastasis of colorectal cancer. Oncotarget. 2017;8(41):70642–70652. doi:10.18632/oncotarget.19956
102. Yamahashi Y, Hatakeyama M. PAR1b takes the stage in the morphogenetic and motogenetic activity of Helicobacter pylori CagA oncoprotein. Cell Adh Migr. 2013;7(1):11–18. doi:10.4161/cam.21936
103. Yang H, Zhang H, Yang Y, et al. Hypoxia induced exosomal circRNA promotes metastasis of colorectal cancer via targeting GEF-H1/RhoA axis. Theranostics. 2020;10(18):8211–8226. doi:10.7150/thno.44419
104. Ozturk N, Singh I, Mehta A, Braun T, Barreto G. HMGA proteins as modulators of chromatin structure during transcriptional activation. Front Cell Dev Biol. 2014;2:5. doi:10.3389/fcell.2014.00005
105. Sgarra R, Rustighi A, Tessari MA, et al. Nuclear phosphoproteins HMGA and their relationship with chromatin structure and cancer. FEBS Lett. 2004;574(1–3):1–8. doi:10.1016/j.febslet.2004.08.013
106. Frost L, Baez MA, Harrilal C, Garabedian A, Fernandez-Lima F, Leng F. The dimerization state of the mammalian high mobility group protein AT-hook 2 (HMGA2). PLoS One. 2015;10(6):e0130478. doi:10.1371/journal.pone.0130478
107. Galdiero F, Romano A, Pasquinelli R, et al. Detection of high mobility group A2 specific mRNA in the plasma of patients affected by epithelial ovarian cancer. Oncotarget. 2015;6(22):19328–19335. doi:10.18632/oncotarget.2896
108. Chen RX, Chen X, Xia LP, et al. N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10(1):4695. doi:10.1038/s41467-019-12651-2
109. Dexiang Z, Li R, Ye W, et al. Outcome of patients with colorectal liver metastasis: analysis of 1613 consecutive cases. Ann Surg Oncol. 2012;19(9):2860–2868. doi:10.1245/s10434-012-2356-9
110. Adam R, Hoti E, Folprecht G, Benson AB. Accomplishments in 2008 in the management of curable metastatic colorectal cancer. Gastrointest Cancer Res. 2009;3(5 Supplement 2):S15–22.
111. Enquist IB, Good Z, Jubb AM, et al. Lymph node-independent liver metastasis in a model of metastatic colorectal cancer. Nat Commun. 2014;5:3530. doi:10.1038/ncomms4530
112. Zhang Y, Davis C, Ryan J, Janney C, Peña MM. Development and characterization of a reliable mouse model of colorectal cancer metastasis to the liver. Clin Exp Metastasis. 2013;30(7):903–918. doi:10.1007/s10585-013-9591-8
113. Lee HS, Kim HO, Hong YS, et al. Prognostic value of metabolic parameters in patients with synchronous colorectal cancer liver metastasis following curative-intent colorectal and hepatic surgery. J Nucl Med. 2014;55(4):582–589. doi:10.2967/jnumed.113.128629
114. Xiao Y, Freeman GJ. The microsatellite instable subset of colorectal cancer is a particularly good candidate for checkpoint blockade immunotherapy. Cancer Discov. 2015;5(1):16–18. doi:10.1158/2159-8290.CD-14-1397
115. Wu T, Dai Y. Tumor microenvironment and therapeutic response. Cancer Lett. 2017;387:61–68. doi:10.1016/j.canlet.2016.01.043
116. Dougan M, Dougan SK. Targeting immunotherapy to the tumor microenvironment. J Cell Biochem. 2017;118(10):3049–3054. doi:10.1002/jcb.26005
117. Wrobel P, Ahmed S. Current status of immunotherapy in metastatic colorectal cancer. Int J Colorectal Dis. 2019;34(1):13–25. doi:10.1007/s00384-018-3202-8
118. Andersen R, Donia M, Westergaard MC, Pedersen M, Hansen M, Svane IM. Tumor infiltrating lymphocyte therapy for ovarian cancer and renal cell carcinoma. Hum Vaccin Immunother. 2015;11(12):2790–2795. doi:10.1080/21645515.2015.1075106
119. Veluchamy JP, Lopez-Lastra S, Spanholtz J, et al. In vivo efficacy of umbilical cord blood stem cell-derived NK cells in the treatment of metastatic colorectal cancer. Front Immunol. 2017;8:87. doi:10.3389/fimmu.2017.00087
