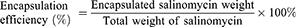Back to Journals » International Journal of Nanomedicine » Volume 16
Synthesis and Characterization of Salinomycin-Loaded High-Density Lipoprotein and Its Effects on Cervical Cancer Cells and Cervical Cancer Stem Cells
Authors Yin X, Lu Y, Zou M, Wang L, Zhou X, Zhang Y, Su M
Received 23 June 2021
Accepted for publication 3 September 2021
Published 17 September 2021 Volume 2021:16 Pages 6367—6382
DOI https://doi.org/10.2147/IJN.S326089
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Mian Wang
Xirui Yin,1 Yuhui Lu,1 Miao Zou,1 Liuli Wang,1 Xuan Zhou,1 Yingyu Zhang,2 Manman Su1
1Department of Regenerative Medicine, School of Pharmaceutical Sciences, Jilin University, Changchun, People’s Republic of China; 2Department of Medical Science, Chang Chun Medical College, Changchun, People’s Republic of China
Correspondence: Manman Su
Department of Regenerative Medicine, School of Pharmaceutical Sciences, Jilin University, 1266 Fujin Road, Changchun, 130021, People’s Republic of China
Tel +86 13843037881
Fax +86 431 85619252
Email [email protected]
Background: Cervical cancer stem cells (CCSCs), a small part of tumor population, are one of the important reasons for metastasis and recurrence of cervical cancer. Targeting CCSCs may be an effective way to eliminate tumors. Salinomycin (Sal) has been proved to be an effective anticancer drug in many studies, especially for cancer stem cells (CSCs). However, the cytotoxicity of salinomycin limits its further research as an anticancer drug. High-density lipoprotein (HDL) nanoparticles are an excellent drug carrier, which can reduce the toxicity of Sal, have a certain targeting effect and improve the clinical benefit of Sal.
Methods: Salinomycin-loaded high-density lipoprotein (S-HDL) was synthesized and characterized by various analytical techniques. CD44highCD24low CCSCs were isolated from HeLa cells by magnetic separation. The uptake of HDL nanoparticles was observed by laser confocal microscopy, and the effect of S-HDL on the proliferation of CCCs and CCSCs was detected by cell viability analysis. Genome-wide analysis was used to analyze the effects of S-HDL on the biological processes of CCCs and then cell apoptosis, cell cycle and cell migration were selected for verification.
Results: S-HDL had a particle size of 38.98 ± 1.78 nm and an encapsulation efficiency of 50.73 ± 4.29%. Cell uptake analysis showed that HDL nanoparticles could enhance the drug uptake of CCCs and CCSCs and may target CCCs and CCSCs. In cell viability analysis, CCCs and CCSCs showed high sensitivity to S-HDL. S-HDL can more efficiently prevent CCSCs from developing tumorspheres than Sal in tumorsphere formation study. S-HDL had stronger ability to induce cell cycle arrest, promote cell apoptosis and inhibit cell migration compared with free Sal, which was consistent with the results of Genome Wide analysis.
Conclusion: S-HDL can effectively target and eliminate CCCs and CCSCs, which is a potential drug for the treatment of cervical cancer.
Keywords: HDL, salinomycin, cervical cancer stem cells, cellular uptake, LPR1
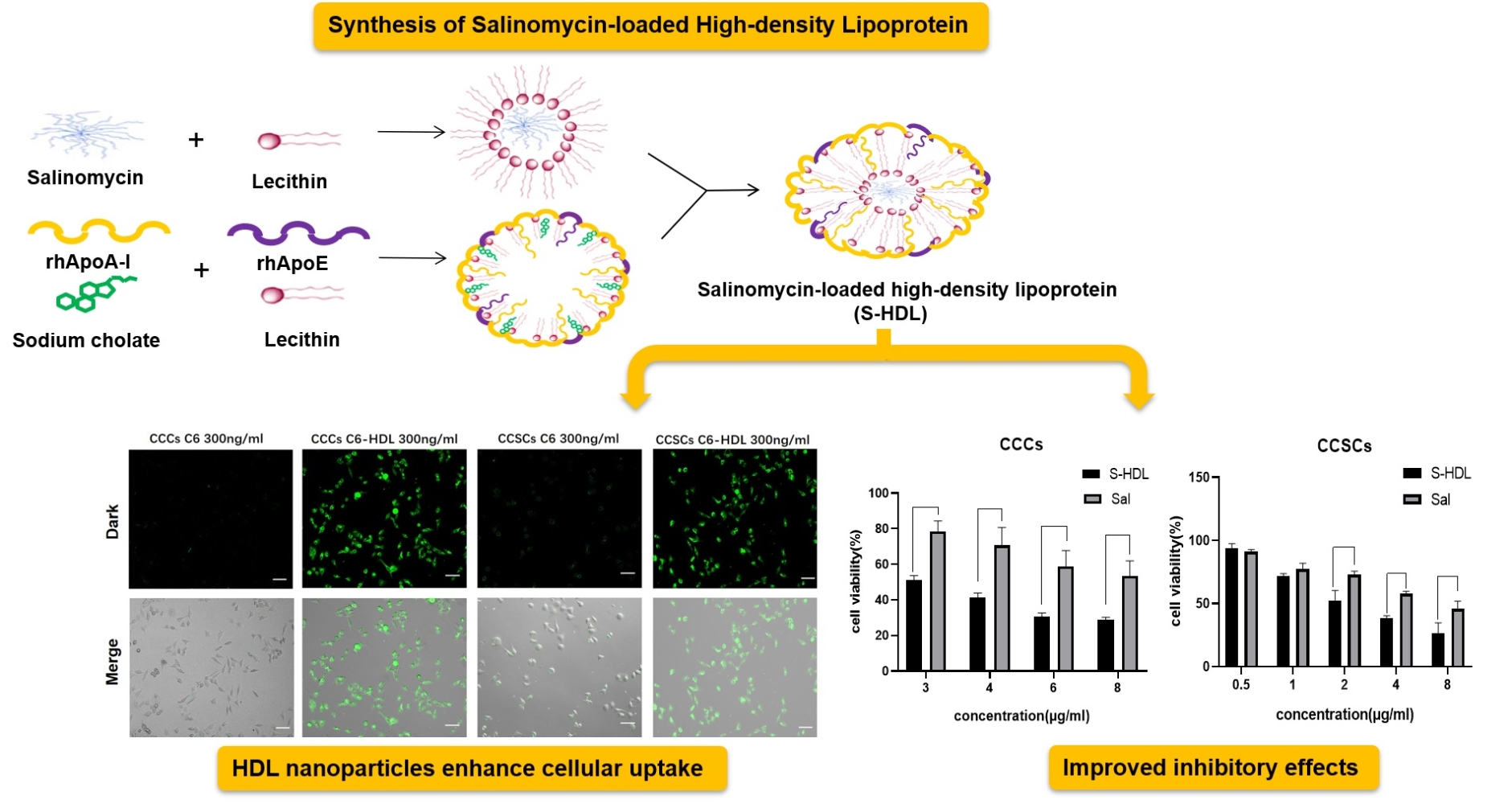
Graphical Abstract:
Introduction
Cervical cancer is a kind of gynecological malignant tumor, which has been a serious threat to the health and life of women all over the world for many years. Although great efforts have been made to develop more effective treatment methods in recent years, clinical trials show that the therapeutic effect is limited due to the metastasis and recurrence of cervical cancer.1 Cervical cancer stem cells (CCSCs), a small part of cervical cancer cells (CCCs), are resistant to conventional cancer treatments, such as chemotherapy and radiotherapy. Residual CCSCs may cause recurrence after treatment.2,3 Therefore, the development of treatment strategies based on CCSCs has become a key goal to achieve the challenge of radical eradication of cervical cancer, especially for patients with metastatic cervical cancer.
