Back to Journals » Infection and Drug Resistance » Volume 13
Coinfection of SARS-CoV-2 and Other Respiratory Pathogens
Authors Ma L, Wang W, Le Grange JM , Wang X, Du S, Li C, Wei J, Zhang JN
Received 11 June 2020
Accepted for publication 8 August 2020
Published 26 August 2020 Volume 2020:13 Pages 3045—3053
DOI https://doi.org/10.2147/IDR.S267238
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Sahil Khanna
Ling Ma,1,* Wenjing Wang,2,* Jehane Michael Le Grange,2 Xiaorong Wang,3 Shuaixian Du,1 Chen Li,1 Jia Wei,4 Jin-Nong Zhang2
1Department of Clinical Laboratory, Wuhan Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, People’s Republic of China; 2Department of Emergency Medicine, Wuhan Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, People’s Republic of China; 3Department of Respiratory and Critical Care Medicine, Union Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430022, People’s Republic of China; 4Department of Hematology, Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, Hubei 430030, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Jin-Nong Zhang; Jia Wei Email [email protected]; [email protected]
Purpose: To differentiate between respiratory infections caused by SARS-CoV-2 and other respiratory pathogens during the COVID-19 outbreak in Wuhan, we simultaneously tested for SARS-CoV-2 and pathogens associated with CAP to determine the incidence and impact of respiratory coinfections in COVID-19 patients.
Patients and Methods: We included 250 patients who were diagnosed with COVID-19. RT-PCR was used to detect influenza A, influenza B and respiratory syncytial viruses. Chemiluminescence immunoassays were used to detect IgM antibodies for adenovirus, Chlamydia pneumoniae and Mycoplasma pneumoniae in the serum of patients. Based on these results, we divided the patients into two groups, the simple SARS-CoV-2-infected group and the coinfected SARS-COV-2 group. Coinfected patients were then further categorized as having a coinfection of viral pathogen (CoIV) or coinfection of atypical bacterial pathogen (CoIaB).
Results: No statistically significant differences were found in age, gender, the time taken to return negative SARS-CoV-2 nucleic acid test results, length of hospital stays, and mortality between the simple SARS-CoV-2 infection group and the coinfection group. Of the 250 hospitalized COVID-19 patients, 39 (15.6%) tested positive for at least one respiratory pathogen in addition to SARS-CoV-2. A third of these pathogens were detected as early as the 1st week after symptom onset and another third were identified after more than three weeks. The most detected CAP pathogen was C. pneumoniae (5.2%), followed by the respiratory syncytial virus (4.8%), M. pneumoniae (4.4%) and adenovirus (2.8%). Patients coinfected with viral pathogens (CoIV) (n=18) had longer hospital stays when compared to patients coinfected with atypical bacterial pathogens (CoIaB) (n=21). Except for one fatality, the remaining 38 coinfected patients all recovered with favourable outcomes.
Conclusion: Coinfections in COVID-19 patients are common. The coinfecting pathogens can be detected at variable intervals during COVID-19 disease course and remain an important consideration in targeted treatment strategies for COVID-19 patients.
Keywords: SARS-CoV-2, COVID-19, viral coinfection, atypical bacterial coinfection
Introduction
Having started in December 2019, the COVID-19 pandemic continues to pose a serious and perilous global health burden. This is the third time that a coronavirus is responsible for widespread human infection, with the current culprit being named by the World Health Organization (WHO) as SARS-CoV-2. Although the number of confirmed global cases of COVID-19 now exceeds 16 million, as of July 29, and several retrospective observational studies have noted that coinfection with other respiratory pathogens is relatively common,1–4 the clinical features of coinfection and its impact on patient outcomes, is yet to be clarified.
