Back to Journals » Cancer Management and Research » Volume 13
Systemic Inflammatory Markers of Resectable Colorectal Cancer Patients with Different Mismatch Repair Gene Status
Authors Li J, Zhang Y, Xu Q, Wang G, Jiang L, Wei Q, Luo C, Chen L, Ying J
Received 5 January 2021
Accepted for publication 10 March 2021
Published 30 March 2021 Volume 2021:13 Pages 2925—2935
DOI https://doi.org/10.2147/CMAR.S298885
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
Jingjing Li,1 Yiwen Zhang,2 Qi Xu,1 Gang Wang,3 Lai Jiang,3 Qing Wei,1 Cong Luo,1 Lei Chen,1 Jieer Ying1
1Department of Hepato-Pancreato-Biliary & Gastric Medical Oncology, Cancer Hospital of the University of Chinese Academy of Sciences (Zhejiang Cancer Hospital), Institute of Cancer and Basic Medicine (IBMC), Chinese Academy of Sciences, Hangzhou, Zhejiang Province, 310022, People’s Republic of China; 2Department of Pharmacy, Zhejiang Provincial People’s Hospital, People’s Hospital of Hangzhou Medical College, Hangzhou, Zhejiang Province, 310000, People’s Republic of China; 3Department of Colorectal Surgery, Cancer Hospital of the University of Chinese Academy of Sciences (Zhejiang Cancer Hospital), Institute of Cancer and Basic Medicine (IBMC), Chinese Academy of Sciences, Hangzhou, Zhejiang Province, 310022, People’s Republic of China
Correspondence: Jieer Ying
Department of Hepato-Pancreato-Biliary & Gastric Medical Oncology, Cancer Hospital of the University of Chinese Academy of Sciences (Zhejiang Cancer Hospital), Institute of Cancer and Basic Medicine (IBMC), Chinese Academy of Sciences, Hangzhou, Zhejiang Province, 310022, People’s Republic of China
Tel +86-571-88122062
Email [email protected]
Background: We aimed to assess the differences in gene expression and systemic inflammatory markers in colorectal cancer (CRC) patients with different mismatch repair (MMR) statuses.
Methods: Bioinformatics analysis was used to identify the different expression genes in patients with CRC at different MMR statuses. A total of 208 patients with resectable colorectal cancer, including 104 deficient mismatch repair (dMMR) patients and 104 matched proficient mismatch repair (pMMR) patients, were retrospectively analyzed.
Results: Bioinformatics analysis showed that chemokine-mediated signaling pathway and inflammatory responses were the main differences in gene expression between dMMR and pMMR CRC patients. In all 208 patients with CRC, those with dMMR frequently had it located on the right side, with more mucinous adenocarcinoma and grade 3 tumors. Patients with dMMR had an earlier American Joint Committee on Cancer (AJCC) stage than pMMR patients. Meanwhile, lymph nodes (LNs) metastasis was more frequently negative in dMMR patients than pMMR patients. Interestingly, patients with CRC with dMMR had more regional lymph nodes removed during surgery, although with less metastatic cancer. Patients with resectable CRC with dMMR were more likely to have higher levels of neutrophil, monocyte, platelet, neutrophil/lymphocyte ratio (NLR), platelet/lymphocyte ratio (PLR), monocyte-to-lymphocyte ratio (MLR), C-reactive protein to albumin ratio (CAR), Glasgow prognostic score (GPS) and C-reactive protein (CRP). In patients with dMMR, those with higher levels of PLR, MLR, CAR, and co-effect present had shorter overall survival (OS) significantly. It was noteworthy that the prognosis of high levels of systemic inflammatory markers did not predict prolonged OS in patients with pMMR CRC.
Conclusion: dMMR CRC has presented a comprehensively distinct systemic inflammatory microenvironment. The systemic inflammatory response can predict oncological outcomes in patients with CRC with dMMR.
