Back to Journals » OncoTargets and Therapy » Volume 13
STAT3-Induced Upregulation of lncRNA CASC9 Promotes the Progression of Bladder Cancer by Interacting with EZH2 and Affecting the Expression of PTEN
Authors Yuan B, Sun R, Du Y, Jia Z, Yao W, Yang J
Received 1 February 2020
Accepted for publication 29 July 2020
Published 16 September 2020 Volume 2020:13 Pages 9147—9157
DOI https://doi.org/10.2147/OTT.S248006
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Arseniy Yuzhalin
Bo Yuan,1 Rongqing Sun,1 Yuming Du,1 Zhankui Jia,2 Wencheng Yao,2 Jinjian Yang2
1Intensive Care Unit, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, Henan Province 450052, People’s Republic of China; 2Urinary Surgery Department, The First Affiliated Hospital of Zhengzhou University, Zhengzhou, Henan Province 450052, People’s Republic of China
Correspondence: Jinjian Yang
Urinary Surgery Department, The First Affiliated Hospital of Zhengzhou University, No. 1 Jianshe East Road, Erqi District, Zhengzhou, Henan Province 450052, People’s Republic of China
Email [email protected]
Objective: Long non-coding RNA (lncRNA) cancer susceptibility candidate 9 (CASC9) has been reported to play a vital role in tumorigenesis. This study explored the biological role of CASC9 and its regulation mechanism in bladder cancer (BC).
Methods: Gene expression was evaluated using quantitative reverse transcription polymerase chain reaction and Western blot. The functional role of CASC9 in BC was studied using Cell Counting Kit-8, colony formation assay, scratch wound healing assay, transwell invasion assay, and xenograft tumor assay. In addition, the mechanism of CASC9 function in BC was determined using RNA immunoprecipitation assay and chromatin immunoprecipitation assay.
Results: CASC9 was upregulated in BC tissues and cell lines, and correlated with the staging and metastasis in BC. Knockdown of CASC9 inhibited the proliferation, migration, and invasion of BC cells. Similarly, silencing of CASC9 inhibited tumor growth in vivo. Signal transducer and activator of transcription 3 (STAT3) was upregulated in BC tissues and cell lines, and positively correlated with CASC9 in BC tissues. Moreover, CASC9 was shown to be regulated by STAT3 in BC cells. Furthermore, CASC9 regulated phosphatase and tensin homolog (PTEN) expression by interacting with enhancer of zeste homolog 2 (EZH2). More significantly, CASC9 silencing-mediated inhibition of BC progression was partly reversed by EZH2 overexpression or PTEN inhibition.
Conclusion: Upregulation of CASC9 induced by STAT3 promoted the progression of BC by interacting with EZH2 and affecting the expression of PTEN, representing a novel regulatory mechanism for BC progression.
Keywords: bladder cancer, cancer susceptibility candidate 9, signal transducer and activator of transcription 3, phosphatase and tensin homolog, enhancer of zeste homolog 2
Introduction
Bladder cancer (BC) is referred to the malignant tumor arising from the epithelial lining of the urinary tract, showing prominent male predominance.1,2 More than 430,000 cases yearly are diagnosed with BC, making it the ninth most common malignancies throughout the world.3 Various factors, such as smoking, obesity, chemical exposure, and bladder infections, may lead to the occurrence of BC.4–7 In the past years, the morbidity and mortality of BC have gradually increased in China.8 Despite the improvement of diagnostic methods and surgical management, the therapeutic effect of BC is still dissatisfactory, with a lower 5-year survival rate.9 Therefore, exploring the genetic regulatory networks implicated in BC progression may help in finding novel therapeutic targets for BC, thereby improving the prognosis of patients with BC.
Long non-coding RNAs (lncRNAs; ≥200 nucleotides), an important member of non-coding RNAs (ncRNAs), have attracted extensive attention in term of their involvement in various biological processes.10 LncRNAs have been documented to participate in cell functions by different mechanisms, including epigenetic modification, transcription modulation, RNA decay, and miRNA decoys.11–13 LncRNAs have been reported to serve as tumor-suppressive or oncogenic roles in carcinogenesis.14 To date, hundreds of lncRNAs have been studied and identified in carcinogenesis.15 Among them, cancer susceptibility candidate 9 (CASC9) reportedly plays a crucial role in several types of human cancers. As an example, CASC9 was upregulated in ovarian cancer and functioned as a sink for miR-758 to regulate the expression of Lin-7 homolog A and then promote the progression of ovarian cancer.16 Similarly, the oncogenic role of CASC9 was discovered in esophageal squamous cell carcinoma.17 Suppression of CASC9 inhibited the proliferation, migration, and invasion of Eca-109 cells by affecting epithelial mesenchymal transition.18 Nevertheless, there are very few studies on the function and mechanism of CASC9 in BC progression.