120. Katz SC, Burga RA, McCormack E, et al. Phase I hepatic immunotherapy for metastases study of intra-arterial chimeric antigen receptor-modified T-cell therapy for CEA+ liver metastases. Clin Cancer Res. 2015;21(14):3149–3159. doi:10.1158/1078-0432.CCR-14-1421
121. Quiroga D, Lyerly HK, Morse MA. Deficient mismatch repair and the role of immunotherapy in metastatic colorectal cancer. Curr Treat Options Oncol. 2016;17(8):41. doi:10.1007/s11864-016-0414-4
122. Yaghoubi N, Soltani A, Ghazvini K, Hassanian SM, Hashemy SI. PD-1 inhibitor bests chemo for colorectal cancer. Cancer Discov. 2020;10(7):OF2. doi:10.1158/2159-8290.CD-NB2020-051
123. Herbst RS, Soria JC, Kowanetz M, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515(7528):563–567. doi:10.1038/nature14011
124. Bu P, Chen KY, Xiang K, et al. Aldolase B-mediated fructose metabolism drives metabolic reprogramming of colon cancer liver metastasis. Cell Metab. 2018;27(6):1249–1262.e4. doi:10.1016/j.cmet.2018.04.003
125. Na H, Liu X, Li X, et al. Novel roles of DC-SIGNR in colon cancer cell adhesion, migration, invasion, and liver metastasis. J Hematol Oncol. 2017;10(1):28.
126. Li Y, Xi Y, Zhu G, et al. Downregulated IGFBP7 facilitates liver metastasis by modulating epithelial‑mesenchymal transition in colon cancer. Oncol Rep. 2019;42(5):1935–1945. doi:10.3892/or.2019.7303
127. Chai WX, Sun LG, Dai FH, Shao HS, Zheng NG, Cai HY. Inhibition of PRRX2 suppressed colon cancer liver metastasis via inactivation of Wnt/β-catenin signaling pathway. Pathol Res Pract. 2019;215(10):152593. doi:10.1016/j.prp.2019.152593
128. Urosevic J, Blasco MT, Llorente A, et al. ERK1/2 signaling induces upregulation of ANGPT2 and CXCR4 to mediate liver metastasis in colon cancer. Cancer Res. 2020;80(21):4668–4680. doi:10.1158/0008-5472.CAN-19-4028
129. Lichao S, Liang P, Chunguang G, Fang L, Zhihua Y, Yuliang R. Overexpression of PTGIS could predict liver metastasis and is correlated with poor prognosis in colon cancer patients. Pathol Oncol Res. 2012;18(3):563–569. doi:10.1007/s12253-011-9478-4
130. Ding YL, Zhou Y, Xiang L, Ji ZP, Luo ZH. Expression of glioma-associated oncogene homolog 1 is associated with invasion and postoperative liver metastasis in colon cancer. Int J Med Sci. 2012;9(5):334–338. doi:10.7150/ijms.4553
131. Yamamoto M, Kikuchi H, Ohta M, et al. TSU68 prevents liver metastasis of colon cancer xenografts by modulating the premetastatic niche. Cancer Res. 2008;68(23):9754–9762. doi:10.1158/0008-5472.CAN-08-1748
132. Min BS, Kim NK, Jeong HC, Chung HC. High levels of serum VEGF and TIMP-1 are correlated with colon cancer liver metastasis and intrahepatic recurrence after liver resection. Oncol Lett. 2012;4(1):123–130. doi:10.3892/ol.2012.691
133. Tokunaga T, Oshika Y, Abe Y, et al. Vascular endothelial growth factor (VEGF) mRNA isoform expression pattern is correlated with liver metastasis and poor prognosis in colon cancer. Br J Cancer. 1998;77(6):998–1002. doi:10.1038/bjc.1998.164
134. Okamoto K, Oshika Y, Fukushima Y, et al. Inhibition of liver metastasis of colon cancer by in vivo administration of anti-vascular endothelial growth factor antibody. Oncol Rep. 1999;6(3):553–556. doi:10.3892/or.6.3.553
135. Warren RS, Yuan H, Matli MR, Gillett NA, Ferrara N. Regulation by vascular endothelial growth factor of human colon cancer tumorigenesis in a mouse model of experimental liver metastasis. J Clin Invest. 1995;95(4):1789–1797. doi:10.1172/JCI117857
136. Ding YL, Fu QY, Tang SF, Zhang JL, Li ZY, Li ZT. [Effect of stromal cell-derived factor-1 and its receptor CXCR4 on liver metastasis of human colon cancer]. Zhonghua Wai Ke Za Zhi. 2009;47(3):210–213. Chinese.
