Back to Journals » Cancer Management and Research » Volume 13
Clinical Response to Traditional Chinese Herbs Containing Realgar (As2S2) is Related to DNA Methylation Patterns in Bone Marrow DNA from Patients with Myelodysplastic Syndrome with Multilineage Dysplasia
Authors Zhou QB , Du Y, Zhang SS, Liu ZT, Ma R, Xu YG
Received 8 September 2020
Accepted for publication 4 December 2020
Published 7 January 2021 Volume 2021:13 Pages 55—63
DOI https://doi.org/10.2147/CMAR.S280886
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
Qing-Bing Zhou,1 Yu Du,2 Shan-Shan Zhang,2 Zheng-Tang Liu,1 Rou Ma,2 Yong-Gang Xu2
1Institute of Geriatric Medicine, Xiyuan Hospital, China Academy of Chinese Medical Sciences, Beijing 100091, People’s Republic of China; 2Department of Hematology, Xiyuan Hospital, China Academy of Chinese Medical Sciences, Beijing 100091, People’s Republic of China
Correspondence: Rou Ma; Yong-Gang Xu
Department of Hematology, Xiyuan Hospital, Xiyuan Playground No. 1, Haidian District, Beijing 100091, People’s Republic of China
Tel/Fax +86 10 6283 5627
Email [email protected]; [email protected]
Purpose: DNA methylation is known to play an important role in myelodysplastic syndrome (MDS). We previously showed that Chinese herbs (CHs) containing realgar (As2S2) were effective at treating MDS with multilineage dysplasia (MDS–MLD). We tested whether the response to CH treatment was related to changes in DNA methylation in MDS–MLD.
Patients and Methods: First, the Illumina methylation 850K array BeadChip assay was used to assess the pretreatment methylation status in bone marrow cells from eight MDS–MLD patients and 3 healthy donors. The eight MDS–MLD patients were then treated with CHs for six months, the arsenic concentration was measured following treatment. The patients were subsequently divided into “effective” and “ineffective” treatment response groups and the DNA methylation patterns of the two groups were compared. Finally, the BeadChip data were validated by pyrosequencing.
Results: Five of the eight MDS–MLD patients showed hematological improvement (effective-treatment group), while three showed disease progression (ineffective-treatment group) (positive response rate: 62.5%). The arsenic concentrations in the patients ranged from 26.60 to 64.16 μg/L (median 48.4 μg/L) and were not significantly different between the two groups (p = 0.27). Compared with the healthy controls, three genes were hypomethylated and 110 were hypermethylated in the ineffective-treatment group. However, in the group showing hematological improvement, 102 genes were markedly hypomethylated and 87 hypermethylated. The effective-treatment group had a higher proportion of hypomethylated sites than the ineffective-treatment group (53.9% vs 2.6%, respectively; chi-square test) (p < 0.0001). Two hypermethylated and two hypomethylated genes were selected for validation by pyrosequencing (all p < 0.05).
Conclusion: MDS–MLD patients may present different DNA methylation subtypes. CHs containing realgar may be effective for treating MDS–MLD patients with the hypomethylation subtype.
