Back to Journals » Journal of Inflammation Research » Volume 14
Altered Monocyte Subsets in Kawasaki Disease Revealed by Single-cell RNA-Sequencing
Authors Geng Z, Tao Y , Zheng F, Wu L, Wang Y, Wang Y, Sun Y, Fu S, Wang W, Xie C, Zhang Y, Gong F
Received 25 November 2020
Accepted for publication 9 March 2021
Published 16 March 2021 Volume 2021:14 Pages 885—896
DOI https://doi.org/10.2147/JIR.S293993
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 4
Editor who approved publication: Professor Ning Quan
Zhimin Geng,* Yijing Tao,* Fenglei Zheng, Linlin Wu, Ying Wang, Yujia Wang, Yameng Sun, Songling Fu, Wei Wang, Chunhong Xie, Yiying Zhang, Fangqi Gong
Department of Cardiology, Children’s Hospital, Zhejiang University School of Medicine, National Clinical Research Center for Child Health, Hangzhou, Zhejiang, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Fangqi Gong
Department of Cardiology, Children’s Hospital, Zhejiang University School of Medicine, National Clinical Research Center for Child Health, Hangzhou, Zhejiang, People’s Republic of China
Tel/Fax +86 571 86670008
Email [email protected]
Background: Kawasaki disease (KD) is characterized by a disorder of immune response, and its etiology remains unknown. Monocyte is an important member of the body’s innate immune system; however its role in KD is still elusive due to its ambiguous heterogeneity and complex functions. We aim to comprehensively delineate monocyte heterogeneity in healthy and KD infants and to reveal the underlying mechanism for KD.
Methods: Peripheral monocytes were enriched from peripheral blood samples of two healthy infants and two KD infants. scRNA-seq was performed to acquire the transcriptomic atlas of monocytes. Bio-information analysis was utilized to identify monocyte subsets and explore their functions and differentiation states. SELL+CD14+CD16- monocytes were validated using flow cytometry.
Results: Three monocyte subsets were identified in healthy infants, including CD14+CD16- monocytes, CD14+CD16+ monocytes, and CD14LowCD16+ monocytes. Cell trajectory analysis revealed that the three monocyte subsets represent a linear differentiation, and possess different biological functions. Furthermore, SELL+CD14+CD16- monocytes, which were poorly differentiated and relating to neutrophil activation, were found to be expanded in KD.
Conclusion: Our findings provide a valuable resource for deciphering the monocyte heterogeneity in healthy infants and uncover the altered monocyte subsets in KD patients, suggesting potential biomarkers for KD diagnosis and treatment.
Keywords: Kawasaki disease, monocyte subsets, scRNA-seq
Introduction
KD is an acute, self-limited disease with systemic vasculitis,1 which is the leading cause of acquired heart disease in children of developed countries due to coronaritis. Although its etiology remains unknown, most studies have shown that KD is characterized by a disorder of immune response to one or more traditional antigens influenced by genetic determinants.2
Studies have confirmed that monocyte plays an important role in vasculitis.3,4 Previously, human monocytes were divided into 3 principal subsets, including CD14+CD16- monocytes named classical monocytes (CM), CD14+CD16+, and CD14LowCD16+ monocytes that were known as intermediate monocytes (IM) and non-classical monocytes (NCM), respectively.5 IM increase in acute KD6 and the activated monocytes result in the synthesis and secretion of inflammatory cytokines and chemokines, which can activate endothelial cells and support the secondary recruitment of neutrophils.7 Hence, activated monocytes play a key role in the pathogenesis and vascular inflammation of KD.8 However, the role of activated monocytes in KD vasculitis is still not clear enough to serve as a therapeutic target, mainly because of the obvious heterogeneity and complex functions of monocytes.9 A possible solution to this problem is to accurately separate the monocyte subsets, so the subsets that promote KD could be precisely targeted while the other subsets can be undisturbed and maintain homeostasis.
With the development of mass cytometry and single-cell RNA-seq (scRNA-seq) techniques, it is possible to accurately separate different monocyte subsets and reveal their distinct functions. Studies have found that 8 monocyte subsets are identified with a panel of 39 antibodies against cell-surface proteins using mass cytometry approach.10 Moreover, 4 monocyte subsets are defined at the single-cell transcriptome level on the basis of 339 monocytes.11 However, identities of monocyte subsets remain unclear, as the subsets were identified by only a restricted set of surface markers and number of monocytes was insufficient to detect rare cell subsets. In addition, these studies mainly focused on the monocytes in adults. As the immune system changes during ageing12 and infants are more prone to KD,1 it is essential to identify heterogeneity of monocytes in infants and figure out the monocyte subset that works in KD patients.
