Back to Journals » International Journal of General Medicine » Volume 14
Step-by-Step Construction of Gene Co-Expression Network Analysis for Identifying Novel Biomarkers of Sepsis Occurrence and Progression
Authors Yu X , Qu C, Ke L , Tong Z, Li W
Received 12 July 2021
Accepted for publication 10 September 2021
Published 24 September 2021 Volume 2021:14 Pages 6047—6057
DOI https://doi.org/10.2147/IJGM.S328076
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Xianqiang Yu,1,* Cheng Qu,2,* Lu Ke,2 Zhihui Tong,2 Weiqin Li1,2
1Medical School, Southeast University, Nanjing, People’s Republic of China; 2Department of Critical Care Medicine, Jinling Hospital, Medical School of Nanjing University, Nanjing, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Weiqin Li Email [email protected]
Background: Sepsis is the leading cause of death in critically ill patients. Although it is well known that the immune system plays a key role in sepsis, exactly how it works remains unknown.
Methods: In our study, we used weighted gene co-expression network analysis (WGCNA) to screen out the immune-related genes that may play a critical role in the process of sepsis.
Results: A total of three sepsis-related hub genes were screened for further verification. Subsequent analysis of immune subtypes suggested their potential predictive effect in the clinic.
Conclusion: Our study shows that three immune-related genes CHMP1A, MED15 and MGAT1 are important biomarkers of sepsis. The screened genes may help to distinguish normal individuals from patients with different degrees of sepsis.
Keywords: sepsis, gene co-expression network, biomarker, immune cell
Background
Sepsis is the leading cause of death in critically ill patients, accounting for 30–50% of hospital mortality in the United States.1–3 From the perspective of pathophysiology, sepsis occurs with complex pro-inflammatory and anti-inflammatory process disorders, further leading to systemic inflammatory response syndrome (SIRS) and multiple organ dysfunction syndrome (MODS).4–7 Although a variety of interventions are currently available for the treatment of sepsis, the lack of targeted drugs leads to limited overall clinical efficacy.8–10 In addition, the complexity and variability of sepsis make it hard to grasp its severity, so early prediction of sepsis progression will be particularly important in guiding treatment.11,12
Biomarkers are increasingly being used to predict the pathophysiological characteristics of diseases, including sepsis.13–15 However, the predictive function of most sepsis-related biomarkers is limited, and it is urgently needed to explore high-efficiency biomarkers in clinical practice. To the best of our knowledge, sepsis is a heterogeneous, life-threatening syndrome that coexists with multiple diseases.7 The innate immune system is the first line of defense against infection when sepsis occurs, and it often collaborates with adaptive immune responses to protect the host. Genetic differences among different individuals contribute to the diversity of immune responses to sepsis.16 This is partly due to the fact that the quantitative characteristics of pathogen-associated molecular patterns (PAMPs) and pattern recognition receptors (PRRs) interactions are not similar across all patients.17 The interaction between PAMPs and PRRs results in the production and subsequent release of pro-inflammatory and anti-inflammatory mediators, coordinating the clinical state of the host. And immune cells are direct participants in this response. Therefore, based on this premise, it is of potential value to search for immune-related markers in the process of sepsis for disease prediction.
With the rapid development and application of gene chip technology, weighted gene co-expression network analysis (WGCNA) is gradually used to analyze the molecular mechanism and network relationships of disease.18,19 Obviously, this revolutionary technological advance allows the search for biomarkers of sepsis at genetic level to guide clinical practice. In our study, we used WGCNA to screen out the immune-related genes that may play a critical role in the process of sepsis. We screened out promising immune-related candidate biomarkers of sepsis in which highly correlated genes clustered. From this perspective, the hub genes we screened also provide us with the possibility to better understand the mechanism and occurrence of sepsis.
Materials and Methods
Data Collection
We downloaded the GEO database to analyze the gene expression profiles of sepsis. Inclusion criteria: (1) the whole-genome expression profiling of whole blood of sepsis patients or healthy control samples were available in the datasets; (2) no less than 15sepsis samples and/or 15 healthy control samples were included in each dataset. Exclusion criteria: (1) a history of treatment prior to sample collection; (2) a clear risk of systemic disease in the control group. In our study, we downloaded the gene expression profiles of sepsis GSE54514 and GSE63042 for further analysis. GSE54514 and GSE63042 contained 53 (normal: 18; sepsis: 35) and 129 (normal: 23; sepsis: 106) samples, respectively.