Keywords: myelodysplastic syndromes, DNA methylation pattern, Chinese herbs, realgar
Introduction
Myelodysplastic syndrome (MDS), a common malignant disease, comprises a heterogeneous group of disorders characterized by specific cytomorphological features of the blood and bone marrow, ineffective hematopoiesis, and a high risk of evolution to acute myeloid leukemia (AML).1 According to the World Health Organization (WHO) classification, myelodysplastic syndromes with multilineage dysplasia (MDS–MLD) represent one of several clinical subtypes of MDS. The pathological mechanism underlying MDS mainly involves abnormal DNA methylation, gene mutations, and chromosomal aberrations.2 Many therapeutic options, including hypomethylating agents (HMAs), are available for the treatment of MDS; however, allogeneic hematopoietic stem cell transplantation (allo-HSCT) represents the only potentially curative option for MDS.3 Nevertheless, the use of allo-HSCT for MDS patients is often limited because of the significant mortality associated with allo-HCST and the lack of suitable donors.3 Consequently, finding effective therapeutic drugs from traditional medicines has become an important strategy for the curing of MDS.4,5
Traditional Chinese medicine (TCM) is characterized by a special theory (see Discussion below) and has been applied in China for more than 3000 years. In 1979, Ting-dong Zhang, a TCM doctor, reported for the first time that acute promyelocytic leukemia (APL) could be treated effectively with an injection, the main constituent drug of which was arsenic trioxide (ATO).6 APL is currently a highly curable disease because of the worldwide use of ATO.7,8 Another arsenic compound is arsenic disulfide (AS2S2), which is isolated from a mined ore known as realgar. Our hospital has been using realgar as the core drug for the treatment of MDS for more than 30 years. Realgar was first recorded in the Shennong Herbal Sutra, a pharmaceutical work written 2000 years ago. We have previously shown that Chinese herbs (CHs) containing realgar can treat MDS–MLD patients effectively and safely. The overall response rate to the realgar-based treatment was 81.53% (106/130) for hematologic improvement in patients with MDS–MLD.4 We have also recently reported that the clinical efficacy of CHs was related to the concentration of arsenic in the peripheral blood, and that increasing the arsenic concentration to over 20 μg/L had potential as a therapeutic strategy for treating MDS–MLD patients.5 However, some MDS–MLD patients do not respond to treatment, even at arsenic concentrations greater than 20 μg/L, which suggested that other factors could also influence the efficacy of CHs.
DNA methylation, the covalent addition of a methyl group from S-adenosylmethionine (SAM) to cytosine mediated by DNA methyltransferases (DNMTs), is an essential epigenetic modification of genomic DNA in mammalian cells.9 Studies have demonstrated that aberrant DNA methylation (including abnormal hypermethylation and hypomethylation) is an important factor in the development of MDS. Dysregulated DNA hypermethylation can lead to gene silencing, including of tumor suppressor genes, while aberrant hypomethylation can result in the activation of oncogene expression.10,11
Given the crucial role of DNA methylation in MDS, we hypothesized that the effect underlying CH treatment was related to the pattern of DNA methylation in MDS–MLD patients, which may provide a new research direction for improving the clinical efficacy of MDS treatment.
Patients and Methods
Patients and Samples
Patients were diagnosed with MDS–MLD according to the 2016 WHO classification12 and a total of eight MDS–MLD patients were enrolled from the outpatient department of Xiyuan Hospital. Pregnant and breastfeeding women were excluded. Exclusion criteria also included New York Heart Association heart failure classes III and IV, active or stable malignancy, and fever with infection. Patients should not have previously taken any therapy with HMAs. Three bone marrow samples from healthy donors served as the control group for the assessment of methylation status. Written informed consent, in accordance with the Declaration of Helsinki, was provided by all the subjects, or by a parent if the participant was under 18 years of age. This study was approved by the Medical Ethics Committee of Xiyuan Hospital (No. 2018XLA005-2), and the clinical registration number is ChiCTR1800015614.
Treatment Regimen
All of the MDS–MLD patients took realgar orally at a dose of 0.1 g daily for six months. Realgar was provided by Xiyuan Hospital. Other herbs including Rhizoma diosscoreae (15 g), Radix rehmanniae (20 g), Fructus corni (20 g), F. psoraleae (15 g), F. jujubae (15 g), Poria (15 g), Semen cuscutae (20 g), Cortex mouta (10 g), Rhizoma atractylodis macrocephalae (20 g), R. alismatis (20 g), Radix pseudostellariae (30 g), R. polygoni multiflori (25 g), R. morindae officinalis (15 g), Zingiberis recens rhizoma (15 g), and Ramulus cinnamomi (15 g) were taken as auxiliary therapy. None of the MDS–MLD patients received other drugs during the treatment period, including erythropoietin or granulocyte colony-stimulating factor (G-CSF). Red blood cells were administered to patients with hemoglobin concentrations <60 g/L. Patients presenting with fever underwent anti-infection treatment.
Efficacy and Safety
Patients were evaluated for therapeutic efficacy and risk stratification via bone marrow evaluation, chromosomal karyotype analysis, and complete blood count according to the International Working Group criteria.13,14 The patients were divided into effective-treatment and ineffective-treatment groups based on treatment efficacy. National Cancer Institute Common Terminology Criteria for Adverse Events (version 5.0) was used to grade the adverse events (AEs) during treatment.