Salinomycin (Sal) is a polyether K+ selective membrane ionophore antibiotic of Streptomyces albus.4 Gupta et al found that Sal had an obvious inhibitory effect on breast cancer stem cells compared with other compounds in 2009.5 Since then, salinomycin began to be studied as an anticancer drug, which was proved to have significant inhibitory effect on pancreatic cancer, lung cancer, colorectal cancer and other cancer stem cells.6–8 It can eliminate cancer stem cells (CSCs) by promoting the differentiation and sensitivity to radiotherapy and chemotherapy of CSCs.9,10 However, the cytotoxicity and hydrophobicity of salinomycin limit its further research as an anticancer drug.11,12 Nanocarrier encapsulation is a good strategy to improve anticancer drugs. It can increase the solubility of anticancer drugs and increase the drug accumulation in tumor by enhancing the permeability and retention effect of drugs, so as to reduce the toxicity and side effects of drugs.13,14 Some studies have shown that using nano carriers to encapsulate salinomycin can effectively improve its anti-cervical cancer effects and reduce its toxic and side effects. Compared with free Sal, Sal-loaded polyethylene glycol-peptide-polycaprolactone nanoparticles showed stronger inhibition of cervical cancer, lower side effects, and had a killing effect on CCSCs.15 Combined delivery of salinomycin and docetaxel by dual-targeting gelatinase nanoparticles can enhance anti-cervical cancer efficacy and reduce side effects by simultaneously suppressing CCCs and CCSC.16
High-density lipoprotein (HDL) is a kind of lipoproteins that carry lipids as multifunctional aggregates in plasma. HDL can be used as an effective carrier for many drugs because its phospholipid core can be combined with hydrophobic drugs. Nanoparticle drug carriers based on HDL can be constructed in vitro by using a combination of the main components of HDL, such as Apolipoprotein E (ApoE), Apolipoprotein A-I (ApoA-I), phospholipid and cholesterol.17 The cells with natural receptors of HDL or its components can be targeted by them.18 ApoE, a component of HDL, is a ligand for low-density lipoprotein receptor-related proteins 1 (LRP1).19,20 ApoE can target LRP1 and be internalized through LRP1 mediated endocytosis in hepatoma cells.21 LRP1 is expressed in some malignant tumors, and its amount of expression is related to the degree of malignancy. Some studies have shown that LRP1 plays a role in regulating the invasion and metastasis of thyroid cancer and breast cancer cells.22–24 In addition, Catasus found that increased LRP1 expression predicted more aggressive tumor behavior and was associated with increased histological grade of endometrial carcinoma.25 Therefore, nanocarrier with ApoE may be able to target the malignant tumors with highly expressed LRP1. ApoA-I is the main protein component of HDL, which is widely used to regulate cholesterol transport and prevent cardiovascular disease, and may also regulate inflammation and immune response.26 In recent years, many studies have reported the antitumor effect of ApoA-I. Recombinant ApoA-I can promote cell cycle arrest and apoptosis of hepatoma cells and reduce the ability of angiogenesis and extracellular matrix remodeling of hepatoma cells, thereby inhibiting their metastatic potential.27 In animal experiments, the application of ApoA-I mimic peptide or overexpression of ApoA-I can reduce the growth of primary tumor and the metastasis of colon cancer cells to the lung.28,29
With a view to synthesizing an anticancer drug that is effective for both CCCs and CCSCs, and reduce the toxicity of normal tissues, we developed a salinomycin-loaded high-density lipoprotein (S-HDL). The shell of S-HDL nanoparticles consists of recombinant human ApoA-I (rh ApoA-I) and recombinant human ApoE (rh ApoE), and Sal is encapsulated in its hydrophobic core. We isolated CCSCs from CCCs and then verified the efficacy of S-HDL on CCCs and CCSCs in vitro by cellular uptake analysis, cell viability assay and tumorsphere formation study. Furthermore, the biological process of S-HDL affecting cervical cancer cells was screened by genome-wide mRNA expression and then verified, which will let us understand the way S-HDL works better.
Materials and Methods
Materials
Lecithin and Coumarin 6 were purchased from Sigma-Aldrich (USA). RhApoA-I and rhApoE were prepared and preserved in our laboratory. Salinomycin was purchased from Selleck (China). Vanillin was purchased from J&K scientific (China). Basic fibroblast growth factor (b-FGF) and epidermal growth factor (EGF) were bought from Peprotech (USA). B27 Supplement, DMEM medium and DMEM/F12 medium were purchased from Gibco (USA). Antihuman CD44 microbeads antibody, antihuman CD24-biotin antibody, anti-biotin microbeads, PE-conjugated antihuman CD44 antibody and FITC-conjugated antihuman CD24 antibody were purchased from Miltenyi Biotec (Germany). Antibodies against C-myc and Sox-2 were purchased from Cell Signaling Technology (USA). Antibody against LRP1 was purchased from Abcam (USA). Horseradish Peroxidase-conjugated secondary antibody, Cell Counting Kit-8 and Cell Cycle Analysis Kit were purchased from Beyotime Institute of Biotechnology (China). Annexin V-FITC/PI apoptosis detection kit was purchased from BD company (USA).
Salinomycin-Loaded High-Density Lipoprotein (S-HDL) Synthesis and Characterization
S-HDL was synthesized using sodium cholate methods.30 An ethanol solution (1 mL) containing 2.7 mg lecithin and 1 mg salinomycin was rapidly injected into 6 mL phosphate buffer (pH = 7.4) via a skin test syringe. After mixing in N2 for 15 min, phosphate buffer (0.75 mL) containing 2.7 mg sodium cholate, 5 mg rhApoA- I and 2.5 mg rhApoE was added to the lipid mixture by stirring. The mixture was incubated at room temperature for 30 min and then was incubated at 4°C for 12 h, and the components polymerized to form S-HDL. Then, the solution was mixed and added to a 10KD dialysis bag for complete dialysis with a phosphate buffer at 4°C (about 2 days) to remove ethanol and sodium cholate. After dialysis, the solution was filtered by 0.22 μm filter membrane and stored at 4°C. After the synthesis of S-HDL, it was characterized using a Zetasizer Nano ZS dynamic light scattering (DLS) and a transmission electron microscope (TEM).