In our study, we found the increased expression of CASC9 in BC tissues and cell lines, and explored the functional role of CASC9 in BC progression through altering its expression in vitro and in vivo. Mechanistically, the upstream and downstream regulation of CASC9 were analyzed in vitro. Our findings revealed that upregulation of CASC9 induced by STAT3 promoted the progression of BC by interacting with enhancer of zeste homolog 2 (EZH2) and affecting the expression of phosphatase and tensin homolog (PTEN). This study expands our understanding of the mechanisms of the genesis and development of BC.
Materials and Methods
Patient Specimens
Thirty-five pairs of BC tissues and adjacent normal tissues were collected from patients with BC who underwent surgery at the First Affiliated Hospital of Zhengzhou University. After resection, samples were snap-frozen and store them at −80°C for further analysis. All patients have not received radiotherapy or chemotherapy before the operation. Written informed consents were obtained from all patients. This study was reviewed and approved by the Ethics Review Committee of the First Affiliated Hospital of Zhengzhou University.
Cell Culture
Human bladder cancer cell lines EJ (transitional cell carcinoma), UMUC3 (high grade III, invasive), 5637 (grade II carcinoma), J82 (transitional cell carcinoma), T24 (transitional cell carcinoma) and normal human urothelial cells (SV-HUC-1) were acquired from American Type Culture Collection (Manassas, VA, USA), and cultured in RPMI-1640 medium plus 10% fetal bovine serum (FBS; Solarbio, Beijing, China) in a humidified incubator at 37°C with 5% CO2.
Cell Transfection
To overexpress CASC9, the full sequence of CASC9 was introduced into the pcDNA-3.1 vector (Invitrogen, Carlsbad, CA, USA) to generate pcDNA-CASC9 constructs (CASC9-OE). Similarly, the sequence of STAT3 was subcloned into the pcDNA-3.1 vector to create pcDNA-STAT3 constructs (STAT3-OE). shRNA-NC, CASC9 shRNA, STAT3 shRNA, EZH2 shRNA, and PTEN shRNA were synthesized by Sangon Biotech (Shanghai, China). CASC9 shRNA sequence: GCC CAG AAG ACA GTG GAA TGA. STAT3 shRNA sequence: GTC ACA CAG ATG AAC TTG GTC TTC AGG T. EZH2 shRNA sequence: AAG ACT CTG AAT GCA GTT GCT.
Construction of Stably Transfected Cells
Transfection was performed using LipofectamineTM 2000 (Invitrogen) following protocols supplied by the manufacturer when cells reached approximately 70% to 80% confluency. To establish shRNA-NC or CASC9 shRNA stably transfected cells, G418 selection was conducted based on the antibiotic resistance of the plasmids. At 48 h after transfection, transiently transfected cells were subjected to 0.5 mg/mL G418 (Sigma-Aldrich, St. Louis, MO, USA). After 4 weeks of incubation of 0.5 mg/mL G418, G418-resistant cell clones were obtained as the stably transfected cells.
Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated using Trizol reagent following the product manual. qRT-PCR assay was performed on an ABI 7600 FAST thermal cycler with One Step TB Green™ PrimeScript™ RT-PCR Kit (Takara, Dalian, China) and the following primers: CASC9 Forward: 5ʹ-TTG GTC AGC CAC ATT CAT GGT-3ʹ and Reverse: 5ʹ-AGT GCC AAT GAC TCT CCA GC-3ʹ; STAT3 Forward: 5ʹ-GGA GGA GGC ATT CGG AAA G-3ʹ and Reverse: 5ʹ-TCG TTG GTG TCA CAC AGA T-3ʹ; PTEN Forward: 5ʹ-AAG GAC AGC AAG AAG AAG G-3ʹ and Reverse: 5ʹ-GAG GAA CAG GTG GAG AGT-3ʹ; EZH2 Forward: 5ʹ-TTC ATG CAA CAC CCA ACA CTT-3ʹand Reverse: 5ʹ-GGT GGG GTC TTT ATC CGC TC-3ʹ; GAPDH Forward: 5ʹ-ACC CAG AAG ACT GTG GAG G-3ʹ and Reverse: 5ʹ-TTC TAG ACG GCA GGT CAG GT-3ʹ. The data were analyzed using 2−ΔΔCt method and normalized against GAPDH.