To better explore the monocyte subset that functions in KD patients, the transcriptomic profiles of a total of 8880 enriched monocytes from 2 healthy infants and 2 KD patients were acquired using scRNA-seq. Using discriminative gene markers, we identified the functions of the defined monocyte subsets and their differentiation relationship. In addition, the monocyte subset that functions in KD could be distinguished by the featured gene expression profile and was associated with neutrophil activation. Thus, our findings will improve the current understanding about the mechanism for monocyte subsets in KD vasculitis, and are potentially valuable in providing novel diagnostic markers for KD.
Materials and Methods
The data and computer code that support the findings of this study are available from the corresponding author on reasonable request.
Human Subjects
We studied 2 patients with acute KD (aged 10 months) and 2 healthy infants (aged 10 months) under well-child checking at Children’s Hospital, Zhejiang University School of Medicine in September 2019. KD was diagnosed according to the diagnosis criteria established by the American Heart Association.13 Two-dimensional echocardiography was used to examine whether there was abnormal cardiac function and coronary artery lesions during the acute and convalescent disease phases. Neither KD patients developed coronary aneurysms 1 month after the disease onset. Clinical data of the 4 subjects are summarized in Supplementary Table 1.
Peripheral blood samples (2 mL each sample) were collected from the four subjects. For the 2 patients with KD, blood samples were collected at acute stage before intravenous immunoglobulin and corticosteroids treatment. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Institutional Review Board of Children’s Hospital, Zhejiang University School of Medicine (IRB number: 2019-IRB-073). Parents or guardians of the participants have provided their informed consent to participate in the study.
Enrichment of Monocytes
Peripheral blood mononuclear cells were isolated using Ficoll-Paque PLUS (GE Healthcare Biosciences AB) according to the manufacturer’s protocol. Briefly, we carefully layered the blood sample diluted with phosphate-buffered saline (4 mL) onto the Ficoll-Paque media solution (3 mL) and obtained mononuclear cell layer after centrifuging at 400 g for 30 minutes at 19°C. The mononuclear cells were cryopreserved in liquid nitrogen after washed twice. Enriched monocytes were purified from the thawed mononuclear cells by negative magnetic sorting techniques using Pan Monocyte Isolation Kit (Miltenyi Biotec, Bergisch Gladbach, Germany) according to the manufacturer’s instructions, and then cryopreserved in liquid nitrogen.
Flow Cytometry Analysis
The enriched monocytes were thawed and suspended in PBS supplemented with 10% fetal bovine serum (Gibco, Invitrogen), and then stained with phycoerythrin [PE]-conjugated anti-human CD14 and fluorescein isothiocyanate [FITC]-conjugated anti-human CD16 or their isotype controls which were PE and FITC-conjugated mouse IgG1, respectively (κ isotype control; BioLegend), for identifying monocytes before scRNA-seq. Fluorescence intensity was examined by a Beckman Gallios Flow Cytometer, and analyzed by flow cytometry software Kaluza analysis 2.0 (Beckman Coulter Life Sciences). For the validation of SELL+ CM in KD infants, the mononuclear cells were thawed, and stained with FITC conjugated anti-human CD14 (BD Pharmingen™), PE-Cy7 conjugated anti-human CD16 (BD Pharmingen™), PE-conjugated SELL (BioLegend), or their isotype controls (BioLegend). Fluorescence intensity was achieved on a Beckman Gallios Flow Cytometer, and analyzed by flow cytometry software FLOWJo (BD Pharmingen™). Student’s t test was used to compare the ratio of SELL+CM to CM between the healthy and KD infants.
Single Cell Multiplexing Labelling and Single Cell Transcriptome Capturation
The enriched monocytes of each sample were firstly stained with two fluorescent dye, Calcein AM (Thermo Fisher Scientific Cat. No. C1430) and Draq7 (Cat. No. 564904), for precise determination of cell concentration and viability via BD Rhapsody™ Scanner before single-cell multiplexing labelling. The cell viability varies from 60% to 80%. Enriched monocytes of each sample were sequentially labeled with BD Human Single-Cell Multiplexing Kit (Cat. No. 633781) which utilizing an innovative antibody-oligo technology14 mainly to provide higher sample throughput and eliminate batch effect for single-cell library preparation and sequencing. Briefly, cells from each sample were labeled by antibodies with different sample tags (sample tag 06–09) and washed with BD Pharmingen™ Stain Buffer (Cat. No. 554656) before pooling all samples together. BD Rhapsody Express system based on Fan et al15 was used for single-cell transcriptomics capturation. Pooled samples were then loaded in one BD Rhapsody™ Cartridge that was primed and treated strictly following the manufacturers protocol (BD Biosciences). Cell capture beads were then loaded excessively onto the cartridge, and the excessive beads were washed away from the cartridge. Viable cells with beads captured in wells were detected in BD Rhapsody™ Scanner. Then, cells were lysed, and cell capture beads were retrieved and washed prior to performing reverse transcription and treatment with exonuclease I.
