Back to Journals » OncoTargets and Therapy » Volume 13
TBRG4 Knockdown Suppresses Proliferation and Growth of Human Osteosarcoma Cell Lines MG63 Through PI3K/Akt Pathway
Authors Huang F, Liao F, Ma G, Hu Y, Zhang C, Xu P, Xu T, Chang J
Received 12 February 2020
Accepted for publication 14 May 2020
Published 27 July 2020 Volume 2020:13 Pages 7271—7281
DOI https://doi.org/10.2147/OTT.S249477
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Carlos E Vigil
Fei Huang,1,* Faxue Liao,1,* Guangwen Ma,1 Yong Hu,2 Chi Zhang,1 Pengfei Xu,1 Tangbing Xu,1 Jun Chang3
1Department of Orthopaedics, The Fourth Affiliated Hospital of Anhui Medical University, Hefei, Anhui, People’s Republic of China; 2Department of Orthopaedics, The First Affiliated Hospital of Anhui Medical University, Hefei, Anhui, People’s Republic of China; 3Clinical Research Centre, Zhujiang Hospital, Southern Medical University, Guangzhou, Guangdong, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Jun Chang
Clinical Research Centre, Zhujiang Hospital, Southern Medical University, Guangzhou, Guangdong, People’s Republic of China
Tel/ Fax + 86 551 66331115
Email [email protected]
Background: The transforming growth factor β regulator 4 (TBRG4) has been proved to be involved in various types of tumor. However, its contribution in human osteosarcoma (OS) is still unclear.
Patients and Methods: In the present study, immunohistochemistry and quantitative real-time PCR were performed to investigate the expression of TBRG4 in OS tissues obtained from patients and three types of cell lines. The effect of TBRG4 knockdown using lentivirus on tumorigenesis was detected by CCK8, high-content screening analysis, colony formation assay and flow cytometric analysis. Bioinformatics analysis was operated to investigate related signaling pathways following TBRG4 knockdown.
Results: The results showed that the expression of TBRG4 increased significantly in OS tissues and MG63 cell line. TBRG4 knockdown inhibited cell proliferation, colony and tumor formation, while activating cell apoptosis. Ingenuity Pathway Analysis and Western blot assay further indicated that TBRG4 knockdown may regulate the proliferation of human MG63 cells through PI3K/Akt signaling pathway.
Conclusion: Our results suggest that TBRG4 may become a promising therapeutic target for the treatment of human OS.
Keywords: TBRG4, osteosarcoma, proliferation, PI3K/Akt, MG63 cells
Introduction
Osteosarcoma (OS) is the most frequent malignant primary bone tumor commonly found in children and adolescents.1–3 It is characterized by high malignancy, early metastasis, poor prognosis and low survival rate, with an annual incidence rate of 4.4 cases per million in the US.2 Although OS accounts for less than 1% of all diagnosed cancers in the US, it is the most common primary malignancy with the 5-year survival rate of <40%.4–6 In addition, treatments with poor prognosis, drug resistance and toxic side effects are still crucial obstacles for radical therapies.7,8 Therefore, novel treatments are required urgently. In recent years, targeted therapy has shown great potential in the treatment of OS, thus, based on the implicated pathogenesis of OS, identification of new effective prognosis markers and key molecular targets for OS will facilitate the accurate diagnosis and therapy for OS patients.
Multiple studies have focused on novel molecular targets to improve OS therapeutic efficacy including RTKs,9 steroid receptor co-activator,10 mTOR,11 and aurora kinases.12 Recently, there has been interest in developing better understanding of tumor genesis mechanisms, especially about transforming growth factorβ (TGF-β). TGF-β is a growth factor which stimulates tumor growth in microenvironment of OS, the expression of TGF-β is higher in patients with OS than that in healthy individuals.13 Latest evidence has reported that TGF-β is a potential target regulator in the metastatic dissemination.14 However, whether TGF-β could be a potential treatment target for OS remains unclear.