Encapsulation Efficiency of S-HDL
Salinomycin encapsulation efficiency was analyzed by measuring with a UV–vis–NIR spectrophotometer following the published method.31 Briefly, S-HDL was mixed with acetonitrile in the ratio of 1:4 and vortex to demulsify. The supernatant was derivatized with vanillin in an acidic medium at 72°C for 40 min. The derivative mixture was determined by UV–vis–NIR spectrophotometer at a wavelength of 528 nm. For the following experiments, the concentration of S-HDL was calculated by the content of salinomycin in it. The encapsulation efficiency was calculated as follows:
Cell Culture and CCSCs Isolation
HeLa cells and human ovarian epithelial cells IOSE80 were purchased from the cell bank of Chinese Academy of Sciences and cultured in Dulbecco modified Eagle medium (DMEM) at 37°C with 10% fetal bovine serum in 5% CO2 humidified atmosphere. CCSCs sub-populations in the HeLa cells were isolated using Auto MACS Pro Separator according to protocols from the manufacturer. In brief, HeLa cells (1 x 107cells/mL) were prepared and incubated with antihuman CD44 microbeads antibody for 15 min and then using auto Macs Pro separator for selection of CD44high cells. Those cells were then mixed with antihuman CD24-biotin antibody and incubated for 15 min. After centrifugation, the cells were resuspended with buffer, and then incubated with anti-biotin microbeads for 15 min and then CD24low cells were selected by auto Macs Pro separator. After washing, CD44highCD24low cells were sorted by magnetic separation.
Identification of CCSCs by Flow Cytometry and Western Blot
HeLa cervical cancer cells (CCCs) and CCSCs were mixed with PE-conjugated antihuman CD44 antibody and FITC-conjugated antihuman CD24 antibody, respectively, and incubated for 15 min at 4°C. After washing those cells twice, the expression of CD44 and CD24 on the surface of CCCs and CCSCs was detected by flow cytometry.
Expressions of Sox-2, C-myc in CCCs and CCSCs were measured by Western blotting. Total cellular proteins of CCCs, CCSCs and IOSE80 cells were extracted by ice-cold lysis buffer and quantified using the bicinchoninic acid protein assay. Cell lysate was mixed with an appropriate amount of SDS sample loading buffer (5x) and boiled at 100°C. 45 μg of protein samples were loaded to sodium dodecyl sulfate-polyacrylamide gel and then were transferred to PVDF membrane and further blocked by 10% non-fat milk. Proteins were incubated with primary antibody (β-actin, C-myc, Sox-2) at a 1:1000 dilution for 4°C overnight, and then were incubated with Horseradish Peroxidase-conjugated secondary antibody (1:2000 dilution) for 1 h at room temperature and washed three times with TBST. Protein bands were visualized using Immobilon Western Chemiluminescent HRP Substrate.
Expression of LRP1 and Cellular Uptake of Coumarin-6-Loaded High-Density Lipoprotein
Expressions of LRP1 in CCCs and CCSCs were measured by Western blotting as mentioned above.
Coumarin-6-loaded high-density lipoprotein (C6-HDL) was synthesized using coumarin 6 (C6) as fluorescent probe. The synthetic method of C6-HDL is similar to S-HDL synthesis except that C6 replaces salinomycin. The concentration of C6-HDL was determined by fluorescence microplate reader and the concentration of C6-HDL was calculated by the C6 content inside it. CCCs, CCSCs and IOSE80 cells were cultured in laser confocal 24-well plate for 24 h; then, CCCs and CCSC were cultured with 300ng/mL C6 or 300ng/mL C6-HDL, respectively, IOSE80 cells were cultured with 300ng/mL C6-HDL. After 2 hours of culture, the cellular uptake was observed by laser confocal microscope (FITC, NIKON C2), and the fluorescence intensity was measured by ImageJ.
Cell Viability Assay
The CCCs and CCSCs were seeded in 96-well plates (1x104cells/well) for 24 h. A series of different concentrations of S-HDL or Sal were added to the cells, respectively, and the cells were cultured for 48 h. The control group was treated with the same amount of phosphate buffer saline (PBS) as S-HDL or Sal. The cell viability was determined by Cell Counting Kit-8 (CCK8) assay. The intensity of the color formed by this method is proportional to the number of living cells. The CCK8 solution was added to the plates at 37°C. The optical density (OD) was measured at 450 nm. The mean values and standard deviations were calculated from three experiments. 50% inhibiting concentration (IC50) was calculated by GraphPad Prism 8.
Tumorsphere Formation
CCSCs were treated with a series of concentrations of S-HDL for 48 h. DMEM/F12 medium containing 20ng/mL EGF, 10ng/mL b-FGF and 2% B27 supplement was used to resuspend CCSCs, and then the cells were seeded into 96-well ultralow attachment plates at density of 100 cells/well. The number of tumorsphere was recorded under the microscope after 7 days of culture in a humidified atmosphere of 5% CO2 at 37°C. Tumor spheres larger than 50μm in diameter were counted as positive. The tumorsphere formation efficiency was calculated by dividing the number of tumor spheres by the initial number of single cells.
Analysis of Genome-Wide mRNA by L1000 Technology
We used high-throughput L1000 technology to estimate genome-wide mRNA expression of CCCs after S-HDL treatment. CCCs treated with 8μg/mL S-HDL for 48 h were collected and then lysed with TCL Lysis Buffer. Cell lysate was added to 384-well plate and then using ligation mediated amplification of RNA sequence-specific probes combined with Luminex-based detection to generate expression profiles of 978 genes per sample. Finally, the 978 genes were used as landmark genes to calculate the expression level of the remaining genes. t-test was used to calculate the gene expression of the experimental group compared with the control group and the positive control group (8 μg/mL Sal treatment), respectively, and the differential genes were obtained. The screening threshold was set as p-value <0.05, Foldchange >1.5. The differential genes were enriched by GO database (http://geneontology.org/). Then, 20 biological processes with the lowest p-value were selected for the following analysis. We selected cell apoptosis, cell cycle and cell migration for subsequent experimental verification based on the results of GO analysis.
Apoptosis Assay
The apoptosis rate was detected by Annexin V-FITC/PI apoptosis detection kit. CCCs or CCSCs were treated with different concentrations of S-HDL for 48 h. The cells were collected and resuspended in 500μL binding buffer. Then, 3 μL Annexin V-FITC and 3 μL PI were added to the buffer and incubated for 15 minutes in the dark at room temperature. The cells were analyzed by flow cytometry (BD FACSCanto II).
Cell-Cycle Analysis
We used Cell Cycle Analysis Kit to detect the distribution of cell cycle of CCCs and CCSCs. CCCs or CCSCs were treated with 8 μg/mL S-HDL for 48 h. The cells were collected and fixed with 70% ethanol at 4°C for 12 hours, then stained with propidium iodide solution containing RNase A for 30 minutes. We detected DNA content by flow cytometry (BD FACSCanto II) and analyzed cell cycle profile using cell quest software.
