Back to Journals » OncoTargets and Therapy » Volume 14
Clinical Significance and Oncogenic Activity of GRWD1 Overexpression in the Development of Colon Carcinoma
Authors Zhou X , Shang J, Liu X, Zhuang JF , Yang YF, Zhang YY, Guan GX
Received 11 November 2020
Accepted for publication 26 January 2021
Published 2 March 2021 Volume 2021:14 Pages 1565—1580
DOI https://doi.org/10.2147/OTT.S290475
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Xin Zhou,1,* Jin Shang,2,* Xing Liu,1,* Jin-Fu Zhuang,1 Yuan-Feng Yang,1 Yi-Yi Zhang,1 Guo-Xian Guan1
1Department of Colorectal Surgery, The First Affiliated Hospital of Fujian Medical University, Fuzhou, 350005, Fujian, People’s Republic of China; 2The First Affiliated Hospital, School of Medicine, Xiamen University, Xiamen, 361102, Fujian, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Guo-Xian Guan
Department of Colorectal Surgery, The First Affiliated Hospital of Fujian Medical University, 20 Chazhong Road, Fuzhou, 350005, Fujian, People’s Republic of China
Tel +8613609592321
Email [email protected]
Objective: GRWD1 (glutamate-rich WD40 repeat containing 1) is a multifunctional protein involved in multiple cellular regulatory pathways, particularly those associated with cell growth control. GRWD1 is represented as a potential oncogene in several cancers, however, the function and mechanism of GRWD1 in the development of colon cancer are still unknown.
Materials and Methods: IHC was used to detect the expression of GRWD1 in colon carcinoma tissues. CCK-8, colony formation, and EdU were used to measure the cell proliferation after GRWD1 knockdown and overexpression. The distribution of the cell cycle was analyzed by flow cytometry. The effect of GRWD1 knockdown on migration and invasion was analyzed by wound healing and transwell assays.
Results: Overexpression of GRWD1 in colon carcinoma tissues was associated with pathological grading, tumor size, N stage, TNM stage, and poor survival. GRWD1 had high sensitivity and specificity in distinguishing colon cancer from noncancerous tissues, and might be served as an independent prognosis in colon carcinoma patients. Knockdown of GRWD1 significantly inhibited the cell proliferation and colony formation, and induced cell cycle arrest and more drug susceptibility, and suppressed the migration and invasion. GRWD1 exhibited these oncogenic activities might be associated with its regulation on the expression of PCNA and Ki67, Cyclin A2 and Cyclin B1, ABCC1 and GSTP1, MTA1 and MTA2.
Conclusion: GRWD1 may play an oncogenic activity in the development of colon carcinoma and its overexpression was associated with malignant characteristics and poor survival outcome of colon carcinoma. GRWD1 might be a potential target for future therapy.
Keywords: proliferation, cell cycle, drug susceptibility, migration and invasion, GRWD1, colon carcinoma
Introduction
Colorectal carcinoma (CRC) is the third commonly diagnosed cancer and one of the leading causes of cancer death. Besides, morbidity and mortality are both increasing in the most recent decade in many countries including China.1 Especially, colon carcinoma incidence is rising among young adults. According to the studies between 1975 and 2010, the incidence of young patients (<50 years old) was increasing by years.2 The mortality rates were predicted based on this trend, in 2030, the incidences for patients 20 to 34 years and 35 to 49 years will increase by 27.7% and 46%, respectively.3 Although early detection of colon cancer and surgery increased survival among patients, the poor early diagnosis and high rates of metastasis have significantly decreased the 5-year survival rates.4,5 Therefore, it is necessary to further investigate the pathogenesis of colon carcinoma so that we have to find some new strategies for diagnosis and treatment to reduce the mortality.
GRWD1 (glutamate-rich WD40 repeat containing 1), highly conserved throughout eukaryotes, is likely to be involved in ribosome biosynthesis.6 Previous studies suggested that p53 could be activated through the RPL11-MDM2 axis. GRWD1 could negatively regulate the expression of p53 through inhibiting the interaction of RPL11 and MDM2. A recent study showed that GRWD1 could also inhibit p53 by directly binding to its DNA-binding domain. Based on these results, GRWD1 could be a potential oncogene.7–9
It has been confirmed that GRWD1 was highly expressed in non-small cell lung cancer (NSCLC) and related to poor differentiation and prognosis.10 PiHL, a transcriptional target of p53, played an important role in CRC cell proliferation by inducing p53 ubiquitination through promoting the complex formation of GRWD1 and RPL11.11 However, the expression and function of GRWD1 in colon carcinoma are still unknown. Here we show that the upregulation of GRWD1 promotes the development of colon carcinoma and predicts a poor prognosis.