The transforming growth factor β regulator 4 (TBRG4), also termed Fas-activated serine/threonine kinase domain-containing protein4, FASTKD4 or cell cycle progression restoration protein2, CPR2,15–17 encodes a regulator for TGF-β.18 It is therefore conceivable that TBRG4 could be a key molecular target in OS because of its regulation for TGF-β. TBRG4 is located on the 7p14-p13 chromosomal region, and participates in the development process of various diseases, especially cancer. Previous studies have demonstrated that TBRG4 is a novel oncogene in breast cancer, with its expression level correlating with breast tumor malignancy. In addition, deficiency of TBRG4 could significantly inhibit tumor cell proliferation and migration through promoting apoptosis.19 Thus, we speculated that TBRG4 may have the same effect on human OS. Notably, Sevcikova et al,18 collected nine samples of bone marrow plasma cells from multiple myeloma (MM) patients to evaluate the related genes associated with the high risk signature of MM patients. They found that TBRG4 may serve as a crucial gene and is highly relatied with MM. Therefore, the hypothesis has been put forward that the altered TBRG4 expression would affect human OS growth and development. However, the effect of TBRG4 on human OS are still poorly understood.
In the present study, we first evaluated the expression of TBRG4 in human osteosarcoma tissues and cell lines. A lentiviral system was used to determine the role of TBRG4 silencing on tumor proliferation, apoptosis, and colony formation. The effect of TBRG4 on differentially expressed genes patterns in human osteosarcoma tissues and cell lines was also evaluated.
Patients and Methods
Patients’ Tissues
In the current study, 40 patients diagnosed with OS at the First Affiliated Hospital of Anhui Medical University between 2013 and 2016 were enrolled. The study was approved by the Medical Ethics Committee of the Anhui Medical University (2013–76). All the patients provided written informed consent before tissue specimens were collected, and the study was conducted in accordance with the Declaration of Helsinki. OS tissues and para-carcinoma tissue which come from pre-chemotherapy of treatment (referred to as normal tissues, at least 1.0 cm apart from the visible cancerous tissues) were collected from distal femur for detecting the levels of TBRG4 expression by immunohistochemistry (IHC).
Cell Culture
Three human OS cells lines, MG63, U2OS, Saos-2 were purchased from the Cell Bank of Chinese Academy of Sciences (Shanghai, People's Republic of China). Cells were cultured in Roswell Park Memorial Institute 1640 medium (Thermo Fisher Scientific, Waltham, MA, USA) with 10% fetal bovine serum and 1% antibiotics (Sangon Biotech, Shanghai, People's Republic of China) in a humidified atmosphere at 37°C under 5% CO2.
Construction and Infection of shTBRG4 Lentivirus
A lentiviral system containing short hairpin RNA (shRNA) was used to manipulate TBRG4 expression. A candidate RNAi for human TBRG4 was designed to target the sequence 5ʹ- GTTCTTCAGCCTGGTACAT-3ʹ. Based on selected target sequences, shRNA interference sequence was designed to synthesize DNA oligo. TBRG4 RNAi lentivirus was generated via ligating Stem-loop DNA oligonucleotides into the lentiviral pGV115-GFP vector (GeneChem, Shanghai, People's Republic of China). The envelope plasmid was infected into MG63 cells, and the scramble shRNA was used as the negative shRNA control. After 72 hours of transfection, the infected cells with GFP proteins were observed using fluorescence microscopy (Olympus XI-71, Tokyo, Japan) with Lentivector Expression System (GeneChem) and fluorescent images were taken. After 72 hours of infection, MG63 cells were harvested and qRT-PCR and Western blot analysis were performed to determine knockdown efficiency.
IHC Staining
The patients’ OS tissue sections (5 μm thick) were dewaxed in xylene for 5 minutes, rehydrated in alcohol gradient and rinsed in deionized water. After incubating in 10 mM citrate buffer at 95°C for 12 minutes, the slides were fixed in fixative and blocked with 5% bovine serum albumin for 60 minutes. Then the slides were incubated with TBRG4 antibody (1:1000, CST) at 4°C overnight, followed by incubation of secondary antibody at room temperature for 1 hour. After extensive washing, the slides were stained with DAB and counterstained with hematoxylin. Finally, the slides were examined with a fluorescence microscope and the positive cells were brown in color.
RNA Extraction and qRT-PCR
Total RNA was extracted from MG63 cells using Trizol reagent (Life Technologies, USA), and reverse transcribed into cDNA using cDNA Synthesis Kit (Sangon, Biotech) according to the manufacturer’s protocol. The resulting cDNA was analyzed by qPCR performed with SYBR Green Supermix (Takara, Dalian, People's Republic of China), and the data were normalized to GAPDH and data analysis was conducted using the 2−ΔΔCt method. The primers used for qPCR were as follows:
GAPDH Forward, 5ʹ-TGACTTCAACAGCGACACCCA-3ʹ
Reverse, 5ʹ-CACCCTGTTGCTGTAGCCAAA-3ʹ(product size: 121bp);
TBRG4 Forward, 5ʹ- CAGCTCACCTGGTAAAGCGAT-3ʹ,
Reverse, 5ʹ-GGGAGTAGATGCTCGTTCCTTC-3ʹ(product size: 194bp).