137. Huang ZL, Liu HY. [Expression of galectin-3 in liver metastasis of colon cancer and the inhibitory effect of modified citrus pectin]. Nan Fang Yi Ke Da Xue Xue Bao. 2008;28(8):1358–1361.
138. Fang YJ, Lu ZH, Wang GQ, et al. Elevated expressions of MMP7, TROP2, and survivin are associated with survival, disease recurrence, and liver metastasis of colon cancer. Int J Colorectal Dis. 2009;24(8):875–884. doi:10.1007/s00384-009-0725-z
139. Wen J, Matsumoto K, Taniura N, Tomioka D, Nakamura T. Hepatic gene expression of NK4, an HGF-antagonist/angiogenesis inhibitor, suppresses liver metastasis and invasive growth of colon cancer in mice. Cancer Gene Ther. 2004;11(6):419–430. doi:10.1038/sj.cgt.7700705
140. Babyatsky M, Lin J, Yio X, et al. Trefoil factor-3 expression in human colon cancer liver metastasis. Clin Exp Metastasis. 2009;26(2):143–151. doi:10.1007/s10585-008-9224-9
141. Yamauchi K, Ogata Y, Nagase H, Shirouzu K. Inhibition of liver metastasis from orthotopically implanted colon cancer in nude mice by transfection of the TIMP-1 gene into KM12SM cells. Surg Today. 2001;31(9):791–798. doi:10.1007/s005950170049
142. Murata S, Yanagisawa K, Fukunaga K, et al. Fatty acid synthase inhibitor cerulenin suppresses liver metastasis of colon cancer in mice. Cancer Sci. 2010;101(8):1861–1865. doi:10.1111/j.1349-7006.2010.01596.x
143. Matsushita Y, Kitajima S, Goto M, et al. Selectins induced by interleukin-1beta on the human liver endothelial cells act as ligands for sialyl Lewis X-expressing human colon cancer cell metastasis. Cancer Lett. 1998;133(2):151–160. doi:10.1016/S0304-3835(98)00220-1
144. Li M, Lin YM, Hasegawa S, et al. Genes associated with liver metastasis of colon cancer, identified by genome-wide cDNA microarray. Int J Oncol. 2004;24(2):305–312.
145. Kitamura T, Fujishita T, Loetscher P, et al. Inactivation of chemokine (C-C motif) receptor 1 (CCR1) suppresses colon cancer liver metastasis by blocking accumulation of immature myeloid cells in a mouse model. Proc Natl Acad Sci U S A. 2010;107(29):13063–13068. doi:10.1073/pnas.1002372107
146. Oba K, Konno H, Tanaka T, et al. Prevention of liver metastasis of human colon cancer by selective matrix metalloproteinase inhibitor MMI-166. Cancer Lett. 2002;175(1):45–51. doi:10.1016/S0304-3835(01)00726-1
147. Oshima T, Kunisaki C, Yoshihara K, et al. Clinicopathological significance of the gene expression of matrix metalloproteinases and reversion-inducing cysteine-rich protein with Kazal motifs in patients with colorectal cancer: MMP-2 gene expression is a useful predictor of liver metastasis from colorectal cancer. Oncol Rep. 2008;19(5):1285–1291.
