Back to Journals » Cancer Management and Research » Volume 13
Circular RNA CircEPB41L2 Functions as Tumor Suppressor in Hepatocellular Carcinoma Through Sponging miR-590-5p
Authors Chen F, He L, Qiu L, Zhou Y, Li Z , Chen G, Xin F , Dong X, Xu H, Wang G, Liu J , Cai Z
Received 14 November 2020
Accepted for publication 1 March 2021
Published 1 April 2021 Volume 2021:13 Pages 2969—2981
DOI https://doi.org/10.2147/CMAR.S291682
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Sanjeev K. Srivastava
Feng Chen,1,2 Lei He,2,3 Liman Qiu,2,3 Yang Zhou,2,3 Zhenli Li,2,3 Geng Chen,2,3 Fuli Xin,2,3 Xiuqing Dong,2,3 Haipo Xu,2,3 Gaoxiong Wang,4 Jingfeng Liu,2,3 Zhixiong Cai2,3
1College of Life Science, Fujian Agriculture and Forestry University, Fuzhou, 350002, People’s Republic of China; 2The United Innovation of Mengchao Hepatobiliary Technology Key Laboratory of Fujian Province, Mengchao Hepatobiliary Hospital of Fujian Medical University, Fuzhou, 350025, People’s Republic of China; 3Mengchao Med-X Center, Fuzhou University, Fuzhou, 350025, People’s Republic of China; 4Department of Hepatobiliary Surgery, The Second Affiliated Hospital of Fujian Medical University, Quanzhou, 362001, People’s Republic of China
Correspondence: Gaoxiong Wang
Department of Hepatobiliary Surgery, The Second Affiliated Hospital of Fujian Medical University, Zhongshan North Road 34, Quanzhou, 362001, People’s Republic of China
Tel/Fax +86 591 8370 5927
Email [email protected]
Zhixiong Cai
The United Innovation of Mengchao Hepatobiliary Technology Key Laboratory of Fujian Province, Mengchao Hepatobiliary Hospital of Fujian Medical University, Xihong Road 312, Fuzhou, Fujian, 350025, People’s Republic of China
Tel/Fax +86 591 8370 5927
Email [email protected]
Background: Circular RNAs (circRNAs) could interact with miRNAs to regulate gene expression, participating in hepatocellular carcinoma (HCC) initiation and development. This work aimed to determine the potential function and molecular mechanism of circEPB41L2 (hsa_circ_0077837) during HCC progression.
Materials and Methods: The expression of circEPB41L2 in HCC tissues and HCC cell lines was quantified using real-time quantitative PCR (qRT-PCR). CCK-8 assays and colony formation assays were utilized to detect the proliferation of HCC cells. Wound healing assay and transwell assay were performed to determine the capability of migration and invasion for HCC cells. Western blot was conducted to determine gene expression on protein levels. The effect of circEPB41L2 on HCC in vivo was investigated via xenograft experiment. Interaction between circEPB41L2 and miR-590-5p was predicted through bioinformatics methods and confirmed via luciferase reporter assay.
Results: Extensive analysis of circRNA profiles in tumor and matched para-tumor tissues collected from 61 HCC patients identified that circEPB41L2 was significantly down-regulated in HCC, which was further confirmed in another HCC group by qRT-PCR analysis. The clinicopathological analysis revealed that down-regulation of circEPB41L2 was negatively associated with tumor size, vascular invasion and alpha-fetoprotein, while positively correlated with HCC prognosis. The biological function experiments showed that overexpression of circEPB41L2 could obviously inhibit the proliferation and metastasis of HCC cells in vitro, while knockdown of circEPB41L2 induced opposite results. Moreover, we also found that circEPB41L2 inhibited HCC migration and invasion though EMT signaling pathway. Similarly, overexpression of circEPB41L2 can also significantly inhibit the proliferation of HCC cells in vivo. Bioinformatic analysis and luciferase reporter assay revealed that circEPB41L2 interacts directly with miR-590-5p and the corresponding biological functions were also verified in miRNA rescue experiments.
Conclusion: Our results suggest that circEPB41L2 might function as a tumor suppressor during HCC progression by sponging miR-590-5p.