Measurement of Arsenic Concentration
To exclude the effect of arsenic concentration on treatment response, the arsenic concentrations of all of the patients were measured following treatment. Briefly, 2 mL of peripheral venous blood was extracted from each patient on an empty stomach at 08:00 hours. The blood samples were then placed at −20 °C within 3 h. Following thawing at room temperature, 0.5 mL of each blood sample was used to determine the whole-blood arsenic concentration by atomic fluorescence spectrometry (PSA 10.055 Millennium Excalibur AFS, PS Analytical, Orpington, Kent, UK) according to the manufacturer’s instructions.
DNA Methylation Microarray Assay
Bone marrow samples were obtained from three healthy individuals and eight MDS–MLD patients before treatment, and genomic DNA was extracted for the analysis of the methylation patterns using the Illumina Human Methylation 850K array (Illumina, Inc., San Diego, CA, USA). First, the EZ DNA Methylation Kit (Zymo Research, Irvine, CA, USA) was used for bisulfite conversion. Second, the bisulfite-converted DNA was hybridized to the 850K BeadChip platform. Chip analysis was performed using an Illumina HiScan SQ fluorescence scanner. Then, quality control was performed at the sample-level and the probe-level. Samples must have passed the Illumina quality control standard. Samples having 1% of sites with a detection p-value greater than 0.01 were removed. Sites having 1% of samples with a detection p-value greater than 0.01 or sites with bead counts < 3 in 5% of samples were removed. If a CpG site was completely unmethylated, then the ratio would approach 0, whereas if a CpG site was fully methylated, the ratio would approach 1. The identified differentially methylated genes (DMGs) were analyzed using the KEGG database, an important bioinformatics tool for screening related functions.15 Finally, the genomic DNA methylation patterns of the two groups were compared with those of the healthy volunteers.
Methylation Validation by Pyrosequencing
Pyrosequencing was used to validate the 850K data. Two hypermethylated and two hypomethylated genes were selected for validation. The sequences of the primers used for the four genes are listed in Table S1. Following bisulfite treatment, the treated DNA and Hot Start High-Fidelity Taq DNA Polymerase (Qiagen, Hilden, Germany) were used for polymerase chain reaction (PCR), performed with the following cycling parameters: 98 °C for 10 s, 55 °C for 30 s, and 72 °C for 30 s for 35 cycles. The amplicons were then subjected to quantitative DNA methylation analysis using pyrosequencing on the PyroMark Q96 system (Qiagen, Germany). The results were analyzed using the Pyro Q-CpG software.
Statistical Analysis
Bayesian and linear regression analysis were applied to assess the statistical significance of differences in the methylation status of the different groups. The CpG sites with either a minimum change of either ± 0.1 or 0.2 in β-values (delta beta), and p-values <0.05 in the different groups were considered significant. The chi-squared test and the t-test were used to examine the differences in methylation sites and methylation levels.
Results
Patient Characteristics
Table 1 lists the clinical characteristics of the eight patients. The ages of the patients ranged from 17 to 51 years. The chromosomal karyotype of all the patients was normal, and two patients had bilineage cytopenia (25%) and six had trilineage cytopenia (75%). All of the patients belonged to the intermediate-1 risk group according to the International Prognostic Scoring System (IPSS).14
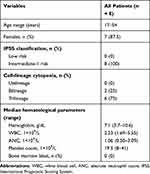 |
Table 1 Patient Demographics and MDS–MLD Disease Characteristics |
Arsenic Concentration, Efficacy, and Safety
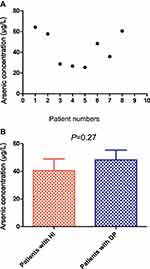
The arsenic concentration of patients included in this study ranged from 26.60 to 64.16 μg/L, with a median concentration of 48.4 μg/L (Figure 1A). The treatment employed in the study was relatively well-tolerated, and none of the patients discontinued treatment due to AEs. All eight patients were evaluated for treatment response after six months of treatment. Overall, five of the eight (62.5%) patients showed hematologic improvement (HI), and three showed disease progression (DP) (46%) (Table 2). No changes in chromosome karyotype or the number of primitive cells in bone marrow were observed following treatment. There was no significant difference in arsenic concentrations between patients with HI and those showing DP (Figure 1B).