Similar to influenza virus pneumonia,5 the pulmonary structure is severely damaged during a SARS-CoV-2 infection. This has been observed as diffuse alveolar damage in several autopsy findings,6,7 the invasion of viral particles in bronchial mucosal and alveolar epithelia, the destruction and shedding of the epithelium and excessive exudate accumulation in the bronchiole lumens and alveolar spaces of the patients. These pathologic findings would explain why COVID-19 patients are predisposed to coinfection with common respiratory pathogens, as discussed in some earlier clinical studies,1–4 and postmortem reports.8 As such, coinfection with common respiratory pathogens in COVID-19 patients could potentially have an impact on their clinical management, disease progression and outcomes.
To differentiate between respiratory infections caused by SARS-CoV-2 and those caused by other respiratory pathogens during the COVID-19 outbreak in Wuhan, we simultaneously tested for SARS-CoV-2, common respiratory viruses, and atypical respiratory bacteria. We paid specific attention to the timing of the detection and identification of the coinfecting pathogens, clinical features of the potential coinfection, and the impact of the coinfection on patient outcomes.
Patients and Methods
Patients and Allocation
We included 250 patients diagnosed with COVID-19, who had visited the fever clinic at Wuhan Union Hospital (WHUH) due to an acute fever or respiratory symptoms between Jan 19, 2020, and Feb 26, 2020. All these patients were tested for SARS-CoV-2, respiratory syncytial virus, influenza A virus, influenza B virus, adenovirus, Chlamydia pneumoniae, and Mycoplasma pneumoniae, using sputum or nasopharyngeal swab specimens collected in the interval between the onset of symptoms, and up to seven days after their hospital admission. The patients were admitted into the isolation wards of the infectious disease department once they tested positive for SARS-CoV-2 or were suspected of having COVID-19 based on the characteristic viral pneumonia pattern on chest CT scans (their diagnoses were later confirmed by either repeated positive ribonucleic acid (RNA) tests for SARS-CoV-2 or positive serological conversion of SARS-CoV-2 IgM and/or IgG antibodies). The diagnosis of COVID-19 and classification of disease severity was conducted in accordance with the American Centers for Disease Control and Prevention Interim Guidance.9
Firstly, we divided the 250 patients into two groups: a simple SARS-CoV-2 infection group (n=211), infected only with SARS-COV-2, and a coinfection group (n=39), infected with at least one additional respiratory pathogen. Then, we further divided the 39 coinfected patients into two groups, based on whether they were coinfected with a viral pathogen (CoIV group, n=18) or coinfected with an atypical bacterial pathogen (CoIaB group, n=21). This was based on the pathogens that we identified in addition to SARS-CoV-2, and we aimed to determine not only the clinical characteristics but also the impact that these coinfections had on disease progression and patient outcomes. The patient demographic data, disease severity and outcome, laboratory investigations, and therapeutic regimens, were collected by two physicians and verified by a senior physician.
The Medical Ethics Committee of Wuhan Union Hospital (WHUH) approved this study, which complies with the Declaration of Helsinki. The Medical Ethics Committee waived written informed consent because the etiological screening test is a clinical routine for respiratory infection in WHUH, we obtained the patients’ oral consent. Furthermore, the study was observational, and all the patients’ identities were concealed.
Laboratory Tests
Routine laboratory tests, including tests for SARS-CoV-2 and other common respiratory viral and atypical bacterial pathogens, routine blood investigations, coagulation studies, organ function tests and inflammatory biomarkers, such as c-reactive protein (CRP) and procalcitonin (PCT), were taken at the time of patient presentation, while the serum interleukin (IL)-2, IL-4, IL-6, IL-10, TNF-α and IFN-γ levels were obtained on the 2nd day of admission. These tests were selectively repeated as needed or at the time of follow-up, usually at 1-week and 2-week intervals post-admission and again prior discharge. In the interest of biosafety, we did not submit respiratory samples for bacterial or viral cultures.