Keywords: circular RNA, circEPB41L2, hepatocellular carcinoma, miR-590-5p
Introduction
Hepatocellular carcinoma (HCC) is the most common liver malignancy worldwide and is considered as the fourth leading cause of cancer-related death in China.1,2 Due to the lack of early and specific clinical manifestations of HCC, more than 75% of HCC patients are usually diagnosed at an advanced stage.3,4 Therefore, it has become an important direction to study the pathogenesis of HCC and seek for potential molecular targets for early diagnosis, prognosis, evaluation and treatment of HCC.
CircRNAs, a special class of non-coding RNAs, characterized by a covalent closed loop and a polyadenylate tail, are thus relatively stable and highly abundant in tissue and blood.5–7 For a long time, circRNAs were regarded as a byproduct of splicing without biological functions.8 However, with the development of bioinformatics technologies, the functional importance of circRNAs in cancer has been gradually recognized.9,10 An increasing number of studies reported that circRNAs could regulate gene expression at the transcriptional or post-transcriptional levels by competing with miRNA response elements to participate in HCC progression.11,12 For example, circRNA_MAN2B2 promotes proliferation of HCC cells through the miRNA-217/MAPK1 axis.13 CircRNA_0000502 promotes HCC metastasis and inhibits apoptosis by targeting miRNA-124.14 CircRNA_5692 enhances the expression of DAB2IP by sponging miR-328-5p, thereby inhibiting the progression of HCC.15 However, due to the complex pathogenesis of HCC, the mechanisms of circRNAs in HCC still needs further investigation.
In this study, to analyze the circRNA profiles and their alteration during the HCC progression, we extensively analyzed the full transcriptome sequencing data of HCC tissues and matched para-tumor tissues from 61 HCC patients, and identified an interesting circRNA (circEPB41L2) that was significantly down-regulated in HCC. Then, the clinical significance, biological functions and corresponding mechanisms of this circRNA were carefully explored. Overall, we revealed the role of circEPB41L2 during HCC progression and identified it as a novel prognostic biomarker and potential therapeutic target for HCC patients.
Materials and Methods
Clinical Samples
Primary tumor samples and matched para-tumor samples were collected from 100 HCC patients at Mengchao Hepatobiliary Hospital of Fujian Medical University. None of the included patients received any chemotherapy or radiation before surgery. The human tissue specimens used in this project have been approved by the Institutional Review Board of Mengchao Hepatobiliary Hospital of Fujian Medical University, and the written informed consents of all patients were obtained. The methodology conforms to the standards set out in the Helsinki Declaration.
CircRNAs Screening
New circRNAs prediction and circRNAs expression analysis were performed on RNA-seq data of 61 HCC tissues and para-tumor tissues. The expression of circRNAs was calculated based on effective circRNA reads, which were normalized using library sizes and normalization factors extracted from linear transcripts as counts per million reads between HCC samples. The differentially expressed circRNAs were screened based on the multiple changes (fold change ≥2.0, p < 0.05) in HCC tissues and matched para-tumor tissues. Volcano plot was used to demonstrate the differential expression of these circRNAs. The data were analyzed by using R software limma.
Cell Culture
Human HCC cell lines including Hep3B, SK-Hep-1, Snu-449 and human embryonic kidney cell line HEK-293T were purchased from American Type Culture Collection (ATCC, Rockefeller, MD, USA). Human HCC cell line SMMC-7721 was purchased from the Chinese Cell Bank of the Chinese Academy of Sciences (Shanghai, China). Cells were cultured in Minimum Essential Medium (MEM; Gibco, Carlsbad, CA, USA), Dulbecco’s modified Eagle medium (DMEM; Gibco, Carlsbad, CA, USA) or RPMI-1640 (Gibco, Carlsbad, CA, USA) with 10% fetal bovine serum (ExCell Bio, Australia). Cells were cultured in an atmosphere of 95% humidified air and 5% CO2 at 37 °C.
Lentivirus Constructs and Cell Transfection
The full-length sequence of circEPB41L2 was cloned into pLO-ciR vector for constructing pLO-ciR-circEPB41L2 overexpression plasmid (termed as ex-circEPB41L2) which was further verified by Sanger sequencing. The corresponding empty pLO-ciR vector was used as a negative control (termed as ex-NC). Moreover, a short hairpin RNA (shRNA) against circEPB41L2 was also designed (shRNA sequence: sense: 5ʹ-GCCCTGAGCAGAAACATAA-3ʹ; anti-sense: 5ʹ-TTATGTTTCTGCTCAGGGC-3ʹ) and cloned into PLKO.1-GFP plasmid to knockdown circEPB41L2 (termed as sh-circEPB41L2) in HCC cell lines. The corresponding empty PLKO.1-GFP vector was used as a negative control (termed as sh-NC). Afterwards, HEK-293T cells were co-transfected with lentivirus packaging plasmid (Invitrogen, Waltham, MA, USA) and corresponding ex-circEPB41L2/ex-NC or sh-circEPB41L2/sh-NC expression plasmids using the Lipofectamine 3000 (Invitrogen) according to the manufacturer’s protocol.