Keywords: gene expression, systemic inflammatory markers, colorectal cancer, bioinformatics analysis, lymph nodes
Introduction
Colorectal cancer (CRC) is the leading malignant tumor worldwide. According to the report in the Global Burden of Disease study, CRC ranks fourth in terms of morbidity but second in terms of mortality in 2017.1 In China, according to 2015 data, CRC ranks fifth in both morbidity and mortality.2 CRC constitutes a great threat to human health. Understanding its clinicopathological characteristics could guide clinical diagnosis and treatment.
The occurrence and development of CRC are complex processes. Microsatellite instability (MSI) is considered as the genetic inducer for hereditary nonpolyposis colorectal cancer (HNPCC). Some studies have shown that MSI is also associated with the formation and development of sporadic colorectal cancer (SCRC), and MSI can be seen in about 15% of all CRC cases.3
MSI is defined as the change of the length of microsatellite DNA, which is caused by the insertion or deletion of repeat units in tumor tissue, and results in the emergence of new microsatellite DNA alleles. Mismatch repair (MMR) refers to the repair of nucleotide sequences in DNA molecules, which contains mismatched bases.4 It has been reported that MLH1, MSH2, MSH6 and PMS2 are dominant proteins of MMR.5 Loss of function of one or more mismatch repair proteins can lead to MSI, known as mismatch repair protein deficiency (dMMR). Therefore, the MSI status of tumor can be shown indirectly by detecting the missing mismatch repair protein.
Due to its prognostic and predictive values in some tumors, the MSI/MMR status has attracted more and more attention in cancer research. Clinical researches showed that CRC patients with MSI high or dMMR had improved clinical outcomes.6 Moreover, clinical data shown that solid tumor patients with MSI-high or dMMR can benefit from programmed cell death 1 (PD-1) inhibitors.7 Understanding the relationship between MMR status and clinicopathological characteristics in CRC patients can help to further guide clinical treatment and explore the pathogenesis of this disease.
Here, we explore the differences in gene expression between dMMR and pMMR patients with colorectal cancer and then investigate the differences in plasma inflammation markers of resectable colorectal cancer patients with different mismatch repair gene statuses.
Materials and Methods
Bioinformatics Analysis
The Gene Expression Omnibus (GEO) database under the National Center for Biotechnology Information (NCBI) platform was used for chip screening. The target chip needs to meet the following screening criteria. First, the specimens were human colon cancer specimens, excluding cell lines and animal specimens. Second, the chip to be selected needs to have MMR status information. Third, gene expression microarray is required, and each probe has a normalized expression value. Finally, the quality of the selected chips should meet the quality requirements, and their quality was determined by regression analysis using the affyPLM package in R language (http://bioconductor.org/biocLite.R). The online tool GEO2R (https://www.ncbi.nlm.nih.gov/geo/gep2r/) was used to analyze the selected chips. Genes with P<0.05 and more than two times difference were selected as candidate differential gene probes, and the probes were converted into standard gene names. Then, DAVID version 6.8 (https://david.ncifcrf.gov) was performed for online analysis, gene ontology (GO), gene function annotation, and KEGG pathway enrichment analysis were conducted to screen out the characteristic gene clusters and pathways between pMMR and dMMR.
Patients
The protocol was approved by a human research ethics committee. A single-center study was conducted in Zhejiang Cancer Hospital, whose database comprises 1679 CRC cases who underwent surgery from January 2013 to June 2014. The last follow-up ended on November 1, 2019. Written informed consent was provided by each patient before enrolment. Among these cases, a total of 1184 underwent MMR testing by IHC, and 170 (14.4%) had the dMMR status. Exclusion criteria were: undefined disease stage, metastasis, and loss to follow-up. Finally, 104 patients with stage I–III sporadic colorectal adenocarcinoma were included in this study. Subsequently, we identified another 104 patients with resectable colorectal cancer with pMMR matched with dMMR patients over the same period (Figure 1). Clinicopathological data, including age, gender, tumor location, histological subtype, tumor grade, tumor stage, regional lymph node (LNs) metastasis, American Joint Committee on Cancer (AJCC) 8th edition stage, perineural invasion, vascular invasion, number of retrieved LNs, number of metastatic LNs, systemic inflammation markers, including C-reactive protein (CRP), lactic dehydrogenase (LDH), albumin, neutrophil, monocyte, lymphocyte, platelet, and survival were retrospectively reviewed. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of IRB-2020-239 (Ke).