Materials and Methods
Tissue Samples and Clinical Data
A tissue microarray (TMA) purchased from Shanghai Xin chao Biotech Co. Ltd was used for this study. The study was approved by the Medical Ethics Committee of The First Affiliated Hospital of Xiamen University in accordance with the ethical guidelines of the 1975 Declaration of Helsinki. This TMA contains 94 formalin-fixed and paraffin-embedded primary colon carcinoma specimens and 86 matched adjacent noncancerous colon specimens. The matched adjacent noncancerous tissues were obtained from a segment of the resected specimens that were >5cm away from the tumor. None of the patients received preoperative radiation or chemotherapy. The clinicopathological data were summarized in Table 2.
 |
Table 1 Expression of GRWD1 Protein in Different Colon Tissues |
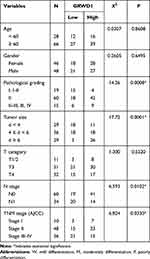 |
Table 2 Associations Between GRWD1 and Clinicopathological Factors of Colon Carcinoma Patients |
Immunohistochemistry (IHC)
4μm-thick sections of Paraffin-embedded tissues were dewaxed in xylene, then rehydrated and heated into 0.01M citrate buffer for 20 min. The samples were quenched the endogenous peroxidase activity by 0.3% H2O2. After blocking with goat serum for 1h, samples were incubated with anti-GRWD1 antibody (1:500 dilution; HPA042643; Sigma, America) at 4°C overnight. The positive signals were visualized via the DAB kit (Maxin Bio, Fuzhou, China) and then scored. The scoring was assessed based on the percentage of positive cells and intensity of the staining in GRWD1. Staining intensity was scored as follows: -, 10%; +, 10%-25%; ++, 26–50%; +++, >50%. All scores were subdivided into two categories: GRWD1 high expression (++, +++) and low expression (-, +). Scoring was performed by two independent individuals to avoid subjectivity.
Cell Culture and Transfection
The human CRC cell lines RKO and SW480 were purchased from the Cell Bank of Type Culture Collection of Chinese Academy of Sciences (Shanghai, China). Cells were cultured at 37°C with 5% CO2 in DMEM medium, supplemented with 10% FBS (Peakserum, US) and 100U/mL penicillin. Plasmids of knockdown or overexpression of GRWD1 were constructed by Public Protein/Plasmid Library. HEK293T cells were transfected by the resulting lentiviral vector and packing plasmids with Lipofectamine 2000 (Invitrogen). Medium was replaced after 8h, later supernatants were collected and filtered after 48h. RKO and SW480 cells were infected by viral supernatant and selected by puromycin.
Western Blot Analysis
Cells were collected and lysed with ice-cold lysis buffer, the extractives were centrifuged and supernatant was diluted 1:4 with protein loading buffer. 10μg of proteins were separated by SDS-PAGE and transferred to PVDF membranes. Following blocking with 5% nonfat milk at room temperature for 1h, membranes were incubated with primary antibodies at 4°C overnight. After washing with PBST for 3 times, the membranes were incubated with the horseradish peroxidase-conjugated secondary antibodies at room temperature for 1h. Finally, an enhanced chemiluminescence system was used to detect the signals. The primary antibody GRWD1 was purchased from Sigma-Aldrich, α-actinin was obtained from Proteintech.
Cell Proliferation Assay and Drug Susceptibility Assay
CCK8 assay (MCE) was used to detect the cell proliferation and cell survival after drug treatment. Cells were seeded with a density of 5×103/well in 96-well plates and cultured for a certain period time. 10μL CCK-8 reagent was added into each well and incubated for 3h.
Xenograft Nude Mouse Model
All experimental procedures involving animals were conducted in accordance with animal protocols of the Laboratory Animal Center of Xiamen University, which were approved by the Animal Ethics Committee of Xiamen University. 4–6 weeks old Balb/c nude mice were separated randomly into two groups including control group and the GRWD1 knockdown group. 1×107 cells were subcutaneously injected into the right side of each mouse. The tumor sizes were measured every two days. All mice were euthanized on the 19th day and tumors were dissected.
Colony Formation Assay
RKO and SW480 cell lines were seeded on 6-well plates with 500 cells/well, respectively. Cells were cultured for two weeks with 5% CO2 at 37°C, then were treated with methanol for 10 minutes and stained with crystal violet for 15 minutes. The cell colonies were photographed under the microscope.
