Back to Journals » Drug Design, Development and Therapy » Volume 10
Synthesis, activity, and docking study of phenylthiazole acids as potential agonists of PPARγ
Authors Ma L , Wang T, Shi M, Ye H
Received 14 February 2016
Accepted for publication 29 March 2016
Published 30 May 2016 Volume 2016:10 Pages 1807—1815
DOI https://doi.org/10.2147/DDDT.S106406
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Prof. Dr. Wei Duan
Liang Ma,1,* Taijin Wang,2,* Min Shi,1 Haoyu Ye2
1Department of Nephrology, Kidney Research Institute, 2State Key Laboratory of Biotherapy and Cancer Center/Collaborative Innovation Center of Biotherapy, West China Hospital, West China Medical School of Sichuan University, Chengdu, People’s Republic of China
*These authors contributed equally to this work
Abstract: Peroxisome proliferator-activated receptor gamma (PPARγ) is a ligand-mediated transcription factor playing key roles in glucose and lipid homeostasis, and PPARγ ligands possess therapeutic potential in these as well as other areas. In this study, a series of phenylthiazole acids have been synthesized and evaluated for agonistic activity by a convenient fluorescence polarization-based PPARγ ligand screening assay. Compound 4t, as a potential PPARγ agonist with half maximal effective concentration (EC50) 0.75±0.20 µM, exhibited in vitro potency comparable with a 0.83±0.14 µM of the positive control rosiglitazone. Molecular docking and molecular dynamics simulations indicated that phenylthiazole acid 4t interacted with the amino acid residues of the active site of the PPARγ complex in a stable manner, consistent with the result of the in vitro ligand assay.
Keywords: PPARγ, phenylthiazole acids, ligand screening assay, docking study
Introduction
Nuclear receptors (NRs) form a superfamily of ligand-dependent transcription factors that are involved in diverse pathophysiological processes, including reproduction, development, homeostasis, and immune regulation.1,2 This superfamily includes receptors for thyroid hormones, steroid hormones, retinoids, and vitamin D, as well as a large number of orphan receptors.3 In the human genome sequence that has been elucidated, ~50 NRs were found to be expressed.4 A large number of studies have focused on peroxisome proliferator-activated receptors (PPARs), a subset of these NRs, mainly because of their pivotal roles in energy homeostasis.5
PPARs are regulated by endogenous fatty acids and synthetic ligands.6–8 So far, three subtypes, PPARα, PPARβ/δ, and PPARγ, have been identified, which are differentially expressed in a tissue-specific manner.9 The most extensively studied isoform, PPARγ, modulates the transcription of genes responsible for adipose differentiation, glucose homeostasis, and lipid metabolism.9,10 Therefore, PPARγ has been considered as a therapeutic target for the treatment of diabetes and dyslipidemia. Furthermore, thiazolidinediones (TZDs) such as rosiglitazone (RSG) and pioglitazone are high-affinity ligands and full agonists of PPARγ that have been used in the treatment of type 2 diabetes mellitus.11 Although TZDs have adverse effects in clinical use, the use of these synthetic ligands has increased the understanding of the PPARγ mechanism of activation and its subsequent biological functions. PPARγ agonists TZDs have also been potential drug leads to develop potent and safe drug candidates, especially antidiabetic drug RSG.5,12–14
This study describes a series of small-molecule phenylthiazole acids that have been synthesized and investigated for agonistic activity by a convenient fluorescence polarization (FP)-based PPARγ ligand screening assay. Compound 4t, as a potential PPARγ agonist with half maximal effective concentration (EC50) 0.75±0.20 μM, exhibited comparable potency to a 0.83±0.14 μM of the positive RSG control. Docking studies indicated that the phenylthiazole acid derivative 4t interacted with the amino acid residues of the active site of the PPARγ complex in a stable manner, in which results are consistent with those of the in vitro ligand assay.