Single-Cell RNA-Seq
Transcriptomic and sample tag information of single cells were obtained through BD Rhapsody System. Microbeads-captured single-cell transcriptome and sample tag sequences were generated into cDNA library and sample tag library separately containing cell labels and unique molecular identifiers (UMI) information, respectively. All the libraries were sequenced in a PE150 mode (Pair-End for 150 bp read) on the NovaSeq platform (Illumina).
Single-Cell Data Analysis
Raw sequencing reads of cDNA library and sample tag library were processed through the BD Rhapsody Whole Transcriptome Assay Analysis Pipeline (Early access), which included filtering by reads quality, annotating reads, annotating molecules, determining putative cells, and generating single-cell expression matrix.
The R package Seurat (version 3.1.5)16 was used for subsequent analysis. Raw gene expression matrices from the cartridge were read into R (version 4.0.1) and converted to Seurat objects. Cells label as “undetermined” and “multiplet” were excluded in the following analysis. Cells with more than 25% mitochondrial genes were also excluded from the analysis. The gene expression matrix was then normalized to the total cellular UMI count. In order to reduce dimensionality, PCA was performed based on the highly variable genes after scaling the data with respect to UMI counts. Principle components were chosen for downstream clustering based on heatmap of principle components, Jackstraw plot, and elbow plot of principle components to further reduce dimensionality using the t-distributed stochastic neighbor embedding (tSNE) algorithm. Marker genes of each cluster were calculated using the FindAllMarkers function with the Wilcoxon Rank-Sum Test under the following criteria: log2 fold change > 0.25; p < 0.01; min. PCT > 0.25. Differential expressed genes between KD-CM (SELL+CD14+CD16- monocytes) and CM were identified by the FindMarkers function with criteria of log2 fold change > 0.25; p < 0.05; min. PCT > 0.25.
Gene Ontology Enrichment Analysis
Gene ontology analysis was used to explore the functions of distinct monocyte subsets using Metascape (http://metascape.org/).17 Gene ontology terms with adjusted P-value < 0.05 were considered significantly enriched.
Construction of Single-Cell Trajectory
Monocle18 uses an algorithm to learn the sequence of gene expression changes each cell must go through as part of a dynamic biological process. Once it has learned the overall “trajectory” of gene expression changes, Monocle can arrange each cell in the trajectory. Monocle 2 was used to construct a single-cell trajectory with genes differentially expressed in the cells collected at the beginning of the process compared with those at the end.
Results
Single Cell Profile of Enriched Monocytes
To examine the heterogeneity of monocytes, scRNA-seq was used to identify cellular signature. To ensure the purity of monocytes used in scRNA-seq, flow cytometry was performed to analyze the proportion of monocytes in the enriched monocytes. The purity of monocytes was 85.24%, in which CM accounted for 78.16%, IM and NCM accounted for about 7.08% (Figure S1), which was in agreement with previous studies.19
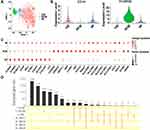
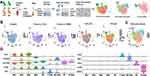
Enriched monocytes from KD patients (n=2) and healthy infants (n=2) were transcriptionally profiled after the isolation process (Figure 1A). In total, 8880 cells were sequenced. On average, a median of 1096 genes and 37075 reads per cell were detected. After quality control, 6283 individual cells, including 2302 cells of healthy infants and 3981 cells of KD patients, were left for further analysis.
The enriched monocytes were clustered in different samples after unsupervised clustering, implying differences among different samples (Figure S2). Therefore, we integrated the 4 samples using Seurat v3 Integration,20 and 9 cell types were clustered in the integrated dataset (Figure 1B). The clustering was not affected by different samples, number of mRNAs, percentage of mitochondrial genes and cell cycle (Figure 1B). The 9 cell types included 4 monocyte subsets (expressing CD14 or FCGR3A), DC (expressing HLA-DRA and ITGAX, neither CD14 nor FCGR3A), T cells (expressing CD3D and CD2), NK cells (expressing FCGR3A and PRF1), B cells (expressing MS4A1 and CD79a) and endothelial cells (expressing CD34) (Figure 1D). Markers used to identify the cell types were consistent with the related cell markers in the website “CellMarker” (http://biocc.hrbmu.edu.cn/CellMarker/index.jsp) and “The Human Protein Atlas” (https://www.proteinatlas.org/). Note that FCGR3B, which is highly homologous to FCGR3A, is expressed in only dozens of cells (Figure S3), so FCGR3A was identified as the classical marker for CD16. In order to elucidate the monocyte subsets, only monocytes were extracted for further analysis (Figure 1C).