The RNA of SARS-CoV-2 in nasopharyngeal swab samples was extracted according to the instructions of the RNA isolation kit (Xi’an Tianlong Science and Technology Co. Ltd, China). The reverse transcription-polymerase chain reaction (RT-PCR) assay (Shanghai Pfizer Biotechnology Co., Ltd., China) for SARS-CoV-2 was conducted by amplifying two target genes, namely the open reading frame 1ab (ORF1ab) and nucleocapsid protein. The swab samples were also tested for influenza A virus, influenza B virus and respiratory syncytial virus (RSV) RNA with the Xpress Flu/RSV Assay (Cepheid, USA) using GeneXpert Dx System (Cepheid, USA). Adenovirus-IgM antibodies, C. pneumoniae-IgG/IgM antibodies, M. pneumoniae-IgG/IgM antibodies, as well as SARS-CoV-2-IgG/IgM antibodies in the patients’ serum, were measured with Chemiluminescence Immunoassays (test kits were purchased from iFLASH3000, YHLO Biotech and Snibe Diagnostic, respectively), and values greater than 1.10 AU/mL were considered positive. Any result deemed positive for the presence of IgM antibodies, was considered as being indicative of a current infection for that pathogen.
Serum interleukin (IL)-2, IL-4, IL-6, IL-10, TNF-α and IFN-γ levels were measured using a cytometric Bead Array TM kit (CBA, BD Biosciences, San Jose, CA, USA). Antibodies against the human surface and intracellular molecules were purchased commercially. The total number of lymphocytes in the peripheral blood was determined using a hemocytometer, and the fractions of the lymphocyte subsets, including CD4+ T-cells, CD8+ T-cells, B-cells and NK-cells, in the peripheral blood, were determined using the FACSCanton II flow cytometric system (BD, USA) and the BD FACS Diva software (BD, USA).
Post-Discharge Follow-Up
Patients were only deemed suitable for hospital discharge when their clinical symptoms were well controlled, and their SARS-CoV-2 nucleic acid test result was negative on two consecutive occasions. Patients were advised to attend a hospital follow-up 2 to 4 weeks after hospital discharge. A chest CT scan, SARS-CoV-2 nucleic acid tests of nasopharyngeal swab samples and serum IgG/IgM antibody tests were recommended.
Statistical Analysis
Descriptive analyses of the variables were expressed as the median and interquartile range (IQR), or number (%), where appropriate. Categorical data were compared using the Fisher’s exact test, or Χ2 test. Non-normal distributed continuous data were compared using the Mann–Whitney U-test. P<0.05 was defined as statistically significant. All analyses were performed with SPSS, version 25.0 (IBM SPSS) and GraphPad Prism8.0.2 (GraphPad Software Inc., San Diego, CA, USA).
Results
Demographic and Baseline Characteristics of the Patients
Two hundred and fifty patients hospitalized with confirmed COVID-19 in WHUH between Jan 19, 2020, and Feb 26, 2020, were included in the study. Among them, 211 patients (84.4%) were infected with only SARS-CoV-2, and 39 patients (15.6%) were coinfected with at least one additional respiratory pathogen. No statistically significant difference was found in gender and age, between the simple SARS-CoV-2 group and the coinfection group, however, the coinfection group included a higher proportion of severe and critically ill patients, and a lower proportion of mild and moderate patients than the simple SARS-CoV-2 infection group (Supplement Table 1). Further analysis of the coinfection subgroups revealed that patients in the CoIV group (n=18) had no significant difference in age, gender, medical history, comorbidities, onset-symptoms (with the exception of fatigue) and disease severity upon initial evaluation, when compared to the CoIaB group (n=21) (Table 1).
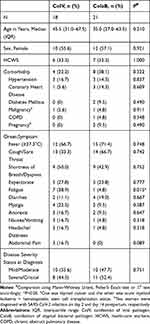 |
Table 1 Demographic and Baseline Characteristics |
CAP Pathogens Identified
Of the 250 COVID-19 patients, an additional respiratory pathogen was detected in 13.6% (34/250) of patients, while in 2.0% (5/250) of patients, two additional respiratory pathogens were detected. The most commonly detected pathogen was C. pneumoniae (5.2%) (13/250), followed by RSV (4.8%), M. pneumoniae (4.4%), adenovirus (2.8%), influenza A virus (0.8%) and influenza B virus (0.4%) (Table 2). In certain patients (16/39), positive test results for the detection of other respiratory pathogens were obtained prior to the patient testing positive for SARS-CoV-2. As shown in Figure 1, the coinfecting pathogens (CoIPs) were often detected at variable intervals from symptom onset. They were detected either early after symptom onset or, in some cases, much later, even during the COVID-19 recovery phase. This variability was illustrated by CoIPs being detected during the 1st week after symptom onset in a third (13/39) of the coinfected patients, while CoIPs were detected after more than three weeks after symptom onset in another third (12/39) of the coinfected patients, when most patients were already in the recovery phase of COVID-19.