Materials and methods
Chemical synthesis
Analytical grade chemical reagents were purchased from Chengdu Changzheng Chemical Factory (Sichuan, People’s Republic of China). Thin-layer chromatography was performed on 0.20 mm silica gel 60 F254 plates (Qingdao Ocean Chemical Factory, QingDao, People’s Republic of China). Proton nuclear magnetic resonance (1H NMR) spectra were recorded at 400 MHz on a Varian spectrometer model Gemini 400 and reported in parts per million (ppm). Chemical shifts (δ) are quoted in ppm relative to tetramethylsilane as internal standard, where (δ) tetramethylsilane =0.00 ppm. The multiplicity of the signal is indicated as s, singlet; d, doublet; t, triplet; q, quartet; and m, multiplet, defined as all multipeak signals where overlap or complex coupling of signals makes definitive descriptions of peaks difficult. Mass spectra were obtained using a Q-TOF Premier mass spectrometer utilizing electrospray ionization (Waters Quattro Premier XE, Milford, MA, USA). The purity of compounds was determined to be ≥97% by high-performance liquid chromatography analysis with a photodiode array detector (Waters, Milford, MA, USA) and an Atlantis C18 chromatographic column (150×4.6 mm2, id 5 μm; Waters).
General procedure for the synthesis of α-bromoacetophenones
Copper (II) bromide (19.5 mmol) was added to a solution of acetophenones (6.5 mmol) in anhydrous ethyl acetate (25 mL) and chloroform (25 mL), and the mixture was stirred at 65°C for 0.5 hours. The reaction was then extracted with chloroform (40 mL) and the combined organic layer was washed with water (2×10 mL) and brine (2×10 mL). The solvent was removed to obtain the crude product without further purification.15
General procedure for the synthesis of phenylthiazol-2-amines
Thiourea (4.51 mmol) was added to a solution of α-bromoacetophenones (4.3 mmol) in anhydrous ethanol (10 mL) and the mixture was refluxed for 2 hours. The reaction was allowed to cool to room temperature when a color precipitate was formed. The precipitated solid was collected by vacuum filtration, washed with water (4×15 mL), ethanol (2×10 mL), and ether (2×10 mL), and finally dried in vacuo at 40°C for 24 hours.
General procedure for the synthesis of phenylthiazole acids
A mixture of phenylthiazol-2-amines (3.5 mmol) and succinic anhydride or diglycolic anhydride or thiodiglycolic anhydride (3.5 mmol) in dimethylformamide (15 mL) was added to pyridine (3.5 mmol) and stirred overnight at room temperature. The solution was poured into a volume of a water/ice mixture and acidified with 1 M HCl. The precipitated solids that were formed were collected by vacuum filtration, washed with water (4×15 mL), ethanol (2×10 mL), and ether (2×10 mL), and finally dried in vacuo at 40°C for 24 hours (Supplementary Materials).
FP-based PPARγ ligand screening assay
A PPARγ Ligand Screening Assay Kit provided a convenient FP-based single step assay for screening PPARγ ligands.16–18 In this assay, a ligand of PPARγ is conjugated to fluorescein and is used as the displacement probe. Ligands, agonists, and antagonists of PPARγ will displace the fluorescent probe, leading to a decrease in FP. The PPARγ Ligand Screening Assay Kit is a robust assay with a Z′ of 0.81 and has a dynamic rangeof >120 mP units. The assay cocktail was first prepared (including FP Assay Buffer, PPARγ FP Assay Fluorescent Probe, and human recombinant PPARγ), and each concentration of ligand was assayed in duplicate. The plate was covered with the plate cover supplied with the kit and incubated for 60–90 minutes at room temperature. The plates were then read at excitation and emission wavelengths of 470 and 530 nm, respectively. The Z′-factor for the PPARγ Ligand Screening Assay Kit was determined to be 0.81. Results were calculated by FP of a formula (polarization mP =1,000× (I║ − I┴)/(I║ + I┴)), and the IC50 was calculated by the mP-concentration displacement curve according to the kit protocol (Cayman).