Western Blot
After treatment, EJ and T24 cells were harvested and lysed in RIPA lysis buffer in the light of the manufacturer’s recommendations. Protein samples were separated by a 14% sodium dodecyl sulfate-polyacrylamide gel and then transferred to polyvinylidene fluoride membranes. After blocking with 5% skim milk, membranes were probed overnight at 4°C with primary antibodies against PTEN (Abcam, Cambridge, MA, USA), EZH2 (Abcam) or GAPDH (Abcam), followed by incubation with secondary horseradish peroxidase-conjugated antibody (Abcam) for 1 h at room temperature. The signals were developed using chemiluminescence (Boster, Wuhan, China), and analyzed using Image J software (National Institutes of Health, NY, USA).
Cell Counting Kit-8 (CCK-8)
EJ and T24 cells were seeded onto 6-well plates and then transfected with CASC9 shRNA, PTEN shRNA or shRNA-NC as described above. At 48 h post-transfection, EJ and T24 cells were plated into a 96-well plate and incubated for 12 h in a 5% CO2 humidified incubator at 37°C. Following this, EJ and T24 cells were treated with CCK-8 solution (Solarbio) for 4 h and then incubated with formazan solution for 10 min on a shaker. The absorbance was determined at the wavelength of 450 nm.
Colony Formation Assay
After treatment, EJ and T24 cells were collected and plated into a 6-well plate. After 2 weeks of incubation at 37°C with 5% CO2, colonies were fixed with methanol for 15 min, followed by staining with crystal violet for 5 min. Colonies (> 50 cells) were pictured and counted under a stereomicroscope.
Scratch Wound Healing Assay
EJ and T24 cells were plated in six-well plates and cultured to 80%–90% confluence in complete medium. Thereafter, a scratch wound was generated with a 10 μL pipette, and the cells were washed twice with PBS. Subsequently, cell culture was continued in medium without FBS. Photos were taken at 0 h and 48 h using a light microscope (Olympus, Tokyo, Japan). The distance of cell migration was analyzed using Image J software (National Institutes of Health, NY, USA).
Transwell Invasion Assay
After treatment, EJ and T24 cells in serum-free medium were placed into the upper chamber pre-coated with Matrigel (Corning, Steuben County, New York, USA). Medium plus 10% FBS were added to the lower chamber to induce directional invasion. After 48 h of incubation, the cells on the upper membrane were gently removed using a cotton swab. The cells on the lower membrane were fixed with methanol, followed by staining with crystal violet. The invaded cells were imaged and counted using a light microscope.
Xenograft Tumor Assay
A total of twelve male BALB/c nude mice (weight, 21–25 g; aged 6 weeks) were purchased from Beijing Vital River Laboratories (Beijing, China) and received sterile rodent chow and water ad libitum and were housed in a facility at 23±1°C with 50% humidity and 12-h light/dark cycles. All animal experiments were performed following the National Institutes of Health Guide for the Care and Use of Laboratory Animals. All experiments performed on animals were approved by the Ethics Committee of The First Affiliated Hospital of Zhengzhou University. T24 cells stably transfected with shRNA-NC or CASC9 shRNA were subcutaneously injected into nude mice (2×106 cells/mice, six in each group). The tumor size was monitored every 5 days and calculated according to the following formula: Volume = 0.52 × length × width2. When the maximum diameter of the transplanted tumor was about 1.4 cm, mice were sacrificed and the tumors were removed and weighed.
RNA Immunoprecipitation (RIP) Assay
RIP assay was conducted using An EZMagna RNA immunoprecipitation (RIP) Kit (Millipore, Bedford, MA, USA) under the guidance of the manual. Briefly, EJ and T24 cells were collected and lysed in complete RIP lysis buffer as directed by manufacturer’s instruction. Afterwards, the cell lysates were incubated with magnetic beads conjugated with anti-ZEH2 antibody or control IgG, followed by incubation with proteinase K. The coprecipitated RNA was prepared for qRT-PCR analysis.