Identification of 3 Monocyte Subsets in Healthy Infants
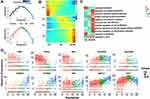
To identify monocyte subsets in healthy infants, the monocytes of healthy infants (1969 cells) were extracted and unsupervised clustered. The monocytes were clustered in 3 subsets (Figure 2A), including CM, IM, and NCM (Figure 2A and B). CM were relatively abundant compared with IM and NCM. Specific markers for each cluster were defined as the 10 genes with the highest differential expression relative to all other cells (Figure 2C). On CM, a pro-inflammatory gene signature (eg, S100A8, S100A9, S100A12, and LYZ) was expressed, while on IM a unique combination of genes that have a potential to present antigen (eg, HLA-DQA1, HLA-DQA2, HLA-DPA1, HLA-DPB1, HLA-DRA, HLA-DQB1, and CD74) were expressed. Specific markers of NCM were mainly associated with phagocytosis (eg, FCGR3A), cell proliferation, and apoptosis (eg, CDKN1C and NAP1L1).
As this is the first study on monocyte subsets in infants based on scRNA-seq, to determine whether the infant monocytes are transcriptionally different from those of adults, the dataset of adult monocytes (between 25 and 40 years of age) in the previous study11 was utilized to compare with the infants. In adults, 4 monocyte subsets were identified, including Mono1 and Mono2 (mainly contained the classical and non-classical monocytes, respectively), Mono3 expressing genes that have the potential to affect cell cycle, differentiation, and trafficking (eg, MXD1, CXCR1, CXCR2, VNN2), and Mono4 expressing a cytotoxic gene signature (eg, PRF1, GNLY, CTSW). After identifying the same marker genes between the clusters of infants and adults, it seemed that CM in infants were similar with Mo1, Mo3, and Mo4 in adults, while IM and NCM in infants were similar with Mo2 in adults (Figure 2D). We conclude that the monocyte subsets in adults and infants are transcriptionally different. Nevertheless, as the number of cells and genes in the datasets of adults and infants were different, so it is difficult to make an intensive comparison between them.
Linear Differentiation of Monocyte Subsets in Healthy Infants
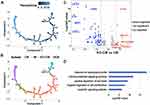
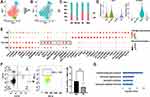
Differentiation continuance of monocyte subsets is not clear yet in infants. Some studies support the origin of NCM from CM in human,21 but whether these findings are based on infants are obscure. To investigate the differentiation continuance of monocyte subsets of infants in healthy homeostasis, we performed the trajectory analysis using the 3 monocyte subsets. Differentiation trajectory with a tree-like structure was constructed (Figure 3A). Based on previous studies,21 CM was chosen as the earliest member of the lineage differentiation followed by IM and NCM (Figure 3A). CM, IM, and NCM represent a linear differentiation state.
We further analyzed the gene expression patterns of marker genes along the trajectory of monocyte differentiation. The marker genes were further clustered into 4 expression patterns (P1–P4), including P1 (expression patterns of CM), P2 and P3 (expression patterns of IM) and P4 (expression patterns of NCM) (Figure 3B). Functional enrichment analysis showed that genes that were highly expressed in P1 were associated with response to bacterium and neutrophil activation (Figure 3C). Function of antigen processing and presentation and response to virus were activated in P2 and P3. Genes with function of cytokine, cell proliferation, and inflammation have higher expression levels in P4. To better elucidate the differentiation process, we assessed the expression of genes regulated during monocyte differentiation. CD36, VCAN, and SELL expression were down-regulated, while HLA-DPA1, HLA-DQA1, and CD74 expression were significantly increased in IM. During the differentiation of CM to NCM, expression of CDKN1C, FCGR3A, TNF, and NAP1L1 was up-regulated (Figure 3D). These changed transcripts may reveal the mechanism of monocytes differentiation.
Altered Monocyte Subsets in KD Patients
In order to explore whether the monocyte subsets in KD patients are the same with those in healthy infants, the monocyte profiles of KD (3616 cells) and healthy infants (1969 cells) were extracted and clustered in 4 subsets (Figure 4A). Subset that mainly appeared in KD patients (Figure 4B, C and Figure S4), which expressed CD14 and not expressed FCGR3A (Figure 4D), were defined as CM in KD patients (KD-CM). To identify the genes that lead to the separation of CM between KD and healthy infants, top 10 highest expressed genes of each subset were presented in dot plot (Figure 4E). KD-CM was featured by high expression of SELL and MALAT, and low expression of CXCL8 and JUN compared with CM. Flow cytometry confirmed the expansion of SELL+ CM in infants with KD (Figure 4F). Gene ontology enrichment analysis demonstrated that marker genes of KD-CM were mainly related with neutrophil activation (Figure 4G).
Poorly Differentiated State of KD-CM
To explore the differentiation state of KD-CM, single-cell trajectory analysis was performed using the 4 monocyte subsets of KD and healthy infants (Figure 5A). The trajectory’s root was mainly populated by KD-CM, and they eventually differentiated into NCM through CM and IM (Figure 5B), suggesting that KD-CM were more poorly differentiated than CM in healthy infants. We then examined gene expression patterns in KD-CM and CM, 79 differentially expressed genes including 15 up-regulated genes and 64 down-regulated genes were identified in KD-CM compared with CM (Figure 5C and Supplementary Table S2). The up-regulated genes in KD-CM related to cell cycle (eg, MALAT1, MPHOSPH8, and TFEC) and production of immunoglobulin (eg, POU2F2). The down-regulated genes in KD-CM were mainly related to response to lipopolysaccharide, cytokine-mediated signaling pathway, positive regulation of cell death and apoptotic signaling pathway (Figure 5D).