148. Suwa D, Konno H, Tanaka T, Urano T. Intraperitoneal infusion of recombinant plasminogen activator inhibitor type 2 induced apoptosis in implanted human colon cancer and inhibited its growth and liver metastasis. Anticancer Res. 2008;28(2A):693–698.
149. Zhang B, Halder SK, Zhang S, Datta PK. Targeting transforming growth factor-beta signaling in liver metastasis of colon cancer. Cancer Lett. 2009;277(1):114–120. doi:10.1016/j.canlet.2008.11.035
150. Ishii M, Iwai M, Harada Y, et al. Soluble TRAIL gene and actinomycin D synergistically suppressed multiple metastasis of TRAIL-resistant colon cancer in the liver. Cancer Lett. 2007;245(1–2):134–143. doi:10.1016/j.canlet.2005.12.040
151. Aykut B, Ochs M, Radhakrishnan P, et al. EMX2 gene expression predicts liver metastasis and survival in colorectal cancer. BMC Cancer. 2017;17(1):555. doi:10.1186/s12885-017-3556-2
152. Teraoku H, Morine Y, Ikemoto T, et al. Role of thrombospondin-1 expression in colorectal liver metastasis and its molecular mechanism. J Hepatobiliary Pancreat Sci. 2016;23(9):565–573. doi:10.1002/jhbp.376
153. Kim CW, Roh SA, Tak KH, et al. ZKSCAN3 facilitates liver metastasis of colorectal cancer associated with CEA-expressing tumor. Anticancer Res. 2016;36(5):2397–2406.
154. Liu H, Xu Y, Zhang Q, et al. Prognostic significance of TBL1XR1 in predicting liver metastasis for early stage colorectal cancer. Surg Oncol. 2017;26(1):13–20. doi:10.1016/j.suronc.2016.12.003
155. Ai X, Wu Y, Zhang W, et al. Targeting the ERK pathway reduces liver metastasis of Smad4-inactivated colorectal cancer. Cancer Biol Ther. 2013;14(11):1059–1067. doi:10.4161/cbt.26427
156. Kawashita Y, Morine Y, Ikemoto T, et al. Loss of Fbxw7 expression is a predictor of recurrence in colorectal liver metastasis. J Hepatobiliary Pancreat Sci. 2017;24(10):576–583. doi:10.1002/jhbp.500
157. Ding Q, Chang CJ, Xie X, et al. APOBEC3G promotes liver metastasis in an orthotopic mouse model of colorectal cancer and predicts human hepatic metastasis. J Clin Invest. 2011;121(11):4526–4536. doi:10.1172/JCI45008
158. Jiang T, Zhu A, Zhu Y, Piao D. Clinical implications of AEG-1 in liver metastasis of colorectal cancer. Med Oncol. 2012;29(4):2858–2863. doi:10.1007/s12032-012-0186-z
159. Huang MY, Yen LC, Liu HC, et al. Significant overexpression of DVL1 in Taiwanese colorectal cancer patients with liver metastasis. Int J Mol Sci. 2013;14(10):20492–20507. doi:10.3390/ijms141020492
160. Georges R, Bergmann F, Hamdi H, et al. Sequential biphasic changes in claudin1 and claudin4 expression are correlated to colorectal cancer progression and liver metastasis. J Cell Mol Med. 2012;16(2):260–272. doi:10.1111/j.1582-4934.2011.01289.x
161. Liu YL, Yang YM, Xu H, Dong XS. Aberrant expression of USP22 is associated with liver metastasis and poor prognosis of colorectal cancer. J Surg Oncol. 2011;103(3):283–289. doi:10.1002/jso.21802
162. Ge Y, Meng X, Zhou Y, Zhang J, Ding Y. Positive MACC1 expression correlates with invasive behaviors and postoperative liver metastasis in colon cancer. Int J Clin Exp Med. 2015;8(1):1094–1100.