Chromatin Immunoprecipitation (ChIP) Assay
The ChIP assay was carried out using the EZ-Magna ChIP Chromatin Immunoprecipitation kit from Millipore, and anti-STAT3 and IgG antibodies from Abcam, according to the manufacturer’s specifications. In briefly, cross-linked chromatin was immunoprecipitated with anti-STAT3 or IgG antibodies after sonication. Thereafter, the chromatin was incubated with protein A/G-Sepharose beads. Finally, ChIP-derived DNA was tested for the enrichment of chromatin using qRT-PCR.
Statistical Analysis
The results were given as the means ± standard error of the mean. Statistical analysis was performed using Student’s t-test or one-way analysis of variance from SPSS 20.0 software (SPSS Inc., Chicago, IL, USA). Statistics with P-value < 0.05 was deemed a criterion for statistical significance.
Results
CASC9 is Overexpressed in BC Tissues and BC Cells
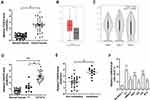
To investigate the role of CASC9 in BC, 35 paired BC tissues and adjacent normal tissues were collected and tested for CASC9 expression using qRT-PCR. The results showed that CASC9 was upregulated in BC tissues compared with that in their adjacent normal tissues (Figure 1A). The differential expression of CASC9 between bladder urothelial carcinoma and normal bladder tissues was verified using the Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/), which is based on the TCGA database (Figure 1B). The differential expression of CASC9 between patients with different stages was also shown in Figure 1C. (Figure 1C). The expression of CASC9 was higher in BC patients with stage T2/T3/T4 than that in BC patients with stage T1, while there was no significant difference in CASC9 expression between stage T1 and adjacent normal tissues (Figure 1D). Moreover, higher expression of CASC9 was found in BC patients with metastasis compared to BC patients without metastasis (Figure 1E). What is more, as shown in Table 1, CASC9 expression was not correlated with patient’s age, and gender, as well as tumor grade. Meanwhile, we also found that CASC9 was overexpressed in BC cell lines, especially in EJ and T24 cells, relative to SV-HUC-1 cells (Figure 1F).
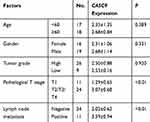 |
Table 1 Association Between CASC9 and Clinical Characteristics |
CASC9 Regulates Cell Proliferation, Migration, and Invasion in vitro
Further, we knockdown CASC9 by CASC9 shRNA in EJ and T24 cells to explore the functional role of CASC9 in BC. The downregulation of CASC9 in EJ and T24 cells was observed following CASC9 shRNA transfection, as determined by qRT-PCR assay (Figure 2A). Transfection of EJ and T24 cells with CASC9 shRNA led to the reduced cell proliferation in CCK-8 assay (Figure 2B) and the decreased colony formation in colony formation assay (Figure 2C and D). In parallel, we found that knockdown of CASC9 markedly inhibited the migration and invasion of EJ and T24 cells, as detected by scratch wound healing assay and transwell invasion assay (Figure 2E–H).
CASC9 Promotes BC Cell Tumorigenesis in vivo
Xenograft tumor assay was performed to demonstrate the function of CASC9 in BC growth. BC cells stably transfected with shRNA-NC or CASC9 shRNA were injected into nude mice. At 30th day after injection, the tumor from CASC9 shRNA-transfecting BC cells was much smaller than that from shRNA-NC-transfecting BC cells (Figure 3A). Consistently, the same tendencies were also discovered in tumor weight (Figure 3B). There was no significant difference in body weight between these two groups (Figure 3C).