Molecular docking
The crystal structure (PDB: 4AME) of PPARγ bound to RSG was used as the docking template. The protein structure was processed by Discovery Studio 3.1 (BIOVIA, San Diego, CA, USA). The chain containing the bound ligand was chosen, and other ligands were deleted. Furthermore, the water molecules were also eliminated. Compounds 4a, 4h, 4i, and 4t were docked into the ligand binding domain using AutoDock 4.2 (The Scripps Research Institute, La Jolla, CA, USA).19 AutodockTools 1.4.6 was implemented to build the autogrid box. The grid center was set based on the known ligand, and the grid contained 40×48×54 autogrid points with 0.375 Å spacing. The number of individuals in the population was 150, and the maximum numbers of energy evaluations and generations were 25 million and 27,000, respectively. Lamarckian GA (4.2) was used to search energetically supported binding modes. In addition, 100 docked poses were generated, with all settings not mentioned left at their default values. In order to obtain further refined information about the interactions between protein residues and agonists, molecular simulations were performed.20,21
Molecular dynamics simulations
The binding model of compound 4t and the X-ray structure of PPARγ in complex with RSG were prepared for molecular dynamics (MD) simulations. MD simulations were performed using GROMACS 4.5.5 (Ohio Supercomputer Center, Columbus, OH, USA)22 with the Amber99sb force field. Compound 4t and RSG were parameterized with general AMBER force (GAF) parameters and AM1-bond charge corrections (BCC) charges by Antechamber python parser interface.23 The system was surrounded by a cubic water box of TIP3P molecules, and periodic boundary conditions were applied in all directions. The distance between the protein and the box edge was 10 Å. Na+ counterions were added by replacing solution to ensure the overall charge neutrality of the simulated system. The steepest descent algorithms were implemented for energy minimization with 50,000 steps. A restraining simulation was performed by fixing the position of the protein and ligand, heating to 300 K, and equilibrating for 100 ps. A 200 ps isothermal-isobaric ensemble and a 25 ns MD were performed in sequence. In carrying out the MD simulations, the particle mesh Ewald method for long-range electrostatics, the Lincs algorithm for covalent bond constraints, a 14 Å cutoff for van der Waals interactions, and a 12 Å cutoff for Coulomb interactions were all applied.
Results and discussion
Chemistry
The preparation of a library of phenylthiazole acid derivatives has been performed in a tandem three-step sequence from commercially available acetophenones as described in Figure 1.15 Treatment of 1 with copper (II) bromide in a mixture of ethyl acetate and chloroform as valid solvent afforded α-bromoacetophenones 2. The pivotal intermediate phenylthiazol-2-amines 3 were synthesized via a simple reaction of thiourea and appropriate 2 in refluxed ethanol. The condensation of 3 with different anhydrides was carried out in the presence of pyridine and acidified with diluted hydrochloric acid (1 M) to give the corresponding phenylthiazole acids (4a–v).24 Following this stage, all final products were fully analyzed and characterized by nuclear magnetic resonance, mass spectroscopy, and high-performance liquid chromatography before being submitted for biological screening.
FP-based PPARγ ligand screening assay
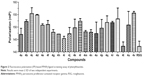
The agonistic potency of 22 small-molecule phenylthiazole acids was assessed using a convenient FP-based PPARγ ligand screening assay.25,26 Here, RSG as a PPARγ agonist was chosen as a positive control. As shown in Figure 2, four compounds (4a, 4h, 4i, and 4t) were found to exhibit potential agonistic activity at a concentration of 10.0 μM. Among them, compound 4t showed the most potent effects of FP, comparable with RSG.
On the basis of primary screening, four selected phenylthiazole acids (4a, 4h, 4i, and 4t) were further investigated. As shown in Table 1, the four compounds possessed remarkable agonistic activity on PPARγ. As anticipated, compound 4t exhibited the greatest agonistic activity on PPARγ (EC50 =0.75±0.20 μM), compared with that of RSG (EC50 =0.83±0.14 μM).
Among the 22 small-molecule phenylthiazole acid derivatives (4a–f and 4g–v), the long chain fatty acid group was found to contribute most to the agonist potency (4h, 4i, 4t > 4a). The introduction of methyl substitution in the thiazole ring was unable to promote agonistic activity (methyl phenylthiazole 4f, 4o, 4u, and 4v), which indicated that the small steric hindrance of thiazole was favorable for agonist potency. Between biphenyl derivatives 4o and 4t, the sulfur atom of the acid chain significantly improved the agonistic effects compared with the oxygen atom. Additionally, the p position of the phenyl group in 4t exhibited potential activity because of structural similarity to the tail group of RSG, identified by the subsequent molecular docking studies and dynamic simulation.
