Back to Journals » OncoTargets and Therapy » Volume 13
Tumor Mutational Burden and PD-L1 Expression in Non-Small-Cell Lung Cancer (NSCLC) in Southwestern China
Authors Ma Y, Li Q, Du Y, Chen W, Zhao G, Liu X, Ye L, Li H, Wang X, Liu J, Shen Z, Ma L, Zhou Y
Received 28 March 2020
Accepted for publication 22 May 2020
Published 8 June 2020 Volume 2020:13 Pages 5191—5198
DOI https://doi.org/10.2147/OTT.S255947
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Sanjeev K. Srivastava
Yuhui Ma,1,* Quan Li,2,* Yaxi Du,2,* Wanlin Chen,1 Guangqiang Zhao,1 Xing Liu,2 Lianhua Ye,1 Hongsheng Li,3 Xiaoxiong Wang,3 Junxi Liu,3 Zhenghai Shen,4 Luyao Ma,2 Yongchun Zhou4
1Department of Thoracic Surgery I, The Third Affiliated Hospital of Kunming Medical University (Yunnan Cancer Hospital), Kunming 650118, People’s Republic of China; 2Key Laboratory of Lung Cancer Research of Yunnan Province, The Third Affiliated Hospital of Kunming Medical University, Kunming 650118, People’s Republic of China; 3International Joint Laboratory on High Altitude Regional Cancer of Yunnan Province, The Third Affiliated Hospital of Kunming Medical University, Kunming 650118, People’s Republic of China; 4Yunnan Cancer Center, The Third Affiliated Hospital of Kunming Medical University, Kunming 650118, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Yongchun Zhou
Yunnan Cancer Center, The Third Affiliated Hospital of Kunming Medical University, Kunming 650118, People’s Republic of China
Tel/ Fax +86-87168172772
Email [email protected]
Purpose: To explore the impact between the tumor mutational burden (TMB) and programmed death ligand-1 (PD-L1) expression on NSCLC in the Yunnan region of southwestern China.
Patients and Methods: Seventy-one NSCLC specimens that were pathologically confirmed were collected at first. The TMB and driver genetic alterations were evaluated accordingly by next-generation sequencing (NGS). Afterwards, clinical parameters and tumor PD-L1 expressions were collected. Finally, the relationship between TMB, PD-L1 expression and clinical outcome was evaluated.
Results: The median TMB was 5 (0.6– 49) mutations/Mb by our NGS panel and the majority of patients (63/71, 88.7%) did not receive immunotherapy. The progression-free survival (PFS) was longer in TMB-low patients versus TMB-high ones (median 18.0 vs. 9.0 months, hazard ratio = 0.34, 95% confidence interval 0.14 to 0.84, p = 0.02) and the cut-off value was 10 mutations/Mb. The overall survival (OS) was longer in TMB-low patients vs. TMB-high ones (median 21.0 vs. 10.0 months, HR = 0.32, 95% CI 0.12 to 0.82, p = 0.02). Notably, our study also found that, excluding the eight patients with immunotherapy, the PFS was longer in patients with TMB-low vs. TMB-high (median 19.0 vs. 8.0 months, HR = 0.11, 95% CI 0.03 to 0.39, p < 0.01) and the OS was longer in TMB-low patients vs. TMB-high (median 21.0 vs 10.0 months, HR = 0.12, 95% CI 0.03 to 0.42, p < 0.01).
Conclusion: TMB was a valid and independent prognostic biomarker for NSCLC patients’ clinical outcome and comprehensive screening of TMB based on NGS is recommended for individualized treatment strategies in Yunnan population.
Keywords: non-small-cell lung cancer, TMB, programmed death ligand-1 expression, next-generation sequencing
Introduction
The most commonly diagnosed cancers in China, in 2018, were lung cancers (774,323 new cases with an incidence of 35.10/100,000) and they were the dominant cause of cancer-related death (690,567 deaths were estimated) in 2018.1 Studies have shown that non-small-cell lung cancer (NSCLC) is the most common subtype of lung cancer.2 Recently, immunotherapy has become a vital treatment strategy in addition to chemotherapy, radiotherapy and targeted therapy. Since 2015, immunological checkpoint inhibitors (ICIs) have been successively authorized by US Food and Drug Administration (FDA) as standard strategy for NSCLC in second-line or first-line treatment.3–7 Although this breakthrough has significantly prolonged progression-free survival (PFS) and overall survival (OS) of NSCLC patients, just 20% to 25% of patients will benefit from ICIs.3–7 PD-L1 expression, evaluated by immunohistochemical analysis, is the reliable biomarker to predict benefit from immunotherapy.8,9 However, the PD-L1 expression as a valid biomarker remains a challenge, as some patients with low/negative PD-L1 expression were also able to effectively response to ICIs.3,4 Besides, some patients with high PD-L1 expression cannot benefit from ICI treatment, experiencing hyperprogressive disease (HPD).10 HPD, the acceleration of tumor growth when receiving immunotherapy, was found in 9% of advanced cancers with the treatment of PD-1/PD-L1 inhibitors.10,11 Moreover, PD-L1 expression occurs both on cancer cells, and the non-cancer cells with the tumor microenvironment.12 Thus, there is an urgent need for identifying novel valid biomarkers to better select patients that will be treated with ICIs.