 |
Table 2 Responses to Realgar Treatment |
Untreated MDS Patients Displayed a Large Number of Abnormally Hypomethylated and Hypermethylated Genes
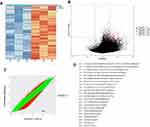
Analysis using the human methylation 850K BeadChip assay showed that a large number of CpG sites were significantly and differentially hypomethylated and hypermethylated in MDS patients compared with those in healthy volunteers (Table S2). A volcano plot comparing the eight MDS patients with the three healthy individuals is shown in Figure 2A. Gene Ontology (GO) analysis showed that many of both the hyper- and hypomethylated genes had roles in cancer-related functions and pathways, including cell differentiation, Rho protein signal transduction, and hippo signaling (Table S3). Pathway analysis showed that many of both the hypo- and hypermethylated genes were involved in important biological signaling pathways, including the mTOR signaling pathway, the Wnt signaling pathway, cancer-related pathways, the cGMP-PKG signaling pathway, the JAK–STAT signaling pathway, and hematopoietic cell lineage-related pathways (Figure 2B and C).
Comparison of the Pretreatment Genomic DNA Methylation Patterns of Two Groups
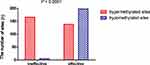
BeadChip analysis revealed that 169 sites were significantly and differentially methylated in the ineffective-treatment group when compared with the healthy controls (methylation differences ± 0.1 and p-value <0.05). Of the 169 sites, 166 were hypermethylated and three were hypomethylated, corresponding to 110 hypermethylated and three hypomethylated genes, respectively (Figure 3A and B, Table S4). In contrast, compared with the healthy donors, the effective-treatment group displayed 198 hypomethylated and 138 hypermethylated sites, corresponding to 102 hypomethylated and 87 hypermethylated genes, respectively (Figure 3C and D; Tables S5 and S6). The effective-treatment group showed a greater proportion of hypomethylated genes than the ineffective-treatment group (χ2 = 153.3, p < 0.0001) (Figure 4).
Validation of the 850K BeadChip DNA Methylation Data
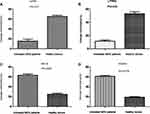
We analyzed the methylation status of the LEPR, LPHN3, WNT6, and HOXA4 genes by pyrosequencing. Before treatment, 850K analysis of these four genes showed that they were significantly and differentially methylated between the effective-treatment group and the healthy donors (p < 0.05). Pyrosequencing revealed that LEPR (p = 0.017) and LPHN3 (p = 0.035) were hypomethylated, whereas WNT6 (p = 0.0027) and HOXA4 (p = 0.0179) were hypermethylated, in the five untreated MDS patients (Figure 5). These results were consistent with those from the BeadChip analysis.
Discussion
To the best of our knowledge, the present study is the first to demonstrate the significant finding that DNA methylation subtypes can differ among MDS–MLD patients. Our study also indicated that the clinical response to the realgar regimen is related to DNA methylation patterns in MDS–MLD patients.
Our study demonstrated that MDS–MLD patients have different DNA methylation patterns, of which we identified two types: a hypermethylation pattern, in which the percentage of hypermethylated sites was 98.22%, and a hypomethylation pattern, in which the percentage of hypomethylated sites was 58.9%. We believe that this novel finding will be very useful for the treatment of MDS. Because of the aberrant hypermethylation observed in MDS, HMAs (decitabine and azacitidine) targeting hypermethylation have represented the greatest progress in treating MDS over the past decade, showing a 40% response rate for high-risk patients.16 However, the therapeutic response is transient and most MDS patients receiving HMAs eventually relapse.17 In addition to hypermethylation, aberrant hypomethylation has also been observed in MDS patients. One study showed that the frequency of spalt-like transcription factor 4 (SALL4) hypomethylation is significantly increased in patients at higher risk of MDS.18 Wu et al reported that MDS patients with hypomethylated microRNA let-7a-3 had a poor prognosis.19 Hypomethylation can result in the transcriptional activation of oncogenes in cancers, including in MDS.20–22 In this study, we also observed that many genes were hypomethylated in MDS–MLD patients, and identified that many of these genes had roles in cancer-related pathways, including the mTOR and Wnt signaling pathways, supporting the hypothesis that aberrantly hypomethylated genes also play a key role in MDS. Moreover, we found that MDS–MLD patients have different DNA methylation patterns, even if they belong to the same WHO classification subtype. HMAs may show therapeutic benefits for MDS patients in the hypermethylation subgroup; however, new treatments are needed for those with a hypomethylation subtype.