 |
Table 2 Coinfections in SARS-CoV-2 Patients and the Additional Respiratory Pathogens Identified |
 |
Figure 1 Timeline of the detection of respiratory pathogens in patients with a coinfection. |
Laboratory Variables on the Day of Hospital Admission
Laboratory values for white blood cells (WBC), lymphocytes, neutrophils, c-reactive protein (CRP), and procalcitonin (PCT), done at the time of admission, were not statistically significant between the CoIV and CoIaB groups (Table 3). Patients in the CoIaB group demonstrated higher serum levels of IL-2, IL-4, and TNF-α than those in the CoIV group (P<0.05) during their disease course (Table 3). The laboratory values of the total T-cells, CD4+ T-cells, CD8+ T-cells, and NK-cells decreased below the normal range in more than 50% (data not shown) of the patients during their disease course, irrespective of whether they were in the CoIV or CoIaB groups (P > 0.05). B-cell values remained within the normal range in most patients, with no significant inter-group variance.
 |
Table 3 Laboratory Findings on Admission |
Treatment and Outcome
In accordance with our previous publication,10 the core therapeutic regimen utilized in our facility for COVID-19 consisted of a combination of Arbidol (Umifenovir) and antimicrobials recommended in the guidelines for managing adult community-acquired pneumonia (CAP).11 When comparing the antiviral treatment between the simple SARS-CoV-2 and coinfection groups, or between the CoIV and CoIaB groups, there was a similar rate of initiation of antiviral treatment (Supplement Table 1 and Table 1). There were no statistically significant differences in the time taken to return negative SARS-CoV-2 nucleic acid test results, length of hospital stays, and SARS-CoV-2 IgG antibody values, between the simple SARS-CoV-2 infection group and the coinfection group (Supplement Table 1). However, lower SARS-CoV-2 IgM values and delayed antibody production from the time of symptom onset were observed in the coinfection group when compared to the simple SARS-CoV-2 infection group (Supplement Table 1). The conversion rate of patients returning negative SARS-CoV-2 nucleic acid test results within 2 weeks, was also similar between the CoIV and CoIaB groups (27.8% versus 28.6%, P>0.05) (Table 4). Although more patients in the CoIV group received corticosteroids (22.2% vs 9.5%) and intravenous immunoglobulin (IVIG) (38.9% vs 9.5%) treatment than in the ColaB group, it was not statistically significant (P>0.05). On average, patients in the CoIV group had a longer hospital stay (median of 24 vs 15 days, P<0.05) than those in the CoIaB group (Table 4).
 |
Table 4 Treatment and Outcomes Between Coinfection of Viral Pathogens and Coinfection of Bacterial Pathogens |
Of the 211 patients in the simple SARS-CoV-2 infection group, there were three fatalities, and of the 39 patients in the co-infection group, there was one fatality, with no statistically significant difference in mortality between the two groups (Supplement Table 1). Except for the one fatality in the CoIV subgroup of coinfected patients, the remaining 38 coinfected patients survived, with 37 of them returning for follow-up consultations, with a median time to follow-up of 38 days (IQR 32.0–50.5 days). SARS-CoV-2 IgM and/or IgG antibodies were detected in 36 of these patients, with no significant difference in antibody levels between the two groups (P>0.05, Table 4).