Tumor mutational burden (TMB) has demonstrated itself as a new estimated biomarker in NSCLC.13,14 Previous study demonstrated that patients with TMB-high had prolonged PFS after they received PD-1/PD-L1 blockade, compared with patients who received chemotherapy.15 Rizvi et al have also reported a whole exome sequencing (WES) analysis that shows that a higher TMB is related to a prolonged PFS in patients that were treated with PD-1/PD-L1 blockade.13 Although the value of WES in detecting TMB and estimating response to anti-PD-1/PDL-1 therapies has been verified by some previous study, it has many limitations.13,16 The use of clinical limitations as well as time consuming procedures, cost and tissue availability make next-generation sequencing (NGS) better for clinical environment than WES.17 The NGS platform can simultaneously capture all classes of DNA alterations and TMB from only one sample.18,19 TMB, assessed by NGS, is related to response to ICIs in NSCLC patients.20,21
Our center has had an established TMB and PD-L1 expression testing facilities (by NGS) since 2017; the tested data can provide guiding information about local NSCLC patients. TMB and PD-L1 expression status in NSCLC patients in Yunnan area are unclear, so we first revealed the TMB and PD-L1 with NSCLC in Yunnan area and further explored the significance of TMB and PD-L1 for NSCLC patients treated with ICIs and advanced the effective guidance for clinical treatment.
Methods
Patient
For this retrospective study population, seventy-one NSCLC patients were recruited for TMB and PD-L1 expression status analysis, from the Third Affiliated Hospital of Kunming Medical University between January 2017 and March 2020. The included patients were provided with written informed consent and the study was approved by the ethics committee of the Third Affiliated Hospital of Kunming Medical University, complying with the Declaration of Helsinki. Enrolment criteria were adult age (>18 years) residency in the Yunnan area, pathologically confirmed NSCLC and sufficient tissue for TMB and PD-L1 analysis. Exclusive criteria were as follows: any specimen with possible contamination. Seventy-one sufficient tissue specimens were provided TMB and PD-L1 analysis, PFS and OS data. In addition, other information was collected including sex, age, tumor stage, smoking history, histology, treatment status, tumor type, demographic, TMB, PD-L1 expression and driver mutations.
DNA Extraction and NGS Library Construction
Genomic DNA samples from formalin-fixed paraffin-embedded (FFPE) were extracted by using the QIAampDNA FFPE Tissue Kit according to the manufacturer’s instructions. The FFPE tissue section was cut into 5 μm in thickness and the percentage of tumor cells was more than 20%. Qubit dsDNA HS Assay Kit detected the concentration of DNA (>300ng and 10 ng/μL). The range of DNA input was from 50 to 200ng. If DNA input was <50ng, an extra PCR cycle was adopted during target enrichment. And then enzyme-shearing extracted DNA and sequencing libraries were prepared by using the TIANSeq DirectFast DNA Library Prep Kit. VAHTS Library Quantification Kit for Illumina was used to quantify nucleic acid when the Ct value of library was lower than 10.