Lentiviruses with a minimum titer of 5×105 infectious units (IFU)/mL were subject to later reprogramming experiments. Together with 5 μg/mL Polybrene (Santa Cruz Biotechnology, Santa Cruz, CA, USA), SMMC-7721 and Hep3B cells were co-transduced with ex-circEPB41L2/ex-NC or sh-circEPB41L2/sh-NC, respectively. After 48h of transfection, the transfected cells were cultured in the medium containing 2 μg/mL puromycin (Sigma-Aldrich, St. Louis, MO, USA) for 2 weeks to gain four positive stable cell lines for in vitro and in vivo experiments: circEPB41L2 overexpression cell line (ex-circEPB41L2 SMMC-7721), the overexpression control cell line (ex-NC SMMC-7721), circEPB41L2 knockdown cell line (sh-circEPB41L2 Hep3B), the knockdown control cell line (sh-NC Hep3B).
Real-Time Quantitative PCR (qRT-PCR)
Total RNA was extracted from HCC tissues, para-tumor tissues and HCC cells using Trizol reagent (TransGen Biotech, Beijing, China) according to the manufacturer’s instructions. Reverse transcription was carried out using a Transcriptor First Strand cDNA Synthesis Kit (Roche, Indianapolis, IN, USA). We specifically designed primers for backsplice junction sites of four circRNAs (hsa_circ_0001727, hsa_circ_0001543, hsa_circ_0002538 and circEPB41L2) to determine their abundance in tissues or cells (Table 1). The SYBR Premix Ex Taq II (Takara, Dalian, China) was used to conduct qRT-PCR in the StepOne Software V2.3 thermal circulation system. The amplification parameters were as follows: pre-denaturation at 95 °C for 10 min, denaturation at 95 °C for 30 s, annealing at 60 °C for 30 s, extension at 72 °C for 30 s, a total of 40 cycles. Relative RNA expression calculated with 2 −ΔΔCt method, using GAPDH or 18S as internal controls. QRT-PCR was performed at least 3 times for each circRNA.
 |
Table 1 Real-Time-PCR Primer Sequence |
Actinomycin D and RNase R Treatment
For actinomycin D treatment, SMMC-7721 cells were treated with actinomycin D (2μg/mL) and total RNA was extracted at 0h, 3h, 6h, 9h and 12h. The dynamic changes of circEPB41L2 and the corresponding linear EPB41L2 mRNA (mEPB41L2) expression levels were detected by qRT-PCR. For RNase R treatment, RNase R (20U/μL) was added to 5 μg total RNA (3U/μg) and incubated at 37°C for 5 min. RNase R− free group was used as negative control. Then, the digested total RNA was transcribed into cDNA for qRT-PCR detection.
Cell Proliferation Assay (CCK-8)
The CCK-8 assays were performed in six repeated wells using the Cell Counting Kit-8 assay kit (TransGen Biotech), following the manufacturer’s guidelines. In brief, 5×103 cells were seeded into a 96-well plate and incubated for 0, 24, 48, 72 and 96 h. Subsequently, 10μL CCK-8 reagents added into each well and incubated for another 2h. The OD450 absorption value was measured using an automatic microplate reader (Molecular Devices LLC, Sunnyvale, CA, USA).
Colony Formation Assay
Cells were seeded in six-well plates at a density of 2 × 103 per well and incubated at 37 °C in 5% CO2 for 10 days. After that, formed colonies were fixed with 4% paraformaldehyde (Beyotime Biotechnology, Shanghai, China) for 30 min and then stained with 0.1% Crystal violet dye (Beyotime Biotechnology) for 15 min. Finally, the colonies were counted and photographed. The experiments were repeated independently three times.
Wound-Healing Test
A wound-healing system was used to determine the migration ability of HCC cells. Ex-circEPB41L2-SMMC-7721 or sh-circEPB41L2-Hep3B cells were seeded on both sides of the scratch dish in a number of 7×104 cells per well. Ex-NC and sh-NC served as negative control, respectively. After overnight cultivation, the scratch dish was removed. The cells were then added with DMEM medium containing 2% FBS and cultured in an incubator at 37°C with 5% CO2. The images were photographed and analyzed by Image J software after 24 hours.