Cell Migration Assay
Cell migration was analyzed using xCELLigence Real Time Cell Analyzer (RTCA) DP instrument (ACEA Biosciences). 165 μL DMEM medium supplemented with 20% FBS was added to the lower chamber of cell migration plate (CIM-plate 16) of RTCA DP instrument, and 50μL DMEM without serum was added to the upper chamber. The CIM-plate was placed on RTCA DP instrument to detect the baseline after being cultured at 37°C and 5% CO2 for 1 h. CCCs or CCSCs with a density of 5×105 cells/mL were collected after being treated with S-HDL for 48 h. 100 μL cell suspension was added to the upper chamber of the CIM-plate. After being stored at room temperature for 30 min, the cells were detected on RTCA DP analyzer every 15 min for 70 hours.
Statistical Analysis
Intergroup t-test comparisons were performed using the mean ± standard deviation (x ± s), p-values <0.05 denoted statistically significant differences.
Results
S-HDL Preparation and Salinomycin Encapsulation
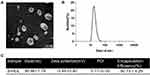
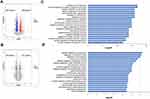
Salinomycin-loaded high-density lipoprotein was synthesized by sodium cholate method for the first time.30 S-HDL was observed by transmission electron microscope, which showed a uniform discoidal shape (Figure 1A). The particle size of S-HDL nanoparticles measured by DLS is between 10nm-50nm, and the peak value is between 20–25 nm (Figure 1B). The size distribution of S-HDL makes it possible to stay in tumor tissues for a longer time by enhanced permeability and retention effect.32,33 The particle size, zeta potential, and polydispersity index (PDI) of S-HDL were then measured using DLS (Figure 1C). The average particle size of S-HDL was 38.98 ± 1.78nm, zeta potential was - 3.44 ± 0.90 mV, PDI was 0.17 ± 0.05 (<0.2), which indicated that the dispersion system of S-HDL was stable. The encapsulation efficiency of salinomycin was 50.73 ± 4.29%, which was determined by UV spectrophotometer (Figure 1C).
Isolation and Identification of CCSCs
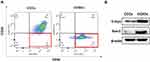
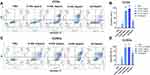
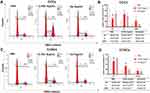
CSCs were isolated and identified by surface markers in vitro. It has been proved that CD44highCD24low cervical cancer cells have the characteristics of CSCs.34–36 CCSCs with CD44highCD24low were separated from CCCs (HeLa cells) by a magnetic separator. The proportion of CD44highCD24low cells in CCSCs was 97.2%, which was much higher than 1.52% in CCCs, indicating that the CD44highCD24low subpopulation has been successfully enriched from CCCS (Figure 2A). In order to further detect whether CD44highCD24low CCCs possess the characteristics of cancer stem cells, we analyzed the protein expression of cancer stem cell markers C-myc and Sox-2 in CD44highCD24low CCCs by Western blotting. The expression of C-myc and Sox-2 in CCSCs was significantly higher than that in CCCs (Figure 2B). It was proved that the CD44highCD24low CCSCs have the characteristics of cancer stem cells.
Expression of LRP1 and Cellular Uptake of HDL Nanoparticles
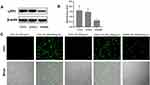
Low-density lipoprotein receptor-related protein 1 (LRP1) is an important member of the low-density lipoprotein receptor gene family.37 We analyzed the expression of LRP1 in CCCs and CCSCs compared with human ovarian epithelial cells IOSE80, in which the expression of LRP1 in CCCs and CCSCs was 2.5 and 2.3 times higher than that in IOSE80, respectively (Figure 3A and B). LRP1 is a natural receptor of ApoE, a component of HDL, which may trigger endocytosis.19 Therefore, the high expression of LRP1 is easier to be targeted by HDL nanoparticles, thereby increasing the cellular uptake of HDL encapsulated drugs and reducing the toxicity to normal tissues.
The uptake of C6 and C6-HDL by CCCs, CCSCs and IOSE80 is shown in Figure 3C. HDL nanoparticles enhanced the cellular uptake of C6. The proportion of C6-HDL uptake by CCCs and CCSCs was higher than that of C6, and the fluorescence intensity of C6-HDL uptake by CCCs and CCSCs was stronger than that of C6. The uptake of C6-HDL by different cells was different. The fluorescence intensity of C6-HDL by IOSE80 was weaker than that of CCCs and CCSCs. The results indicated that C6-HDL has a potential targeting effect on CCCs and CCSCs, which may be related to the expression of LRP1.
Inhibitory Effects of S-HDL on CCCs and CCSCs Proliferation
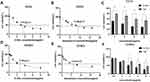
In order to evaluate the effects of S-HDL on the proliferation of CCCs and CCSCs, we used CCK-8 analysis to detect the cell viability of CCCs and CCSCs after S-HDL treatment, and used GraphPad Prism 8 software to calculate the IC50 of S-HDL on CCCs and CCSCs. The IC50 of S-HDL on CCCs was 3.43μ/mL, which was significantly lower than that of Sal on CCCs (9.18μg/mL) (Figure 4A and B). At the same concentration, the cell viability of CCCs treated with 3μg/mL, 4μg/mL, 6μg/mL and 8μg/mL S-HDL was significantly lower than that treated with Sal (Figure 4C). The IC50 of S-HDL on CCSCs was 2.70μg/mL, which was significantly lower than that of Sal on CCSCs (6.12μg/mL) (Figure 4D and E). At the same concentration, the cell viability of CCSCs treated with 2μg/mL, 4μg/mL and 8μg/mL S-HDL was significantly lower than that treated with Sal (Figure 4F). Therefore, our findings suggested that using HDL encapsulated salinomycin can reduce the IC50 on CCCs and CCSCs, and significantly enhance the inhibitory effects on the proliferation of CCCs and CCSCs.
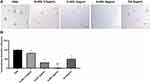
S-HDL Inhibits Tumorsphere Formation
The tumorsphere formation ability is regarded as reflecting the self-renewal characteristics of CSCs in vitro and is used to screen anti-CSCs drugs.5,38 The tumorsphere formation of CCSCs treated with S-HDL and Sal was imaged and quantified in Figure 5. The tumorsphere formation efficiency of S-HDL decreased in a concentration-dependent manner, which was significantly lower than that of PBS. Meanwhile, the tumorsphere formation efficiency of 8μg/mL S-HDL was significantly lower than that of 8μg/mL Sal (Figure 5B). Thus, our data demonstrated that S-HDL can efficiently prevent CCSCs from developing tumorspheres and has a stronger inhibitory effect on CCSCs regeneration in vitro compared with Sal.