163. Huang MY, Wang HM, Chang HJ, Hsiao CP, Wang JY, Lin SR. Overexpression of S100B, TM4SF4, and OLFM4 genes is correlated with liver metastasis in Taiwanese colorectal cancer patients. DNA Cell Biol. 2012;31(1):43–49. doi:10.1089/dna.2011.1264
164. Cho YB, Lee WY, Choi SJ, et al. CC chemokine ligand 7 expression in liver metastasis of colorectal cancer. Oncol Rep. 2012;28(2):689–694. doi:10.3892/or.2012.1815
165. Hu XT, Chen W, Wang D, et al. The proteasome subunit PSMA7 located on the 20q13 amplicon is overexpressed and associated with liver metastasis in colorectal cancer. Oncol Rep. 2008;19(2):441–446.
166. Sun Y, Li Z, Xu Z. [Association of expression of HGF and its receptor c-Met to primary colorectal cancers with synchronous liver metastasis]. Zhonghua Wei Chang Wai Ke Za Zhi. 2014;17(6):598–601. Chinese.
167. Komatsu K, Kobune-Fujiwara Y, Andoh A, et al. Increased expression of S100A6 at the invading fronts of the primary lesion and liver metastasis in patients with colorectal adenocarcinoma. Br J Cancer. 2000;83(6):769–774. doi:10.1054/bjoc.2000.1356
168. Nakayama N, Yamashita K, Tanaka T, et al. Genomic gain of the PRL-3 gene may represent poor prognosis of primary colorectal cancer, and associate with liver metastasis. Clin Exp Metastasis. 2016;33(1):3–13. doi:10.1007/s10585-015-9749-7
169. Tanaka T, Kaida T, Yokoi K, et al. Critical relevance of genomic gains of PRL-3/EGFR/c-myc pathway genes in liver metastasis of colorectal cancer. Oncol Lett. 2019;17(1):1257–1266. doi:10.3892/ol.2018.9728
170. Huang CJ, Teng HW, Chien CC, Lin JK, Yang SH. Prognostic significance of C-reactive protein polymorphism and KRAS/BRAF in synchronous liver metastasis from colorectal cancer. PLoS One. 2014;8(6):e65117. doi:10.1371/journal.pone.0065117
171. Shoji Y, Takamura H, Ninomiya I, et al. The embryonic stem cell-specific transcription factor ZFP57 promotes liver metastasis of colorectal cancer. J Surg Res. 2019;237:22–29. doi:10.1016/j.jss.2018.11.014
172. Liang L, Wei Y, Ren L, Zhong YS, Xu JM. [A study of the relationship between the mutation of PIK3CA, PTEN and the occurrence of liver metastasis of colorectal cancer: survival analysis]. Zhonghua Wai Ke Za Zhi. 2012;50(11):1007–1010. Chinese.