 |
Figure 1 Flowchart of patient selection. |
Inflammation-Based Prognostic Score
The NLR, PLR, MLR were calculated respectively as NLR = neutrophil ratio (%) (or number of neutrophils)/Lymphocyte ratio (%) (or number of lymphocytes), PLR = platelet ratio (%) (or number of platelet)/Lymphocyte ratio (%) (or number of lymphocytes), MLR = monocyte ratio (%) (or number of monocyte)/Lymphocyte ratio (%) (or number of lymphocytes). GPS was calculated as: patients showing both elevated levels of CRP (>10 mg/dl) and hypo-albuminemia (<3.5 g/dl) were given a score of 2, while patients with only one or none of these biochemistry abnormalities were given a score of 1 or 0, respectively. The CAR was calculated as: CAR = CRP level (mg/dl)/albumin level (g/dl).
Immunohistochemistry
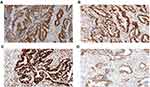
The procedures were performed according to the operation steps of immunohistochemistry, just as described in our previous study.8 As shown in Figure 2, the above proteins were all detected in the nucleus. Two blinded investigators separately assessed signal intensities under a light microscope at 200×. Discrepancies required discussion until a consensus was reached or a third investigator reviewed. Non-neoplastic colonic tissue, stroma and infiltrating lymphocytes usually show positive haematoxylin staining and serve as internal positive controls.
Statistical Analysis
SPSS v19.0 (SPSS, USA) was employed for data analysis. The independent samples t-test or χ2 test was employed for assessing associations of MMR statuses with various clinicopathologic features and systemic inflammation markers. Survival was estimated by Kaplan-Meier curves. Two-sided P<0.05 indicated statistical significance.
Results
Differential Expression Genes in Patients with Different MMR Status
Through screening and chip quality analysis, one chip met the inclusion criteria mentioned above, which was GSE39084. This chip data contained 16 patients with CRC with dMMR and 54 patients with CRC with pMMR. After data preprocessing and normalization, we used Limma packages in R language analysis to identify differential expression genes (DEG) between the two groups. According to threshold criterion (P<0.05 and |log FC|>1), we obtained 531 DEG, including 230 up-regulated and 301 down-regulated, and we conducted cluster analysis on the top 50 DEG (Figure 3A). Moreover, these 50 genes were inputted into DAVID for functional annotation, including biological process, cellular components, molecular function, and KEGG pathway enrichment analysis. As shown in Figure 3B–E, these overlapping genes were significantly enriched in “chemokine-mediated signaling pathway” and “inflammatory response”, which is characterized by complex series of events of changes in various blood cells, such as neutrophil leucocytes and mononuclear phagocytes. Based on this result, we further explored the blood cells and other inflammation markers in the serum of patients with resectable colorectal cancer with different mismatch repair gene statuses.
Baseline Patient Features
Table 1 summarizes the clinicopathological features of all 208 CRC patients, including 104 dMMR patients and 104 pMMR patients. Among them, patients with dMMR were younger than those with pMMR (56.1 v 59.7 years, P=0.003). There was no significant association between patients with different mismatch repair gene statuses and their sex, tumor stage, perineural invasion, and vascular invasion (P>0.05). Compared to patients with CRC with pMMR, those with dMMR were more frequently located on right side (41.3% v 18.3%, P<0.001), were with more mucinous adenocarcinoma (58.7% v 5.8%, P<0.001), and were with more grade 3 for tumor grade (35.6% v 15.4%, P=0.003). Patients with dMMR had earlier AJCC stage than pMMR patients (AJCC stage 3:26% v 57.7%, P<0.001). Meanwhile, LNs metastasis was more frequently negative in dMMR patients than pMMR patients (26% v 58.6%, P<0.001). Interestingly, CRC patients with dMMR had more regional lymph nodes removed during surgery (28.4 v 18.8, P<0.001), although with less metastatic cancer (1.0 v 2.3, P=0.001).