Transwell Assay
Cell metastatic properties were detected by a transwell plate containing 8-μm pores membrane filter. To detect cell migration, 1×105 cells in 200 μL serum-free medium were placed in the upper chamber. For invasion assay, 1×105 cells in 200μL serum-free medium were placed in the top compartment with matrigel coating. In both assays, 600 μL of medium with 10% FBS were supplied into the bottom units. After 24 hours of incubation, the penetrated cells through the pores of inserts were fixed and stained. Penetrating cells were captured and calculated in five randomly selected fields. Each experiment was repeated three times.
Double Luciferase Reporting Experiment
The interaction between circEPB41L2 and miR-590-5p was verified using the PMIR-ReportTM system. The full-length circEPB41L2 sequence was inserted into the downstream of luciferase expression sequence to construct the report plasmid. According to the manufacturer’s instructions, HEK-293T cells were co-transfected with a reporter plasmid containing circEPB41L2, miR-590-5p mimic (GENERAL BIOSYSTEMS) and pRL-Renilla plasmid in a 24-well plate. The cells were collected after 36 hours, and the luciferase activity was detected by TransDetect Double-Luciferase Reporter Assay Kit. PRL-renilla activity as an internal control.
Western Blotting
Total protein was extracted from cells using RIPA lysis buffer, and quantitatively analyzed using a BCA kit (TransGen Biotech). Then western blot analysis was carried out following the manufacturer’s instructions. In brief, proteins were separated by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and transferred onto nitrocellulose filter membrane. Then, the membranes were blocked in 5% FBS at room temperature for 2 hours. After washing three times with TBST, membranes were incubated with primary antibody purchased from cell signaling technology [rabbit anti-GAPDH (1:5000; 5174S), rabbit anti-E-cadherin (1:1000; 3195S), rabbit anti-N-cadherin (1:1000; 13116S), rabbit anti-vimentin (1:1000; 5741S), rabbit anti-Snail (1:1000; 3879S), rabbit anti-Slug (1:1000; 9585S)] at 4°C overnight and were then incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies (1:5000) at room temperature for 2h. Finally, membranes were washed three times before protein bands were detected using enhanced chemiluminescence substrate on ChemiDocTM MP Imaging system.
Xenograft Tumor Mouse Model
Four-week-old male BALB/C athymic nude mice were used for in vivo experiments. Animal experiments were conducted in strict accordance with the procedures approved by the animal management committee of Mengchao Hepatobiliary Hospital of Fujian Medical University. And, 1×10^7 of ex-circEPB41L2-SMMC-7721 cells and the corresponding negative control cells were subcutaneously injected into the forelimb armpit of nude mice, respectively. Tumor volumes were measured every 3 days, and the mice were killed after 3 weeks. The tumor was removed and photographed, and immediately frozen at −80°C for future use.
Statistic Analysis
SPSS 19.0 software was used for statistical analysis, and the data were expressed as mean ± standard deviation. The two-tailed unpaired student's t-test was applied to assess the statistical significance between two groups, the statistical significance between more than two groups was analyzed by one-way variance (ANOVA) analysis. Fisher’s precise test was used to assess the relationship between circEPB41L2 expression and clinical characteristics. The prognostic correlation of circEPB41L2 was analyzed by univariate and multivariate Cox analyses. The difference was deemed as statistically significant at p < 0.05. All the experiments were repeated more than three times.
Results
Down-Regulated Expression of CircEPB41L2 is in HCC Tissues
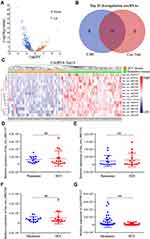
Firstly, we identified the dysregulation of circRNA profiles during HCC progression in the transcriptome data of 61 pairs of HCC tumor and matched para-tumor tissues from Mengchao Hepatobiliary Hospital of Fujian Medical University. CircRNAs were first identified using CIRI and then the Bioconductor package “limma” was used to compare the expression level of circRNA between HCC tissues and matched para-tumor tissues. A total of 881 circRNAs, which were dysregulated in HCC tumor tissue when compared with matched para-tumor tissues (fold change ≥2 and p < 0.05), are shown in the volcano plot (Figure 1A). Among the 881 dysregulation circRNAs, 156 and 725 of them were significantly up-regulated and down-regulated in HCC tissues, respectively. To provide a more reliable results, we performed another differential analysis using circRNAs identified by circTest. The intersection of the top 20 differentially expressed circRNAs from the above analysis contains 14 circRNAs for downstream analysis (Figure 1B and C).