Sequencing and TMB Calculation
Before sequencing, capture libraries were manufactured at a concentration of 2ng/μL. Captured probes were designed and synthesized by Integrated DNA Technologies (IDT, xGen Lockdown Probes), which contained 547 genes to assess TMB and the detection sites were spread at an average of genomes in human 23 chromosomes. Hybrid capture-selected libraries with captured DNA segments were performed sequencing by Illumina® HiSeq X-TEN and libraries were diluted to 78pM. PhiX libraries were mixed with diluted captured libraries in a ratio of 1:1. Raw data were processed on Illumina® HiSeq X-TEN (>1000X depth in tissue, sensitivity <1%) and aligned to the hg19 reference genome.21 The subsequent analysis was performed by our own program KEYseq V2.0. TMB was counted by summing all variants present at 5% allele frequency or more, which was shown with mutations per megabase (mut/Mb) unit. Notably, non-synonymous variants were included in the variants. Germline alterations and known/likely driver genes were filtered out by a somatic-germline/zygosity (SGZ) algorithm. The TMB calculation is dependent on variables such as percentage of tumor, filtering of alterations included in the score, the read depth and so on. Some studies15,22 showed that the definition of high TMB was not one universal number in various tumor types. The optimal cutpoint seems to change in various cancers. Based on tumor type and clinical practicability, there were low and high mutational burden types for TMB according to the previous studies.15,23 Therefore, TMB was split into two levels in our study: low (0–9 mutations/mb), and high (≥10 mutations/mb).15,23
PD-L1 Immunohistochemical Analysis
PD-L1 expression was evaluated with Ventana SP263 assay on the BenchMark platform (Ventana, Tucson, AZ, USA). The PD-L1 expression positivity means over 1% of tumor cells based on NCCN Guidelines.7 We required the samples to include at least 100 viable cancer cells, assessed by two experienced pathologists for PD-L1 immunohistochemical analysis. Thirty-nine patients’ samples were available for evaluation of PD-L1 expression.
Outcomes Assessment
OS was measured as from the initiation of the TMB test until patient death due to any reasons. PFS was measured as from the initiation of the TMB test to clinical/radiographic progression or death. If patients did not progress or die, they would be censored at the date of the last follow-up. We evaluated survivals every 3 months after the first TMB test, and updated the last follow-up time on March 5, 2020.
Statistical Analysis
The relationship between TMB, PD-L1 expression and the clinical parameters was analyzed by χ2 or Fisher’s exact tests. The PFS and OS were calculated by Kaplan–Meier curves (P values by Log-rank test). The method of mantel-Haenszel test was calculated hazard ratios (HRs). Two-sided, p < 0.05 was deemed to be statistically significant. The whole statistics were performed by Graph-Pad Prism version 7.0 (San Diego, CA, USA) and SPSS 22.0 software (SPSS Inc., Chicago, IL, USA).
Results
Patient Characteristics
There were 71 consecutive patients, including 27 (38.1%) females and 44 (61.9%) males, enrolled in our study. These patients were between the age of 31 and 80, with the median age of 56. The most common histology was adenocarcinoma (n=53, 74.6%), and the others were squamous carcinoma (n=18, 25.4%). In addition, 71.8% patients were made a diagnosis of stage IV disease. Forty-two patients (59.1%) were non-smokers. Forty-eight patients (67.6%) received first line of chemotherapy. Eight patients were treated in the second-line immunotherapy. In a total of 39 (54.9%) patients with available tumor PD-L1 immunohistochemical data, 19 (48.7%) expressed PD-L1 ≥1%, and negative results were identified in 20 (52.6%).
However, no evident differences were found in characteristics such as sex, smoking history, stage, tumor type, chemotherapy, immunotherapy. Table 1 summarizes the primary patients’ characteristics.
 |
Table 1 Characteristics of 71 Patients with NSCLC |
Driver Alterations and Mutation Burden
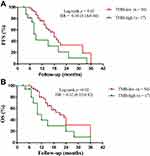
Ten (14.0%) patients harbored KRAS mutations, Thirty-seven (52.1%) harbored TP53 mutations, four (5.6%) harbored EGFR mutations, and one (1.4%) harbored ALK mutation. No evident differences were found in KRAS gene status, TP53 gene status, EGFR gene status and ALK gene status (Table 1). The median TMB was 5 (0.6–49) mutations/Mb by NGS. We divided TMB into two groups based on previous studies:22 TMB-low and TMB-high groups. These patients were between the age of 31 and 80, with the median age of 56. Fifty-four patients (TMB-low group) were between TMB 0.6 and 9, with a median TMB of 4.2 mutations/Mb. Seventeen patients (TMB-high group) were between 10 and 49, median TMB 18.0 mutations/Mb. In addition, the PFS was longer in TMB-low patients versus TMB-high ones (median 18.0 vs. 9.0 months, hazard ratio = 0.34, 95% CI 0.14 to 0.84, p = 0.02, Figure 1A). Similarly, we found that the OS was longer in TMB-low patients vs. TMB-high ones (median 21.0 vs. 10.0 months, HR = 0.32, 95% CI 0.12 to 0.82, p = 0.02, Figure 1B).