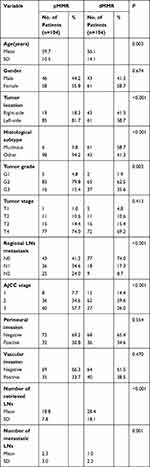 |
Table 1 Clinicopathological Characteristics of CRC Patients with Different MMR Gene Status |
Systemic Inflammatory Markers and Tumor Markers
By bioinformatics analysis, we found that unlike pMMR patients, patients with CRC with dMMR showed significant differences in gene expression in inflammatory and chemokine-related signaling pathways. Therefore, we investigated the differences in systemic inflammation markers and tumor markers between the two groups of patients. Various blood cells, including neutrophil, monocyte, lymphocyte, platelet, biochemical index including albumin, LDH, CRP, and tumor markers including CEA, CA125, CA199, AFP were collected retrospectively, then NLR, PLR, MLR, and GPS were calculated. As shown in Table 2, patients with resectable CRC with dMMR were more likely to have higher levels of neutrophil (4.37 v 3.57, P<0.001) (×109/L), monocyte (0.56 v 0.49, P=0.024) (×109/L) and platelet (287.15 v 226.65, P<0.001) (×109/L), but there were no differences of lymphocyte (P=0.764). In addition, the level of serum LDH was not different between CRC patients with dMMR or pMMR (P=0.526). Meanwhile, the level of serum CRP presented high levels in patients with dMMR (14.21 v 7.79, P=0.018) (mg/L), and the levels of albumin were lower in the dMMR patients (39.97 v 42.54, P=0.001) (g/L) than in pMMR patients. Similarly, there were significant differences between the two groups in other systemic inflammatory makers including NLR (2.8 v 2.28, P=0.005), PLR (186.94 v 143.48, P=0.002), MLR (0.36 v 0.31, P=0.022) and CAR (0.41 v 0.21, P=0.016). In addition, GPS was also different between the two groups of patients. In patients with dMMR, it was significantly higher than in patients with pMMR (P<0.001) (shown in Table 3). However, our study has shown that the tumor markers were not different between patients with dMMR and pMMR (P>0.05).
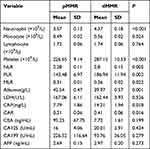 |
Table 2 Systemic Inflammation Markers of CRC Patients with Different MMR Gene Status |
 |
Table 3 GPS of Patients with Different Mismatch Repair Gene Status |
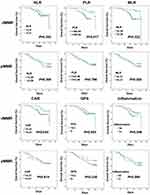
Impact of Inflammatory Markers on the Survival of Patients with dMMR and pMMR CRC
Survival analysis showed that patients with CRC with dMMR had a longer overall survival (OS) than those with pMMR (1746 days v 1523 days, P=0.047). Subsequently, to determine whether the inflammatory markers, including NLR, PLR, MLR, CAR, and GPS, have an impact on the survival of patients with CRC, we conducted the Kaplan-Meier analysis in our patients with different MMR statuses. We also explored the co-effect of the five inflammatory markers. We defined that patient with any inflammatory marker increased was high inflammation (HI), otherwise low inflammation (LI). Each inflammatory marker was divided into a high-level group and the low-level group according to the mean value, then the overall survival of the patients with different levels of inflammatory markers was compared in populations with different MMR states. OS curves are displayed in Figure 4. In patients with CRC with dMMR, higher levels of PLR (P=0.017), MLR (P=0.022), CAR (P=0.044), and co-effect (P=0.049) let to shorter OS significantly. In addition, OS was not significantly different for NLR and GPS in patients with dMMR CRC (P>0.05); however, survival was better with lower levels of these two inflammatory markers. It was noteworthy that the prognosis of high levels of systemic inflammatory markers did not predict long OS in patients with pMMR CRC (P>0.05).