CASC9 is Upregulated by the Transcription Activator STAT3 in BC Cells
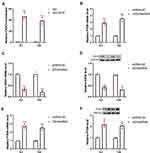
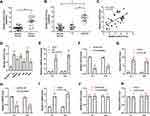
As shown in Figure 4A, STAT3 expression was obviously increased in BC tumors as compared to normal tissues. A positive correlation between CASC9 and STAT3 expression was also observed in BC tissues (Figure 4B). Additionally, The expression of STAT3 was higher in BC patients with stage T2/T3/T4 than that in BC patients with stage T1, while there was no significant difference in CASC9 expression between stage T1 and adjacent normal tissues (Figure 4C). Similarly, the upregulation of STAT3 was identified in BC cell lines (Figure 4D). We further explore if CASC9 was upregulated by STAT3 in BC. A ChIP assay revealed that STAT3 directly bound to the promoter region of CASC9 to induce its expression (Figure 4E). To further identify the interaction between CASC9 and STAT3, we altered the expression of STAT3 in EJ and T24 cells by transfecting STAT3 shRNA, STAT3-OE or their matched controls. Compared to the control group, the downregulation of STAT3 in STAT3 shRNA-transfected BC cells and the upregulation of STAT3 in STAT3-OE-transfected BC cells were confirmed by qRT-PCR (Figure 4F and G). Knockdown of STAT3 markedly reduced the expression of CASC9 in EJ and T24 cells (Figure 4H). In contrast, overexpression of STAT3 exhibited the opposite effects in EJ and T24 cells (Figure 4I). Interestingly, in both EJ and T24 cells, the expression of STAT3 was unaltered following CASC9 shRNA or CASC9 transfection (Figure 4J and K).
CASC9 Represses PTEN by Interacting with EZH2
Previous studies have shown that lncRNAs affect the expression of downstream targets by recruiting EZH2 or LSD1 proteins.19,20 To determine whether CASC9 regulates target genes through a similar mechanism, we explored the relationship between CASC9 and EZH2. As shown in Figure 5A, RIP experiments showed that CASC9 could directly bind to EZH2 in EJ and T24 cells (Figure 5A). Silencing of CASC9 markedly increased the mRNA levels of PTEN in EJ and T24 cells (Figure 5B). Further, we knockdown EZH2 in EJ and T24 cells by transfecting EZH2 shRNA. The downregulation of EZH2 was identified by qRT-PCR and Western blot (Figure 5C and D). Silencing of EZH2 markedly increased the mRNA and protein levels of PTEN in EJ and T24 cells (Figure 5E and F).
EZH2 and PTEN are Involved in CASC9-Mediated BC Progression in vitro
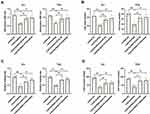
To determine whether EZH2 and PTEN participated in CASC9-mediated promotion of BC growth, EJ and T24 cells were co-transfected with CASC9 shRNA and PTEN shRNA or EZH2 overexpression plasmid. As a result, silencing of CASC9 markedly inhibited the proliferation of EJ and T24 cells, which was blocked by PTEN inhibition or EZH2 overexpression (Figure 6A). Likewise, CASC9 silencing-mediated inhibition of cell colony formation was mitigated by PTEN silence or EZH2 overexpression (Figure 6B). In parallel, CASC9 silencing-mediated inhibition of cell migration and invasion was also abrogated following transfection of PTEN shRNA or EZH2 overexpression plasmid (Figure 6C and D).
Discussion
The significance of lncRNAs has been discussed in various human cancers, including BC. Abnormal expression of lncRNAs has been recognized as a crucial mechanism for BC progression.21–23 However, till now, there are only several lncRNAs have been mechanistically characterized in BC. LncRNA CASC9 has been demonstrated to serve as an oncogenic factor in plenty of human cancers, such as breast cancer,24 colorectal cancer,25 and lung adenocarcinoma.26 A pan-cancer analysis revealed that CASC9 was strikingly overexpressed in bladder cancers, especially in urothelial carcinomas with squamous differentiation or pure squamous bladder cancers, indicating the potential role of CASC9 in BC.27 However, no study has been reported regarding the contribution of CASC9 in BC. In our work, the upregulation of CASC9 was identified in BC tissues and cell lines, and CASC9 was correlated with the staging and metastasis in BC. Functionally, knockdown of CASC9 inhibited the proliferation, migration, and invasion of BC cells, as well as inhibited tumor growth in vivo. These findings suggested that CASC9 acts as an oncogene in BC progression and targeting CASC9 may be a potential therapeutic approach for BC.