Molecular docking
Primarily, the binding mode of RSG has been analyzed according to the X-ray structure of the PPARγ complex. The TZD of RSG forms a H-bonding interaction with the three amino acid residues of Tyr473 (2.6 Å), His323 (3.2 Å), and Ser289 (2.2 Å). The hydrophobic tail of RSG interacts with the residues of Leu330, Val339, Ile341, Leu353, Phe363, Met364, and Phe368. Therefore, docking poses with a similar binding mode and low binding energy were chosen for the study. In Table 2, the in vitro agonistic activity and the molecular properties showed good correlation. The potent compounds displayed high values of AlogP and LogD, which proved useful for selecting the phenylthiazoles for predicting molecular properties. A redocking experiment was performed to identify whether the complex could be used for the docking study. The root mean square deviation (RMSD) was 0.9 Å, ie, sufficient for the docking template.
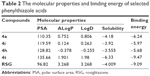  | Table 2 The molecular properties and binding energy of selected phenylthiazole acids |
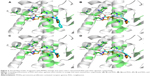
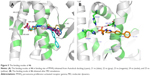
The binding energy of RSG was -9.09 kcal/mol, while that of 4t was -9.47 kcal/mol. There was a difference relationship between the in vitro PPARγ agonistic activity and biological activity and the binding energy to PPARγ of 4t, and because the receptor was adjusted significantly to the presence of the ligand, RSG was better able to bind in the current phase. Among the four agonists (4a, 4h, 4i, and 4t), compound, 4t exhibited the lowest binding energy and the highest activity. The carboxylate group of 4t interacted with residues of Tyr473 (2.0 Å), His323 (3.2 Å), His449 (3.5 Å), and Ser289 (1.9 Å) to form hydrogen bonds to the PPARγ active sites, and its hydrophobic tail was encompassed by a surface formed from Leu330, Phe368, Met364, Ile341, Val339, Phe282, Ile281, and Leu353. Additionally, the N atom of the amide group of 4t formed a H-bonding interaction with the Cys285 (2.1 Å) of the PPARγ complex (Figure 3A). However, the other three agonists only interacted with part of the abovementioned amino acid residues (Figure 3B–D).
MD simulations
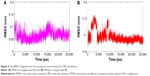
Due to the fact that the potential ligand-induced conformational changes were not fully taken into consideration in the docking study, the MD simulations were performed on the RSG/4t-PPARγ complex to check whether they were stable in aqueous solution and if any induced fit occurred. Therefore, the RMSDs of RSG and 4t were analyzed to assess the degree of the conformation drift. In the simulation of RSG-bound PPARγ, the RMSD of RSG was ~<1 Å (Figure 4A). It was reasonable to suppose that the pose of the RSG-PPARγ complex was taken from the crystal structure and its conformation was relatively stable. Unlike RSG-bound PPARγ, the RMSD of 4t was stable after 10 ns simulations (Figure 4B). Compared with the initial docking pose of 4t, the structure closed to the helix and formed the stronger H-bonding interaction with the nearest residues in the stable phases (Figure 5A). At t 25 ns, the carboxylate group of 4t formed a H-bonding interaction with His449 (2.0 Å), His323 (2.0 Å, 2.7 Å), and Ser289 (2.6 Å). In addition, its hydrophobic tail fully contacted a surface formed by Leu330, Phe368, Met364, Ile341, Val339, Phe282, Ile281, and Leu353.
In order to predict binding affinities in stable phases, 4t in these conformations was redocked into the active site. The binding energy of the pose whose RMSD value was among the lowest in the 100 docking poses was retained (Figure 5B). At 25 ns, the lowest value of RMSD was 0.71 Å, and the corresponding binding energy of the pose was −9.47 kcal/mol. The result indicated that 4t could bind to the active site of PPARγ, consistent with the data of the in vitro screening assay.
Conclusion
PPARγ modulates the transcription of the genes responsible for adipose differentiation, glucose homeostasis, and lipid metabolism and has been considered as a therapeutic target for the treatment of diabetes and dyslipidemia. In this study, we describe 22 phenylthiazole acid derivatives that have been synthesized and evaluated for agonistic activity and to which an easy, a quick, and a commercial FP-based PPARγ ligand screening assay was applied to obtain in vitro data for the primary structure–activity relationship study. Among the derivatives tested, a potent PPARγ ligand 4t exhibited comparable agonistic activity to RSG. Docking and dynamics simulation studies showed that 4t docked in a stable manner to the active site of the PPARγ complex, and the result was consistent with that of the in vitro ligand assay. Further chemical modifications of 4t, as well as studies of in vivo activity and reaction mechanisms, are in progress.