Discussion
The data from KEYNOTE-1647 and KEYNOTE-1589 confirm that patients with solid cancer with MSI-H/dMMR can achieve an extended overall survival with pembrolizumab treatment, which is a PD-1 inhibitor. The result of the NICHE study suggested that neoadjuvant ipilimumab (targeting cytotoxic T lymphocyte antigen 4, CTLA4) plus nivolumab (targeting PD-1) in early-stage colon cancer had amazing treatment effects. Contrarily, most studies showed that patients with MSS/pMMR do not benefit from the treatment of PD-1 inhibitor, single drug, and combination with other therapy. We speculate that gene expression differences may cause the different responses of these two groups of patients to PD-1 inhibitor treatment. To prove our hypothesis, we found one gene chip contained in 16 patients with colorectal cancer with dMMR and 54 patients CRC with pMMR from the GEO database. Bioinformatics analysis has shown that compared to pMMR patients, gene changes in patients with CRC with dMMR were concentrated in chemokine-mediated signaling pathway and inflammatory response. Our result was consistent with data from Chaoran Yu’s analysis that chemokine receptor binding was top‑ranked in biological processes, cellular components, and molecular function terms between MSI-H and MSS CRC groups.10
As is well known, chemokines are small peptides with 8–12 kDa, classified into four families, namely C, CC, CXC, and CX3C, according to the number and spacing of cysteine residues within their N-terminal regions. CC and CXC families are the main components of chemokines, and they perform the cellular effects by binding to G-protein coupled cell-surface receptors. Several previous studies suggested that leukocytes including neutrophils and monocytes will express specific chemokine receptors, and will migrate along a gradient of homologous chemokine ligands to sites of infection or tissue damage.11,12 In patients with tumors, it was also found that some chemokines can mediate the migration of leukocytes to tumor sites, and participate in the initiation and development of tumor and anti-tumor effects.13 A recent study has shown that chemotherapy-resistant breast cancer cells can secrete high levels of chemokines, such as CXCR2 ligands, and can recruit vast numbers of neutrophils to the tumor site, which induce cancer-associated inflammation.14 Lymphocytes are essential cells in the process of tumor immunity. The present study proved that lymphocyte could be recruited through IL-6-regulated macrophage polarization and chemokine CC-chemokine-ligand-20/CC-chemokine-receptor-6.15 Many studies have shown that some immune cells including CD8+ tumor-infiltrating lymphocytes (TILs), T helper 1 (TH1) CD4+ TILs and macrophages are rich in tumors with MSI-H/dMMR compared to other tumors.16,17 Flow cytometry experiment demonstrated that the frequencies of cells expressing CCR2, CCR4, CXCR3, and CXCR6 differed significantly between CD4+ and CD8+ T lymphocytes, which was suggested that lymphocytes recruitment and transformation were controlled by potential chemokines and their receptors.18 Platelets are also considered as immune cells and affect all phases of immune responses.19 Platelets have direct contact and secret soluble chemokines, such as CD40L, CXCL4 and CCL5 to maintain homeostasis of the immune environment.20 Similarly, other chemokines such as CX3CL1/CX3CR1 and CCL2/CCR2 axes can affect the function and migration ability of platelets.21 According to the above analysis, the chemokine-mediated signaling pathway can regulate neutrophils, monocytes, lymphocytes, and platelets, respectively.