Our research has been identified the oncogenic role of CASC9 in BC, however, the mechanism for upstream regulation of CASC9 involved in BC progression is still almost blank. As with the protein-coding transcripts, the transcription of lncRNAs is also controlled by the transcriptional factors.28,29 As an example, lncRNA HAS2-AS1 was upregulated by transcription factor cAMP responsive element-binding protein1 in epithelial ovarian cancer, and then promoted the tumorigenesis of epithelial ovarian cancer by regulating miR-466/RUNX family transcription factor 2 axis.30 Recently, STAT3 has been reported to play an important role in various malignancies.31 Remarkably, besides its traditional role in tumorigenesis, STAT3 as a transcriptional factor participates in carcinogenesis by regulating the transcription of lncRNAs.32 For instance, the upregulation of lncRNA double homeobox A pseudogene 8 induced by STAT3 enhanced cell proliferation, migration, and invasion and induced cell apoptosis in HCT116 and LOVO cells by regulating miR-577/ras-related protein 14 axis.33 In gastric cancer, STAT3 reportedly upregulated the expression of lncRNA HAGLROS, which could increase the proliferation, migration, and invasion of AGS cells via mTOR signal-mediated inhibition of autophagy.34 Herein, STAT3 was upregulated in BC tissues and cell lines, indicating the oncogenic role of STAT3 in BC. Moreover, STAT3 bound with the promoter region of CASC9 and negatively regulated the expression of CASC9, illustrating the upstream and origination of CASC9 in BC.
A great many of lncRNAs has been documented to interact with chromatin modifying enzymes to induce epigenetic activation or silencing of gene expression.35 Herein, we found that CASC9 could bind to EZH2, which serves as a key regulator in a range of cellular functions. Previous studies have been shown that EZH2 can epigenetically modulate the expression of PTEN through catalyzing trimethylation of histone H3 lysine 27.36–38 In addition, lncRNAs were shown to regulate tumorigenesis through epigenetically silencing PTEN via EZH2.39 As an example, Sox2 overlapping transcript could bind to PTEN, thereby epigenetically silencing PTEN expression, thus facilitating the progression of laryngeal squamous cell carcinoma.40 In accordance with the above statements, it seems likely that CASC9 may regulate the tumorigenesis of BC through regulating PTEN via EZH2. Our mechanistic experiments revealed that CASC9 could silence PTEN expression by interacting with EZH2. Furthermore, PTEN inhibition and upregulation of EZH2 could reverse CASC9 silencing-mediated inhibition of BC progression, supporting the idea that upregulation of CASC9 induced by STAT3 promoted the progression of BC by interacting with EZH2 and affecting the expression of PTEN. Similarly, a previous study in HepG2 and LM3 cells demonstrated that CASC11 induced by STAT3 promoted cell metastasis and epithelial-mesenchymal transition by epigenetically silencing PTEN via EZH2.41 Interestingly, Gan et al38 suggested that EHZ2 binds to PTEN promoter to regulate the PTEN expression. Here, we demonstrated that CASC9 can bind to EZH2, thus regulating the PTEN expression.
Conclusions
Collectively, our data revealed that CASC9, induced by STAT3, functioned as an oncogenic role in BC by interacting with EZH2 and affecting the expression of PTEN, which provided a novel mechanism for BC progression. Our findings suggested that CASC9 might be a potential therapeutic target for BC.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Czerniak B, Dinney C, McConkey D. McConkey D Origins of Bladder Cancer. Annu Rev Pathol. 2016;11(1):149–174. doi:10.1146/annurev-pathol-012513-104703
2. Fajkovic H, Halpern JA, Cha EK, et al. Impact of gender on bladder cancer incidence, staging, and prognosis. World J Urol. 2011;29(4):457–463. doi:10.1007/s00345-011-0709-9