Then, we selected four circRNAs (hsa_circ_0001727, hsa_circ_0001543, hsa_circ_0002538 and circEPB41L2) with the most significant different expression between HCC and para-tumor to further validate their expression in another 15 HCC patients. Interestingly, only circEPB41L2 was significantly down-regulated in HCC compared with matched para-tumor tissue samples (p = 0.0018, Figure 1D–G). Taken together, these results indicated that the dysregulation of circEPB41L2 might be associated with HCC progression.
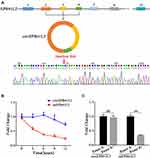
Characteristics of the Circular RNA CircEPB41L2
The erythrocyte protein band 4.1 family has been reported to regulate cancer cell proliferation and metastasis in HCC.16 CircEPB41L2 was generated by reverse splicing between splicing acceptor of exons 2, 3, and 4 from erythrocyte protein band 4.1 family member EPB41L2. The expected sequences of circEPB41L2 were amplified using divergent primers and confirmed by sanger sequencing (Figure 2A). To further evaluate the stability of circEPB41L2 in HCC, we quantified the level of circEPB41L2 and the corresponding mEPB41L2 by actinomycin D (an inhibitor of transcription) intervention and RNase R treatment. As expected, qRT-PCR assays revealed that circEPB41L2 was highly stable with a half-life of >12 h, whereas mEPB41L2 exhibited a half-life of <6 h (Figure 2B). Moreover, anti-digestion experiments also confirmed that circEPB41L2 was resistant to RNase R, while mEPB41L2 was easily degraded (Figure 2C). In summary, our results showed the high structural stability of circEPB41L2, which is consistent with the general characteristics of circRNAs.
Correlation of CircEPB41L2 with Clinical Features
To investigate the clinical role of circEPB41L2 during HCC progression, we next analyzed the association between the expression levels of circEPB41L2 and the clinical characteristics in 100 HCC patients. Using the median circEPB41L2 expression level as cut-off values, 100 HCC patients were divided into two groups with low circEPB41L2 expression or high circEPB41L2 expression. As summarized in Table 2, HCC patients with low circEPB41L2 expression in tumor tissues were associated with bigger tumor size (p = 0.009), more possibility of vascular invasion (p = 0.016) and higher alpha-fetoprotein (AFP, p = 0.002). Moreover, the multivariable Cox proportional hazards model demonstrated that the expression of circEPB41L2 in HCC could serve as an independent prognostic factor for overall survival (OS, p = 0.039, Table 3). Meanwhile, Kaplan–Meier analysis showed that the HCC patients with low circEPB41L2 expression showed significant worse prognosis in both recurrence-free survival (RFS, p = 0.036) and overall survival (OS, p < 0.001, Figure 3A and B). In summary, these results suggested that down-regulated circEPB41L2 expression in HCC tissues is well associated with poor prognosis of HCC patients.
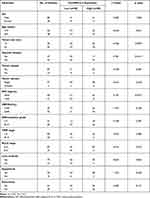 |
Table 2 Correlation Between CircEPB41L2 Expression and Clinical Data |
 |
Table 3 Univariate and Multivariate Analyses of Factors Associated with OS of HCC Patients |
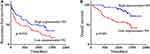 |
Figure 3 Correlation of circEPB41L2 expression with clinical features. (A) and (B) Kaplan–Meier analysis of the association between circEPB41L2 expression level and RFS (A)/OS (B) of HCC patients. |
The Function and Molecular Mechanism of CircEPB41L2 in HCC
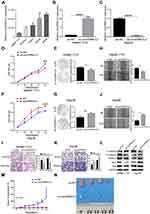
To further explore the functional roles of circEPB41L2, the expression of circEPB41L2 was firstly evaluated in five HCC cell lines (SMMC-7721, SK-Hep-1, HepG2, Snu449 and Hep3B). As shown in Figure 4A, circEPB41L2 was relatively low expressed in SMMC-7721 and relatively highly expressed in Hep3B cells. Therefore, we selected SMMC-7721 and Hep3B cells as models for lentivirus transduction to establish stable circEPB41L2-overexpression and circEPB41L2-knockdown cell lines, respectively. As shown in Figure 4B and C, the qRT-PCR results indicated that circEPB41L2 was ~10 times overexpressed in ex-circEPB41L2-SMMC7721 cell lines and 92% of knockdown efficiency was achieved in sh-circEPB41L2-Hep3B cell lines.