Due to the significant differences in chemokine-mediated signaling pathway and inflammatory response between patients with dMMR and pMMR CRC, it is reasonable to believe that there may be some differences in blood-cells related inflammatory markers between the two populations. Previous research described the systemic inflammation status and tumor immune cell infiltrates in patients with dMMR CRC.22,23 In this study, the levels of NLR, PLR, MLR were significantly elevated in patients with dMMR CRC. Some scholars found that inflammatory reaction markers such as GPS and NPS were significantly different between patients with MSI and MSS CRC.24,25 Consistently, our study also has shown that GPS, CRP, and CAR were elevated in patients with dMMR. Meanwhile, the tumor markers including CEA, CA125, CA199, AFP, and LDH were not different between the two populations. Since immunotherapy is closely related to inflammation response, we need more clinical and basic studies to confirm whether the increase of inflammatory markers in patients with dMMR CRC is associated with the benefit of the PD-1 inhibitor.
However, most studies reported that inflammatory markers could predict the prognosis of patients with tumors. In patients with resectable hepatocellular carcinoma (HCC), high NLR and PLR were independent risk factors predicting poor survival.26 In patients with CRC, low levels of preoperative lymphocyte-CRP ratio is an independent prognostic factor for both disease-free survival and overall survival.27 Patients with CRC with preoperative high NLR and mGPS have worse OS; however, only elevated levels of NLR are associated with frequent recurrences.28 A systematic review including 10,259 patients with CRC demonstrated that elevated preoperative NLR was associated with poor long-term survival in both patients with localized CRC and those with liver metastasis.29 However, there are few studies on the prognostic effects of inflammatory markers in patients with CRC with different MMR states. Kong et al found that patients with CRC with dMMR were more likely to present a high level of NLR and CAR. In addition, low levels of circulating blood lymphocytes (<1.12×109/L) predicted short OS and disease-free survival only in patients with dMMR CRC, although not in those with pMMR CRC.25 Our results are concordant with this finding, and we explored the prognostic role of additional systemic inflammatory markers in patients with CRC with different MMR statuses. In patients with dMMR CRC, the elevated levels of PLR, MLR, and CAR predicted short OS. Interestingly, there was no significant difference between the OS in patients with pMMR CRC and the different systemic inflammatory markers, and this might be due to the limited period of the follow-up. In other words, the prognostic effect of systemic inflammatory markers on survival was more significant in patients with dMMR CRC patients than those with pMMR. About 20 years ago, some scholars noted that there were leukocytes in tumor tissues.30 Correlation between inflammation and tumorigenesis has been gradually identified.31 Inflammation plays a critical role in all phases of tumor development, not only the acceleration of tumor cells’ proliferation and metastasis but also resistance to cytotoxic drugs.32,33 Thus, we hypothesized that there was a causal relationship between inflammation and tumor prognosis in different genetic settings.
Conclusion
There were differences in gene expression between patients with dMMR CRC and pMMR, mainly focusing on the chemokine-mediated signaling pathway and inflammatory response. Clinical data analysis suggested that compared to patients with pMMR, those with dMMR CRC had elevated systemic inflammatory markers including NLR, PLR, MLR, CRP, and CAR. Interestingly, survival analysis has shown that the prognostic effect of inflammatory markers on survival was more pronounced in patients with dMMR CRC. However, due to potential limitations in this study, bias was inevitable First, this was a single-center retrospective trial. In addition, the sample size was relatively small, and the follow-up time was short. Further mechanistic assessments are required to determine whether the chemokine-mediated signaling pathway or the increase of inflammatory markers in patients with dMMR CRC is associated with the benefit of the PD-1 inhibitor.
Abbreviations
ICBM, Institute of Cancer and Basic Medicine; LNs, lymph node; CRC, Colorectal cancer; CIN, chromosomal instability; MSI, microsatellite instability; HNPCC, hereditary nonpolyposis colorectal cancer; SCRC, sporadic colorectal cancer; MMR, Mismatched repair; PD-1, programmed cell death 1; NCBI, National Center for Biotechnology Information; GO, gene ontology; AJCC, American Joint Committee on Cancer; CRP, C-reactive protein; LDH, lactic dehydrogenase; IHC, immunohistochemical; DAB, diaminobenzidine; DEG, differential expression genes; OS, overall survival; HI, high inflammation; LI, low inflammation.