Genome-Wide mRNA Expression After Treatment with S-HDL
To understand the biological processes by which S-HDL affects CCCs and CCSCs, we carried out Genome-wide mRNA analysis using RNA extracts from CCCs after treatment with S-HDL for 48 h. S-HDL treatment resulted in up-regulation of 496 genes and down-regulation of 667 genes compared with the control group, meanwhile up-regulation of 101 genes and down-regulation of 138 genes compared with Sal group (Figure 6A and B). GO enrichment analysis revealed that S-HDL treatment altered the expression of genes that are involved in many biological processes, such as cell migration, regulation of cell adhesion, apoptosis pathway and cell death compared with control group (Figure 6C). Compared with Sal, S-HDL treatment also altered the expression of genes that are involved in many biological processes, such as proteolysis, cell cycle, cell adhesion, DNA metabolic process and apoptosis pathway (Figure 6D).
Cervical cancer cells promote their own proliferation by down regulating apoptosis and activating cell cycle, which is closely related to the occurrence and development of tumor.39 Migration of cervical cancer cells from the original tissue to surrounding or distant organs is an essential process of tumor development.40 Therefore, the biological processes of cell cycle, cell apoptosis and cell migration are the key to the occurrence and development of cervical cancer. Therefore, we explored the effects of S-HDL on cell apoptosis, cell cycle and cell migration of CCCs and CCSCs in the following experiments according to the results of Genome-wide mRNA expression.
S-HDL Promotes Apoptosis of CCCs and CCSCs
Annexin V-FITC/PI apoptosis detection kit was used to detect the apoptosis rate of CCCs and CCSCs treated with different concentrations of S-HDL. S-HDL significantly promoted the apoptosis of CCCs and CCSCs in a concentration-dependent manner. The apoptosis rates of CCCs treated with 4μg/mL, 8μg/mL S-HDL and the apoptosis rates of CCSCs treated with 2μg/mL, 8μg/mL S-HDL were significantly higher than those treated with 8μg/mL Sal (Figure 7). Interestingly, the apoptosis rate of CCSCs treated with 0.5μg/mL S-HDL (38.34±2.16%) was higher than that of CCCs treated with 2μg/mL S-HDL (21.63±2.15%), which was consistent with the result that the IC50 of S-HDL on CCSCs (2.70μg/mL) was lower than that on CCCs (3.43μg/mL) in cell proliferation, suggesting that S-HDL could exert effect on CCSCs at a lower concentration.
S-HDL Induces S-Phase and G2/M-Phase Arrest in CCCs and CCSCs
In order to investigate the effects of S-HDL on the cell cycle progression of CCCs and CCSCs, the DNA content of the cells treated with S-HDL was determined by flow cytometry. Compared with PBS-treated cells, S-HDL-treated CCCs and CCSCs showed increased S-phase and G2/M-phase subpopulations and decreased G1-phase subpopulation (Figure 8). Sal-treated CCCs showed an increased S-phase subpopulation, a decreased G1-phase subpopulation, but no significant changes in G2/M-phase subpopulation and Sal-treated CCSCs showed an increased G2/M-phase subpopulation but no significant changes in G1-phase and S-phase subpopulations (Figure 8). The results suggest that S-HDL can effectively induce the S-phase and G2/M-phase arrest of CCCs and CCSCs, and the cell cycle arrest ability of S-HDL was higher than that of Sal, which is consistent with the previous results of cell proliferation and apoptosis.
S-HDL Inhibits Migration of CCCs and CCSCs
To explore the effect of S-HDL on the migration ability of CCCs and CCSCs, RTCA technique was used to measure the migration ability of CCCs and CCSCs after S-HDL treatment for 48 h. S-HDL significantly decreased the migration ability of CCCs and CCSCs compared with PBS-treatment and Sal-treatment (Figure 9). S-HDL can effectively inhibit the migration of CCCs and CCSCs, thereby inhibiting tumor metastasis.
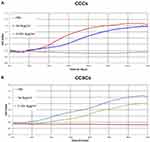 |
Figure 9 The effects of S-HDL on cell migration ability of CCCs and CCSCs. Cell migration ability of CCCs (A) and CCSCs (B) analyzed by xCELLigence Real Time Cell Analyzer DP instrument. |
Discussion
In recent years, nanoparticles have been widely studied as a drug delivery system because of their unique characteristics.15,41 HDL has unique physicochemical properties, including naturally synthesized physiological components, amphiphilic apolipoproteins, specific protein–protein interactions, hydrophobic agent-incorporating and small size.18 It has been proved that the phospholipid core of HDL can be combined with hydrophobic drugs to improve the water solubility of drugs and promote the effective release of drugs.17,42 Nanoparticles with smaller particle size can target the tumor sites through enhanced permeability and retention (EPR) effect, reducing the systemic toxicity.43,44 Salinomycin is an effective drug for the treatment of CSCs, which is one of the main causes of cancer recurrence and metastasis. However, the hydrophobicity and toxicity of Sal limit its development as an anticancer drug. Therefore, we use HDL for the delivery of Sal to improve its water solubility, which further contributes to target CCCs and CCSCs to enhance the anticancer efficacy and reduce the systemic toxicity.
In previous studies, we successfully synthesized recombinant human high-density lipoprotein (rhHDL) by sodium cholate method and proved that rhHDL has the characteristics of natural HDL.30 In this study, we successfully synthesized salinomycin-loaded high-density lipoprotein for the first time. S-HDL was a homogeneous and discoidal-shaped nanoparticle system with small particle size. The absence of vasculature supportive tissues promotes the formation of a diameter of 100 nm ~ 2 μm leaky vessels and pores in tumor tissues and this phenomenon is called EPR effect.43 Nanoparticles with diameter smaller than pore size can penetrate into tumor tissue through leaky blood vessels to achieve passive targeting of tumor tissue and retain it for a longer time.45 Sykes et al found that gold nanoparticles possessing size ˂ 45 nm can easily penetrate into the tumor tissue without affecting the tumor size.46 The particle size of S-HDL makes it possible to accumulate in tumor tissue through EPR effect.