Western Blot Analysis
To evaluate TBRG4 knockdown efficiency in MG63 cells, Western blot analysis was employed to determine the TBRG4 expression. The MG63 cells were sonicated in RIPA lysis buffer. The total protein concentration of cell lysates was measured by the BCA method using BCA protein kit (Sangon Biotech). Forty micrograms of proteins were resolved by 812% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to polyvinylidene fluoride membranes (Sango Biotech). The membranes were blocked with 5% non-fat milk for 2 hours, incubated with primary antibody for 3 hours at room temperature, followed by 4°C overnight. Afterward, the membranes were incubated with secondary antibody at room temperature for 1 hour and visualized using the ECL system (GE Healthcare Bio-Sciences AB, Image Quan LAS 500, Sweden).
Cell Proliferation Assay
MG63 cells transfected with TBRG4 lentivirus were seeded in 96-well black assay plates with a clear bottom and lid at a density of 2000 cells per well. After transfection, the plate was imaged and analyzed once a day for 4 days continuously using a Celigo Cell Imaging Cytometer (Nexcelom Bioscience, Boston, MA, USA). Images were processed and quantified using the Cell Analyzer software.
The effect of shTBRG4 on MG63 cells proliferation was also evaluated by CCK8 assay using the cell viability assay kit. First, 2000 cells per well were seeded in 96-well plates and incubated for 24 hours. The cell-free group was set as the blank control group, and three repeating pipettes were set in each group. Then, 10 µL CCK8 reagent was added into each well and incubated for 4 hours at 37°C. The absorbance was read at 490 nm using ELx800 Absorbance Microplate Reader (BioTek Instruments, Winooski, VT, USA) for 4 consecutive days.
Colony Formation Assay
MG63 cells transfected with shTBRG4 or shCtrl lentivirus were cultured and incubated in complete medium for 2 days until the logarithmic phase was reached. Then, cells were collected and reseeded at a density of 600 cells per well in 6-well plates and incubated for 14 days at 37°C with 5% CO2, in which three duplicates were set for each group. The culture medium was changed every 3 days and the state of cells were observed continuously. The cells were fixed in 4% paraformaldehyde for 60 minutes at room temperature after washing twice with PBS, then the colonies were stained with Giemsa solution (Dingguo Biotech, Shanghai, People's Republic of China) for 20 minutes. The colonies were imaged and counted by a fluorescence microscope (Olympus XI-71, Tokyo, Japan).
Tumor Sphere Formation Assay
Cells were trypsinized and seeded at a density of 1×103/well in a 24-well ultra-low attachment round bottom plate, cultured in complete medium with DMEM/F12 media with 10 ng/mL basal fibroblast growth factor, 5 µg/mL insulin, 0.4% bovine serum albumin, 20 ng/mL EGF. The spheres were photographed and counted on day 1, day 6, and day 11 using a fluorescence microscope.
High-Content Screening Analysis
To evaluate the effect of shTBRG4 on cell proliferation, 1×104 cells/well were seeded in 96-well assay plates that were black with a clear bottom and lid. After incubation of 24 hours, the cells were transfected with shTBRG4 and shCtrl for 72 h. EGFP were added and the live-cell imaging was performed on the IN Cell analyzer 2200 Cell Imaging System for 4 consecutive days. Cell images were analyzed by the IN Cell Investigator software.
Apoptotic Assay
For determination of shTBRG4 on MG63 cell apoptosis, the Annexin V-FITC and PI staining kit (AAT bioquest, People's Republic of China) was used. MG63 cells were infected with shTBRG4 or shCtrl lentivirus, incubated for 48 hours, the cells were then collected and washed in 1 mL of binding buffer and centrifuged at 3000 g for 10 minutes. Then the cell pellets were resuspended in binding buffer, and 10 µL of Annexin V-FITC was added to each group incubating for 15 minutes at room temperature while protecting from light. After which, 5 µL of PI solution was added immediately prior to analysis using flow cytometry (Beckman Coulter).