173. Watanabe T, Kobunai T, Yamamoto Y, et al. Prediction of liver metastasis after colorectal cancer using reverse transcription-polymerase chain reaction analysis of 10 genes. Eur J Cancer. 2010;46(11):2119–2126. doi:10.1016/j.ejca.2010.04.019
174. Ito D, Yogosawa S, Mimoto R, et al. Dual-specificity tyrosine-regulated kinase 2 is a suppressor and potential prognostic marker for liver metastasis of colorectal cancer. Cancer Sci. 2017;108(8):1565–1573. doi:10.1111/cas.13280
175. Lin Q, Wei Y, Zhong Y, et al. Aberrant expression of sphingosine-1-phosphate receptor 1 correlates with metachronous liver metastasis and poor prognosis in colorectal cancer. Tumour Biol. 2014;35(10):9743–9750. doi:10.1007/s13277-014-2267-4
176. Kee JY, Ito A, Hojo S, et al. Chemokine CXCL16 suppresses liver metastasis of colorectal cancer via augmentation of tumor-infiltrating natural killer T cells in a murine model. Oncol Rep. 2013;29(3):975–982. doi:10.3892/or.2012.2185
177. Oshima T, Kawasaki T, Ohashi R, et al. Downregulated P1 promoter-driven hepatocyte nuclear factor-4alpha expression in human colorectal carcinoma is a new prognostic factor against liver metastasis. Pathol Int. 2007;57(2):82–90. doi:10.1111/j.1440-1827.2006.02061.x
178. Chu PC, Lin PC, Wu HY, et al. Mutant KRAS promotes liver metastasis of colorectal cancer, in part, by upregulating the MEK-Sp1-DNMT1-miR-137-YB-1-IGF-IR signaling pathway. Oncogene. 2018;37(25):3440–3455. doi:10.1038/s41388-018-0222-3
179. Nakamura M, Miyamoto S, Maeda H, et al. Low levels of insulin-like growth factor type 1 receptor expression at cancer cell membrane predict liver metastasis in Dukes’ C human colorectal cancers. Clin Cancer Res. 2004;10(24):8434–8441. doi:10.1158/1078-0432.CCR-04-0430
180. Tóth C, Sükösd F, Valicsek E, Herpel E, Schirmacher P, Tiszlavicz L. Loss of CDX2 gene expression is associated with DNA repair proteins and is a crucial member of the Wnt signaling pathway in liver metastasis of colorectal cancer. Oncol Lett. 2018;15(3):3586–3593. doi:10.3892/ol.2018.7756
181. Xu H, Wang C, Song H, Xu Y, Ji G. RNA-Seq profiling of circular RNAs in human colorectal Cancer liver metastasis and the potential biomarkers. Mol Cancer. 2019;18(1):8. doi:10.1186/s12943-018-0932-8
182. Wang Q, Shi L, Shi K, et al. CircCSPP1 functions as a ceRNA to promote colorectal carcinoma cell EMT and liver metastasis by upregulating COL1A1. Front Oncol. 2020;10:850. doi:10.3389/fonc.2020.00850
183. Shi L, Tao C, Tang Y, Xia Y, Li X, Wang X. Hypoxia-induced hsa_circ_0000826 is linked to liver metastasis of colorectal cancer. J Clin Lab Anal. 2020;34(9):e23405. doi:10.1002/jcla.23405
184. Zhi Q, Wan D, Ren R, et al. Circular RNA profiling identifies circ102049 as a key regulator of colorectal liver metastasis. Mol Oncol. 2020;15:623–641. doi:10.1002/1878-0261.12840
185. Ye LC, Ren L, Qiu JJ, et al. Aberrant expression of long noncoding RNAs in colorectal cancer with liver metastasis. Tumour Biol. 2015;36(11):8747–8754. doi:10.1007/s13277-015-3627-4
186. Shan Y, Ma J, Pan Y, Hu J, Liu B, Jia L. LncRNA SNHG7 sponges miR-216b to promote proliferation and liver metastasis of colorectal cancer through upregulating GALNT1. Cell Death Dis. 2018;9(7):722. doi:10.1038/s41419-018-0759-7
187. Zhang P, Shi L, Song L, et al. LncRNA CRNDE and lncRNA SNHG7 are promising biomarkers for prognosis in synchronous colorectal liver metastasis following hepatectomy. Cancer Manag Res. 2020;12:1681–1692. doi:10.2147/CMAR.S233147