Co-Expression Network Construction
We used “WGCNA” package in R to establish the co-expression network based on the profile of differentially expressed genes (DEGs).20 Furthermore, we performed sample clustering to plot the sample tree and detect outliers.
Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis
We used STRING (search tool for the retrieval of interacting genes/proteins) database for GO Enrichment and KEGG pathway analysis. P < 0.05 was considered as the cut-off value of enriched GO terms and KEGG pathway.
Identification and Validation of Hub Gene
Pearson’s correlation of module membership >0.2 and P < 0.05 was used to evaluate the connectivity of hub modules. Furthermore, hub module genes were carried out to establish a protein‐protein interaction (PPI) network. Finally, we obtained the real hub genes in the overlapping part of this PPI network.
Evaluation of Immune Cell Subtype Distribution
CIBERSORT is a deconvolution algorithm that converts normalized gene expression matrix into components of infiltrating immune cells. During CIBERSORT calculations, we quantified the abundance of specific cell types in whole blood and verified CIBERSORT results by fluorescent-activated cell sorting (FACS). P value <0.05 of CIBERSORT output was defined as more accurate prediction of immune cell composition. Then, the samples satisfying the constraints were further analyzed.
Identification of Differentially Expressed Immune-Related Gene
GSE63042 data set was used for WGCNA analysis, and key modules and their genes were selected. The GSE54514 data set was classified, and immune infiltration analysis was conducted to explore the differences in the proportion of immune cells in individuals with three different states. Finally, the correlation between the biomarkers and immune cells was calculated and the credibility of the correlation was verified by significance analysis. We examined the expression of real hub genes in these samples. P value <0.05 was considered statistically significant.
Statistical Analysis
Heat-maps were conducted by using the R software. Statistical analyses were conducted with the R package. P value <0.05 was considered statistically significant.
Results
WGCNA Analysis
Establishment of Co-Expression Matrix
The gene was divided into dozens of modules, and the co-expression matrix was established. The minimum number of genes in each gene module was set to 50, and a total of 32 modules were aggregated (Figure 1A). Grey defaults were genes that could not be classified into any modules, further merging the previous 32 modules into 13 (Figure 1B).
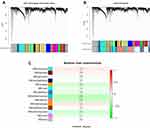 |
Figure 1 Modular clustering and its analysis correlation with clinical characteristics. (A) Module cluster tree diagram. (B) Cluster dendrogram. (C) Module–trait relationships. |
Correlation Analysis
Correlation analysis was conducted between gene modules and clinical features, and the selected clinical features were the severity of the disease. During the analysis, phenotypic traits were transformed into quantitative traits (0 represents normal samples; the disease ranges from 1 to 4, with a higher value indicating a more severe disease). Among the 13 co-expression modules, MEdarkturquoise had the maximal relevance to sepsis (Figure 1C).
Hub Module Enrichment
The R software cluster Profiler package was used for enrichment analysis to find the common functions and related pathways in differentially expressed gene set. Through enrichment analysis of the 901 hub module genes obtained above, the bubble chart of top 10 term related to biological processes (BP) was obtained (Figure 2).
 |
Figure 2 The bubble diagram of BP-related TOP10 term obtained by GO enrichment of HUB module gene. |
PPI Network Construction
We analyzed the protein network interaction relationship of 901 module genes (confidence = 0.7) and obtained 410 hub genes. We selected the top 10 proteins and used Cytoscape software to beautify the protein network, and screened out the proteins with the top 10 proteins and extended to the outer layer (Figure 3).
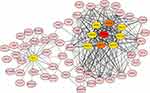 |
Figure 3 The protein interaction network of key genes. |
Screening of Biomarkers
We selected a total of four gene sets: 410 PPI hub genes, 68 key genes of hub module and the set of two DEGs. The intersection of these four gene sets was used to obtain the biomarkers of this study. Finally, three biomarkers were selected from the intersection part, namely CHMP1A, MED15 and MGAT1 (Figure 4).
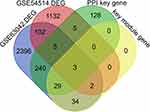 |
Figure 4 Screening of biomarkers. |
Correlation Between Biomarkers and Survival
In order to explore whether these three genes have an impact on survival and death of patients, we collated the data of these two datasets into different groups and visualized their significance through box plots. The disease samples were divided into survival and non-survival samples (normal samples were not used). MED15 gene had a significant effect on survival and death in GSE54514 dataset (Figure 5A). CHMP1A and MGAT1 gene had a significant effect on survival and death in GSE63042 dataset (Figure 5B).