EdU Incorporation Assay
EdU assay (RiboBio, China) was used to measure cell proliferation. EdU is an analog of thymidine, it could replace thymidine to insert in synthetic DNA molecule during DNA duplicate phase. Briefly, cells were seeded at a density of 1×104/well onto 96-well plates, after cultured for 24h, cells were treated with 50μM EdU for 2h with 5% CO2 at 37°C. Then, cells were washed with PBS twice to remove residual EdU and fixed with 4% formaldehyde for 30 min. PBS with 0.5% Triton X-100 was used to permeate cells. Next, cells were treated with 1× Apollo® fluorochrome for 30 min at a dark place. After that, each well was added with DAPI and 1× Hoechst 3342 reaction cocktail. Results were observed with a fluorescence microscope.
Cell Cycle Analysis
RKO and SW480 cells were treated with serum-free medium for 24h, then cells were softly collected and fixed with cold 70% ethanol at 4°C overnight. After that, cells were washed in cold PBS twice, and then resuspended in Cell Staining Buffer containing 2.5μg/mL propidium iodide solution and 100μg/mL RNase for 30 minutes at 37°C。DNA content were analyzed by Flow Cytometry. The percentage of cells at each cell cycle phase was analyzed using the FlowJo software.
Migration and Invasion Assays
Wound-healing assay and transwell assay were used to measure the activities of migration and invasion. For wound-healing assay, SW480 and RKO cells were seeded into 35-mm dishes and cultured for 24h to 95% confluence. 200μL pipette tip was used to scratch the monolayer and the well was washed with PBS for three times to wipe off the detached cells. After that, cells were cultured with DMEM medium containing 1% FBS. The same vision was captured under the microscope at 0h and 48h. For transwell assay, Corning® BioCoat™ Matrigel® Invasion Chambers with 8.0 µm PET Membrane were used according to the manufacturer’s instruction. SW480 and RKO cells were seeded in the upper chamber with a serum-free medium. The lower chamber was supplemented with DMEM containing 10% FBS. After 48h, invaded cells were stained with crystal violet and counted. The migration assay was performed without Matrigel and other procedures were the same with invasion assay.
Statistical Analysis
SPSS 16.0 statistic software (SPSS Inc., Chicago, IL, United States) was used to analyze all data. All survival rate was analyzed by the Kaplan-Meier method using a long-rank test. Student’s t-test was employed to compare the differences between the two groups. The Χ2 was used to analyze the relationship between GRWD1 expression and clinical features. The value of P < 0.05 was considered statistically significant.
Results
Overexpression of GRWD1 in Colon Cancer Tissues
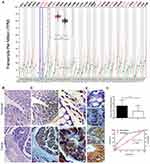
Evaluation from the online GEPIA database indicated that GRWD1 mRNA was widely expressed in many cancer tissues, but it was significantly highly expressed in COAD tissues compared with normal tissues (P < 0.05) (Figure 1A). In order to confirm the expression of GRWD1 in the colon cancer tissues, IHC was applied to detect the GRWD1 protein level in clinical colon cancer tissues and noncancerous tissues. The representative H&E staining images of noncancerous tissue and colon cancer tissue were shown in Figure 1B. IHC analysis showed that GRWD1 protein was weakly expressed in noncancerous tissues, while markedly expressed in the nucleus of colon cancer tissues (Figure 1C). To further investigate the expression of GRWD1 protein, the staining intensity was separated into four different grades including negative expression (-), weakly positive (+), moderately positive (2+), and highly positive (3+) (Figure 1D). The results from Table 1 showed that the rate of high expression (2+, 3+) of GRWD1 in colon cancer tissues was 58.6%, which was significantly higher than noncancerous tissues with 18.7%. The histogram of the score also showed staining intensity of GRWD1 in colon cancer tissues was pretty higher than that of noncancerous tissues (Figure 1E, upper). In addition, we assessed the sensitivity and specificity of GRWD1 expression to discriminate cancer tissues from noncancerous tissues. The value of area under the curve (AUC) was 0.7332 (95% CI = 0.6596 to 0.8067, P < 0.0001) (Figure 1E, bottom), and the sensitivity was 82.97% and the specificity was 94.19%, which presented that GRWD1 had high sensitivity and specificity in distinguishing colon cancer from noncancerous tissues.