Recently, CSCs subpopulations in cancer cells are mainly sorted by surface markers in vitro. CD44 is a common marker of CSCs. It plays an important role in cell adhesion and migration, and also has the ability to bind with hyaluronic acid in the extracellular matrix.47 However, the results of immunohistochemistry showed that CD44 was expressed in both normal and cancerous cervix.48 Therefore, CD44 should be used in combination with other surface markers to identify CCSCs. CD44 is usually used in combination with CD24 as a marker of CSCs and CSCs are characterized by CD44highCD24low.49,50 CD24 is a kind of cell surface protein, which is associated with low expression and poor differentiation.51 We used the expression levels of C-myc and Sox-2 to further verify the CSCs characteristics of CD44highCD24low CCCs. C-myc gene is an important member of myc gene family, which is related to cell proliferation and cell division and it is also involved in the occurrence and development of a variety of tumors. As a result, the C-myc gene can be used as a marker for CCSCs.52 Sox-2 is a key transcription factor, which plays a role in embryonic development and determines the fate of stem cells.53 Some studies have shown that CCCs overexpressing Sox-2 have stronger proliferation ability, clonogenicity and tumorigenicity.54
The main components of the shell of HDL nanoparticles include ApoA-I and ApoE. ApoA-I has been reported to have antitumor effect by many studies, which can promote cell cycle arrest and inhibit metastatic potential of hepatoma cells.27 Apolipoprotein E is a ligand of LRP1, which can target LRP1 and be internalized through LRP1 mediated endocytosis.21 The expression of LRP1 in cancer may be positively correlated with the degree of malignancy. Studies have shown that LRP1 can support the migration and invasion of breast cancer cells and thyroid cancer cells.22–24,55 In addition, the increased expression of LRP1 can predict more aggressive tumor behavior and is related to higher histological grade in endometrial carcinoma.25 Therefore, nanocarrier with ApoE may be able to target the malignant tumors with highly expressed LRP1. The relatively high expression of LRP1 of CCCs and CCSCs not only indicated the malignant degree of tumor tissues but also increased the possibility of being targeted by S-HDL. In the cellular uptake experiment, HDL nanoparticles were more absorbed by CCCs and CCSCs compared with the free drug, which related to the enhancement of the solubility of the drug-loaded and the endocytosis of the nanoparticles. The cellular uptake of HDL nanoparticles by CCCs and CCSCs was stronger than that of IOSE80, which was consistent with our hypothesis that HDL nanoparticles increase cellular uptake by targeting LRP1.
The toxicity of chemotherapeutic drugs will bring considerable discomfort and risk to cancer patients. Nanoparticles, as the carrier of concentrate anticancer drugs, have been proved in clinical studies to have the effect of reducing toxicity.56 S-HDL can significantly reduce the IC50 of CCCs and CCSCs, indicating that S-HDL can kill cervical cancer cells at a lower concentration, which may be related to the phenomenon that CCCs and CCSCs can take in more HDL encapsulated drugs than free drugs. Lower drug concentration can reduce the systemic toxicity of anticancer drugs.
Genome-wide mRNA analysis can only show the changes at the gene level in the cell, and there is a certain deviation in its accuracy due to the processing of enormous information. The possible biological processes can be inferred according to the changes of genes in the cell, and the corresponding biological processes should be verified in combination with corresponding experiments. Genome-wide mRNA analysis showed that S-HDL could significantly affect the expression of multiple genes in CCCs. We compared the CCCs gene expression of S-HDL-treatment group with control group, and selected the genes with significant changes. Based on these changes, we predicted the possible cellular biological processes induced by S-HDL. Apoptosis is an ordered and orchestrated cellular process. One of the main reasons of malignant transformation is that cells hardly get into the process of apoptosis.57 Cell cycle is a molecular event during which DNA damage is detected and repaired to prevent uncontrolled cell division, and occurs in an orderly sequential irreversible fashion.58 The deregulation of cell cycle is one of the causes of tumor formation, which promotes the further proliferation of tumor cells. The proportion of cancer cells with active division (G0 phase) is significantly higher than that of normal tissues. Cancer cells, which are actively undergoing cell cycle, are the targets of cancer treatment, as DNA is relatively exposed in the process of cell division and hence susceptible to damage by drugs. In addition, it has been confirmed that the arrest of G1/G0-phase arrest may promote the migration of cancer cells.59 S-HDL reduced G1 phase of cell cycle, which did not cause tumor cell migration and was also confirmed by migration analysis. The migration of cancer cells is the initial step of tumor metastasis. Through a series of cellular events, including the change of phenotype caused by cytoskeleton remodeling and the degradation of extracellular matrix, cancer cells can detach from the primary tumor and metastasize to distant sites.60 Apoptosis, cell cycle and cell migration are important biological processes that affect the occurrence and development of tumors. S-HDL can affect these biological processes of CCCs and CCSCs simultaneously to eliminate tumor cells.
Conclusion
In this study, we successfully synthesized S-HDL by loading salinomycin into HDL, which had good encapsulation efficiency and stability. The shell of S-HDL was mainly composed of ApoA-I and ApoE. ApoE can be combined with LRP1, which has been proved to be highly expressed in CCCs and CCSCs in our study. This may realize the targeting of S-HDL to CCCs and CCSCs in order to reduce the side effects of drugs. In addition, HDL nanoparticles can enhance the uptake of drugs by CCCs and CCSCs and kill CCCs and CCSCs at lower concentrations, thus improving the antitumor efficacy and drug safety. Meanwhile, S-HDL can promote cell apoptosis, induce cell cycle arrest and inhibit cell migration. Thus, S-HDL treatment may become a potential treatment for cervical cancer in the future.
Abbreviations
CSCs, cancer stem cells; Sal, salinomycin; S-HDL, salinomycin-loaded high-density lipoprotein; CCCs, cervical cancer cells; CCSCs, cervical cancer stem cells; HDL, high-density lipoprotein; ApoA-I, Apolipoprotein A-I; ApoE, Apolipoprotein E; rhApoA-I, recombinant human ApoA-I; rhApoE, recombinant human ApoE; LRP1, lipoprotein receptor-related proteins 1; EE,
Encapsulation efficiency; DMEM, Dulbecco modified Eagle medium; C6, Coumarin 6; C6-HDL, Coumarin-6-loaded high-density lipoprotein; PDI, polydispersity index.
Acknowledgments
This work was supported by grants from Jilin Science and Technology Funds (20190201062JC), Education Department of Jilin Province (JJKH20190100KJ). And thanks to the laser confocal microscope provided by Jilin Zhongke Bio-engineering Joint Stock Co., Ltd. and the genome-wide analysis provided by Jilin Prochance Biomedical Co., Ltd.
Disclosure
Professor Manman Su report grants from the National Natural Science Foundation of China, during the conduct of the study. The authors report no other conflicts of interest in this work.