Ethics Approval and Consent to Participate
The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of IRB-2020-239(Ke).
Acknowledgments
We are grateful to all clinical investigators and the included patients.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This study was funded by the Project Supported by Zhejiang Provincial Natural Science Foundation of China (LQ20H160005).
Disclosure
The authors report no potential conflicts of interest.
References
1. Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
3. Boland CR, Thibodeau SN, Hamilton SR, et al. A National Cancer Institute Workshop on microsatellite instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res. 1998;58(22):5248–5257.
4. Sameer AS, Nissar S, Fatima K. Mismatch repair pathway: molecules, functions, and role in colorectal carcinogenesis. Eur J Cancer Prev. 2014;23(4):246–257. doi:10.1097/CEJ.0000000000000019
5. Pawlik TM, Raut CP, Rodriguez-Bigas MA. Colorectal carcinogenesis: MSI-H versus MSI-L. Dis Markers. 2004;20(4–5):199–206. doi:10.1155/2004/368680
6. Sargent DJ, Marsoni S, Monges G, et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J Clin Oncol. 2010;28(20):3219–3226. doi:10.1200/JCO.2009.27.1825
7. Le DT, Kim TW, Van Cutsem E, et al. Phase II open-label study of pembrolizumab in treatment-refractory, microsatellite instability-high/mismatch repair-deficient metastatic colorectal cancer: KEYNOTE-164. J Clin Oncol. 2020;38(1):11–19. doi:10.1200/JCO.19.02107
8. Li J, Xu Q, Luo C, et al. Clinicopathologic characteristics of resectable colorectal cancer with mismatch repair protein defects in Chinese population: retrospective case series and literature review. Medicine (Baltimore). 2020;99(24):e20554. doi:10.1097/MD.0000000000020554
9. Marabelle A, Le DT, Ascierto PA, et al. Efficacy of pembrolizumab in patients with noncolorectal high microsatellite instability/mismatch repair-deficient cancer: results from the phase II KEYNOTE-158 Study. J Clin Oncol. 2020;38(1):1–10. doi:10.1200/JCO.19.02105
10. Yu C, Hong H, Zhang S, et al. Identification of key genes and pathways involved in microsatellite instability in colorectal cancer. Mol Med Rep. 2019;19(3):2065–2076. doi:10.3892/mmr.2019.9849
11. Stadtmann A, Block H, Volmering S, et al. Cross-talk between Shp1 and PIPKIgamma controls leukocyte recruitment. J Immunol. 2015;195(3):1152–1161. doi:10.4049/jimmunol.1500606
12. Scanzello CR. Chemokines and inflammation in osteoarthritis: insights from patients and animal models. J Orthop Res. 2017;35(4):735–739. doi:10.1002/jor.23471
13. Hurt B, Schulick R, Edil B, et al. Cancer-promoting mechanisms of tumor-associated neutrophils. Am J Surg. 2017;214(5):938–944. doi:10.1016/j.amjsurg.2017.08.003
14. Wu L, Awaji M, Saxena S, et al. IL-17–CXC chemokine receptor 2 axis facilitates breast cancer progression by up-regulating neutrophil recruitment. Am J Pathol. 2020;190(1):222–233. doi:10.1016/j.ajpath.2019.09.016
15. Wunderlich CM, Ackermann PJ, Ostermann AL, et al. Obesity exacerbates colitis-associated cancer via IL-6-regulated macrophage polarisation and CCL-20/CCR-6-mediated lymphocyte recruitment. Nat Commun. 2018;9(1):1646. doi:10.1038/s41467-018-03773-0
16. Llosa NJ, Cruise M, Tam A, et al. The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov. 2015;5(1):43–51. doi:10.1158/2159-8290.CD-14-0863
17. Gajewski TF, Schreiber H, Fu YX. Innate and adaptive immune cells in the tumor microenvironment. Nat Immunol. 2013;14(10):1014–1022. doi:10.1038/ni.2703
18. Lofroos AB, Kadivar M, Resic Lindehammer S, et al. Colorectal cancer-infiltrating T lymphocytes display a distinct chemokine receptor expression profile. Eur J Med Res. 2017;22(1):40. doi:10.1186/s40001-017-0283-8
19. Morrell CN, Pariser DN, Hilt ZT, et al. The platelet napoleon complex—small cells, but big immune regulatory functions. Annu Rev Immunol. 2019;37(1):125–144. doi:10.1146/annurev-immunol-042718-041607
20. Ribeiro LS, Migliari Branco L, Franklin BS. Regulation of innate immune responses by platelets. Front Immunol. 2019;10:1320.