PathArray Analysis
The genome-wide effect of TBRG4 knockdown was analyzed by the PrimeView Human Gene Expression Array. Three replicates of MG63 cells transfected with shTBRG4 or shCtrl lentiviruses were analyzed. Total RNA of MG63 cells infected with shTBRG4 or shCtrl for 72 hours was extracted by Trizol, reversed transcription into cDNA, which was further underwent in vitro transcriptional transcription and synthesis into antisense RNA (aRNA). Then the aRNA was fragmented into cDNA and hybrid with GeneChip, which was performed with GeneChip Hybridization Wash and scanned directly post-hybridization using a GeneChip Scanner.
Ingenuity Pathway Analysis (IPA)
Results of differentially expressed genes were analyzed by IPA tool. The p-value (<0.05) and Z-score (>2) were used to interpret the differentially expressed data, in which the “regulator effects,” “diseases and functions,” “canonical pathway,” “upstream analysis,” and “molecular network” were involved. The differentially expressed genes were mapped onto genetic networks and ranked based on the Z-score to measure biological networks, functional signaling pathways and upstream or downstream target genes according to IPA program.
Statistical Analysis
In the present study, all results are from at least three independent experiments and data were expressed as Mean±Standard (SD). Statistical analyses were performed by SPSS v. 20.0 with one-way ANOVA to test the significance of normal group and TBRG4 knockdown group. A p-value of less than 0.05 was considered statistically significant.
Results
The Role of TBRG4 Expression in Human OS Tissues and Cell Lines
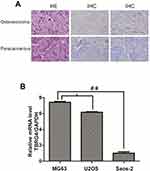
To investigate the role of TBRG4 in OS, we analyzed the expression levels of TBRG4 in human osteosarcoma tissues and cells. As shown in Figure 1A, all specimens emerged as osteosarcoma by pathological examinations with HE staining (HE images on the left). High TBRG4 immunoreactivity was observed in OS tissues compared with para-carcinoma (IHC images in the middle and on the right). The mRNA expression levels of TBRG4 was detected in malignant human OS cells, TBRG4 was widely expressed in MG63 and U2OS cells, rather than Saos-2 cells (Figure 1B) (p<0.05). Thus, MG63 cells would be selected for subsequent experimental analysis. Taken together, these results demonstrated that TBRG4 exhibit high expression levels at both mRNA and protein levels in OS tissues and cells compared to the normal group (p<0.05).
The Efficiency of Lentivirus-Mediated TBRG4 in MG63 Cells
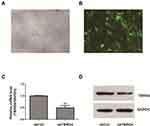
To evaluate the efficiency of knocking down TBRG4 expression mediated by lentivirus, the MG63 cells were transfected with shTBRG4 or shCtrl lentivirus. After transfection with lentivirus for 72 hours, the proportions of infected MG63 cells expressing GFP accounted for more than 80%, which was confirmed as high infection efficiency and could be used for further analysis (p<0.05) (Figure 2A and B). TBRG4 mRNA expression and protein level was determined by qPCR and Western blot assay respectively. As shown in Figure 2C, OS cells transfected with shTBRG4 lentivirus showed a 50.3% decrease in TBRG4 mRNA expression (p<0.05). Consistent with qPCR data, the protein expression level of the shTBRG4 lentivirus transfected group showed the similar trend compared to control group (p<0.05) (Figure 2D). Therefore, these results demonstrated that TBRG4 expression was downregulated by shTBRG4 lentivirus infection successfully at both mRNA and protein levels.
TBRG4 Knockdown Inhibited Cell Proliferation in MG63 Cells
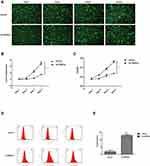
The effect of TBRG4 knockdown on MG63 cells proliferation was determined by both High-content screening analysis and CCK8 assay. To evaluate the effect of shTBRG4 on cell proliferation, high-content screening analysis was used to observe the cell count transfected with shTBRG4 or shCtrl lentivirus for 4 days continuously. The number of cells in the control group increased in a time-dependent manner, while only a slight increase was observed in the shTBRG4 group (Figure 3A), and the difference was statistically significant (Figure 3B). Furthermore, the effect of shTBRG4 on cell proliferation was also detected using CCK8 assay. After transfecting with shTBRG4 or shCtrl lentivirus for 72 h, CCK8 reagent was used to figure up the OD450 ratio, the result showed that TBRG4 knockdown inhibited cell proliferation significantly compared with the control group (Figure 3C) (p<0.05). All these results showed that TBRG4 knockdown could inhibit MG63 cells proliferation significantly.