Acknowledgments
This research was supported by the Natural Science Foundation of China (no 81402782) and the China Postdoctoral Science Foundation (no 2015T80985).
Disclosure
The authors report no conflicts of interest in this work.
References
Evans RM, Mangelsdorf DJ. Nuclear receptors, RXR, and the big bang. Cell. 2014;157(1):255–266. | ||
Bensinger SJ, Tontonoz P. Integration of metabolism and inflammation by lipid-activated nuclear receptors. Nature. 2008;454(7203):470–477. | ||
Gronemeyer H, Gustafsson J-A, Laudet V. Principles for modulation of the nuclear receptor superfamily. Nat Rev Drug Discov. 2004;3(11):950–964. | ||
Chawla A, Repa JJ, Evans RM, Mangelsdorf DJ. Nuclear receptors and lipid physiology: opening the X-files. Science. 2001;294(5548):1866–1870. | ||
Grygiel-Górniak B. Peroxisome proliferator-activated receptors and their ligands: nutritional and clinical implications – a review. Nutr J. 2014;13(1):1. | ||
Kliewer SA, Sundseth SS, Jones SA, et al. Fatty acids and eicosanoids regulate gene expression through direct interactions with peroxisome proliferator-activated receptors α and γ. Proc Natl Acad Sci U S A. 1997;94(9):4318–4323. | ||
Berger J, Moller DE. The mechanisms of action of PPARs. Annu Rev Med. 2002;53(1):409–435. | ||
Wright HM, Clish CB, Mikami T, et al. A synthetic antagonist for the peroxisome proliferator-activated receptor γ inhibits adipocyte differentiation. J Biol Chem. 2000;275(3):1873–1877. | ||
Marion-Letellier R, Savoye G, Ghosh S. Fatty acids, eicosanoids and PPAR gamma. Eur J Pharmacol. Epub 2015 Nov 26. | ||
Ahmadian M, Suh JM, Hah N, et al. PPARγ signaling and metabolism: the good, the bad and the future. Nat Med. 2013;99(5):557–566. | ||
Tripathi AC, Gupta SJ, Fatima GN, Sonar PK, Verma A, Saraf SK. 4-thiazolidinones: the advances continue. Eur J Med Chem. 2014;72:52–77. | ||
da Silva FMC, dos Santos JC, Campos JLO, et al. Structure-based identification of novel PPAR gamma ligands. Bioorg Med Chem Lett. 2013;23(21):5795–5802. | ||
Ohashi M, Gamo K, Oyama T, Miyachi H. Peroxisome proliferator-activated receptor gamma (PPARγ) has multiple binding points that accommodate ligands in various conformations: structurally similar PPARγ partial agonists bind to PPARγ LBD in different conformations. Bioorg Med Chem Lett. 2015;25(14):2758–2762. | ||
Ohashi M, Oyama T, Miyachi H. Different structures of the two peroxisome proliferator-activated receptor gamma (PPARγ) ligand-binding domains in homodimeric complex with partial agonist, but not full agonist. Bioorg Med Chem Lett. 2015;25(13):2639–2644. | ||
Xu Q, Huang L, Liu J, et al. Design, synthesis and biological evaluation of thiazole- and indole-based derivatives for the treatment of type II diabetes. Eur J Med Chem. 2012;52:70–81. | ||
Sakamoto J, Kimura H, Moriyama S, et al. Activation of human peroxisome proliferator-activated receptor (PPAR) subtypes by pioglitazone. Biochem Biophys Res Commun. 2000;278(3):704–711. | ||