Single-Gene Analysis of Biomarkers
CHMP1A Gene
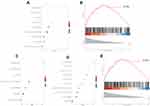
We calculated the correlation coefficient between the expression of all genes and CHMP1A gene, and performed gene set enrichment analysis (GSEA) according to the sequence of correlation coefficient, which was not enriched in the pathway but enriched to 9 GO terms (Figure 6A). One of the immune-related GO was visualized separately (Figure 6B).
MED15 Gene
We calculated the correlation coefficient between the expression of all genes and MED15 gene, and performed GSEA according to the sequence of correlation coefficient, which was not enriched in the pathway but enriched to 7 GO terms (Figure 6C).
MGAT1 Gene
We calculated the correlation coefficient between the expression of all genes and MGAT1 gene, and performed GSEA according to the sequence of correlation coefficient, which was not enriched in the pathway but enriched to 46 GO terms (Figure 6D). One of the immune-related GO was visualized separately (Figure 6E).
Immune Cell Infiltration in Different Individuals
In our study, 21 immune cells in the samples were screened out by using CIBERSORT algorithm and LM22 gene marker. Among these immune cells, T cells CD4 memory activated, T cells regulatory (Tregs), NK cells, macrophages (M0), mast cells and neutrophils had most significant differences in the proportion of normal/survival/non-survival individuals (Figure 7).
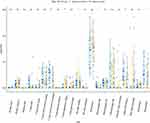 |
Figure 7 Immune cell infiltration in normal/living/dead individuals.*P<0.05, **P<0.01,***P<0.001. Abbreviation: ns, not significant. |
Correlation Analysis Between Biomarkers and Immune Cells
The above three biomarkers were used for correlation analysis of each immune cell. CHMP1A gene was positively correlated with T cells CD4 naive and macrophages (M2) but negatively correlated with macrophages (M0) and T cells CD8 (Figure 8A). MED15 gene was positively correlated with mast cells activated and T cells CD4 naive but negatively correlated with macrophages (M0) and neutrophils (Figure 8B). MGAT1 gene was positively correlated with T cells CD4 naïve and monocytes but negatively correlated with neutrophils and T cells CD8 (Figure 8C).
Discussion
Early prediction of sepsis biomarkers is important for effective clinical intervention.21 In our study, WGCNA was used to explore the pathological process and marker genes in whole blood samples of sepsis. After data preprocessing and weighted gene network construction, the modules were associated with feature and function enrichment analysis. According to the module recognition heat-map and scatter diagram, the module related to sepsis onset was turquoise module (p < 0.05). Finally, we selected three sepsis-related biomarkers: CHMP1A, MED15 and MGAT1.
Chromatin modifying protein 1A (CHMP1A) is a member of the endosomal sorting complex required for transport (ESCRT-III) family found in both cytoplasmic and nuclear matrix fractions, which identified as roles of chromatin modification, fundamental proteins required for multivesicular sorting in eukaryotes and regulation of cell-cycle progression.22,23 To the best of our knowledge, CHMP1A is encoded by PRSM1 gene and found in mitotic chromosome scaffold and nuclear matrix.24–26 Over-expression of CHMP1A interferes with nuclear structure and DNA replication, etc.23 Recent studies showed that CHMP1A acts as a tumor suppressor through the p53 signaling pathway in human embryonic kidney and ductal pancreatic tumor cells.27,28 Subsequently, some scholars further discovered that CHMP1A plays a role of regulating the proliferation and differentiation of retinal progenitor cells.22 For the first time, our analysis suggested that CHMP1A is associated with sepsis and is positively associated with T cells CD4 naïve and macrophages (M2), but a strong negative correlation with macrophages (M0) and T cells CD8.
The mediator complex is a four module transcriptional co-activator among eukaryotes, in which the tail module primarily recruits multiple transcriptional regulators to the transcription unit.29,30 As one of the tail subunits, MED15 acts as a linker between regulatory proteins and RNA polymerase II.31–33 Abnormal expression of MED15 is associated with a variety of human malignant tumors. The expression of MED15 was up-regulated to varying degrees in different parts of head and neck squamous cell carcinoma tissue.34,35 In addition, compared with androgen-sensitive prostate cancer (PCA) and benign tissue, MED15 was over-expressed in approximately 70% of locally recurrent and distantly metastatic castrated resistant PCA.36 Some scholars indicated that the degree of malignancy of renal cell carcinoma was significantly reduced by knocking down MED15.29 For the first time, our study proved that MED15 is significantly involved in the immune regulation process of sepsis. MED15 has a strong positive correlation with mast cells activated and T cells CD4 naive, but a strong negative correlation with macrophages (M0) and neutrophils.