Clinical Significance of GRWD1 Overexpression in Colon Carcinoma Patients
To explore the clinical significance of GRWD1 overexpression in colon carcinoma patients, the associations between GRWD1 expression level and clinical pathological factors were analyzed in Table 2. The analysis revealed a marked association between GRWD1 overexpression with pathological grading, tumor size, N stage and TNM stage (P < 0.05). However, the correlation between GRWD1 overexpression and patients' age, gender, T category was not found (P > 0.05). These results suggest that GRWD1 overexpression might be served as a predictor for the progression of colon carcinoma patients.
The Prognostic Value of GRWD1 Overexpression in Colon Carcinoma Patients
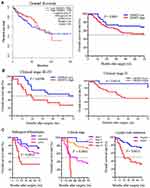
Firstly, A survival analysis was performed using data from GEPIA database, but the Kaplan-Meier survival curves and Log rank test demonstrated that patients with overexpression of GRWD1 had no significant association with overall survival (OS) of all patients (P > 0.05). However, data from clinical colon carcinoma samples indicated that patients with overexpression of GRWD1 had a shorter OS rate than low expression (P < 0.05) (Figure 2A). Besides, the OS of colon patients with GRWD1 overexpression is poor both in clinical stage (III–IV) and clinical stage (II) (P <0.05) (Figure 2B). Furthermore, Univariate Cox regression analysis showed that pathological differentiation, tumor size, clinical stage, lymph node metastasis, and GRWD1 expression were significantly associated with OS (P <0.05). Pathological differentiation, clinical stage, lymph node metastasis, and GRWD1 expression were finally confirmed to be independent factors with prognostic value for OS in patients with colon carcinoma via Multivariate Cox analysis (P < 0.05) (Table 3). Kaplan-Meier survival curves confirmed that pathological differentiation, clinical stage, and lymph node metastasis were significantly associated with poor OS (P < 0.05) (Figure 2C). These results indicated that GRWD1 overexpression predicted a poor prognosis of colon carcinoma patients.
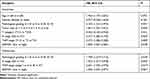 |
Table 3 Cox Model Analysis of Prognostic Factors and GRWD1 on Overall Survival |
Knockdown of GRWD1 Inhibits the Proliferation of Colon Cancer Cells
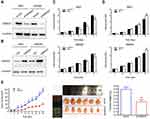
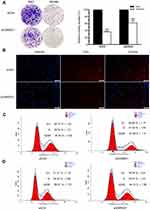
To explore the functions of GRWD1 on colon cancer cells, the cell models of stable GRWD1 knockdown and overexpression were established, separately. The Western blot analyses of GRWD1 protein expression were presented in Figure 3A and B. The results of cell proliferation detected by CCK8 assays showed that GRWD1 knockdown inhibited cell growth compared with control groups in both RKO and SW480 cells (Figure 3C). Conversely, the proliferation of both RKO and SW480 cells was enhanced after GRWD1 overexpression (Figure 3D). We also established the nude mouse model to verify the effect of GRWD1 on proliferation in vivo, the results showed that knockdown of GRWD1 inhibited tumor growth. The tumor weight of the GRWD1 knockdown group was much lower than the control group (Figure 3E).
Next, colony formation assays were used to detect the in-vitro tumorigenesis in RKO and SW480 cells. The results showed that knockdown of GRWD1 could significantly inhibit the colony formation both of RKO and SW480 cells (Figure 4A). Furthermore, EdU assay was performed to measure DNA synthesis, which could also represent the proliferative ability of cells. The results showed a remarkable decrease in the numbers of EdU-positive cells in shGRWD1 group compared with shCtrl group (Figure 4B).
These findings indicated that GRWD1 knockdown could significantly inhibit the proliferation of colon cancer cells in vitro and in vivo.
Knockdown of GRWD1 Induces G1 Phase Cell Cycle Arrest
Cell cycle disorder is one of the malignant characteristics in the development of colon carcinoma.12 Results from flow cytometry showed that GRWD1 knockdown inhibited cell cycle progression and caused the G1 phase distribution to increase from 40.12 ± 1.02% to 56.89 ± 1.17% (P < 0.05), and it caused a slightly decreasing number of cells in S and G2/M phase compared with control group in RKO cells (Figure 4C). In addition, GRWD1 knockdown also increased the G1 phase distribution from 41.31 ± 1.18% to 55.36 ± 0.91% (P < 0.05), whereas the phase of S and G2/M were decreased (Figure 4D). Similar results from RKO and SW480 cells indicated that GRWD1 played a key role in the procession of colon cancer cell cycle.