Discussion
Monocytes are important innate immune cells involving in vasculitis.4,22,23 However, it is difficult to translate the research findings of monocytes into effective clinical therapies, mainly because of its obvious heterogeneity.10 scRNA-seq can reveal the overall transcriptome characteristics of single cells, which allow for the identification of molecular drivers of disease in pathogenic cell subsets.24 In the present study, 3 monocyte subsets in healthy infants were revealed and KD-CM associated with neutrophil activation were identified in KD using scRNA-seq analysis. These findings shed light on the role of monocytes in KD.
Monocytes in healthy infants were divided into 3 subsets, which was consistent with previous classification of human monocytes.19 The distribution of CD14 and CD16 is continuous on different monocyte subsets,25 so it is difficult to classify distinct subsets using the traditional CD14/CD16-based biaxial gating strategies. Therefore, we analyzed the relatively specific gene markers for the 3 monocyte subsets (Figure 2C), which made it possible to find the new markers of different subsets. CM highly expressed S100A8, S100A9 and S100A12, IM expressed HLA-DQA1 and HLA-DPA1, and NCM mainly expressed CD16. Combination of the above expressed marker genes could help us classify the 3 monocyte subsets.
Studies have confirmed that human monocyte subsets have different functions: CM exhibit a more pro-inflammatory phenotype including anti-microbial responses, IM are specialized in antigen presentation and NCM are mainly responsible for the anti-viral responses.19,26 In our results, CM was associated with response to bacterium and IM were responsible for antigen presentation, which was consistent with previous finding.19 Studies have confirmed that monocyte subsets could secrete different cytokines to balance the activation of endothelial cells and promote secondary leukocyte recruitment in vitro.7 Ly6Chi monocyte in mice, which resembles CM in human, could activate neutrophil in defense against candidiasis.27 Therefore, we infer that CM possess the function of neutrophil activation for immune balance in healthy homeostasis. NCM possess not only the function of anti-viral responses, cytokine-mediated signaling pathway, leukocyte migration and adhesion, and inflammatory responses as previous foundings19 but also the function of negative regulation of cell proliferation implying terminally differentiated monocyte subsets.
Several studies have discovered that human monocyte subsets represent a single lineage at different stages of differentiation in blood.21,28 Moreover, numerous studies in human have shown that monocytes circulating in the blood can differentiate into monocyte-derived macrophages or monocyte-derived dendritic cells at steady state.29 Our results support the previous findings of a linear monocyte subsets differentiation. As for some branches appeared during the differentiation of CM to IM, we speculate that some CM may leave the bloodstream to infiltrate tissues and differentiate into monocyte-derived macrophages or monocyte-derived dendritic cells.
Katayama et al observed that IM increase in acute KD using flow cytometry. However, the ratio of IM in healthy and KD infants was similar (Figure 4C) in our results, which were mainly due to that the scRNA-seq technology is more accurate in cell classification or the sample size was too small in our research. KD-CM, which was featured by the expression of SELL and MALAT (Figure 4E), was identified in KD patients based on the overall characteristics of monocyte transcriptome. SELL encodes selectin L, which is a cell surface adhesion molecule and is required for binding and subsequent rolling of leucocytes on endothelial cells, facilitating their migration into secondary lymphoid organs and inflammation sites. Studies have confirmed that CD62L could regulate recruitment of monocytes to lymphoid tissue from the blood during inflammation30 and the flux of rolling inflammatory monocytes.31 MALAT, metastasis-associated lung adenocarcinoma transcript 1, acts as a transcriptional regulator for numerous genes, including genes involved into cancer metastasis,32 cell migration,33 and cell cycle regulation;34 however, its role in monocytes still needs further exploration.
Studies have demonstrated that neutrophil-to-lymphocyte ratio may be used as a biomarker for detecting IVIG-resistant KD, especially after the initial treatment of IVIG, implying the key functions of neutrophils.35 Moreover, neutrophils were involved in the pathogenesis of KD,36,37 and suppressing neutrophil activation prove effective.38 Neutrophils have a restricted set of pro-inflammatory functions,39 suggesting that neutrophil/myeloid activation may cause endothelial cells damage in KD. In our results, marker genes of KD-CM were mainly related to neutrophils activation, so we infer that KD-CM may regulate KD vasculitis through activation of neutrophils. Recently, the link between neutrophil inflammasome dysfunction and neutrophil-mediated human diseases attracted the attention of researchers.40 Moreover, studies have revealed that inositol-triphosphate 3-kinase C mediates inflammasome activation and increase the production of IL-1b and IL-18 in KD patients.41 It is possible that neutrophil-activating monocytes are involved in vasculitis through neutrophil inflammasome dysfunction.