As the correlation between the circEPB41L2 expression and clinical features suggested that circEPB41L2 function is related to the HCC proliferation and invasion, we further verified its biological functions in stable circEPB41L2 overexpression and knockdown HCC cells. As shown in Figure 4D–G, both CCK-8 assays and colony formation assays showed that circEPB41L2 overexpression significantly inhibited cell proliferation of SMMC-7721 cells, while cell proliferation was markedly promoted in sh-circEPB41L2-Hep3B cells. Moreover, wound healing assays and trans-well assays showed that circEPB41L2 overexpression could slow down the migration and invasion of HCC cells, while circEPB41L2 knockdown produced opposite results (Figure 4H–K). Surprisingly, we noticed that overexpression of circEPB41L2 could also increase the expression of E-cadherin and inhibit the expression of N-cadherin, vimentin, Snail, and Slug proteins, while the opposite trend was observed in sh-circEPB41L2-Hep3B cells (Figure 4L). These genes are all important players in epithelial-mesenchymal transformation (EMT) process, which has been reported as one of the most important processes in HCC metastasis. The regulatory interaction between circEPB41L2 and these genes supported that circEPB41L2 inhabit HCC migration and invasion though EMT pathways.
Moreover, to determine the efficacy of circEPB41L2 in tumor growth in vivo, HCC xenograft mouse model was conducted for ex-circEPB41L2-SMMC-7721 cells. Mouse injected with ex-circEPB41L2-SMMC-7721 cells showed a significant suppressed cell growth in vivo, compared with those injected with negative control cells (p = 0.031, Figure 4M and N). These results suggested that circEPB41L2 functions as a suppressor in HCC cell proliferation, migration, and invasion, which was well consistent with previous clinical analysis results.
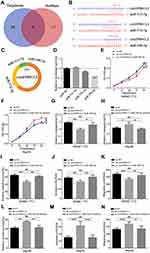
CircEPB41L2 Serves as a miR-590-5p Sponge During HCC Progression
Current studies have shown that an important function of circRNA is as a “sponge” to bind functional miRNA, therefore we speculated that circEPB41L2 may also serve as miRNA sponges for HCC progression. To identify the potential miRNAs that could be bound to circEPB41L2, we analyzed two independent miRNA database (TargetScan and starBase). The Venn diagram in Figure 5A and B showed that a total of 3 miRNAs (miR-515-5p, miR-576-5p and miR-590-5p) were predicated to have strong binding affinity, along with miRNA seed sequence matching circEPB41L2 sequence. The binding sites of these three miRNAs on circEPB41L2 are shown in Figure 5C. Then, luciferase screening assays were performed to validate the binding affinity for these three miRNAs. As shown in Figure 5D, only miR-590-5p showed reduced luciferase reporter activities of circEPB41L2 in HEK-293T at ~50%, when comparing with miR-NC. These results indicated that circEPB41L2 could directly bind miR-590-5p. To further determine whether the circEPB41L2 could affect cell proliferation and migration in HCC via sponging miR-590-5p, we used miR-590-5p mimics and miR-590-5p inhibitor to treat the ex-circEPB41L2-SMMC-7721 and sh-circEPB41L2-Hep3B cells, respectively. As shown in Figure 5E and G, the CCK-8 assays and colony formation assays showed that the decreased cell activity induced by circEPB41L2 could be rescued by miR-590-5p (Figure S1A). Meanwhile, analogously miR-590-5p inhibitor restored the cell activity that was increased by sh-circEPB41L2 (Figure 5F and H, Figure S1B). The similar dynamic trends of cell invasion and migration were also confirmed by wound healing assays and trans-well assays (Figure 5I–N, Figure S1C-F). Overall, these results demonstrated that circEPB41L2 could serve as a tumor suppressor by regulating miR-590-5p during HCC progression.