MGAT1 is a microsomal enzyme that catalyzes the synthesis of diacylglycerol (DAG) and triacylglycerol (TAG).37,38 Mammalian TAG synthesis occurs in two ways, with the MGAT mentioned directly catalyzing the synthesis of DAG by monoacylglycerol via an alternative pathway. Finally, the action of DGAT1 or DGAT2 converts DAG to TAG.39 In addition, MGAT1 is a novel transcriptional target of Wnt/β-Catenin signaling pathway. This signaling pathway plays a key role in controlling tumor progression.40–42 From this perspective, MGAT1 functions in promoting the synthesis of triglycerides and the development of tumors. Our study indicated that MGAT1 has a strong positive correlation with T cells CD4 naive and monocytes but a strong negative correlation with neutrophils and T cells CD8.
Many scholars have attempted to find a group of biomarkers to better identify patients with sepsis at adverse risk. However, none of the biomarkers can fully reflect the potential deterioration of the disease in patients with sepsis. Currently, the three best predictors of sepsis are IL-1 receptor antagonist (IL-1ra), protein C and neutrophil gelatinase associated lipocalin (NGAL).43 Each of them may serve as a potential biomarker for sepsis, either as an anti-inflammatory protein (IL-1ra), an important component of coagulation (protein C), or as a marker of organ damage (NGAL). However, it is difficult to predict whether these three biomarkers are superior to traditional biomarkers.
Genome-wide association studies (GWASs) have provided valuable insights by pinpointing associations to both innate and adaptive immune response loci, as well as novel unexpected risk factors for infection susceptibility.44,45 Recently, some scholars used WGCNA to screen regulatory factors related to sepsis, and finally found that transcription factors CEBPB and ETV6 were the main regulatory factors.46 Meanwhile, our study provided effective biomarkers for the prediction of sepsis progression, and we found that CHMP1A, MED15 and MGAT1 play an important role in the prediction and immune response state of sepsis.
We visualized the samples and screened out 21 types of immune cells. Among them, T cells CD4 memory activated, T cells regulatory (Tregs), NK cells, macrophages (M0), mast cells and neutrophils had most significant differences in the proportion of normal/survival/non-survival individuals. We speculated that the analyzed genes may distinguish between normal individuals and patients with varying degrees of sepsis. From another perspective, new therapeutic approaches of sepsis by rebalancing multiple immune cell subset homeostasis may become potential targeted therapies. The limitation of this study is that we did not conduct an experimental study. Therefore, a further study on immune cells can identify ideal immunotherapy targets and improve the autoimmune regulation ability of patients with sepsis.
Conclusion
In conclusion, our study is the first to discover the predictive role of CHMP1A, MED15 and MGAT1 in the immunologic process of sepsis through WGCNA. The screened genes may help to distinguish normal individuals from patients with different degrees of sepsis.
Data Sharing Statement
The datasets used or analysed during the current study are available at: http://www.ncbi.nlm.nih.gov/ geo.
Consent for Publication
All authors have agreed to publish this article.
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agreed to be accountable for all aspects of the work.
Funding
National Natural Science Foundation of China (No. 82070669).
Disclosure
Dr Cheng Qu is now affiliated at General Surgery Department, Drum Tower Hospital, Nanjing University. The authors declare that they have no competing interests.
References
1. Binnie A, Tsang J, Hu P, et al. Epigenetics of sepsis. Crit Care Med. 2020;48(5):745–756. doi:10.1097/CCM.0000000000004247
2. Levy MM, Dellinger RP, Townsend SR, et al. The surviving sepsis campaign: results of an international guideline-based performance improvement program targeting severe sepsis. Intensive Care Med. 2010;36:222–231. doi:10.1007/s00134-009-1738-3
3. Teggert A, Datta H, Ali Z. Biomarkers for point-of-care diagnosis of sepsis. Micromachines. 2020;11(3):286.
4. Gullo A, Bianco N, Berlot G. Management of severe sepsis and septic shock: challenges and recommendations. Crit Care Clin. 2006;22:489–501. doi:10.1016/j.ccc.2006.03.006
5. Hotchkiss RS, Karl IE. The pathophysiology and treatment of sepsis. N Engl J Med. 2003;348:138–150. doi:10.1056/NEJMra021333
6. Hawiger J, Veach RA, Zienkiewicz J. New paradigms in sepsis: from prevention to protection of failing microcirculation. J ThrombHaemost. 2015;13:1743–1756. doi:10.1111/jth.13061
7. Singer M, Deutschman CS, Seymour CW, et al. The third international consensus definitions for sepsis and septic shock (sepsis-3). JAMA. 2016;315:801–810.