Knockdown of GRWD1 Enhances Drug Susceptibility of Colon Cancer Cells
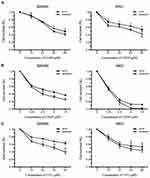
Resistance to chemotherapy is a main obstacle in the tumor clinical therapy including colon carcinoma.13 To further investigate whether GRWD1 knockdown affected drug susceptibility in colon cancer cells, CCK8 assays were performed to detect drug susceptibility of RKO and SW480 cells to three common chemotherapy drugs including LOHP, CDDP and 5-FU after GRWD1 knockdown. As shown in Figure 5A–C, the sensitivity to LOHP, CDDP, and 5-FU was increased after GRWD1 knockdown in both SW480 cells and RKO cells, compared with control groups. The results suggested that GRWD1 overexpression might contribute to the drug resistance of colon cancer cells.
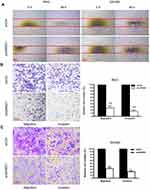
Knockdown of GRWD1 Suppresses Migration and Invasion of Colon Cancer Cells
In the clinical data analysis, it indicated that GRWD1 overexpression was associated with lymph node metastasis of colon carcinoma. Therefore, we detected the effect of GRWD1 knockdown on the migration and invasion by wound-healing assay and transwell assay. As shown in Figure 6A, the healing ability of RKO and SW480 cells was impaired after GRWD1 knockdown. Furthermore, transwell assays indicated that the migration and invasion activities of GRWD1 knockdown group cells were markedly decreased compared to the control group cells in both RKO (Figure 6B) and SW480 (Figure 6C) (P < 0.001). These two results suggested that GRWD1 played a vital promoting role in the migration and invasion of colon carcinoma cells.
The Potential Mechanisms of GRWD1 in Regulating the Development of Colon Carcinoma
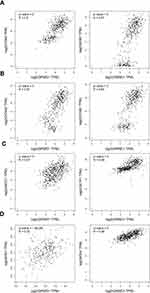
Those above results showed that GRWD1 played important roles in the proliferation, cell cycle, drug sensitivity, and migration and invasion of colon cancer cells. However, the potential mechanisms under the regulation of colon carcinoma development by GRWD1 need to be investigated. The primary analysis from online GEPIA database showed that GRWD1 expression was positively related to proliferation-related genes: PCNA and Ki67 (Figure 7A), cell cycle-related genes: Cyclin A2 and Cyclin B1 (Figure 7B), drug resistance-related genes: ABCC1 and GSTP1 (Figure 7C) and metastasis-related genes: MTA1 and MTA2 (Figure 7D).
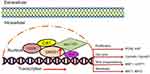
Based on the above research results, we proposed a working model about the tumorigenic function of GRWD1. As a novel Cdt1-binding protein in MCM helicase complexes,14 GRWD1 upregulation promotes the combination with CDC6 and Cdt1, further increases MCM loading and induces DNA replication, and then activates the transcriptions of several tumorigenesis-related genes (Figure 8)
Discussion
Our results revealed that GRWD1 was overexpressed in tissues of clinical CRC patients through the TCGA database and IHC analyses. GRWD1 upregulation was also correlated with several clinical- pathological factors including pathological grading, tumor size, N stage, and TNM stage. We constructed GRWD1 knockdown and overexpression CRC cell models to detect whether GRWD1 had effects on the development of CRC. We found that GRWD1 knockdown inhibited cell proliferation, on the contrary, the growth of CRC cells was promoted by GRWD1 overexpression. Besides, GRWD1 knockdown arrested cell cycle on the G1 phase, increased drug sensitivity, and reduced the ability of invasion and migration. From these results, we concluded that GRWD1 played an important role in tumorigenesis of colon carcinoma.
Many previous studies reported that incidence rates of CRC were increasing every year, especially among young people.15 Scientists dedicated to find the molecular mechanisms and possible therapeutic targets of cancer in the past few years, studies on potential genes that are evolved in colon carcinoma tumorigenesis may provide good prospects for diagnosis and therapy.16 Our study found GRWD1 was highly expressed in several cancers including breast carcinoma, lung squamous cell carcinoma, skin cutaneous melanoma, and also markedly evaluated in colon carcinoma. A recent study also showed that GRWD1 was highly expressed in non-small cell lung cancer (NSCLC). Besides, it could activate the Notch signaling pathway which induces cell proliferation in NSCLC.10 We also confirmed GRWD1 had a significant correlation with pathological grading, tumor size, and TNM stage. Although GRWD1 expression was not shown an obvious relationship with the overall rate of survival in all CRC patients, the overall survival rates of the months after surgery in all patients and patients with different clinical stages were shown significant relations with GRWD1 expression. These results further demonstrated that GRWD1 was an independent risk factor for poor development of colon carcinoma.