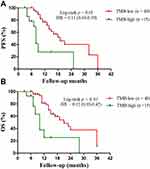
Six patients received ICIs in the TMB-low group and two patients received ICIs in the TMB-high group. Therefore, we designed a new survival analysis to compare the two groups without these 8 patients who received immunotherapy. We found that the PFS was longer in TMB-low patients vs. TMB-high ones (median 19.0 months vs. 8.0 months, HR = 0.11, 95% CI 0.03 to 0.39, p < 0.01, Figure 2A). Meanwhile, the OS was longer in patients with TMB-low vs. TMB-high group (median 21.0 vs. 10.0 months, HR = 0.12, 95% CI 0.03 to 0.42, p < 0.01, Figure 2B).
Outcome by PD-L1 Expression
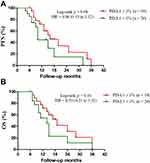
PD-L1 expression was divided into two groups based on former studies:7,24 PD-L1 < 1% and PD-L1 ≥ 1% groups. These patients were between the age of 36 and 80, with the median age of 61 years. Twenty patients come from the PD-L1 < 1% group and 19 patients come from the PD-L1 ≥ 1% group. Furthermore, the median PFS in the PD-L1 ≥ 1% vs. the PD-L1 < 1% group is 16.0 months vs. 10.0 months (HR = 0.46, 95% CI 0.19 to 1.12, p = 0.09, Figure 3A). Similarly, the median OS in the PD-L1 ≥ 1% vs. the PD-L1 < 1% group is 19.0 months vs. 14 months (HR = 0.53, 95% CI 0.21 to 1.32, p = 0.17, Figure 3B). There is no evident difference in these two groups.
Discussion
TMB is the whole mutations number of a tumor specimen. Some previous studies13,25 showed that highly mutated tumor cells can lead to the neoantigens production, which enable the immune cell to identify and attack cancer cells. A number of retrospective studies demonstrated that TMB levels influenced drug response to NSCLC patients receiving ICIs and TMB-high patients had significantly longer PFS and OS than TMB-low ones.23,26 A recently published prospective study (CheckMate 227) demonstrated that NSCLC patients, who received ICIs, with TMB-high (≥10 mutations/Mb) had evidently improved PFS, but OS was not predicted with the same reliability.27
Most of the studies reported the relationship between TMB levels and NSCLC patients’ clinical outcome during Immunotherapy. However, relationship between TMB levels and NSCLC patients’ clinical outcome without ICIs, especially in the Yunnan region of southwestern China, is unclear. A higher TMB is normally related to an adverse outcome, since TMB-high usually exists in advanced tumors.28 Herein, for the first time, our study demonstrated that NSCLC patients with TMB-high were related to adverse outcome in the Yunnan region, having a median PFS of 9.0 months and OS of 10.0 months. These results are partly similar to previous study.29 In addition, our other survival analysis, excluding patients with a history of ICI treatment, also showed that high TMB was related to adverse outcome in the Yunnan area. These observations are also similar to former study.22 In summary, it may suggest that TMB levels are significant for predicting NSCLC patients’ clinical outcome. However, the difference in patients’ clinical outcome between our results (patients with TMB-low showed longer PFS and OS) and the CheckMate-227 (patients with TMB-high showed longer PFS) may result from three reasons.27 First, the retrospective nature of this study may limit the interpretation of the results. Second, the difference in primary patients’ characteristics between our study (only eight patients were treated in the second-line immunotherapy) and the CheckMate-22727 (most of the patients were treated in the first-line immunotherapy) may lead to a discrepancy in assessment of clinical outcome. Finally, our study included 5 patients with known EGFR or ALK mutations, but CheckMate-22727 did not include any patients with known EGFR or ALK mutations. This may also lead to a bias in assessment of clinical outcome. Therefore, due to the number of patients receiving immunotherapy is limited, the relationship between TMB levels and NSCLC patients’ clinical outcome during Immunotherapy, in the Yunnan region of southwestern China, is unclear.
The median TMB in our NSCLC patients was 5 mutations/Mb (ranged from 0.6 to 49). According to other reports of TMB, the median value is lower.19,21 We discuss three reasons for this issue. First, because the research recruited patients only in a single center and the number of NSCLC patient samples was limited, our sample data may not accurately reflect the status of TMB in Yunnan area. Second, in most of our TMB tests by sequencing single sample, only little biopsy material is available, hence intratumoral heterogeneity may influence the results. Finally, advanced NSCLC patients with EGFR or ALK mutations are mostly with a low-TMB level.30 In our data, 5 TMB-low patients who harbored EGFR or ALK mutations may influence the results. In the clinical process of treatment, TMB testing is essential for selecting the suitable individualized treatment strategies, and therefore it needs sufficient qualified tumor samples. Although the role of blood TMB test in NSCLC is not yet clear, it might have potential to show a new integral landscape of the present mutations. Further prospective studies are expected.