3. Antoni S, Ferlay J, Soerjomataram I, et al. Bladder Cancer Incidence and Mortality: A Global Overview and Recent Trends. Eur Urol. 2017;71(1):96–108. doi:10.1016/j.eururo.2016.06.010
4. Masaoka H, Matsuo K, Ito H, et al. Cigarette smoking and bladder cancer risk: an evaluation based on a systematic review of epidemiologic evidence in the Japanese population. Jpn J Clin Oncol. 2016;46(3):273–283. doi:10.1093/jjco/hyv188
5. Turati F, Bosetti C, Polesel J, et al. Family history of cancer and the risk of bladder cancer: A case-control study from Italy. Cancer Epidemiol. 2017;48:29–35. doi:10.1016/j.canep.2017.03.003
6. Sun JW, Zhao LG, Yang Y, et al. Obesity and risk of bladder cancer: a dose-response meta-analysis of 15 cohort studies. PLoS One. 2015;10(3):e0119313. doi:10.1371/journal.pone.0119313
7. Volanis D, Kadiyska T, Galanis A, et al. Environmental factors and genetic susceptibility promote urinary bladder cancer. Toxicol Lett. 2010;193(2):131–137. doi:10.1016/j.toxlet.2009.12.018
8. Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66(2):115–132. doi:10.3322/caac.21338
9. Guidance, N.I.C.E. Bladder cancer: diagnosis and management of bladder cancer: (c) NICE (2015) Bladder cancer: diagnosis and management of bladder cancer. BJU Int. 2017;120(6):755–765. doi:10.1111/bju.14045
10. Batista PJ, Chang H. Chang HY Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152(6):1298–1307. doi:10.1016/j.cell.2013.02.012
11. Rinn JL, Kertesz M, Wang JK, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007;129(7):1311–1323. doi:10.1016/j.cell.2007.05.022
12. Guttman M, Rinn JL. Rinn JL Modular regulatory principles of large non-coding RNAs. Nature. 2012;482(7385):339–346. doi:10.1038/nature10887
13. Schmitt AM, Chang HY. Chang HY Long Noncoding RNAs in Cancer Pathways. Cancer Cell. 2016;29(4):452–463. doi:10.1016/j.ccell.2016.03.010
14. Jiang MC, Ni JJ, Cui WY, et al. Emerging roles of lncRNA in cancer and therapeutic opportunities. Am J Cancer Res. 2019;9(7):1354–1366.
15. Chi Y, Wang J, Wang J, et al. Long Non-Coding RNA in the Pathogenesis of Cancers. Cells. 2019;8(9):9. doi:10.3390/cells8091015
16. Hu X, Li Y, Kong D, et al. Long noncoding RNA CASC9 promotes LIN7A expression via miR-758-3p to facilitate the malignancy of ovarian cancer. J Cell Physiol. 2019;234(7):10800–10808. doi:10.1002/jcp.27903
17. Pan Z, Mao W, Bao Y, et al. The long noncoding RNA CASC9 regulates migration and invasion in esophageal cancer. Cancer Med. 2016;5(9):2442–2447. doi:10.1002/cam4.770
18. Gao GD, Liu XY, Lin Y, et al. LncRNA CASC9 promotes tumorigenesis by affecting EMT and predicts poor prognosis in esophageal squamous cell cancer. Eur Rev Med Pharmacol Sci. 2018;22(2):422–429. doi:10.26355/eurrev_201801_14191
19. Chen QN, Chen X, Chen ZY, et al. Long Intergenic Non-Coding RNA 00152 Promotes Lung Adenocarcinoma Proliferation via Interacting With EZH2 and Repressing IL24 Expression. Mol Cancer. 2017;16(1):17. doi:10.1186/s12943-017-0581-3
20. Zhang F, Peng H. LncRNA-ANCR Regulates the Cell Growth of Osteosarcoma by Interacting With EZH2 and Affecting the Expression of p21 and p27. J Orthop Surg Res. 2017;12(1):103. doi:10.1186/s13018-017-0599-7
21. Gulia C, Baldassarra S, Signore F, et al. Role of Non-Coding RNAs in the Etiology of Bladder Cancer. Eur urol. 2017;8(11). doi:10.3390/genes8110339
22. Pop-Bica C, Gulei D, Cojocneanu-Petric R, et al. Understanding the Role of Non-Coding RNAs in Bladder Cancer: from Dark Matter to Valuable Therapeutic Targets. Genes. 2017;18:7. doi:10.3390/genes8110339
23. Terracciano D, Ferro M, Terreri S, et al. Urinary long noncoding RNAs in nonmuscle-invasive bladder cancer: new architects in cancer prognostic biomarkers. Transl Res. 2017;184:108–117. doi:10.3390/ijms18071514