Usui S, Suzuki T, Hattori Y, et al. Design, synthesis, and biological activity of novel PPARγ ligands based on rosiglitazone and 15d-PGJ2. Bioorg Med Chem Lett. 2005;15(6):1547–1551. | ||
Vidal-Puig A, Jimenez-Liñan M, Lowell BB, et al. Regulation of PPAR gamma gene expression by nutrition and obesity in rodents. J Clin Invest. 1996;97(11):2553. | ||
Goodsell DS, Morris GM, Olson AJ. Automated docking of flexible ligands: applications of AutoDock. J Mol Recognit. 1996;9(1):1–5. | ||
Cosconati S, Forli S, Perryman AL, Harris R, Goodsell DS, Olson AJ. Virtual screening with AutoDock: theory and practice. Expert Opin Drug Discov. 2010;5(6):597–607. | ||
Morris GM, Huey R, Olson AJ. Using autodock for ligand-receptor docking. Curr Protoc Bioinformatics. 2008;8(14):1–40. | ||
Pronk S, Páll S, Schulz R, et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics. 2013;29(7):845–854. | ||
da Silva AWS, Vranken WF. ACPYPE-antechamber python parser interface. BMC Res Notes. 2012;5(1):367. | ||
Cai H, Yan G, Zhang X, Gorbenko O, Wang H, Zhu W. Discovery of highly selective inhibitors of human fatty acid binding protein 4 (FABP4) by virtual screening. Bioorg Med Chem Lett. 2010;20(12):3675–3679. | ||
Fang M, Webster TF, Ferguson PL, Stapleton HM. Characterizing the peroxisome proliferator-activated receptor (PPARγ) ligand binding potential of several major flame retardants, their metabolites, and chemical mixtures in house dust. Environ Health Perspect. 2015;123(2):166–172. | ||
Rossi AM, Taylor CW. Analysis of protein-ligand interactions by fluorescence polarization. Nat Protoc. 2011;6(3):365–387. |
Supplementary materials
The chemical structure data of reported compounds in this study are described in full.
4-Oxo-4-((5-(p-tolyl)thiazol-2-yl)amino)butanoic acid (4a)
Yield 62.8%. 1H NMR (DMSO-d6, 400 MHz) δ 12.27 (s, 1H), 12.15 (s, 1H), 7.77 (d, J=8.0 Hz, 2H), 7.52 (s, 1H), 7.22 (d, J=8.0 Hz, 2H), 2.68 (t, J=6.4 Hz, 3H), 2.56 (t, J=6.4 Hz, 2H), 2.30 (s, 3H); MS (ESI), m/z 289.3 [M-H]−.
4-((5-(4-Chlorophenyl)thiazol-2-yl)amino)-4-oxobutanoic acid (4b)
Yield 72.5%. 1H NMR (DMSO-d6, 400 MHz) δ 12.32 (s, 1H), 7.90 (d, J=8.4 Hz, 1H), 7.79 (d, J=8.8 Hz, 1H), 7.67 (s, 1H), 7.51–6.48 (m, 2H), 2.69 (t, J=6.8 Hz, 2H), 2.57 (t, J=6.8 Hz, 2H); MS (ESI), m/z 309.3 [M-H]−.
4-Oxo-4-((5-(4-(trifluoromethyl)phenyl)thiazol-2-yl)amino)butanoic acid (4c)
Yield 75.2%. 1H NMR (DMSO-d6, 400 MHz) δ 12.38 (s, 1H), 8.10 (d, J=8.0 Hz, 2H), 7.85 (s, 1H), 7.79 (d, J=8.4 Hz, 2H), 2.70 (t, J=6.4 Hz, 3H), 2.58 (t, J=6.4 Hz, 3H); MS (ESI), m/z 343.3 [M-H]−.
4-((5-(3,4-Difluorophenyl)thiazol-2-yl)amino)-4-oxobutanoic acid (4d)
Yield 48.5%. 1H NMR (DMSO-d6, 400 MHz) δ 12.34 (s, 1H), 12.24 (s, 1H), 7.88–7.93 (m, 1H), 7.77–7.74 (m, 1H), 7.72 (s, 1H), 7.48 (q, J=8.4, 10.4 Hz, 1H), 2.69 (t, J=6.4 Hz, 2H), 2.57 (t, J=6.4 Hz, 2H); MS (ESI), m/z 311.3 [M-H]−.
4-((5-(Naphthalen-2-yl)thiazol-2-yl)amino)-4-oxobutanoic acid (4e)
Yield 76.4%. 1H NMR (DMSO-d6, 400 MHz) δ 12.39 (s, 1H), 12.26 (s, 1H), 8.00 (d, J=8.4 Hz, 1H), 7.98–7.92 (m, 3H), 7.77 (s, 1H), 7.56–7.50 (m, 2H), 2.71 (t, J=6.4 Hz, 2H), 2.59 (t, J=6.4 Hz, 2H); MS (ESI), m/z 325.3 [M-H]−.