Deficiency of TBRG4 Promoted Apoptosis in MG63 Cells
To evaluate whether TBRG4 knockdown affect MG63 cells apoptosis, flow cytometry was used to determine the cell apoptosis. The result observed was 1.88% of the apoptotic rate in cells transfected shCtrl lentivirus, while, in OS cells infected with shTBRG4, apoptotic cell rate reached 12.95% (Figure 3D). The difference was statistically significant (Figure 3E). The results revealed that TBRG4 deficiency could lead to induction of apoptosis in MG63 cells.
TBRG4 Knockdown Suppressed the Ability of MG63 Cells to Form Colonies and Tumor
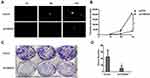
Tumor sphere formation assay was performed to inspect the effect of shTBRG4 lentivirus on tumor sphere formation capacity of MG63 cells. As shown in Figure 4A, deficiency of TBRG4 significantly reduced the size and number of spheres than that formed by the control lentivirus cells, and the difference was statistically significant (Figure 4B). Colony formation assay was further performed to evaluate the ability of colony formation following TBRG4 knockdown. The result showed significant cell proliferation suppression (p<0.05). As shown in Figure 4C and D, an average count of colony was 112 in cells with shCtrl infection, however, only 46 colonies were found on the cells transfected with shTBRG4 lentivirus. These results suggest that depletion of TBRG4 can inhibit the ability of tumor proliferation of MG63 cells.
Knockdown of TBRG4 Can Alter Profiles of Signaling Molecules in MG63 Cells
Gene expression profiling platform with a PrimeView Human Gene Expression Array was used to compare differential gene expression profiles between three biological replicates of shCtrl and shTBRG4 cells. A total of 2,268 differentially expressed genes were identified, in which 1,134 were upregulated and 1,134 were downregulated (Figure 5A). To verify the accuracy of the gene chip, we selected 30 genes with significant differences to be checked using qPCR. The results showed that all the 30 genes expression was consistent with the results of gene expression array (Figure 5B). An IPA program was performed to analyze the regulator effect, diseases and functions, canonical pathway, upstream analysis, and molecular network in the 2,268 differently regulated genes. The enrichment analysis of differential genes using IPA database revealed 13 significantly altered ingenuity canonical pathways, in which seven pathways were significantly inhibited (Z-score≤-2) and six were activated (Z-score≥2). 3-Phosphoinositide Biosynthesis Pathway was significantly inhibited with a Z-score of −2.401 and Superpathway of Cholesterol Pathway was significantly activated with a Z-score of 2.449 (Figure 5C). To investigate the signaling pathways in shTBRG4-treated MG63 cells, we evaluated the PI3K/Akt signaling pathway expression involved in 3-Phosphoinositide Biosynthesis Pathway using Western blot assay. The data showed that the protein expression of PI3K and Akt was significantly decreased in shTBRG4-treated MG63 cells compared to shCtrl-treated MG63 cells (Figure 5D). These results provide foundation for the subsequent molecular mechanisms study of TBRG4 in MG63 cells.
Discussion
OS is highly aggressive and is characterized by early dissemination and poor prognosis with no significant therapeutic advances since the 1970s.20 Despite the effective limitation of micro-metastatic and improvement of OS rate by the incorporation of adjuvant systemic chemotherapy including doxorubicin, cisplatin, and high-dose methotrexate,21 the patients with metastatic disease remain a considerable challenge with a 5-year survival rate of only 20% or less.22 In addition, high-dose chemotherapy induces several adverse effects.23 Therefore, there is an increasing sense of urgency to develop specific molecular targeted therapeutic approaches for OS patients with poor prognosis. Accumulating evidences have demonstrated that TBRG4 is implicated in numerous types of cancer including lung cancer,19 glioblastoma,24 Sézary syndrome,25 and multiple myeloma.18 These findings implied that TBRG4 is closely related to human tumorigenesis, which provides evidence to speculate that TBRG4 might be a potential biomarker in OS. However, the contribution of TBRG4 to OS has not yet been evaluated.
The present study described that the expression of TBRG4 in human OS tissues and MG63 cells was significantly higher than in paracancerous tissues (Figure 1A, IHC assay) and other cancer cell lines (Figure 1B). To further investigate its effect on tumorigenesis in MG63 cells, the lentivirus was applied to efficiently knockdown TBRG4 expression and the results suggested that silence of TBRG4 suppressed cell proliferation, the ability to form colonies and tumor, while activating apoptosis in MG63 cells. The results above indicate that TBRG4 plays a crucial role in the promoting proliferation of MG63 cells and may participate in the pathological process of OS.