188. Huang L, Lin H, Kang L, et al. Aberrant expression of long noncoding RNA SNHG15 correlates with liver metastasis and poor survival in colorectal cancer. J Cell Physiol. 2019;234(5):7032–7039. doi:10.1002/jcp.27456
189. Zhu X, Bu F, Tan T, et al. Long noncoding RNA RP11-757G1.5 sponges miR-139-5p and upregulates YAP1 thereby promoting the proliferation and liver, spleen metastasis of colorectal cancer. J Exp Clin Cancer Res. 2020;39(1):207. doi:10.1186/s13046-020-01717-5
190. Wang L, Wei Z, Wu K, et al. Long noncoding RNA B3GALT5-AS1 suppresses colon cancer liver metastasis via repressing microRNA-203. Aging (Albany NY). 2018;10(12):3662–3682. doi:10.18632/aging.101628
191. Bleau AM, Redrado M, Nistal-Villan E, et al. miR-146a targets c-met and abolishes colorectal cancer liver metastasis. Cancer Lett. 2018;414:257–267. doi:10.1016/j.canlet.2017.11.008
192. Iguchi T, Nambara S, Masuda T, et al. miR-146a polymorphism (rs2910164) predicts colorectal cancer patients’ susceptibility to liver metastasis. PLoS One. 2016;11(11):e0165912. doi:10.1371/journal.pone.0165912
193. Lai H, Zhang J, Zuo H, et al. Overexpression of miR-17 is correlated with liver metastasis in colorectal cancer. Medicine (Baltimore). 2020;99(9):e19265. doi:10.1097/MD.0000000000019265
194. Zhang Y, He X, Liu Y, et al. microRNA-320a inhibits tumor invasion by targeting neuropilin 1 and is associated with liver metastasis in colorectal cancer. Oncol Rep. 2012;27(3):685–694. doi:10.3892/or.2011.1561
195. Sun L, Liu X, Pan B, et al. Serum exosomal miR-122 as a potential diagnostic and prognostic biomarker of colorectal cancer with liver metastasis. J Cancer. 2020;11(3):630–637. doi:10.7150/jca.33022
196. Vychytilova-Faltejskova P, Pesta M, Radova L, et al. Genome-wide microRNA expression profiling in primary tumors and matched liver metastasis of patients with colorectal cancer. Cancer Genomics Proteomics. 2016;13(4):311–316.
197. Iino I, Kikuchi H, Miyazaki S, et al. Effect of miR-122 and its target gene cationic amino acid transporter 1 on colorectal liver metastasis. Cancer Sci. 2013;104(5):624–630. doi:10.1111/cas.12122
198. Takeyama H, Yamamoto H, Yamashita S, et al. Decreased miR-340 expression in bone marrow is associated with liver metastasis of colorectal cancer. Mol Cancer Ther. 2014;13(4):976–985. doi:10.1158/1535-7163.MCT-13-0571
199. Cristóbal I, Rincón R, Manso R, et al. Deregulation of miR-200b, miR-200c and miR-429 indicates its potential relevant role in patients with colorectal cancer liver metastasis. J Surg Oncol. 2014;110(4):484–485. doi:10.1002/jso.23661
200. Yin J, Bai Z, Song J, et al. Differential expression of serum miR-126, miR-141 and miR-21 as novel biomarkers for early detection of liver metastasis in colorectal cancer. Chin J Cancer Res. 2014;26(1):95–103. doi:10.3978/j.issn.1000-9604.2014.02.07
201. Luo HY, Yang Z, Wei W, et al. Enzymatically synthesized poly(amino-co-ester) polyplexes for systemic delivery of pcDNA-miRNA-214 to suppress colorectal cancer liver metastasis. J Mater Chem B. 2018;6(40):6365–6376. doi:10.1039/C8TB01932K
202. Cristóbal I, Caramés C, Madoz-Gúrpide J, Rojo F, Aguilera O, García-Foncillas J. Downregulation of miR-214 is specific of liver metastasis in colorectal cancer and could play a role determining the metastatic niche. Int J Colorectal Dis. 2014;29(7):885. doi:10.1007/s00384-014-1872-4