8. Rhodes A, Evans LE, Alhazzani W, et al. Surviving sepsis campaign. Intensive Care Med. 2017;43:304–377.
9. Boomer JS, To K, Chang KC, et al. Immunosuppression in patients who die of sepsis and multiple organ failure. JAMA. 2011;306:2594–2605. doi:10.1001/jama.2011.1829
10. Hotchkiss RS, Monneret G, Payen D. Immunosuppression in sepsis: a novel understanding of the disorder and a new therapeutic approach. Lancet Infect Dis. 2013;13:260–268. doi:10.1016/S1473-3099(13)70001-X
11. Kumar A, Roberts D, Wood KE, et al. Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit Care Med. 2006;34:1589–1596. doi:10.1097/01.CCM.0000217961.75225.E9
12. Zambon M, Ceola M, Almeida-de-Castro R, Gullo A, Vincent JL. Implementation of the surviving sepsis campaign guidelines for severe sepsis and septic shock: we could go faster. J Crit Care. 2008;23:455–460. doi:10.1016/j.jcrc.2007.08.003
13. Kustan P, Horvath-Szalai Z, Muhl D. Nonconventional markers of sepsis. Ejifcc. 2017;28:122–133.
14. Cho SY, Choi JH. Biomarkers of sepsis. Infect Chemother. 2014;46(1):1–12. doi:10.3947/ic.2014.46.1.1
15. Pierrakos C, Vincent JL. Sepsis biomarkers: a review. Crit Care. 2010;14(1):R15. doi:10.1186/cc8872
16. Giamarellos-Bourboulis EJ, Opal SM. The role of genetics and antibodies in sepsis. Ann Transl Med. 2016;4(17):328. doi:10.21037/atm.2016.08.63
17. Kimura A, Kishimoto T. IL-6: regulator of Treg/Th17 balance. Eur J Immunol. 2010;40:1830–1835. doi:10.1002/eji.201040391
18. Miller JA, Woltjer RL, Goodenbour JM, Horvath S, Geschwind DH. Genes and pathways underlying regional and cell type changes in Alzheimer’s disease. Genome Med. 2013;5:48. doi:10.1186/gm452
19. Radulescu E, Jaffe AE, Straub RE, et al. Identification and prioritization of gene sets associated with schizophrenia risk by co-expression network analysis in human brain. Mol Psychiatry. 2018;25:791–804. doi:10.1038/s41380-018-0304-1
20. Pearson GS. Reviewing manuscripts with problematic language issues. J Am Psychiatr Nurses Assoc. 2019;25:251–252. doi:10.1177/1078390319857707
21. Shetty A, Macdonald SPJ, Keijzers G, et al. Review article: sepsis in the emergency department – part 2: investigations and monitoring. EMA-Emerg Med Australas. 2018;30:4–12. doi:10.1111/1742-6723.12924
22. Sun N, Zhang D, Ni N, et al. miR-17 regulates the proliferation and differentiation of retinal progenitor cells by targeting CHMP1A. Biochem Biophys Res Commun. 2020;523(2):493–499. doi:10.1016/j.bbrc.2019.11.108
23. Stauffer DR, Howard TL, Nyun T, et al. CHMP1 is a novel nuclear matrix protein affecting chromatin structure and cell-cycle progression. J Cell Sci. 2001;114(Pt 13):2383–2393.