References
1. Small W
2. Huang R, Rofstad EK. Cancer stem cells (CSCs), cervical CSCs and targeted therapies. Oncotarget. 2017;8(21):35351–35367. doi:10.18632/oncotarget.10169
3. Yao T, Lu R, Zhang Y, et al. Cervical cancer stem cells. Cell Prolif. 2015;48(6):611–625. doi:10.1111/cpr.12216
4. Antoszczak M. A medicinal chemistry perspective on salinomycin as a potent anticancer and anti-CSCs agent. Eur J Med Chem. 2019;164:366–377. doi:10.1016/j.ejmech.2018.12.057
5. Gupta PB, Onder TT, Jiang G, et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell. 2009;138(4):645–659. doi:10.1016/j.cell.2009.06.034
6. de Aberasturi AL, Redrado M, Villalba M, et al. TMPRSS4 induces cancer stem cell-like properties in lung cancer cells and correlates with ALDH expression in NSCLC patients. Cancer Lett. 2016;370(2):165–176. doi:10.1016/j.canlet.2015.10.012
7. Schenk M, Aykut B, Teske C, Giese NA, Weitz J, Welsch T. Salinomycin inhibits growth of pancreatic cancer and cancer cell migration by disruption of actin stress fiber integrity. Cancer Lett. 2015;358(2):161–169. doi:10.1016/j.canlet.2014.12.037
8. Zhang C, Tian Y, Song F, Fu C, Han B, Wang Y. Salinomycin inhibits the growth of colorectal carcinoma by targeting tumor stem cells. Oncol Rep. 2015;34(5):2469–2476. doi:10.3892/or.2015.4253
9. Liu L, Wang Q, Mao J, et al. Salinomycin suppresses cancer cell stemness and attenuates TGF-beta-induced epithelial-mesenchymal transition of renal cell carcinoma cells. Chem Biol Interact. 2018;296:145–153. doi:10.1016/j.cbi.2018.09.018
10. Yue W, Hamai A, Tonelli G, et al. Inhibition of the autophagic flux by salinomycin in breast cancer stem-like/progenitor cells interferes with their maintenance. Autophagy. 2013;9(5):714–729. doi:10.4161/auto.23997
11. Ojo OO, Bhadauria S, Rath SK. Correction: dose-dependent adverse effects of salinomycin on male reproductive organs and fertility in mice. PLoS One. 2019;14(12):e0226872. doi:10.1371/journal.pone.0226872
12. Plumlee KH, Johnson B, Galey FD. Acute salinomycin toxicosis of pigs. J Vet Diagn Invest. 1995;7(3):419–420. doi:10.1177/104063879500700327
13. Mo L, Zhao Z, Hu X, et al. Smart nanodrug with nuclear localization sequences in the presence of MMP-2 to overcome biobarriers and drug resistance. Chemistry. 2019;25(8):1895–1900. doi:10.1002/chem.201805107
14. Zhang ZT, Huang-Fu MY, Xu WH, Han M. Stimulus-responsive nanoscale delivery systems triggered by the enzymes in the tumor microenvironment. Eur J Pharm Biopharm. 2019;137:122–130. doi:10.1016/j.ejpb.2019.02.009
15. Wang Q, Liu F, Wang L, et al. Enhanced and prolonged antitumor effect of salinomycin-loaded gelatinase-responsive nanoparticles via targeted drug delivery and inhibition of cervical cancer stem cells. Int J Nanomedicine. 2020;15:1283–1295. doi:10.2147/IJN.S234679
16. Wang Q, Yen YT, Xie C, et al. Combined delivery of salinomycin and docetaxel by dual-targeting gelatinase nanoparticles effectively inhibits cervical cancer cells and cancer stem cells. Drug Deliv. 2021;28(1):510–519. doi:10.1080/10717544.2021.1886378
17. Oda MN, Hargreaves PL, Beckstead JA, Redmond KA, van Antwerpen R, Ryan RO. Reconstituted high density lipoprotein enriched with the polyene antibiotic amphotericin B. J Lipid Res. 2006;47(2):260–267. doi:10.1194/jlr.D500033-JLR200
18. Mo ZC, Ren K, Liu X, Tang ZL, Yi GH. A high-density lipoprotein-mediated drug delivery system. Adv Drug Deliv Rev. 2016;106(Pt A):132–147. doi:10.1016/j.addr.2016.04.030
19. Beisiegel U, Weber W, Ihrke G, Herz J, Stanley KK. The LDL-receptor-related protein, LRP, is an apolipoprotein E-binding protein. Nature. 1989;341(6238):162–164. doi:10.1038/341162a0
20. Shinohara M, Tachibana M, Kanekiyo T, Bu G. Role of LRP1 in the pathogenesis of Alzheimer’s disease: evidence from clinical and preclinical studies. J Lipid Res. 2017;58(7):1267–1281. doi:10.1194/jlr.R075796
21. Laatsch A, Panteli M, Sornsakrin M, Hoffzimmer B, Grewal T, Heeren J. Low density lipoprotein receptor-related protein 1 dependent endosomal trapping and recycling of apolipoprotein E. PLoS One. 2012;7(1):e29385. doi:10.1371/journal.pone.0029385
22. Chazaud B, Ricoux R, Christov C, Plonquet A, Gherardi RK, Barlovatz-Meimon G. Promigratory effect of plasminogen activator inhibitor-1 on invasive breast cancer cell populations. Am J Pathol. 2002;160(1):237–246. doi:10.1016/S0002-9440(10)64367-2
23. Dedieu S, Langlois B, Devy J, et al. LRP-1 silencing prevents malignant cell invasion despite increased pericellular proteolytic activities. Mol Cell Biol. 2008;28(9):2980–2995. doi:10.1128/MCB.02238-07
24. Fayard B, Bianchi F, Dey J, et al. The serine protease inhibitor protease nexin-1 controls mammary cancer metastasis through LRP-1-mediated MMP-9 expression. Cancer Res. 2009;69(14):5690–5698. doi:10.1158/0008-5472.CAN-08-4573
25. Catasus L, Llorente-Cortes V, Cuatrecasas M, Pons C, Espinosa I, Prat J. Low-density lipoprotein receptor-related protein 1 (LRP-1) is associated with high grade, advanced stage and p53 and p16 alterations in endometrial carcinomas. Histopathology. 2011;59(3):567–571. doi:10.1111/j.1365-2559.2011.03942.x
26. Gordon SM, Hofmann S, Askew DS, Davidson WS. High density lipoprotein: it’s not just about lipid transport anymore. Trends Endocrinol Metab. 2011;22(1):9–15. doi:10.1016/j.tem.2010.10.001
27. Ma XL, Gao XH, Gong ZJ, et al. Apolipoprotein A1: a novel serum biomarker for predicting the prognosis of hepatocellular carcinoma after curative resection. Oncotarget. 2016;7(43):70654–70668. doi:10.18632/oncotarget.12203
28. Chattopadhyay A, Yang X, Mukherjee P, et al. Treating the intestine with oral ApoA-I mimetic tg6f reduces tumor burden in mouse models of metastatic lung cancer. Sci Rep. 2018;8(1):9032. doi:10.1038/s41598-018-26755-0
29. Zamanian-Daryoush M, Lindner D, Tallant TC, et al. The cardioprotective protein apolipoprotein A1 promotes potent anti-tumorigenic effects. J Biol Chem. 2013;288(29):21237–21252. doi:10.1074/jbc.M113.468967
30. Su M, Chang W, Shi K, et al. Preparation and activity analysis of recombinant human high-density lipoprotein. Assay Drug Dev Technol. 2012;10(5):485–491. doi:10.1089/adt.2012.467