In our study, only 55% (39/71) patients’ samples were available to evaluate PD-L1 expression. According to other reports of PD-L1 expression assessment, the value is lower.24 We discuss two reasons for this issue. First, there is no universal standard for the antibodies used for detecting PD-L1 expression in China. Herein, in order not to influence the authenticity of the results, the samples processed with non SP263 antibody are not included. Second, although some samples were detected by 22C3 and 28–8 antibodies with high consistency, the detected results of different antibodies are still different, so the detected results of this part of samples were not included in our study. Therefore, we suggest that the same type of antibody be utilized for detecting PD-L1 expression, which is more likely to acquire real results.
Although NSCLC PD-L1 expression proves to predict up to 3-fold improvement for ICIs, in contrast to negative expressers, it is controversial whether the PD-L1 can be a valid indicator for assessing patients’ clinical outcome. In our study, we found that both median PFS and OS were not significantly different in the group PD-L1 < 1% comparing with the group PD-L1 > 1%. It may suggest that the PD-L1 value in predicting the clinical outcome of patients is not reliable. However, due to the number of patients receiving immunotherapy is limited, the relations between PD-L1 expression and NSCLC patients’ clinical outcome during Immunotherapy, in the Yunnan region of southwestern China, are unclear. Consequently, further studies are expected.
In conclusion, our retrospective analysis shows a clear panoramic perspective of TMB and PD-L1 expression status from 71 NSCLC patients and their associations with clinical parameters in the Yunnan region. For the first time, we observed that PFS and OS were substantially prolonged with TMB-low than TMB-high among NSCLC patients, regardless of the level of tumor PD-L1 expression. Therefore, TMB is a valid and independent prognostic biomarker for NSCLC patients’ clinical outcome in the Yunnan region of southwestern China.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 81860513), the Project of Basic Applied Research in Yunnan Province (2016FB145) and the Project of Basic Applied Research in Yunnan Province (2017FA037).
Disclosure
The authors declare no conflicts of interest.
References
1. Feng RM, Zong YN, Cao SM, Xu RH. Current cancer situation in China: good or bad news from the 2018 global cancer statistics? Cancer Commun. 2019;39(1):22. doi:10.1186/s40880-019-0368-6
2. Pao W, Chmielecki J. Rational, biologically based treatment of EGFR-mutant non-small-cell lung cancer. Nat Rev Cancer. 2010;10(11):760–774. doi:10.1038/nrc2947
3. Brahmer J, Reckamp KL, Baas P, et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med. 2015;373(2):123–135. doi:10.1056/NEJMoa1504627
4. Borghaei H, Paz-Ares L, Horn L, et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med. 2015;373(17):1627–1639. doi:10.1056/NEJMoa1507643
5. Reck M, Rodriguez-Abreu D, Robinson AG, et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N Engl J Med. 2016;375(19):1823–1833. doi:10.1056/NEJMoa1606774
6. Socinski MA, Jotte RM, Cappuzzo F, et al. Atezolizumab for first-line treatment of metastatic nonsquamous NSCLC. N Engl J Med. 2018;378(24):2288–2301. doi:10.1056/NEJMoa1716948
7. National comprehensive cancer network: NCCN clinical practice guidelines in oncology: non-small cell lung cancer (version 2.2020). 2019;17(12):1529–1554.
8. Herbst RS, Soria J-C, Kowanetz M, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515:563–567. doi:10.1038/nature14011
9. Taube JM, Klein AP, Brahmer JR, et al. Association of PD-1, PD-1 ligands, and other features of the tumor immune microenvironment with response to anti-PD-1 therapy. Clin Cancer Res. 2014;20:5064–5074. doi:10.1158/1078-0432.CCR-13-3271
10. Ferrara R, Mezquita L, Texier M, et al. Hyperprogressive disease in patients with advanced non-small cell lung cancer treated with PD-1/PD-L1 inhibitors or with single-agent chemotherapy. JAMA Oncol. 2018;4(11):1543–1552. doi:10.1001/jamaoncol.2018.3676
11. Champiat S, Dercle L, Ammari S, et al. Hyperprogressive disease is a new pattern of progression in cancer patients treated by anti–PD-1/PD-L1. Clin Cancer Res. 2017;23(8):1920–1928. doi:10.1158/1078-0432.CCR-16-1741
12. Santarpia M, Karachaliou N. Tumor immune microenvironment characterization and response to anti-PD-1 therapy. Cancer Biol Med. 2015;12(2):74–78.