24. Zhang J, Wang Q, Quan Z. Long non-coding RNA CASC9 enhances breast cancer progression by promoting metastasis through the meditation of miR-215/TWIST2 signaling associated with TGF-beta expression. Biochem Biophys Res Commun. 2019;515(4):644–650. doi:10.1016/j.bbrc.2019.05.080
25. Luo K, Geng J, Zhang Q, et al. LncRNA CASC9 interacts with CPSF3 to regulate TGF-beta signaling in colorectal cancer. J Exp Clin Cancer Res. 2019;38(1):249. doi:10.1186/s13046-019-1263-3
26. Zhou J, Xiao H, Yang X, et al. Long noncoding RNA CASC9.5 promotes the proliferation and metastasis of lung adenocarcinoma. Sci Rep. 2018;8(1):37. doi:10.1038/s41598-017-18280-3
27. Sassenberg M, Droop J, Schulz WA, et al. Upregulation of the long non-coding RNA CASC9 as a biomarker for squamous cell carcinoma. BMC Cancer. 2019;19(1):806. doi:10.1186/s12885-019-6021-6
28. Chang S, Sun L, Feng G. Feng G SP1-mediated long noncoding RNA POU3F3 accelerates the cervical cancer through miR-127-5p/FOXD1. Biomed Pharmacother. 2019;117:109133. doi:10.1016/j.biopha.2019.109133
29. Wang C, Yang Y, Zhang G, et al. Long noncoding RNA EMS connects c-Myc to cell cycle control and tumorigenesis. Proc Natl Acad Sci U S A. 2019;116(29):14620–14629. doi:10.1073/pnas.1903432116
30. Tong L, Wang Y, Ao Y, et al. CREB1 induced lncRNA HAS2-AS1 promotes epithelial ovarian cancer proliferation and invasion via the miR-466/RUNX2 axis. Biomed Pharmacother. 2019;115:108891. doi:10.1016/j.biopha.2019.108891
31. Avalle L, Camporeale A, Camperi A, et al. STAT3 in cancer: A double edged sword. Cytokine. 2017;98:42–50. doi:10.1016/j.cyto.2017.03.018
32. Yuan J, Zhang F, Niu R. Niu R Multiple regulation pathways and pivotal biological functions of STAT3 in cancer. Sci Rep. 2015;5(1):17663. doi:10.1038/srep17663
33. Du C, Wang HX, Chen P, et al. STAT3-induced upregulation of lncRNA DUXAP8 functions as ceRNA for miR-577 to promote the migration and invasion in colorectal cancer through the regulation of RAB14. Eur Rev Med Pharmacol Sci. 2019;23(14):6105–6118. doi:10.26355/eurrev_201907_18424
34. Chen JF, Wu P, Xia R, et al. STAT3-induced lncRNA HAGLROS overexpression contributes to the malignant progression of gastric cancer cells via mTOR signal-mediated inhibition of autophagy. Mol Cancer. 2018;17(1):6. doi:10.1186/s12943-017-0756-y
35. Marchese FP, Huarte M. Huarte M Long non-coding RNAs and chromatin modifiers: their place in the epigenetic code. Epigenetics. 2014;9(1):21–26. doi:10.4161/epi.27472
36. Cao R, Wang L, Wang H, et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science. 2002;298(5595):1039–1043. doi:10.1126/science.1076997
37. Jarome TJ, Perez GA, Hauser RM, et al. EZH2 Methyltransferase Activity Controls Pten Expression and mTOR Signaling during Fear Memory Reconsolidation. J Neurosci. 2018;38(35):7635–7648. doi:10.1523/jneurosci.0538-18.2018
38. Gan L, Xu M, Hua R, et al. The polycomb group protein EZH2 induces epithelial-mesenchymal transition and pluripotent phenotype of gastric cancer cells by binding to PTEN promoter. J Hematol Oncol. 2018;11(1):9. doi:10.1186/s13045-017-0547-3
39. Li W, Zhang T, Guo L, et al. Regulation of PTEN expression by noncoding RNAs. J Exp Clin Cancer Res. 2018;37(1):223. doi:10.1186/s13046-018-0898-9
40. Tai Y, Ji Y, Liu F, et al. Long noncoding RNA SOX2-OT facilitates laryngeal squamous cell carcinoma development by epigenetically inhibiting PTEN via methyltransferase EZH2. IUBMB Life. 2019;71(9):1230–1239. doi:10.1002/iub.2026
41. Han Y, Chen M, Wang A, et al. STAT3-induced upregulation of lncRNA CASC11 promotes the cell migration, invasion and epithelial-mesenchymal transition in hepatocellular carcinoma by epigenetically silencing PTEN and activating PI3K/AKT signaling pathway. Biochem Biophys Res Commun. 2019;508(2):472–479. doi:10.1016/j.bbrc.2018.11.092
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.