The TBRG4 gene is located on 7p14-p13 and encodes a regulator of TGF-β. Evidences have reported that increased expression of TBRG4 had a direct connection to MM, and TBRG4 could facilitate risk stratification of MM patients.26 Wang et al19 demonstrated that TBRG4 increased significantly in lung cancer tissues compared to cancer-adjacent normal tissues. Also, previous studies showed that TBRG4 is a novel oncogene in breast cancer, with its expression level correlating with breast tumor malignancy. In line with these findings, our study identified that TBRG4 expression increased both in human OS tissues and MG63 cell lines compared to the control group, which suggests that TBRG4 plays an important role in OS and may participate in the pathological mechanisms of human OS.
Multiple studies have reported a role of TBRG4 in various diseases. TBRG4 was recognized as a cell cycle-related gene which is located in mitochondria. The depletion of TBRG4 leads to alteration of cell functions. Ali et al15 reported the potential role of TBRG4 encoded in the mitochondrial genome for variable mitochondrial-encoded gene expression in complex disease. Disrupting TBRG4 results in decreased ND3 mRNA and ND5-CYB precursor accumulation, suggesting the regulation of mitochondrial RNA homeostasis linked to mitochondrial disease.27 Additionally, it has been reported that nearly 30% of cell lines in head and neck squamous cell carcinomas have chromosome abnormalities on 7p13-p22, the region where TBRG4 is located.28 For instance, Sézary syndrome is a leukemic form of cutaneous T-cell lymphoma with chromosomal abnormalities and mutations in some genes. TBRG4, involved in the pathogenesis of Sézary syndrome, was disrupted due to deletions and duplications of large segments of chromosome 17 in Sézary syndrome patients.25 Recently, the effect of TBRG4 depletion on lung cancer was reported to inhibit tumor cell proliferation and tumor formation and to induce apoptosis in H1299 cell line. Furthermore, deficiency of TBRG4 could significantly reduce tumor cell proliferation and migration in a list of breast cancer cell lines.19 In accordance with those studies, we found that depletion of TBRG4 inhibited viability and proliferation of human OS cells. In addition, our results show that knockdown of TBRG4 induced significant cell apoptosis, and we speculated that the growth suppression may partially contribute to activated apoptosis.
To further investigate the mechanisms of TBRG4 in numerous types of tumor, researchers have predicted potential downstream or upstream target genes using bioinformatics analysis methods, although the study of pathologic mechanisms of TBRG4 in cancers are still limited. Wang et al,19 demonstrated a significant increase in the level of DDIT3 and a decrease in the levels of CAV1 and RRM2 genes, and came to the conclusion that TBRG4 knockdown regulated the tumorigenesis of H1299 cells by upregulation of DDIT3 and downregulating of CAV1 and RRM2. In our study, the whole-genome sequencing analysis of human OS tumors with TBRG4 depletion was performed to obtain an understanding of molecular network related to TBRG4 and OS. According to the results of IPA, 2,268 differentially expressed genes were categorized into a number of diseases. Further analysis showed that 3-phosphoinositide biosynthesis pathway was significantly inhibited with a Z-score of −2.401.
Phosphoinositides play a key role in signal transduction, transmitting signals from the cell surface to modulate intracellular processes. The 3-phosphoinositide pathway is one of the major pathways of intracellular signal transduction, in which the phosphatidylinositol 3-kinases (PI3K) plays a central role in this pathway. Genetic activation of the PI3K pathway is very common in cancer, and it has been shown to regulate cellular activity through stimulation of receptor tyrosine kinases and the concomitant assembly of receptor-PI3K complexes.29 In addition, evidence has shown that targeting PI3K and mTOR were effective at inducing apoptosis in primary OS cell cultures in vitro, suggesting that PI3K/mTOR pathway is a sensitive target in human OS.30 The present study indicates that knockdown of TBRG4 significantly decreased expression of PI3K and Akt, which may be responsible for increased apoptosis and inhibition of tumor proliferation in MG63 cells. However, the exact mechanism of PI3K/Akt pathway involved in shTBRG4-treated human OS cells is currently unclear and requires further investigations.