Pathological conditions can affect the function and differentiation of monocytes. Under pathological conditions, classical monocytes give rise to distinct monocyte-derived cells with functions of pro-inflammatory activities, antigen-presentation, tissue remodeling, or anti-inflammatory abilities.3 In our scRNA-seq results, KD-CM are more poorly differentiated than CM which exists both in KD and healthy infants, which revealing the complexity of monopoiesis under emergency. Moreover, KD-CM increased in KD patients who had inflammation compared with healthy infants. In order to explain the different differentiation state of KD-CM and CM, we analyzed the differentially expressed genes between KD-CM and CM. The up-regulated genes in KD-CM compared with CM related to cell cycle and production of immunoglobulin, which may explain the poorly differentiation state of KD-CM and effectiveness of immunoglobulin in treating KD to a certain degree.
As for etiology of KD, most studies accepted the theory that genetically predisposed individuals are exposed to one or more infectious agents that trigger an inflammatory response. Recently, some studies proposed a hypothesis that a relationship may exist between KD and coronavirus disease 2019 (COVID-19) due to their shared genetic susceptibilities, post-infectious vasculitides and hypersecretion of cytokines.42 Furthermore, scRNA-seq of peripheral blood mononuclear cells of COVID-19 infection revealed an increased ratio of IL-1β+ CM, which is the evidence of macrophage polarization toward pro-inflammatory M1 macrophages.43,44 However, the expression of IL1B and CXCL8 and CXCL2 decreased in KD-CM compared with CM in our results (Figure 5C), implying different phenotype of CM in KD and COVID-19. The phenotype and function of KD-CM in KD still needs further exploration. If KD-CM promotes the development of KD, it would be blocked as a target, otherwise, it would be promoted.
There are some limitations in our study. Only 4 samples were used for scRNA-seq, which may lead to poor statistical power. Although some hypotheses about pathogenesis of KD were proposed, large data, including replication of the results in older children and KD patients with coronary aneurysms are lacking. In addition, only monocytes were analyzed in our study, which limited the assessment of entire immune system in KD. We will study the entire immune system and find the key immune cells that cause damage to endothelial cells in KD patients in the further study.
Conclusions
A comprehensive map of circulating monocyte subsets in healthy infants was plotted and their relationship of linear differentiation were determined in this study. Moreover, KD-CM with featured gene expression profile was revealed, their poorly differentiated state and function of neutrophils activation were identified. The present study will help us clarify the mechanism for monocyte subsets in KD vasculitis, and find new targets of clinical treatment for KD.
Highlights
- Three monocyte subsets are identified in healthy infants using scRNA-seq
- Monocyte subsets in healthy infants represent a linear differentiation and possess different biological functions
- SELL+CD14+CD16- monocytes expand in KD patients
- SELL+CD14+CD16- monocytes are poorly differentiated and related to neutrophil activation
Abbreviations
KD, Kawasaki disease; CM, classical monocytes; IM, intermediate monocytes; NCM, non-classical monocytes; scRNA-seq, single-cell RNA-seq; KD-CM, classical monocytes in KD patients; UMI, unique molecular identifiers; tSNE, t-distributed stochastic neighbor embedding algorithm.
Data Sharing Statement
scRNA-seq data that support this study have been deposited in GEO (GSE152450).
Acknowledgment
We thank Sinotech Genomic Co. LTD. for assistance with single-cell RNA sequencing.
Funding
This research was funded by the National Natural Science Foundation of China. [grant numbers: 81670251 and 81970434].
Disclosure
The authors declare no conflicts of interest for this work.