31. Wang Q, Wu P, Ren W, et al. Comparative studies of salinomycin-loaded nanoparticles prepared by nanoprecipitation and single emulsion method. Nanoscale Res Lett. 2014;9(1):351. doi:10.1186/1556-276X-9-351
32. Li QL, Sun Y, Sun YL, et al. Mesoporous silica nanoparticles coated by layer-by-layer self-assembly using cucurbituril for in vitro and in vivo anticancer drug release. Chem Mater. 2014;26(22):6418–6431. doi:10.1021/cm503304p
33. Nakamura S, Otsuka N, Yoshino Y, Sakamoto T, Yuasa H. Predicting the occurrence of sticking during tablet production by shear testing of a pharmaceutical powder. Chem Pharm Bull (Tokyo). 2016;64(5):512–516. doi:10.1248/cpb.c15-00992
34. Gu W, Yeo E, McMillan N, Yu C. Silencing oncogene expression in cervical cancer stem-like cells inhibits their cell growth and self-renewal ability. Cancer Gene Ther. 2011;18(12):897–905. doi:10.1038/cgt.2011.58
35. Zamulaeva I, Selivanova E, Matchuk O, Kiseleva V, Mkrtchyan L, Krikunova L. Radiation response of cervical cancer stem cells is associated with pretreatment proportion of these cells and physical status of HPV DNA. Int J Mol Sci. 2021;22(3):1445. doi:10.3390/ijms22031445
36. Zhang J, Chen X, Bian L, Wang Y, Liu H. CD44+/CD24+-expressing cervical cancer cells and radioresistant cervical cancer cells exhibit cancer stem cell characteristics. Gynecol Obstet Invest. 2019;84(2):174–182. doi:10.1159/000493129
37. Lillis AP, Van Duyn LB, Murphy-Ullrich JE, Strickland DK. LDL receptor-related protein 1: unique tissue-specific functions revealed by selective gene knockout studies. Physiol Rev. 2008;88(3):887–918. doi:10.1152/physrev.00033.2007
38. Zhang S, Balch C, Chan MW, et al. Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res. 2008;68(11):4311–4320. doi:10.1158/0008-5472.CAN-08-0364
39. Balasubramaniam SD, Balakrishnan V, Oon CE, Kaur G. Key molecular events in cervical cancer development. Medicina. 2019;55(7):384. doi:10.3390/medicina55070384
40. Libra M, Scalisi A, Vella N, et al. Uterine cervical carcinoma: role of matrix metalloproteinases (review). Int J Oncol. 2009;34(4):897–903. doi:10.3892/ijo_00000215
41. Fan L, Wang J, Meng F, et al. Delivering the acetylcholine neurotransmitter by nanodrugs as an effective treatment for Alzheimer’s disease. J Biomed Nanotechnol. 2018;14(12):2066–2076. doi:10.1166/jbn.2018.2649
42. Burgess BL, Cavigiolio G, Fannucchi MV, Illek B, Forte TM, Oda MN. A phospholipid-apolipoprotein A-I nanoparticle containing amphotericin B as a drug delivery platform with cell membrane protective properties. Int J Pharm. 2010;399(1–2):148–155. doi:10.1016/j.ijpharm.2010.07.057
43. Kalyane D, Raval N, Maheshwari R, Tambe V, Kalia K, Tekade RK. Employment of enhanced permeability and retention effect (EPR): nanoparticle-based precision tools for targeting of therapeutic and diagnostic agent in cancer. Mater Sci Eng C Mater Biol Appl. 2019;98:1252–1276. doi:10.1016/j.msec.2019.01.066
44. Kang H, Rho S, Stiles WR, et al. Size-dependent EPR effect of polymeric nanoparticles on tumor targeting. Adv Healthc Mater. 2020;9(1):e1901223. doi:10.1002/adhm.201901223
45. Byrne JD, Betancourt T, Brannon-Peppas L. Active targeting schemes for nanoparticle systems in cancer therapeutics. Adv Drug Deliv Rev. 2008;60(15):1615–1626. doi:10.1016/j.addr.2008.08.005
46. Sykes EA, Dai Q, Sarsons CD, et al. Tailoring nanoparticle designs to target cancer based on tumor pathophysiology. Proc Natl Acad Sci USA. 2016;113(9):E1142–E1151. doi:10.1073/pnas.1521265113
47. Kim WT, Ryu CJ. Cancer stem cell surface markers on normal stem cells. BMB Rep. 2017;50(6):285–298. doi:10.5483/bmbrep.2017.50.6.039
48. Shimabukuro K, Toyama-Sorimachi N, Ozaki Y, et al. The expression patterns of standard and variant CD44 molecules in normal uterine cervix and cervical cancer. Gynecol Oncol. 1997;64(1):26–34. doi:10.1006/gyno.1996.4530
49. Jaggupilli A, Elkord E. Significance of CD44 and CD24 as cancer stem cell markers: an enduring ambiguity. Clin Dev Immunol. 2012;2012:708036. doi:10.1155/2012/708036
50. Li W, Ma H, Zhang J, Zhu L, Wang C, Yang Y. Unraveling the roles of CD44/CD24 and ALDH1 as cancer stem cell markers in tumorigenesis and metastasis. Sci Rep. 2017;7(1):13856. doi:10.1038/s41598-017-14364-2
51. Routray S, Mohanty N. Cancer stem cells accountability in progression of head and neck squamous cell carcinoma: the most recent trends! Mol Biol Int. 2014;2014:375325. doi:10.1155/2014/375325
52. Feng D, Peng C, Li C, et al. Identification and characterization of cancer stem-like cells from primary carcinoma of the cervix uteri. Oncol Rep. 2009;22(5):1129–1134. doi:10.3892/or_00000545
53. Adachi K, Suemori H, Yasuda SY, Nakatsuji N, Kawase E. Role of SOX2 in maintaining pluripotency of human embryonic stem cells. Genes Cells. 2010;15(5):455–470. doi:10.1111/j.1365-2443.2010.01400.x
54. Ji J, Zheng PS. Expression of Sox2 in human cervical carcinogenesis. Hum Pathol. 2010;41(10):1438–1447. doi:10.1016/j.humpath.2009.11.021
55. Montel V, Gaultier A, Lester RD, Campana WM, Gonias SL. The low-density lipoprotein receptor-related protein regulates cancer cell survival and metastasis development. Cancer Res. 2007;67(20):9817–9824. doi:10.1158/0008-5472.CAN-07-0683
56. Maranhao RC, Vital CG, Tavoni TM, Graziani SR. Clinical experience with drug delivery systems as tools to decrease the toxicity of anticancer chemotherapeutic agents. Expert Opin Drug Deliv. 2017;14(10):1217–1226. doi:10.1080/17425247.2017.1276560
57. Wong RS. Apoptosis in cancer: from pathogenesis to treatment. J Exp Clin Cancer Res. 2011;30(1):87. doi:10.1186/1756-9966-30-87
58. Vermeulen K, Van Bockstaele DR, Berneman ZN. The cell cycle: a review of regulation, deregulation and therapeutic targets in cancer. Cell Prolif. 2003;36(3):131–149. doi:10.1046/j.1365-2184.2003.00266.x
59. Kohrman AQ, Matus DQ. Divide or conquer: cell cycle regulation of invasive behavior. Trends Cell Biol. 2017;27(1):12–25. doi:10.1016/j.tcb.2016.08.003
60. Tochhawng L, Deng S, Pervaiz S, Yap CT. Redox regulation of cancer cell migration and invasion. Mitochondrion. 2013;13(3):246–253. doi:10.1016/j.mito.2012.08.002
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.