Discussion
With the development of next-generation sequencing technology, the biological functions and molecular mechanisms of circRNAs in solid tumors have been gradually recognized.17,18 In recent years, more and more studies have demonstrated that circRNAs played important roles during HCC progression and provided new insights for the diagnosis, treatment and the study of the pathogenesis of HCC.19,20 Several circRNAs, such as circADAMTS13, circBACH1 and circMET, have already been suggested to play important roles during HCC progression by acting as sponges of oncogenic miRNA and could be potential targets for HCC treatment.21–23 In our study, we also identified a circRNA named as circEPB41L2. Through comprehensive investigation of the circRNA profiles in 61 pairs of HCC tumor and matched para-tumor tissues, we found that circEPB41L2 was specifically lowly expressed in HCC tissues. Combined with clinical parameters of HCC patients and in vivo and in vitro experiments, we confirmed that circEPB41L2 plays an important role in the proliferation and metastasis of HCC.
CircEPB41L2 has been previously reported to be down-regulated in human lung adenocarcinoma and bladder cancer, and could predict poor prognosis for patients.24,25 In this study, we also found that the downregulation of circEPB41L2 expression in HCC tissues could promote cell proliferation and metastasis, and thus was significantly correlated with worse RFS time and OS time for HCC patients. Taken together, those results suggested that circEPB41L2 might function as a suppressor role in many solid tumors, including HCC.
Of note, circRNA/miRNA axis is a classical regulatory mechanism in current circRNA function studies, which can regulate gene expressions at both transcription and post transcription levels. miR-590-5p was reported to be a crucial factor for the precancerous process of HCC. Previous studies showed that miR-590-5p regulates the proliferation and invasion of HCC cells by targeting TGF-β RII.26 Although YAP1 could serve as a regulator for miR-590-5p expression,27 the direct upstream player of miR-590-5p remained unclear. Undoubtedly, our study firstly showed that circEPB41L2 could directly inhibit the activity of oncogenic miR-590-5p and further exert its influence in HCC cell proliferation and metastasis, providing new insights into the regulatory mechanisms of miR-590-5p in HCC.
In summary, our study demonstrates that low expression of circEPB41L2 in HCC tissue was significantly associated with HCC prognosis. Moreover, circEPB41L2 could serve as a key modulator of HCC cell proliferation and migration by efficiently sponging miR-590-5p. Therefore, circEPB41L2 has the potential to be a diagnostic biomarker and a therapeutic target for HCC therapy.
Abbreviations
EPB41L2, erythrocyte protein band 4.1-like 2; circRNA, circular RNA; HCC, hepatocellular carcinoma; miRNA, microRNA; CCK-8, Cell Counting Kit-8; OS, overall survival; RFS, recurrence-free survival; SD, standard deviation; qRT-PCR, real-time quantitative PCR; AFP, alpha-fetoprotein; HBV, hepatitis B virus; TNM, tumor-node-metastasis; HR, hazard ratio.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grant No. 81802413), the Medical innovation project of Fujian Province (2018-CX-35), the Scientific Foundation of Quanzhou City (2018Z133), the Natural Science Foundation of Fujian Province (2019J01139).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Ferlay J, Soerjomataram I, Dikshit R, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136(5):E359–E386. doi:10.1002/ijc.29210
3. Marrero JA. Surveillance for Hepatocellular Carcinoma. Clin Liver Dis. 2020;24:611–621. doi:10.1016/j.cld.2020.07.013
4. Marrero JA, Kulik LM, Sirlin CB, et al. Diagnosis, staging, and management of hepatocellular carcinoma: 2018 Practice Guidance by the American Association for the Study of Liver Diseases. Hepatology. 2018;68(2):723–750. doi:10.1002/hep.29913
5. Barrett SP, Salzman J. Circular RNAs: analysis, expression and potential functions. Development. 2016;143(11):1838–1847. doi:10.1242/dev.128074
6. Shang Q, Yang Z, Jia R, et al. The novel roles of circRNAs in human cancer. Mol Cancer. 2019;18(1):6. doi:10.1186/s12943-018-0934-6
7. Lasda E, Parker R. Circular RNAs: diversity of form and function. RNA. 2014;20(12):1829–1842. doi:10.1261/rna.047126.114
8. Qu S, Yang X, Li X, et al. Circular RNA: a new star of noncoding RNAs. Cancer Lett. 2015;365:141–148. doi:10.1016/j.canlet.2015.06.003
9. Jakobi T, Dieterich C. Computational approaches for circular RNA analysis. Wiley Interdiscip Rev RNA. 2019;10(3):e1528. doi:10.1002/wrna.1528
10. Bach DH, Lee SK, Sood AK. Circular RNAs in Cancer. Mol Ther Nucleic Acids. 2019;16:118–129. doi:10.1016/j.omtn.2019.02.005
11. Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333–338. doi:10.1038/nature11928
12. Guo JU, Agarwal V, Guo H, et al. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014;15(7):409. doi:10.1186/s13059-014-0409-z
13. Fu X, Zhang J, He X, et al. Circular RNA MAN2B2 promotes cell proliferation of hepatocellular carcinoma cells via the miRNA-217/MAPK1 axis. J Cancer. 2020;11(11):3318–3326. doi:10.7150/jca.36500
14. Zhang S, Liu Y, Liu Z, et al. CircRNA_0000502 promotes hepatocellular carcinoma metastasis and inhibits apoptosis through targeting microRNA-124. J BUON. 2019;24:2402–2410.