203. Chen DL, Wang ZQ, Zeng ZL, et al. Identification of microRNA-214 as a negative regulator of colorectal cancer liver metastasis by way of regulation of fibroblast growth factor receptor 1 expression. Hepatology. 2014;60(2):598–609. doi:10.1002/hep.27118
204. Cui Z, Tang J, Chen J, Wang Z. Hsa-miR-574-5p negatively regulates MACC-1 expression to suppress colorectal cancer liver metastasis. Cancer Cell Int. 2014;14:47. doi:10.1186/1475-2867-14-47
205. Ji D, Chen Z, Li M, et al. MicroRNA-181a promotes tumor growth and liver metastasis in colorectal cancer by targeting the tumor suppressor WIF-1. Mol Cancer. 2014;13:86. doi:10.1186/1476-4598-13-86
206. Li W, Chang J, Wang S, et al. miRNA-99b-5p suppresses liver metastasis of colorectal cancer by down-regulating mTOR. Oncotarget. 2015;6(27):24448–24462. doi:10.18632/oncotarget.4423
207. Cristóbal I, Torrejón B, González-Alonso P, Manso R, Rojo F, García-Foncillas J. Downregulation of miR-138 as a contributing mechanism to Lcn-2 overexpression in colorectal cancer with liver metastasis. World J Surg. 2016;40(4):1021–1022. doi:10.1007/s00268-015-3241-z
208. Jiang H, Liu J, Chen Y, Ma C, Li B, Hao T. Up-regulation of mir-10b predicate advanced clinicopathological features and liver metastasis in colorectal cancer. Cancer Med. 2016;5(10):2932–2941. doi:10.1002/cam4.789
209. Kingham TP, Nguyen H, Zheng J, et al. MicroRNA-203 predicts human survival after resection of colorectal liver metastasis. Oncotarget. 2017;8(12):18821–18831. doi:10.18632/oncotarget.13816
210. Murakami T, Kikuchi H, Ishimatsu H, et al. Tenascin C in colorectal cancer stroma is a predictive marker for liver metastasis and is a potent target of miR-198 as identified by microRNA analysis. Br J Cancer. 2017;117(9):1360–1370. doi:10.1038/bjc.2017.291
211. Makondi PT, Wei PL, Huang CY, Chang YJ. Development of novel predictive miRNA/target gene pathways for colorectal cancer distance metastasis to the liver using a bioinformatic approach. PLoS One. 2019;14(2):e0211968. doi:10.1371/journal.pone.0211968
212. Liu J, Li H, Sun L, et al. Epigenetic alternations of microRNAs and DNA methylation contribute to liver metastasis of colorectal cancer. Dig Dis Sci. 2019;64(6):1523–1534. doi:10.1007/s10620-018-5424-6
213. Sahu SS, Dey S, Nabinger SC, et al. The role and therapeutic potential of miRNAs in colorectal liver metastasis. Sci Rep. 2019;9(1):15803. doi:10.1038/s41598-019-52225-2
214. Wang D, Wang X, Si M, et al. Exosome-encapsulated miRNAs contribute to CXCL12/CXCR4-induced liver metastasis of colorectal cancer by enhancing M2 polarization of macrophages. Cancer Lett. 2020;474:36–52. doi:10.1016/j.canlet.2020.01.005
215. Sakai H, Sato A, Aihara Y, et al. MKK7 mediates miR-493-dependent suppression of liver metastasis of colon cancer cells. Cancer Sci. 2014;105(4):425–430. doi:10.1111/cas.12380
216. Geng L, Chaudhuri A, Talmon G, et al. MicroRNA-192 suppresses liver metastasis of colon cancer. Oncogene. 2014;33(46):5332–5340. doi:10.1038/onc.2013.478
217. Zhao S, Mi Y, Guan B, et al. Tumor-derived exosomal miR-934 induces macrophage M2 polarization to promote liver metastasis of colorectal cancer. J Hematol Oncol. 2020;13(1):156. doi:10.1186/s13045-020-00991-2
218. Machackova T, Vychytilova-Faltejskova P, Souckova K, et al. MiR-215-5p reduces liver metastasis in an experimental model of colorectal cancer through regulation of ECM-receptor interactions and focal adhesion. Cancers (Basel). 2020;12(12):3518. doi:10.3390/cancers12123518
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