24. Williams RL, Urbe S. The emerging shape of the ESCRT machinery. Nat Rev Mol Cell Biol. 2007;8:355–368. doi:10.1038/nrm2162
25. Scott A, Gaspar J, Stuchell-Brereton MD, Alam SL, Skalicky JJ, Sundquist WI. Structure and ESCRT-III protein interactions of the MIT domain of human VPS4A. Proc Natl Acad Sci USA. 2005;102:13813–13818. doi:10.1073/pnas.0502165102
26. Hurley JH, Emr SD. The ESCRT complexes: structure and mechanism of a membrane trafficking network. Annu Rev Biophys Biomol Struct. 2006;35:277–298. doi:10.1146/annurev.biophys.35.040405.102126
27. Li J, Belogortseva N, Porter D, Park M. Chmp1A functions as a novel tumor suppressor gene in human embryonic kidney and ductal pancreatic tumor cells. Cell Cycle. 2008;7:2886–2893. doi:10.4161/cc.7.18.6677
28. You Z, Xin Y, Liu Y, et al. Chmp1A acts as a tumor suppressor gene that inhibits proliferation of renal cell carcinoma. Cancer Lett. 2012;319(2):190–196. doi:10.1016/j.canlet.2012.01.010
29. Weiten R, Muller T, Schmidt D, et al. The mediator complex subunit MED15, a promoter of tumour progression and metastatic spread in renal cell carcinoma. Cancer Biomark. 2018;21(4):839–847. doi:10.3233/CBM-170757
30. Nakatsubo T, Nishitani S, Kikuchi Y, et al. Human mediator subunit MED15 promotes transcriptional activation. Drug DiscovTher. 2014;8(5):212–217.
31. Orphanides G, Lagrange T, Reinberg D. The general transcription factors of RNA polymerase II. Genes Dev. 1996;10:2657–2683. doi:10.1101/gad.10.21.2657
32. Compe E, Egly JM. TFIIH: when transcription met DNA repair. Nat Rev Mol Cell Biol. 2012;13:343–354. doi:10.1038/nrm3350
33. Thomas MC, Chiang CM. The general transcription machinery and general cofactors. Crit Rev Biochem Mol Biol. 2006;41:105–178.
34. Syring I, Klümper N, Offermann A, et al. Comprehensive analysis of the transcriptional profile of the mediator complex across human cancer types. Oncotarget. 2016;7(17):23043–23055. doi:10.18632/oncotarget.8469
35. Shaikhibrahim Z, Offermann A, Halbach R, et al. Clinical and molecular implications of MED15 in head and neck squamous cell carcinoma. Am J Pathol. 2015;185(4):1114–1122. doi:10.1016/j.ajpath.2014.12.010
36. Offermann A, Vlasic I, Syring I, et al. MED15 overexpression in prostate cancer arises during androgen deprivation therapy via PI3K/mTOR signalling. Oncotarget. 2017;8(5):7964–7976. doi:10.18632/oncotarget.13860
37. Lee YJ, Yu JH, Kim WH, Kim JW. New mechanisms contributing to hepatic steatosis: glucose, insulin, and lipid signaling. Anim Cells Syst. 2014;18:77–82. doi:10.1080/19768354.2014.906502
38. Phan CT, Tso P. Intestinal lipid absorption and transport. Front Biosci. 2001;6:D299–319. doi:10.2741/Phan
39. Lee YJ, Kim JW. Monoacylglycerol O-acyltransferase 1 (MGAT1) localizes to the ER and lipid droplets promoting triacylglycerol synthesis. BMB Rep. 2017;50(7):367–372. doi:10.5483/BMBRep.2017.50.7.036
40. Takayama H, Ohta M, Iwashita Y, et al. Altered glycosylation associated with dedifferentiation of hepatocellular carcinoma: a lectin microarray-based study. BMC Cancer. 2020;20(1):192. doi:10.1186/s12885-020-6699-5
41. Akiva I, Birgul IN. MGAT1 is a novel transcriptional target of Wnt/beta-catenin signaling pathway. BMC Cancer. 2018;18(1):60. doi:10.1186/s12885-017-3960-7
42. Polakis P. Wnt signaling and cancer. Genes Dev. 2000;14:1837–1851.
43. Faix JD. Biomarkers of sepsis. Crit Rev Clin Lab Sci. 2013;50(1):23–36. doi:10.3109/10408363.2013.764490
44. Finucane HK, Reshef YA, Anttila V, et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat Genet. 2018;50:621–629. doi:10.1038/s41588-018-0081-4
45. Chen L, Ge B, Casale FP, et al. Genetic drivers of epigenetic and transcriptional variation in human immune cells. Cell. 2016;167:1398–1414. doi:10.1016/j.cell.2016.10.026
46. Zhang Z, Chen L, Xu P, et al. Gene correlation network analysis to identify regulatory factors in sepsis. J Transl Med. 2020;18(1):381. doi:10.1186/s12967-020-02561-z
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.