4-((5-(4-Chlorophenyl)-4-methylthiazol-2-yl)amino)-4-oxobutanoic acid (4f)
Yield 54.8%. 1H NMR (DMSO-d6, 400 MHz) δ12.25 (s, 1H), 12.15 (s, 1H), 7.65 (d, J=8.4 Hz, 2H), 7.49 (d, J=8.8 Hz, 2H), 2.65 (t, J=6.4 Hz, 2H), 2.55 (t, J=6.4 Hz, 2H), 2.46 (s, 3H); MS (ESI), m/z 323.3 [M-H]−.
2-(2-Oxo-2-((5-(p-tolyl)thiazol-2-yl)amino)ethoxy)acetic acid (4g)
Yield 45.8%. 1H NMR (DMSO-d6, 400 MHz) δ 12.17 (s, 1H), 7.79 (d, J=8.1 Hz, 2H), 7.57 (s, 1H), 7.24 (d, J=8.0 Hz, 2H), 4.33 (s, 2H), 4.20 (s, 2H), 2.32 (s, 3H). MS (ESI), m/z 305.0 [M-H]−.
2-(2-((5-(4-Methoxyphenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4h)
Yield 54%. 1H NMR (DMSO-d6, 400 MHz) δ 12.12 (s, 1H), 7.83 (d, J=8.8 Hz, 2H), 7.48 (s, 1H), 6.99 (d, J=8.8 Hz, 2H), 4.34 (s, 2H), 4.21 (s, 2H), 2.53–2.48 (m, 3H). MS (ESI), m/z 321.0 [M-H]−.
2-(2-((5-(2-Chlorophenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4i)
Yield 52%.1H NMR (DMSO-d6, 400 MHz) δ 12.28 (s, 1H), 8.57 (s, 1H), 7.84 (dd, J=7.6, 1.9 Hz, 1H), 7.64 (s, 1H), 7.55 (dd, J=7.8, 1.4 Hz, 1H), 7.41 (m, 2H), 4.34 (s, 2H), 4.20 (s, 2H). MS (ESI), m/z 325.8 [M-H]−.
2-(2-((5-(4-Chlorophenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4j)
Yield 48%. 1H NMR (DMSO-d6, 400 MHz) δ 12.21 (s, 1H), 7.92 (d, J=8.5 Hz, 2H), 7.72 (s, 1H), 7.50 (d, J=8.5 Hz, 2H), 4.34 (s, 2H), 4.20 (s, 2H). MS (ESI), m/z 325.0 [M-H]−.
2-(2-((5-(3,4-Dichlorophenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4k)
Yield 51.8%.1H NMR (DMSO-d6, 400 MHz) δ 12.31 (s, 1H), 8.15 (d, J=2.0 Hz, 1H), 7.97–7.82 (m, 2H), 7.70 (d, J=8.4 Hz, 1H), 4.34 (s, 2H), 4.20 (s, 2H). MS (ESI), m/z 359.0 [M-H]−.
2-(2-((5-(3,4-Difluorophenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4l)
Yield 58%. 1H NMR (DMSO-d6, 400 MHz) δ 12.19 (s, 1H), 7.91 (ddd, J=12.1, 7.9, 1.9 Hz, 1H), 7.76 (s, 2H), 7.50 (dd, J=19.2, 8.6 Hz, 1H), 4.35 (s, 2H), 4.21 (s, 2H). MS (ESI), m/z 327.0 [M-H]−.
2-(2-oxo-2-((5-(4-(Trifluoromethyl)phenyl)thiazol-2-yl)amino)ethoxy)acetic acid (4m)
Yield 44%. 1H NMR (DMSO-d6, 400 MHz) δ 12.26 (s, 1H), 8.62 (s, 1H), 8.12 (d, J=8.2 Hz, 2H), 7.90 (s, 1H), 7.80 (d, J=8.3 Hz, 2H), 4.35 (s, 2H), 4.21 (s, 2H). MS (ESI), m/z 359.0 [M-H]−.