15. Liu Z, Yu Y, Huang Z, et al. CircRNA-5692 inhibits the progression of hepatocellular carcinoma by sponging miR-328-5p to enhance DAB2IP expression. Cell Death Dis. 2019;10(12):900. doi:10.1038/s41419-019-2089-9
16. Yang X, Yu D, Ren Y, et al. Integrative functional genomics implicates EPB41 dysregulation in hepatocellular carcinoma risk. Am J Hum Genet. 2016;99(2):275–286. doi:10.1016/j.ajhg.2016.05.029
17. Liu H, Liu Y, Bian Z, et al. Circular RNA YAP1 inhibits the proliferation and invasion of gastric cancer cells by regulating the miR-367-5p/p27 (Kip1) axis. Mol Cancer. 2018;17:151. doi:10.1186/s12943-018-0902-1
18. Wang M, Yu F, Li P. Circular RNAs: characteristics, function and clinical significance in hepatocellular carcinoma. Cancers. 2018;10(8):258. doi:10.3390/cancers10080258
19. Yao R, Zou H, Liao W. Prospect of circular RNA in hepatocellular carcinoma: a novel potential biomarker and therapeutic target. Front Oncol. 2018;8:332. doi:10.3389/fonc.2018.00332
20. Xu L, Feng X, Hao X, et al. CircSETD3 (Hsa_circ_0000567) acts as a sponge for microRNA-421 inhibiting hepatocellular carcinoma growth. J Exp Clin Cancer Res. 2019;38(1):98. doi:10.1186/s13046-019-1041-2
21. Qiu L, Huang Y, Li Z, et al. Circular RNA profiling identifies circADAMTS13 as a miR-484 sponge which suppresses cell proliferation in hepatocellular carcinoma. Mol Oncol. 2019;13:441–455. doi:10.1002/1878-0261.12424
22. Liu B, Yang G, Wang X, et al. CircBACH1 (hsa_circ_0061395) promotes hepatocellular carcinoma growth by regulating p27 repression via HuR. J Cell Physiol. 2020;235(10):6929–6941. doi:10.1002/jcp.29589
23. Huang XY, Zhang PF, Wei CY, et al. Circular RNA circMET drives immunosuppression and anti-PD1 therapy resistance in hepatocellular carcinoma via the miR-30-5p/snail/DPP4 axis. Mol Cancer. 2020;19:92. doi:10.1186/s12943-020-01213-6
24. Shen C, Wu Z, Wang Y, et al. Downregulated hsa_circ_0077837 and hsa_circ_0004826, facilitate bladder cancer progression and predict poor prognosis for bladder cancer patients. Cancer Med. 2020;9(11):3885–3903. doi:10.1002/cam4.3006
25. Zhang SJ, Ma J, Wu JC, et al. CircRNA EPB41L2 inhibits tumorigenicity of lung adenocarcinoma through regulating CDH4 by miR-211-5p. Eur Rev Med Pharmacol Sci. 2020;24(7):3749–3760. doi:10.26355/eurrev_202004_20839
26. Jiang X, Xiang G, Wang Y, et al. MicroRNA-590-5p regulates proliferation and invasion in human hepatocellular carcinoma cells by targeting TGF-beta RII. Mol Cells. 2012;33:545–551. doi:10.1007/s10059-012-2267-4
27. Chen M, Wu L, Tu J, et al. miR-590-5p suppresses hepatocellular carcinoma chemoresistance by targeting YAP1 expression. EBioMedicine. 2018;35:142–154. doi:10.1016/j.ebiom.2018.08.010
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.