References
1. Marchesi A, Tarissi de Jacobis I, Rigante D, et al. Kawasaki disease: guidelines of the Italian Society of Pediatrics, part I - definition, epidemiology, etiopathogenesis, clinical expression and management of the acute phase. Ital J Pediatr. 2018;44(1):102. doi:10.1186/s13052-018-0536-3
2. Marrani E, Burns JC, Cimaz R. How Should We Classify Kawasaki Disease? Front Immunol. 2018;9:2974. doi:10.3389/fimmu.2018.02974
3. Guilliams M, Mildner A, Yona S. Developmental and Functional Heterogeneity of Monocytes. Immunity. 2018;49(4):595–613. doi:10.1016/j.immuni.2018.10.005
4. Watanabe R, Maeda T, Zhang H, et al. MMP (Matrix Metalloprotease)-9-Producing Monocytes Enable T Cells to Invade the Vessel Wall and Cause Vasculitis. Circ Res. 2018;123(6):700–715. doi:10.1161/CIRCRESAHA.118.313206
5. Ziegler-Heitbrock L, Ancuta P, Crowe S, et al. Nomenclature of monocytes and dendritic cells in blood. Blood. 2010;116(16):e74–e80. doi:10.1182/blood-2010-02-258558
6. Katayama K, Matsubara T, Fujiwara M, Koga M, Furukawa S. CD14+CD16+ monocyte subpopulation in Kawasaki disease. Clin Exp Immunol. 2000;121(3):566–570. doi:10.1046/j.1365-2249.2000.01321.x
7. Chimen M, Yates CM, McGettrick HM, et al. Monocyte Subsets Coregulate Inflammatory Responses by Integrated Signaling through TNF and IL-6 at the Endothelial Cell Interface. J Immunol. 2017;198(7):2834–2843. doi:10.4049/jimmunol.1601281
8. Ikeda K, Yamaguchi K, Tanaka T, et al. Unique activation status of peripheral blood mononuclear cells at acute phase of Kawasaki disease. Clin Exp Immunol. 2010;160(2):246–255. doi:10.1111/j.1365-2249.2009.04073.x
9. Merah-Mourah F, Cohen SO, Charron D, Mooney N, Haziot A. Identification of Novel Human Monocyte Subsets and Evidence for Phenotypic Groups Defined by Interindividual Variations of Expression of Adhesion Molecules. Sci Rep. 2020;10(1):4397. doi:10.1038/s41598-020-61022-1
10. Hamers AAJ, Dinh HQ, Thomas GD, et al. Human monocyte heterogeneity as revealed by high-dimensional mass cytometry. Arterioscler Thromb Vasc Biol. 2019;39(1):25–36. doi:10.1161/ATVBAHA.118.311022
11. Villani AC, Satija R, Reynolds G, et al. Single-cell RNA-seq reveals new types of human blood dendritic cells, monocytes, and progenitors. Science. 2017;356(6335):eaah4573. doi:10.1126/science.aah4573
12. Valiathan R, Ashman M, Asthana D. Effects of Ageing on the Immune System: infants to Elderly. Scand J Immunol. 2016;83(4):255–266. doi:10.1111/sji.12413
13. McCrindle BW, Rowley AH, Newburger JW, et al. Diagnosis, treatment, and long-term management of Kawasaki disease: a scientific statement for health professionals from the American heart association. Circulation. 2017;135(17):e927–e999. doi:10.1161/CIR.0000000000000484
14. Stoeckius M, Zheng S, Houck-Loomis B, et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 2018;19(1):224. doi:10.1186/s13059-018-1603-1
15. Fan HC, Fu GK, Fodor SP. Expression profiling. Combinatorial labeling of single cells for gene expression cytometry. Science. 2015;347(6222):1258367. doi:10.1126/science.1258367
16. Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36(5):411–420. doi:10.1038/nbt.4096
17. Zhou Y, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10(1):1523. doi:10.1038/s41467-019-09234-6
18. Trapnell C, Cacchiarelli D, Grimsby J, et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat Biotechnol. 2014;32(4):381–386. doi:10.1038/nbt.2859
19. Wong KL, Tai JJ-Y, Wong W-C. Gene expression profiling reveals the defining features of the classical, intermediate, and nonclassical human monocyte subsets. Blood. 2011;118(5):e16–e31. doi:10.1182/blood-2010-12-326355
20. Stuart T, Butler A, Hoffman P, et al. Comprehensive Integration of Single-Cell Data. Cell. 2019;177(7):1888–1902 e21. doi:10.1016/j.cell.2019.05.031
21. Patel AA, Zhang Y, Fullerton JN, et al. The fate and lifespan of human monocyte subsets in steady state and systemic inflammation. J Exp Med. 2017;214(7):1913–1923. doi:10.1084/jem.20170355
22. Wenzel P. Monocytes as immune targets in arterial hypertension. Br J Pharmacol. 2019;176(12):1966–1977. doi:10.1111/bph.14389
23. Kuriakose J, Redecke V, Guy C, et al. Patrolling monocytes promote the pathogenesis of early lupus-like glomerulonephritis. J Clin Invest. 2019;129(6):2251–2265. doi:10.1172/JCI125116
24. Proserpio V, Lonnberg T. Single-cell technologies are revolutionizing the approach to rare cells. Immunol Cell Biol. 2016;94(3):225–229. doi:10.1038/icb.2015.106