TCM has been employed in China for more than 3000 years. According to TCM theory, “toxin,” “blood stasis,” and “deficiency” are the main pathologies of MDS. Based on this, we designed a CH-related regimen for the treatment of MDS with realgar as the core drug. Realgar can activate blood and remove blood stasis. The main active component of realgar is arsenic(II) sulfide, As2S2, which has been shown to exert antitumor effects and enhance cell apoptosis and erythroid differentiation.23–25 Importantly, we previously showed that AS2S2 can promote hypomethylation by increasing the expression of DNA methyltransferases in MDS.26 Herbs in this regimen, including Rhizoma diosscoreae, Radix rehmanniae, and Fructus corni, can strengthen the spleen and kidney.
In this study, all eight patients received CH treatment for six months. The response rates were 62.5% (5/8) for hematological improvement. Three patients did not respond to the treatment, even at an arsenic concentration greater than 20 μg/L. The methylation patterns in the two groups before treatment were compared by means of the 850K BeadChip assay. Interestingly, the effective-treatment group exhibited a greater proportion of hypomethylated genes than the ineffective-treatment group (53.9% vs 2.6%, respectively; p < 0.0001). These results strongly indicated that the CHs containing realgar can treat MDS–MLD patients with the hypomethylation pattern effectively, which is an innovative finding. Studies involving larger samples are needed to confirm these results.
Conclusions
Our study showed that MDS–MLD patients have different DNA methylation subtypes. Furthermore, the clinical response to the realgar regimen was related to the patterns of DNA methylation. Realgar treatment is more suitable for MDS–MLD patients with the hypomethylation subtype.
Data-Sharing Statement
We are happy to share all of the data generated in this study. The data can be obtained via Email: [email protected].
Acknowledgments
We thank Hong-zhi Wang for the analysis of the bone marrow samples. This project was financially supported by the China Academy of Chinese Medicine Scientific Foundation (ZZ13-YQ-010) and the National Natural Scientific Foundation of China (81603490, 81774140) for funding.
Disclosure
The authors declare that they have no conflict of interest.
References
1. Lwenberg B. Introduction to a review series on myelodysplastic syndromes. Blood. 2019;133(10):1001. doi:10.1182/blood-2018-12-886549
2. Hosono N. Genetic abnormalities and pathophysiology of MDS. Int J Clin Oncol. 2019;24(8):885–892. doi:10.1007/s10147-019-01462-6
3. Bewersdorf JP, Zeidan AM. Evolving therapies for lower-risk myelodysplastic syndromes. Ann Hematol. 2020;99:677–692.
4. Zhou Q, Zhu Q, Wang H, et al. Traditional Chinese medicine containing arsenic treated MDS patients effectively through regulating aberrant hypomethylation. Evid Based Complement Alternat Med. 2020;7469809.
5. Zhu Q, Deng Z, Zhu S, et al. Study on the clinical safe and effective methods of arsenic-containing compound-Qinghuang powder in the treatment of myelodysplastic syndrome. Evid Based Complement Alternat Med. 2017. doi:10.1155/2017/2095682
6. Zhang TD, Rong FX. Cancer 1 injection and differentiation in the treatment of acute myelocytic leukemia. Heilongjiang Medi J. 1979;(4):7–10.