Discussion
Considering that even physiologically normal lungs are not sterile,12,13 it is important to note that the detection of a pathogen in patient’s respiratory secretions does not necessarily constitute an infection. Despite defining the detection of additional respiratory pathogens in COVID-19 patients as a coinfection in our study, we believe it would be more appropriate to consider these cases as potential coinfections. To truly define the role that these coinfecting pathogens play in the pathogenesis of COVID-19 is exceedingly difficult, despite bacterial coinfections commonly occurring during other viral respiratory infections. This type of coinfection is not only very difficult to prevent or control but also aggravates the underlying viral infection, as is often seen in influenza pneumonia.14,15
It has been determined that upper respiratory symptoms, caused by one pathogen, may enhance the transmission of another pathogen through aerosol production, and that disease transmission may be altered by the interaction between two infections.16–18 Whether there is a similar type of interaction between SARS-CoV-2 and other respiratory pathogens remains unclear. In our study, coinfections were not associated with adverse mortality rates when compared to simple SARS-CoV-2 infections alone, which is consistent with previous studies.3,19 However, the detection rate of other pathogens was 15.6% in our study, while other studies20,21,23 had a higher detection rate, suggesting that coinfection may be a common feature during the COVID-19 pandemic. When managing COVID-19 patients, being aware of the presence of another respiratory pathogen causing coinfection plays an important role in assisting health-care workers in their use of targeted medications and therapies, aimed at treating these potential pathogens.
The rate of the positive detection of CoIPs in our COVID-19 patient population is approximately 16% (39/250). This is slightly less than what has been reported in other comparable literature, such as the study conducted by Kim in the USA, where out of 116 SARS-CoV-2 positive patients, 20% were also positive for other respiratory viruses.3
In comparison to Kim’s USA study, where neither M. pneumoniae nor C. pneumoniae were detected, the patients in Wuhan seemed to be predisposed to M. pneumonia and C. pneumonia coinfection. In our study, the positive rate for anti-C. pneumoniae IgM and anti-M. pneumoniae IgM detection was 5.2% and 4.4%, respectively, while in a retrospective study, based on fatal COVID-19 cases in Wuhan,22 the rates of detection for these specific pathogens were determined to be even greater, at 34.1% and 26.5%, respectively. RSV is another noteworthy pathogen. The positive rate of RSV detection was 5.2% in Kim’s study and 4.8% in our study. The positive rate of adenovirus detection was 2.8% in our study, while it remained at zero in Kim’s study. We did not detect any other coronaviridae during our study; however, in Kim’s study, a detection rate of 4.3% was observed for other coronaviridae pathogens. Coinfection with influenza A and B virus was relatively low (less than 1%), not only in our study but also in Kim’s study. However, in the study on fatal COVID-19 cases in Wuhan, rates of 9.1% and 5.3% for influenza A and B virus were observed, respectively. During an investigation conducted in the Jiangsu province of China, aimed at detecting 39 different respiratory pathogens among 257 confirmed COVID-19 patients, bacterial detection was far more prevalent than viral detection (96.2% versus 14.1%), and included C. pneumonia (2.5%), M. pneumonia (1.6%), adenovirus (3.9%), influenza B virus (1.9%) and influenza A virus (0.8%); however, RSV was not detected in this specific study.23
The timing of these co-infections could vary from very early in the onset of respiratory symptoms, to later in the recovery stage of COVID-19, at approximately 2–3 weeks. This should serve as a reminder that a non-SARS-CoV-2 pathogen infection could be detected both prior to a SARS-CoV-2 infection, or after a SARS-CoV-2 infection, with the causal relationship between the two, yet to be determined.
PCT test results were unable to distinguish between viral coinfection and atypical bacterial coinfection in our study, and this conclusion is consistent with previous findings.24 Though coinfection with a respiratory virus or atypical bacteria did not demonstrate a preferential decrease in the number of T-, B- and NK-cells, patients coinfected with a viral pathogen exhibited a less remarkable inflammatory response when compared to patients with an atypical bacterial coinfection, illustrated by the relatively lower expression of IL-2, IL-4 and TNF-α in the CoIV group. Interestingly, patients in the CoIaB group had higher IL-2, IL-4 and TNF-α levels on admission than those in the CoIV group, yet patients in the CoIaB group had shorter hospital stays than those in the CoIV group. Although difficult to determine the exact reason for this, there was a statistically significant difference in azithromycin use between the CoIV and CoIaB groups, to control the atypical bacterial infection, which could perhaps account for this difference in hospital stay.