25. Liebetrau C, Hoffmann J, Dorr O, et al. Release kinetics of inflammatory biomarkers in a clinical model of acute myocardial infarction. Circ Res. 2015;116(5):867–875. doi:10.1161/CIRCRESAHA.116.304653
26. Gren ST, Rasmussen TB, Janciauskiene S, et al. Profile Reveals Inter-Cellular Heterogeneity within Human Monocyte Subsets. PLoS One. 2015;10(12):e0144351. doi:10.1371/journal.pone.0144351
27. Dominguez-Andres J, Feo-Lucas L, Minguito de la Escalera M, Gonzalez L, Lopez-Bravo M, Ardavin C. Inflammatory Ly6C(high) Monocytes Protect against Candidiasis through IL-15-Driven NK Cell/Neutrophil Activation. Immunity. 2017;46(6):1059–1072e4. doi:10.1016/j.immuni.2017.05.009
28. Zawada AM, Rogacev KS, Rotter B, et al. SuperSAGE evidence for CD14++CD16+ monocytes as a third monocyte subset. Blood. 2011;118(12):e50–e61. doi:10.1182/blood-2011-01-326827
29. Patel VI, Booth JL, Duggan ES, et al. Transcriptional classification and functional characterization of human airway macrophage and dendritic cell subsets. J Immunol. 2017;198(3):1183–1201. doi:10.4049/jimmunol.1600777
30. Xu H, Manivannan A, Crane I, Dawson R, Liversidge J. Critical but divergent roles for CD62L and CD44 in directing blood monocyte trafficking in vivo during inflammation. Blood. 2008;112(4):1166–1174. doi:10.1182/blood-2007-06-098327
31. Zuchtriegel G, Uhl B, Hessenauer ME, et al. Spatiotemporal expression dynamics of selectins govern the sequential extravasation of neutrophils and monocytes in the acute inflammatory response. Arterioscler Thromb Vasc Biol. 2015;35(4):899–910. doi:10.1161/ATVBAHA.114.305143
32. Cheng Y, Imanirad P, Jutooru I, et al. Role of metastasis-associated lung adenocarcinoma transcript-1 (MALAT-1) in pancreatic cancer. PLoS One. 2018;13(2):e0192264. doi:10.1371/journal.pone.0192264
33. Zhuo M, Yuan C, Han T, Cui J, Jiao F, Wang L. A novel feedback loop between high MALAT-1 and low miR-200c-3p promotes cell migration and invasion in pancreatic ductal adenocarcinoma and is predictive of poor prognosis. BMC Cancer. 2018;18(1):1032. doi:10.1186/s12885-018-4954-9
34. Yao W, Bai Y, Li Y, et al. Upregulation of MALAT-1 and its association with survival rate and the effect on cell cycle and migration in patients with esophageal squamous cell carcinoma. Tumour Biol. 2016;37(4):4305–4312. doi:10.1007/s13277-015-4223-3
35. Wu G, Yue P, Ma F, Zhang Y, Zheng X, Li Y. Neutrophil-to-lymphocyte ratio as a biomarker for predicting the intravenous immunoglobulin-resistant Kawasaki disease. Medicine. 2020;99(6):e18535. doi:10.1097/MD.0000000000018535
36. Ko TM, Chang JS, Chen SP, et al. Genome-wide transcriptome analysis to further understand neutrophil activation and lncRNA transcript profiles in Kawasaki disease. Sci Rep. 2019;9(1):328. doi:10.1038/s41598-018-36520-y
37. Pan Y, Fan Q. Identification of potential core genes in immunoglobulin-resistant Kawasaki Disease using bioinformatics analysis. Crit Rev Eukaryot Gene Expr. 2020;30(1):85–91. doi:10.1615/CritRevEukaryotGeneExpr.2020028702
38. Popper SJ, Shimizu C, Shike H, et al. Gene-expression patterns reveal underlying biological processes in Kawasaki disease. Genome Biol. 2007;8(12):R261. doi:10.1186/gb-2007-8-12-r261
39. Kolaczkowska E, Kubes P. Neutrophil recruitment and function in health and inflammation. Nat Rev Immunol. 2013;13(3):159–175. doi:10.1038/nri3399
40. Tyrkalska SD, Candel S, Mulero V. The neutrophil inflammasome. Dev Comp Immunol. 2021;115:103874. doi:10.1016/j.dci.2020.103874
41. Alphonse MP, Duong TT, Shumitzu C, et al. Inositol-Triphosphate 3-Kinase C mediates inflammasome activation and treatment response in Kawasaki Disease. J Immunol. 2016;197(9):3481–3489. doi:10.4049/jimmunol.1600388
42. AbdelMassih AF, AbdelAzeam AS, Ayad A, et al. Unleashing the mysterious link between COVID-19 and a famous childhood vasculitis: kawasaki disease. Egypt Pediatr Assoc Gazette. 2020;68(1). doi:10.1186/s43054-020-00029-9
43. AbdelMassih AF, Fouda R, Kamel A, et al. Single cell sequencing unraveling genetic basis of severe COVID19 in obesity. Obes Med. 2020;20:100303. doi:10.1016/j.obmed.2020.100303
44. Rodriguez L, Pekkarinen PT, Lakshmikanth T, et al. Systems-level immunomonitoring from acute to recovery phase of severe COVID-19. Cell Rep Med. 2020;1(5):100078. doi:10.1016/j.xcrm.2020.100078
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.