7. Chen SJ, Chen Z. Targeting agents alone to cure acute promyelocytic leukemia. N Engl J Med. 2013;369(2):186–187. doi:10.1056/NEJMe1304762
8. Wang ZY, Chen Z. Acute promyelocytic leukemia: from highly fatal to highly curable. Blood. 2008;111(5):2505–2515. doi:10.1182/blood-2007-07-102798
9. Jones PA, Liang G. Rethinking how DNA methylation patterns are maintained. Nat Rev Genet. 2009;10(11):805–811. doi:10.1038/nrg2651
10. Van Tongelen A, Loriot A, De Smet C. Oncogenic roles of DNA hypomethylation through the activation of cancer-germline genes. Cancer Lett. 2017;396(28):130–137. doi:10.1016/j.canlet.2017.03.029
11. Jaroslav J, Shoudan L, Frank N, et al. Cancer drivers affected by aberrant DNA methylation in MDS and AML. Blood. 2011;118(21):1716. doi:10.1182/blood.V118.21.1716.1716
12. Arber DA, Orazi A, Hasserjian R, et al. The 2016 revision to the World Health Organization (WHO) classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–2405. doi:10.1182/blood-2016-03-643544
13. Tefferi A, Barosi G, Mesa RA, et al. International working group (IWG) consensus criteria for treatment response in myelofibrosis with myeloid metaplasia: on behalf of the IWG for myelofibrosis research and treatment (IWGMRT). Blood. 2006;108(5):1497–1503. doi:10.1182/blood-2006-03-009746
14. Greenberg P, Cox C, LeBeau MM, et al. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood. 1997;89(6):2079–2088. doi:10.1182/blood.V89.6.2079
15. Aoki-Kinoshita KF, Kanehisa M. Gene annotation and pathway mapping in KEGG. Methods Mol Biol. 2007;396:71–91.
16. Gonzalez-Fierro A, Dueñas-González A. Emerging DNA methylation inhibitors for cancer therapy: challenges and prospects. Expert Rev Precis Med Drug Dev. 2019;4(1):27–35. doi:10.1080/23808993.2019.1571906
17. Santini V. How I treat MDS after hypomethylating agent failure. Blood. 2019;133(1):521–529. doi:10.1182/blood-2018-03-785915
18. Lin J, Qian J, Yao DM, et al. Aberrant hypomethylation of SALL4 gene in patients with myelodysplastic syndrome. Leuk Res. 2013;37(1):71–75. doi:10.1016/j.leukres.2012.10.014
19. Wu DH, Yao DM, Yang L, et al. Hypomethylation of let-7a-3 is associated with poor prognosis in myelodysplastic syndrome. Leuk Lymphoma. 2017;58(1):96–103. doi:10.1080/10428194.2016.1187273
20. Eden A, Gaudet F, Waghmare A, et al. Chromosomal instability and tumors promoted by DNA hypomethylation. Science. 2003;300(5618):455. doi:10.1126/science.1083557
21. Pogribny IP, Beland FA. DNA hypomethylation in the origin and pathogenesis of human diseases. Cell Mol Life Sci. 2009;66(14):2249–2261. doi:10.1007/s00018-009-0015-5
22. Wu H, Chen Y, Liang J, et al. Hypomethylation-linked activation of PAX2 mediates tamoxifen-stimulated endometrial carcinogenesis. Nature. 2005;438(7070):981–987. doi:10.1038/nature04225
23. Wang S, Zhang C, Li Y, Li P, Zhang D, Li C. Anti-liver cancer effect and the mechanism of arsenic sulfide in vitro and in vivo. Cancer Chemother Pharmacol. 2019;83(3):519–530. doi:10.1007/s00280-018-3755-9
24. Hu XM, Yuan B, Tanaka S, et al. Arsenic disulfide-triggered apoptosis and erythroid differentiation in myelodysplastic syndrome and acute myeloid leukemia cell lines. Hematology. 2014;19(6):352–360. doi:10.1179/1607845413Y.0000000138
25. Xu M, Ren JY, Guo YC, et al. Effects of arsenic disulfide on apoptosis, histone acetylation, toll like receptor 2 activation, and erythropoiesis in bone marrow mononuclear cells of myelodysplastic syndromes patients in vitro. Leuk Res. 2017;62:4–11. doi:10.1016/j.leukres.2017.09.010
26. Zhou QB, Liu ZT, Wang HZ, Guo XQ, Xu YG, Hu XM. Arsenic disulfide promoted hypomethylation by increasing DNA methyltransferases expression in myelodysplastic syndrome. Drug Des Devel Ther. 2020;14:1641–1650. doi:10.2147/DDDT.S239158
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.