Conclusion
In conclusion, our results highlighted that TBRG4 expression is increased in OS tissues and cell lines. We identified the role of TBRG4 in MG63 cells by silencing its expression with shTBRG4 lentivirus. The current study showed that TBRG4 depletion inhibited cell proliferation, the ability of colony formation and tumor formation, while activating apoptosis in MG63 cells. In addition, TBRG4 knockdown inhibits the protein expression of PI3K and Akt, which suggests that PI3K/Akt signaling pathway may participate in the role of TBRG4 knockdown on tumor proliferation. These findings imply that TBRG4 knockdown plays an essential role in decreased tumor proliferation of OS cell lines and offers a broad range of understanding for pathogenic cause of OS. However, the exact underlying molecular mechanisms of TBRG4 in OS are yet to be further studied.
Highlights
1. The high expression of TBRG4 exhibited in human osteosarcoma tissues and MG63 cell line.
2. The inhibition of cell viability and colony formation shown in TBRG4 knockdown cells, with an increasing trend of cell apoptosis.
3. PI3K/Akt pathway played a crucial role in the process of shTBRG4 on proliferation of MG63 cells.
4. TBRG4 will probably become a therapeutic target and biomarker in osteosarcoma treatment.
Abbreviations
TBRG4, transforming growth factor β regulator 4; OS, osteosarcoma; qPCR, quantitative real-time PCR; IPA, ingenuity pathway analysis; TGF-β, transforming growth factor β; MM, multiple myeloma; IHC, immunohistochemistry; shRNA, short hairpin RNA; PI3K, phosphatidylinositol 3-kinases; RTKs, receptor tyrosine kinases; aRNA, antisense RNA.
Acknowledgments
The authors would like to thank the anonymous reviewers for their comments.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Durfee RA, Mohammed M, Luu HH. Review of Osteosarcoma and current management. Rheumatol Ther. 2016;3(2):221–243. doi:10.1007/s40744-016-0046-y
2. Saraf AJ, Fenger JM, Rayan D, et al. Osteosarcoma: accelerating progress makes for a hopeful future. Front Oncol. 2018;8:4. doi:10.3389/fonc.2018.00004
3. Kim M, Kim DJ, Kim JIJOMS. GFRA1: a novel molecular target for the prevention of Osteosarcoma chemoresistance. Int J Mol Sci. 2018;19(4):1078. doi:10.3390/ijms19041078
4. Fu Y, Lan T, Cai H, et al. Meta-analysis of serum lactate dehydrogenase and prognosis for osteosarcoma. Medicine. 2018;97(19):e0741. doi:10.1097/MD.0000000000010741
5. Chen R, Wang G, Zheng Y, et al. Drug resistance related microRNAs in osteosarcoma: translating basic evidence into therapeutic strategies. J Cell Mol Med. 2019;23(4):2280–2292. doi:10.1111/jcmm.14064
6. Janeway KA, Grier HEJLO. Sequelae of osteosarcoma medical therapy: a review of rare acute toxicities and late effects. Lancet Oncol. 2010;11(7):670–678. doi:10.1016/S1470-2045(10)70062-0
7. Zhang Y, Rosenberg AE. Bone-forming tumors. Surg Pathol Clin. 2017;10(3):513–535. doi:10.1016/j.path.2017.04.006
8. Harrison DJ, Geller DS, Gill JD, et al. Current and future therapeutic approaches for osteosarcoma. Expert Rev Anticancer Ther. 2018;18(1):39.
9. Chaiyawat P, Klangjorhor J, Settakorn J, et al. Activation status of receptor tyrosine kinases as an early predictive marker of response to chemotherapy in Osteosarcoma. Transl Oncol. 2017;10(5):846–853. doi:10.1016/j.tranon.2017.08.005
10. Hu C, Deng Z, Zhang Y, et al. The prognostic significance of Src and p-Src expression in patients with osteosarcoma. Med Sci Monit. 2015;21:638–645. doi:10.12659/MSM.892803
11. Zhou Q, Deng Z, Zhu Y, et al. mTOR/p70S6K signal transduction pathway contributes to osteosarcoma progression and patients’ prognosis. Med Oncol. 2010;27(4):1239–1245. doi:10.1007/s12032-009-9365-y
12. Maris JM, Morton CL, Gorlick R, et al. Initial testing of the aurora kinase A inhibitor MLN8237 by the Pediatric Preclinical Testing Program (PPTP). Pediatr Blood Cancer. 2010;55(1):26–34. doi:10.1002/pbc.22430
13. Lamora A, Talbot J, Mullard M, et al. TGF-β signaling in bone remodeling and Osteosarcoma progression. J Clin Med. 2016;5(11):96. doi:10.3390/jcm5110096
14. Rodriguez Calleja L, Jacques C, Lamoureux F, et al. DeltaNp63alpha silences a miRNA program to aberrantly initiate a wound-healing program that promotes TGFbeta-induced metastasis. Cancer Res. 2016;76(11):3236–3251. doi:10.1158/0008-5472.CAN-15-2317
15. Ali AT, Boehme L, Carbajosa G, et al. Nuclear genetic regulation of the human mitochondrial transcriptome. Elife. 2019;8.