GRWD1, a WD-repeat-containing protein, is related to ribosome biosynthesis. GRWD1 overexpression is mainly localized in nuclear and slightly localized in cytoplasmic which is consistent with our findings.6 The IHC analysis was shown that GRWD1 was both expressed in nuclear and cytoplasmic while the staining intensity of GRWD1 in nuclear was higher than cytoplasmic. It was confirmed that GRWD1 was associated with p53. p53 is well known as a tumor suppresser and a transcriptional factor that regulates a series of genes expression including related genes of metabolic homeostasis, DNA repair, cell cycle, and DNA repair.17–19 Ubiquitin ligase MDM2 is involved in p53 degradation.20 Activated ribosomal proteins including RPL11 and RPL23 could interact with MDM2 and result in an accumulation of p53.21 Therefore, p53 is reduced when GRWD1 overexpression. Besides the RPL11/RPL23-MDM2-p53 axis, a recent study reported that GRWD1 is directly bound to the DNA-domain of p53 and negatively regulated p53 expression.7 All these results indicated that GRWD1 is a potential oncogene.
Our results showed that GRWD1 overexpression induced the growth in CRC cells in vitro and in vivo, while GRWD1 knockdown inhibited the colony formation and DNA replicate capacity. The expressions of proliferation-related genes: PCNA and ki67 were positively related to GRWD1 expression. These findings demonstrated that GRWD1 indeed had a markable effect on cell proliferation.
We further analyzed the cell cycle after the GRWD1 knockdown. Previous studies showed that DNA replication is important for cell cycle. Pre-replication complex (pre-RC) contained CDC6, CDC1 and MCM helicase formation is one procedure of DNA replication. GRWD1 could interact with CDC6 and Cdt1, sequentially increase MCM loading and induce DNA replication.22 Our results indicated that GRWD1 knockdown arrested cell cycle at the G1 phase, and slighted increased S and G2/M phase which was consistent with the previous study. Cells of the G2/M phase were slightly increased after GRWD1 depletion.22 We assessed the cell cycle-related genes: cyclin A2 and cyclin B1. Cyclin A2 was proved to be related with G1 and S phase, cyclin B1 inhibition caused G2/M phase arrest and increased cell growth.23,24 Both two genes had a significant correlation with GRWD1 expression. We still need to investigate the expression of cell-cycle related proteins after GRWD1 knockdown in further study.
Oxaliplatin (L-OHP), 5-fluorouracil (5-FU) and cisplatin (CDDP) are widely used in the treatment of colon carcinoma. A combination of different chemotherapy agents is an efficient treatment for cancer patients. Meanwhile, drug resistance also gradually occurs.25–27 In our study, we found that GRWD1 knockdown effectively increased the drug sensitivity in CRC cells. Moreover, the analyses between GRWD1 with MRP1 and GSTP1 showed a positive correlation. MRP1 belongs to the human multidrug resistance-associated protein (MRP) family. This membrane protein mainly results in clinical drug resistance.28,29 GSTP1 belongs to the glutathione S-transferase (GST) family.30 GSTP1 overexpression in cancer cells was found that it promoted the resistance to multiple drugs.31 Therefore, GRWD1 may be a potential therapeutic target in the treatment of colon carcinoma.
Metastasis is the main cause of cancer death and it is responsible for morbidity and mortality.32 Our study found that GRWD1 had a significant correlation with lymph node metastasis, besides GRWD1 knockdown inhibited the ability of migration and invasion of colon carcinoma cells. Analysis of GEPIA database showed that GRWD1 was related to metastasis-related genes: metastasis-associated gene 1 (MTA1) and metastasis-associated gene 2 (MTA2). MTA1, a transcriptional coregulator, played an important role in cancerous procedure. Many studies demonstrated that MTA1 overexpression facilitated invasion and migration in several cancer cells including laryngeal squamous cell carcinoma (LSCC), esophageal squamous cell carcinoma, and melanoma.33–35 MTA2 had a similar function with MTA1. Previous study also showed that MTA2 promoted the invasion of gastric cancer cells.36 In conclusion, GRWD1 had an important effect on invasion and migration in colon carcinoma cells.