13. Rizvi NA, Hellmann MD, Snyder A, et al. Mutational landscape determines sensitivity to PD-1 blockade in non–small cell lung cancer. Science. 2015;348(6230):124–128. doi:10.1126/science.aaa1348
14. Hellmann MD, Callahan MK, Awad MM, et al. Tumor mutational burden and efficacy of nivolumab monotherapy and in combination with ipilimumab in small-cell lung cancer. Cancer Cell. 2019;35(2):329. doi:10.1016/j.ccell.2019.01.011
15. Hellmann MD, Ciuleanu TE, Pluzanski A, et al. Nivolumab plus ipilimumab in lung cancer with a high tumor mutational burden. N Engl J Med. 2018;378(22):2093–2104. doi:10.1056/NEJMoa1801946
16. Johnson DB, Frampton GM, Rioth MJ, et al. Targeted next generation sequencing identifies markers of response to PD-1 blockade. Cancer Immunol Res. 2016;4(11):959–967. doi:10.1158/2326-6066.CIR-16-0143
17. Frampton GM, Fichtenholtz A, Otto GA, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol. 2013;31(11):1023–1031. doi:10.1038/nbt.2696
18. Frampton GM, Fabrizio D, Chalmers ZR, et al. Assessment of tumor mutation burden from >60,000 clinical cancer patients using comprehensive genomic profiling. J Clin Oncol. 2016;11(1):34.
19. Kowanetz M, Zou W, Shames DS, et al. Tumor mutation load assessed by FoundationOne (FM1) is associated with improved efficacy of atezolizumab (atezo) in patients with advanced NSCLC. Ann Oncol. 2016;27:77P. doi:10.1093/annonc/mdw363.25
20. Chalmers ZR, Connelly CF, Fabrizio D, et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017;9(1):34. doi:10.1186/s13073-017-0424-2
21. Kou T, Kanai M, Yamamoto Y, et al. Clinical sequencing using a next-generation sequencing-based multiplex gene assay in patients with advanced solid tumors. Cancer Sci. 2017;108:1440–1446. doi:10.1111/cas.13265
22. Samstein RM, Lee CH, Shoushtari AN, et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat Genet. 2019;51(2):202–206. doi:10.1038/s41588-018-0312-8
23. Huang D, Zhang F, Tao H, et al. Tumor mutation burden as a potential biomarker for PD-1/PD-L1 inhibition in advanced non-small cell lung cancer. Targ Oncol. 2020;15:93–100. doi:10.1007/s11523-020-00703-3
24. Alborelli I, Leonards K, Rothschild SI, et al. Tumor mutational burden assessed by targeted NGS predicts clinical benefit from immune checkpoint inhibitors in non-small cell lung cancer. J Pathol. 2020;250(1):19–29. doi:10.1002/path.5344
25. Snyder A, Makarov V, Merghoub T, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371(23):2189–2199. doi:10.1056/NEJMoa1406498
26. Chan TA, Yarchoan M, Jaffee E, et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol. 2019;30:44–56. doi:10.1093/annonc/mdy495
27. Peters S. Abstract LBA7128 ‘Nivolumab (nivo) + low-dose ipilimumab (ipi) vs platinum-doublet chemotherapy (chemo) as first-line (1L) treatment (tx) for advanced non-small cell lung cancer (NSCLC): checkMate-227 part 1 final analysis’. Ann Oncol. 2019;30. doi:10.1093/annonc/mdz394.075
28. Smolle E, Leithner K, Olschewski H. Oncogene addiction and tumor mutational burden in non-small-cell lung cancer: clinical significance and limitations. Thorac Cancer. 2020;11(2):205–215. doi:10.1111/1759-7714.13246
29. Brahmer JR, Tykodi SS, Chow LQ, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366(26):2455–2465. doi:10.1056/NEJMoa1200694
30. Gainor JF, Shaw AT, Sequist LV, et al. EGFR mutations and ALK rearrangements are associated with low response rates to PD-1 pathway blockade in non-small cell lung cancer: a retrospective analysis. Clin Cancer Res. 2016;22(18):4585–4593. doi:10.1158/1078-0432.CCR-15-3101
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.