It remains important to note that our study has several limitations. Firstly, this is a retrospective observational study with a limited sample size, particularly in the case of the coinfection subgroups. Secondly, due to the medical demand and surge in the number of patients in the early stages of the outbreak, 34 patients who were only infected with SARS-CoV-2 were transferred to other designated hospitals or facilities, which were temporary hospitals that only accepted COVID-19 patients. As a result, only a brief follow-up was achieved, with long-term follow-up not being possible in these patients. A prospective study with a larger sample size, conducted in a COVID-19 pandemic-affected area, is warranted, so as to gain a better understanding of the relationship between SARS-CoV-2 and coinfections with other CAP-associated pathogens.
Conclusion
Coinfections in COVID-19 patients are common, yet no significant difference in patient outcome was observed between the simple SARS-CoV-2 and coinfection groups. Patients coinfected with viral pathogens experienced longer hospital stays than those coinfected with atypical bacteria. The coinfecting pathogens can be detected at variable intervals during COVID-19 disease course and remain an important consideration in targeted treatment strategies for COVID-19 patients.
Abbreviations
COVID-19, coronavirus disease 2019; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; CoIPs, coinfecting pathogens; CoIV, coinfection of viral pathogen; CoIaB, coinfection of atypical bacteria pathogen; CAP, community-acquired pneumonia; RSV, respiratory syncytial virus; M. pneumoniae, Mycoplasma pneumoniae; C. pneumoniae, Chlamydia pneumoniae.
Data Sharing Statement
The data supporting the findings of the article are available from the corresponding authors upon request.
Ethics approval
The Medical Ethics Committee of Wuhan Union Hospital (WHUH) approved this study which complies with the Declaration of Helsinki.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflict of interest in this work.
References
1. Duployez C, Le Guern R, Tinez C, et al. Panton-valentine leukocidin-secreting Staphylococcus aureus pneumonia complicating COVID-19. Emerg Infect Dis. 2020;26(8):1939–1941. doi:10.3201/eid2608.201413
2. Blaize M, Mayaux J, Nabet C, et al. Fatal invasive aspergillosis and coronavirus disease in an immunocompetent patient. Emerg Infect Dis. 2020;26(7):1636–1637. doi:10.3201/eid2607.201603
3. Kim D, Quinn J, Pinsky B, Shah NH, Brown I. Rates of co-infection between SARS-CoV-2 and other respiratory pathogens. JAMA. 2020;323(20):2085–2086. doi:10.1001/jama.2020.6266
4. Jiang S, Liu P, Xiong G, et al. Coinfection of SARS-CoV-2 and multiple respiratory pathogens in children. Clin Chem Lab Med. 2020;58(7):1160–1161. doi:10.1515/cclm-2020-0434
5. Shieh WJ, Blau DM, Denison AM, et al. 2009 pandemic influenza A (H1N1): pathology and pathogenesis of 100 fatal cases in the United States. Am J Pathol. 2010;177(1):166–175. doi:10.2353/ajpath.2010.100115
6. Hanley B, Lucas SB, Youd E, Swift B, Osborn M. Autopsy in suspected COVID-19 cases. J Clin Pathol. 2020;73(5):239–242. doi:10.1136/jclinpath-2020-206522
7. Yao XH, Li TY, He ZC, et al. [A pathological report of three COVID-19 cases by minimal invasive autopsies]. Zhonghua Bing Li Xue Za Zhi. 2020;49(5):411–417.