Our study firstly confirmed GRWD1 promoted cell proliferation, drug resistance, cell cycle procedure, invasion and migration in colon carcinoma cells. We also put forward a potential mechanism of GRWD1 in oncogenesis. However, these predicted mechanisms of GRWD1 tumor-promoting functions are still unconfirmed. Therefore, we will next focus on these in further studies.
Funding
This study was supported by the Science Foundation of the Fujian Province, (No. 2016J01602; 2019J0105), Special Financial Foundation of Fujian Provincial (No.2015-1297), Young and middle-aged backbone training project in the health system of Fujian province (2016-ZQN-26), the Startup Fund for Scientific Research, Fujian Medical University (2017XQ1029, 2018QH2027), and the Professor Development Foundation of Fujian Medical University (No.JS11006).
Disclosure
The authors declare that there are no competing interests.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Levine O, Zbuk K. Colorectal cancer in adolescents and young adults: defining a growing threat. Pediatr Blood Cancer. 2019;66(11):e27941. doi:10.1002/pbc.27941
3. Bailey CE, Hu CY, You YN, et al. Increasing disparities in the age-related incidences of colon and rectal cancers in the United States, 1975–2010. JAMA Surg. 2015;150(1):17–22. doi:10.1001/jamasurg.2014.1756
4. Subramanian S, Amonkar MM, Hunt TL. Use of colonoscopy for colorectal cancer screening: evidence from the 2000 National Health Interview Survey. Cancer Epidemiol Biomarkers Prev. 2005;14(2):409–416. doi:10.1158/1055-9965.EPI-03-0493
5. Li H, Chen YX, Wen JG, Zhou HH. Metastasis-associated in colon cancer 1: a promising biomarker for the metastasis and prognosis of colorectal cancer. Oncol Lett. 2017;14(4):3899–3908. doi:10.3892/ol.2017.6670
6. Gratenstein K, Heggestad AD, Fortun J, Notterpek L, Pestov DG, Fletcher BS. The WD-repeat protein GRWD1: potential roles in myeloid differentiation and ribosome biogenesis. Genomics. 2005;85(6):762–773. doi:10.1016/j.ygeno.2005.02.010
7. Fujiyama H, Tsuji T, Hironaka K, Yoshida K, Sugimoto N, Fujita M. GRWD1 directly interacts with p53 and negatively regulates p53 transcriptional activity. J Biochem. 2020;167(1):15–24. doi:10.1093/jb/mvz075
8. Kayama K, Watanabe S, Takafuji T, et al. GRWD1 negatively regulates p53 via the RPL11-MDM2 pathway and promotes tumorigenesis. EMBO Rep. 2017;18(1):123–137. doi:10.15252/embr.201642444
9. Takafuji T, Kayama K, Sugimoto N, Fujita M. GRWD1, a new player among oncogenesis-related ribosomal/nucleolar proteins. Cell Cycle. 2017;16(15):1397–1403. doi:10.1080/15384101.2017.1338987
10. Wang Q, Ren H, Xu Y, et al. GRWD1 promotes cell proliferation and migration in non-small cell lung cancer by activating the Notch pathway. Exp Cell Res. 2020;387(2):111806. doi:10.1016/j.yexcr.2019.111806
11. Deng X, Li S, Kong F, et al. Long noncoding RNA PiHL regulates p53 protein stability through GRWD1/RPL11/MDM2 axis in colorectal cancer. Theranostics. 2020;10(1):265–280. doi:10.7150/thno.36045
12. Dash BC, El-Deiry WS. Cell cycle checkpoint control mechanisms that can be disrupted in cancer. Methods Mol Biol. 2004;280:99–161. doi:10.1385/1-59259-788-2:099
13. Aparo S, Goel S. Evolvement of the treatment paradigm for metastatic colon cancer. From chemotherapy to targeted therapy. Crit Rev Oncol Hematol. 2012;83(1):47–58. doi:10.1016/j.critrevonc.2011.08.006
14. Aizawa M, Sugimoto N, Watanabe S, Yoshida K, Fujita M. Nucleosome assembly and disassembly activity of GRWD1, a novel Cdt1-binding protein that promotes pre-replication complex formation. Biochim Biophys Acta. 2016;1863(11):2739–2748. doi:10.1016/j.bbamcr.2016.08.008
15. Puccini A, Lenz HJ, Marshall JL, et al. Impact of patient age on molecular alterations of left-sided colorectal tumors. Oncologist. 2019;24(3):319–326. doi:10.1634/theoncologist.2018-0117