2-(2-((5-(4-Nitrophenyl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4n)
Yield 58.7%. 1H NMR (DMSO-d6, 400 MHz) δ 12.30 (s, 1H), 8.32 (d, J=8.8 Hz, 2H), 7.18 (d, J=8.8 Hz, 2H), 8.03 (s, 1H), 4.36 (s, 2H), 4.21(s, 2H). MS (ESI), m/z 336.0 [M-H]−.
2-(2-((5-([1,1′-Biphenyl]-4-yl)thiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4o)
Yield 62%. 1H NMR (DMSO-d6, 400 MHz) δ 12.18 (s, 1H), 8.00 (d, J=8.3 Hz, 2H), 7.76 (s, 1H), 7.73 (dd, J=6.9, 3.8 Hz, 4H), 7.48 (t, J=7.6 Hz, 2H), 7.39 (d, J=7.4 Hz, 1H), 4.36 (s, 2H), 4.22 (s, 2H). MS (ESI), m/z 367.1 [M-H]−.
2-(2-((4-Methyl-5-phenylthiazol-2-yl)amino)-2-oxoethoxy)acetic acid (4p)
Yield 44%. 1H NMR (DMSO-d6, 400 MHz) δ 11.97 (s, 1H), 7.64 (d, J=7.2 Hz, 2H), 7.45 (t, J=7.6 Hz, 2H), 7.34 (t, J=7.4 Hz, 1H), 4.31 (s, 2H), 4.19 (s, 2H), 2.48 (s, 3H). MS (ESI), m/z 305.0 [M-H]−.
2-((2-((5-(4-Chlorophenyl)thiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4q)
Yield 62%. 1H NMR (DMSO-d6, 400 MHz) δ 12.42 (s, 1H), 7.91 (d, J=8.5 Hz, 2H), 7.72 (s, 1H), 7.50 (d, J=8.6 Hz, 2H), 3.55 (s, 2H), 3.43 (s, 2H). MS (ESI), m/z 340.9 [M-H]−.
2-((2-((5-(4-Fluorophenyl)thiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4r)
Yield 68%. 1H NMR (DMSO-d6, 400 MHz) δ 12.41 (s, 1H), 7.93 (dd, J=8.8, 5.7 Hz, 2H), 7.63 (s, 1H), 7.27 (t, J=8.9 Hz, 2H), 3.54 (s, 2H), 3.43 (s, 2H). MS (ESI), m/z 325.0 [M-H]−.
2-((2-((5-(3,4-Dichlorophenyl)thiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4s)
Yield 65%. 1H NMR (DMSO-d6, 400 MHz) δ 12.45 (s, 1H), 8.14 (d, J=2.1 Hz, 1H), 7.87 (d, J=4.0 Hz, 2H), 7.70 (d, J=8.5 Hz, 1H), 3.55 (s, 2H), 3.43 (s, 2H). MS (ESI), m/z 375.0 [M-H]−.
2-((2-((5-([1,1′-Biphenyl]-4-yl)thiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4t)
Yield 72%. 1H NMR (DMSO-d6, 400 MHz) δ 12.28 (s, 1H), 7.83 (dd, J=7.6, 1.5 Hz, 2H), 7.63 (s, 2H), 7.56 (d, J=7.8 Hz, 2H), 7.46–7.36 (m, 4H), 3.55 (s, 2H), 3.41(s, 2H). MS (ESI), m/z 383.1 [M-H]−.
2-((2-((4-Methyl-5-phenylthiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4u)
Yield 65%. 1H NMR (DMSO-d6, 400 MHz) δ 12.26 (s, 1H), 7.64 (d, J=7.6 Hz, 2H), 7.45 (t, J=7.6 Hz, 2H), 7.35 (t, J=7.4 Hz, 1H), 3.51 (s, 2H), 3.43 (s, 2H), 2.47 (s, 3H). MS (ESI), m/z 321.0 [M-H]−.
2-((2-((5-(4-Chlorophenyl)-4-methylthiazol-2-yl)amino)-2-oxoethyl)thio)acetic acid (4v)
Yield 60%. 1H NMR (DMSO-d6, 400 MHz) δ 12.32 (s, 1H), 7.66 (d, J=8.5 Hz, 2H), 7.51 (d, J=8.5 Hz, 2H), 3.51 (s, 2H), 3.42 (s, 2H), 2.47 (s, 3H). MS (ESI), m/z 355.0 [M-H]−.
 © 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2016 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.