21. Marques P, Collado A, Martinez-Hervas S, et al. Systemic inflammation in metabolic syndrome: increased platelet and leukocyte activation, and key role of CX3CL1/CX3CR1 and CCL2/CCR2 axes in arterial platelet-proinflammatory monocyte adhesion. J Clin Med. 2019;8(5):708. doi:10.3390/jcm8050708
22. De Smedt L, Lemahieu J, Palmans S, et al. Microsatellite instable vs stable colon carcinomas: analysis of tumour heterogeneity, inflammation and angiogenesis. Br J Cancer. 2015;113(3):500–509. doi:10.1038/bjc.2015.213
23. Pine JK, Morris E, Hutchins GG, et al. Systemic neutrophil-to-lymphocyte ratio in colorectal cancer: the relationship to patient survival, tumour biology and local lymphocytic response to tumour. Br J Cancer. 2015;113(2):204–211. doi:10.1038/bjc.2015.87
24. Park JH, Powell AG, Roxburgh CSD, et al. Mismatch repair status in patients with primary operable colorectal cancer: associations with the local and systemic tumour environment. Br J Cancer. 2016;114(5):562–570. doi:10.1038/bjc.2016.17
25. Kong P, Wang J, Song Z, et al. Circulating lymphocytes, PD-L1 expression on tumor-infiltrating lymphocytes, and survival of colorectal cancer patients with different mismatch repair gene status. J Cancer. 2019;10(7):1745–1754. doi:10.7150/jca.25187
26. Wang D, Bai N, Hu X, et al. Preoperative inflammatory markers of NLR and PLR as indicators of poor prognosis in resectable HCC. PeerJ. 2019;7:e7132. doi:10.7717/peerj.7132
27. Okugawa Y, Toiyama Y, Yamamoto A, et al. Lymphocyte-C-reactive protein ratio as promising new marker for predicting surgical and oncological outcomes in colorectal cancer. Ann Surg. 2020;272(2):342–351. doi:10.1097/SLA.0000000000003239
28. Climent M, Ryan ÉJ, Stakelum Á, et al. Systemic inflammatory response predicts oncological outcomes in patients undergoing elective surgery for mismatch repair-deficient colorectal cancer. Int J Colorectal Dis. 2019;34(6):1069–1078. doi:10.1007/s00384-019-03274-6
29. Haram A, Boland MR, Kelly ME, et al. The prognostic value of neutrophil-to-lymphocyte ratio in colorectal cancer: a systematic review. J Surg Oncol. 2017;115(4):470–479. doi:10.1002/jso.24523
30. Balkwill F, Mantovani A. Inflammation and cancer: back to Virchow? Lancet. 2001;357(9255):539–545. doi:10.1016/S0140-6736(00)04046-0
31. Grivennikov SI, Greten FR, Karin M. Immunity, inflammation, and cancer. Cell. 2010;140(6):883–899. doi:10.1016/j.cell.2010.01.025
32. Keller DS, Windsor A, Cohen R, et al. Colorectal cancer in inflammatory bowel disease: review of the evidence. Tech Coloproctol. 2019;23(1):3–13. doi:10.1007/s10151-019-1926-2
33. Mirzaei SA, Dinmohammadi F, Alizadeh A, et al. Inflammatory pathway interactions and cancer multidrug resistance regulation. Life Sci. 2019;235:116825
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.