16. Tomlinson I, Ilyas M, Novelli M. Molecular genetics of colon cancer. Cancer Metastasis Rev. 1997;16(1–2):67–79. doi:10.1023/A:1005744323215
17. Vousden KH, Prives C. Blinded by the light: the growing complexity of p53. Cell. 2009;137(3):413–431. doi:10.1016/j.cell.2009.04.037
18. Riley T, Sontag E, Chen P, Levine A. Transcriptional control of human p53-regulated genes. Nat Rev Mol Cell Biol. 2008;9(5):402–412. doi:10.1038/nrm2395
19. Levine AJ, Oren M. The first 30 years of p53: growing ever more complex. Nat Rev Cancer. 2009;9(10):749–758. doi:10.1038/nrc2723
20. Toledo F, Wahl GM. Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nat Rev Cancer. 2006;6(12):909–923. doi:10.1038/nrc2012
21. Zhang YP, Lu H. Signaling to p53: ribosomal proteins find their way. Cancer Cell. 2009;16(5):369–377. doi:10.1016/j.ccr.2009.09.024
22. Sugimoto N, Maehara K, Yoshida K, et al. Cdt1-binding protein GRWD1 is a novel histone-binding protein that facilitates MCM loading through its influence on chromatin architecture. Nucleic Acids Res. 2015;43(12):5898–5911. doi:10.1093/nar/gkv509
23. Yuan J, Kramer A, Matthess Y, et al. Stable gene silencing of cyclin B1 in tumor cells increases susceptibility to taxol and leads to growth arrest in vivo. Oncogene. 2006;25(12):1753–1762. doi:10.1038/sj.onc.1209202
24. Pagano M, Pepperkok R, Verde F, Ansorge W, Draetta G. Cyclin-a is required at 2 points in the human cell-cycle. EMBO J. 1992;11(3):961–971. doi:10.1002/j.1460-2075.1992.tb05135.x
25. Longley DB, Harkin DP, Johnston PG. 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer. 2003;3(5):330–338. doi:10.1038/nrc1074
26. Kim GP, Erlichman C. Oxaliplatin in the treatment of colorectal cancer. Expert Opin Drug Metab Toxicol. 2007;3(2):281–294. doi:10.1517/17425255.3.2.281
27. Kohno K, Wang KY, Takahashi M, et al. Mitochondrial transcription factor A and mitochondrial genome as molecular targets for cisplatin-based cancer chemotherapy. Int J Mol Sci. 2015;16(8):19836–19850. doi:10.3390/ijms160819836
28. Ishikawa T, Kuo MT, Furuta K, Suzuki M. The human multidrug resistance-associated protein (MRP) gene family: from biological function to drug molecular design. Clin Chem Lab Med. 2000;38(9):893–897. doi:10.1515/CCLM.2000.130
29. Borst P, Evers R, Kool M, Wijnholds J. A family of drug transporters: the multidrug resistance-associated proteins. J Natl Cancer I. 2000;92(16):1295–1302. doi:10.1093/jnci/92.16.1295
30. Tew KD. Glutathione-associated enzymes in anticancer drug-resistance. Cancer Res. 1994;54(16):4313–4320.
31. Townsend DM, Shen HX, Staros AL, Gate L, Tew KD. Efficacy of a glutathione S-transferase pi-activated prodrug in platinum-resistant ovarian cancer. Mol Cancer Ther. 2002;1(12):1089–1095.
32. Guan X. Cancer metastases: challenges and opportunities. Acta Pharm Sin B. 2015;5(5):402–418. doi:10.1016/j.apsb.2015.07.005
33. Zhang H, Yang D, Wang H, et al. Metastasis-associated gene 1 promotes invasion and migration potential of laryngeal squamous cell carcinoma cells. Oncol Lett. 2014;7(2):399–404. doi:10.3892/ol.2013.1729
34. Qian H, Lu N, Xue L, et al. Reduced MTA1 expression by RNAi inhibits in vitro invasion and migration of esophageal squamous cell carcinoma cell line. Clin Exp Metastasis. 2005;22(8):653–662. doi:10.1007/s10585-006-9005-2
35. Qian H, Yu J, Li Y, et al. RNA interference of metastasis-associated gene 1 inhibits metastasis of B16F10 melanoma cells in a C57BL/6 mouse model. Biol Cell. 2007;99(10):573–581. doi:10.1042/BC20060130
36. Zhou C, Ji J, Cai Q, et al. MTA2 promotes gastric cancer cells invasion and is transcriptionally regulated by Sp1. Mol Cancer. 2013;12(1):102. doi:10.1186/1476-4598-12-102
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.