16. Jourdain AA, Popow J, Ma DLF, et al. The FASTK family of proteins: emerging regulators of mitochondrial RNA biology. Nucleic Acids Res. 2017;45:19. doi:10.1093/nar/gkx772
17. Simarro M, Gimenez-Cassina A, Kedersha N, et al. Fast kinase domain-containing protein 3 is a mitochondrial protein essential for cellular respiration. Biochem Biophys Res Commun. 2010;401(3):440–446. doi:10.1016/j.bbrc.2010.09.075
18. Sevcikova S, Pascekova H, Besse L, et al. Extramedullary relapse of multiple myeloma defined as the highest risk group based on deregulated gene expression data. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2015;159(2):288–293. doi:10.5507/bp.2015.014
19. Wang A, Zhao C, Liu X, et al. Knockdown of TBRG4 affects tumorigenesis in human H1299 lung cancer cells by regulating DDIT3, CAV1 and RRM2. Oncol Lett. 2018;15(1):121–128. doi:10.3892/ol.2017.7328
20. Harrison DJ, Geller DS, Gill JD, et al. Current and future therapeutic approaches for osteosarcoma. Expert Rev Anticancer Ther. 2017;18(1):39–50. doi:10.1080/14737140.2018.1413939
21. Luetke A, Meyers PA, Lewis I, et al. Osteosarcoma treatment - where do we stand? A State of the Art Rev. 2014;40(4):523–532.
22. Hagleitner MM, Bont ED, Loo, DTJS. Survival trends and long-term toxicity in pediatric patients with Osteosarcoma. Sarcoma. 2012;2012(24):636405. doi:10.1155/2012/636405
23. Marco W, B W. Targets for cancer therapy in childhood sarcomas. Sch?fer, Cancer Treatment Rev. 2010;36(4):318–327. doi:10.1016/j.ctrv.2010.02.007
24. Varghese RT, Liang Y, Guan T, et al. Survival kinase genes present prognostic significance in glioblastoma. Oncotarget. 2016;7(15):20140–20151. doi:10.18632/oncotarget.7917
25. Prasad A, Rabionet R, Espinet B, et al. Identification of gene mutations and fusion genes in patients with Sezary syndrome. J Invest Dermatol. 2016;136(7):1490–1499. doi:10.1016/j.jid.2016.03.024
26. Sarasquete ME, Martinez-Lopez J, Chillon MC, et al. Evaluating gene expression profiling by quantitative polymerase chain reaction to develop a clinically feasible test for outcome prediction in multiple myeloma. Br J Haematol. 2013;163(2):223–234. doi:10.1111/bjh.12519
27. Boehm E, Zaganelli S, Maundrell K, et al. FASTKD1 and FASTKD4 have opposite effects on expression of specific mitochondrial RNAs, depending upon their endonuclease-like RAP domain. Nucleic Acids Res. 2017;45(10):6135–6146. doi:10.1093/nar/gkx164
28. Carey TE, Van Dyke DL, Worsham MJ. Nonrandom chromosome aberrations and clonal populations in head and neck cancer. Anticancer Res. 1993;13(6b):2561–2567.
29. Vanhaesebroeck B, Bilanges B, Madsen RR, et al. Perspective: potential impact and therapeutic implications of oncogenic PI3K activation on chromosomal instability. Biomolecules. 2019;9:8. doi:10.3390/biom9080331
30. Gupte A, Baker EK, Wan SS, et al. Systematic screening identifies dual PI3K and mTOR inhibition as a conserved therapeutic vulnerability in Osteosarcoma. Clin Cancer Res. 2015;21(14):3216–3229. doi:10.1158/1078-0432.CCR-14-3026
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.