8. Barton LM, Duval EJ, Stroberg E, Ghosh S, Mukhopadhyay S. COVID-19 Autopsies, Oklahoma, USA. Am J Clin Pathol. 2020;153(6):725–733. doi:10.1093/ajcp/aqaa062
9. American centers for disease control and prevention (CDC) [homepage on the Internet]. Interim clinical guidance for management of patients with confirmed coronavirus disease (COVID-19). 2020. Available from: https://www.cdc.gov/coronavirus/2019-ncov/hcp/clinical-guidance-management-patients.html.
10. Zhang J, Zhou L, Yang Y, Peng W, Wang W, Chen X. Therapeutic and triage strategies for 2019 novel coronavirus disease in fever clinics. Lancet Respir Med. 2020;8(3):e11–e12. doi:10.1016/S2213-2600(20)30071-0
11. Metlay JP, Waterer GW, Long AC, et al. Diagnosis and treatment of adults with community-acquired pneumonia. An official clinical practice guideline of the American thoracic society and infectious diseases society of America. Am J Respir Crit Care Med. 2019;200(7):e45–e67. doi:10.1164/rccm.201908-1581ST
12. Lee KH, Gordon A, Foxman B. The role of respiratory viruses in the etiology of bacterial pneumonia: an ecological perspective. Evol Med Public Health. 2016;2016(1):95–109. doi:10.1093/emph/eow007
13. Beasley V, Joshi PV, Singanayagam A, Molyneaux PL, Johnston SL, Mallia P. Lung microbiology and exacerbations in COPD. Int J Chron Obstruct Pulmon Dis. 2012;7:555–569. doi:10.2147/COPD.S28286
14. McCullers JA. The co-pathogenesis of influenza viruses with bacteria in the lung. Nat Rev Microbiol. 2014;12(4):252–262. doi:10.1038/nrmicro3231
15. Cauley LS, Vella AT. Why is coinfection with influenza virus and bacteria so difficult to control? Discov Med. 2015;19(102):33–40.
16. Lee N, Chan PK, Yu IT, et al. Co-circulation of human metapneumovirus and SARS-associated coronavirus during a major nosocomial SARS outbreak in Hong Kong. J Clin Virol. 2007;40(4):333–337. doi:10.1016/j.jcv.2007.08.015
17. Bassetti S, Bischoff WE, Sherertz RJ. Are SARS superspreaders cloud adults? Emerg Infect Dis. 2005;11(4):637–638. doi:10.3201/eid1104.040639
18. Bassetti S, Sherertz RJ, Pfaller MA. Airborne dispersal of Staphylococcus aureus associated with symptomatic rhinitis allergica. Ann Intern Med. 2003;139(3):W60. doi:10.7326/0003-4819-139-3-200308050-00021-w1
19. Ding Q, Lu P, Fan Y, Xia Y, Liu M. The clinical characteristics of pneumonia patients coinfected with 2019 novel coronavirus and influenza virus in Wuhan, China. J Med Virol. 2020. doi:10.1002/jmv.25781
20. Ma S, Lai X, Chen Z, Tu S, Qin K. Clinical characteristics of critically ill patients co-infected with SARS-CoV-2 and the influenza virus in Wuhan, China. Int J Infect Dis. 2020;96:683–687. doi:10.1016/j.ijid.2020.05.068
21. Yue H, Zhang M, Xing L, et al. The epidemiology and clinical characteristics of co-infection of SARS-CoV-2 and influenza viruses in patients during COVID-19 outbreak. J Med Virol. 2020. doi:10.1002/jmv.26163
22. Du Y, Tu L, Zhu P, et al. Clinical features of 85 fatal cases of COVID-19 from Wuhan. A retrospective observational study. Am J Respir Crit Care Med. 2020;201(11):1372–1379. doi:10.1164/rccm.202003-0543OC
23. Zhu X, Ge Y, Wu T, et al. Co-infection with respiratory pathogens among COVID-2019 cases. Virus Res. 2020;285:198005. doi:10.1016/j.virusres.2020.198005
24. Metlay JP, Waterer GW. Treatment of community-acquired pneumonia during the coronavirus disease 2019 (COVID-19) pandemic. Ann Intern Med. 2020. doi:10.7326/M20-2189
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
