Back to Journals » International Journal of Nanomedicine » Volume 16
Selenium Nanoparticles by Moderating Oxidative Stress Promote Differentiation of Mesenchymal Stem Cells to Osteoblasts
Authors Fatima S , Alfrayh R , Alrashed M, Alsobaie S , Ahmad R, Mahmood A
Received 7 October 2020
Accepted for publication 27 November 2020
Published 13 January 2021 Volume 2021:16 Pages 331—343
DOI https://doi.org/10.2147/IJN.S285233
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Thomas Webster
Sabiha Fatima,1 Rawan Alfrayh,1 May Alrashed,2 Sarah Alsobaie,1 Rehan Ahmad,3 Amer Mahmood4
1Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, King Saud University, Riyadh 11433, Saudi Arabia; 2Chair of Medical and Molecular Genetics Research, Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, King Saud University, Riyadh 11433, Saudi Arabia; 3Colorectal Research Chair, Department of Surgery, King Saud University, College of Medicine, Riyadh 11472, Saudi Arabia; 4Stem Cell Unit, Department of Anatomy, College of Medicine, King Khalid University Hospital, King Saud University, Riyadh 11461, Kingdom of Saudi Arabia
Correspondence: Sabiha Fatima
Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, King Saud University, Riyadh, Saudi Arabia
Tel +966-11-8052621
Fax +966-11-4693738
Email [email protected]
Amer Mahmood
Stem Cell Unit, Department of Anatomy, College of Medicine, King Saud University, Riyadh 11461, Kingdom of Saudi Arabia
Tel +966-11-4679421
Fax +966-11-4693738
Email [email protected]
Purpose: Redox homeostasis plays an important role in the osteogenic differentiation of human mesenchymal stem cells (hMSCs) for bone engineering. Oxidative stress (OS) is believed to induce osteoporosis by changing bone homeostasis. Selenium nanoparticles (SeNPs), an antioxidant with pleiotropic pharmacological activity, prevent bone loss. However, the molecular mechanism underlying the osteogenic activity during hMSC–SeNP interaction is unclear.
Methods: This study assessed the effects of different concentrations (25, 50, 100, and 300 ng/mL) of SeNPs on the cell viability and differentiation ability of human embryonic stem cell-derived hMSCs. In addition, we analyzed OS markers and their effect on mitogen-activated protein kinase (MAPK) and Forkhead box O3 (FOXO3) during osteogenesis.
Results: SeNPs increased the cell viability of hMSCs and induced their differentiation toward an osteogenic over an adipogenic lineage by enhancing osteogenic transcription and mineralization, while inhibiting Nile red staining and adipogenic gene expression. By preventing excessive reactive oxygen species accumulation, SeNPs increased antioxidant levels in hMSCs undergoing osteogenesis compared to untreated cells. In addition, SeNPs significantly upregulated the gene and protein expression of phosphorylated c-Jun N-terminal kinase (JNK) and FOXO3a, with no significant change in the expression levels of extracellular signal-related kinase (ERK) and p38 MAPK.
Conclusion: The results approved that low concentrations of SeNPs might enhance the cell viability and osteogenic potential of hMSCs by moderating OS. Increased JNK and FOXO3a expression shows that SeNPs might enhance osteogenesis via activation of the JNK/FOXO3 pathway. In addition, SeNP co-supplementation might prevent bone loss by enhancing osteogenesis and, thus, can be an effective candidate for treating osteoporosis through cell-based therapy.
Keywords: selenium nanoparticles stem cells, antioxidant, osteogenic differentiation
Introduction
Osteoporosis is one of the most common bone diseases related to aging. It is characterized by structural deterioration of bone tissue.1 Oxidative stress (OS) plays an important role in decreasing osteogenic differentiation in cell lines and the bone mineral density during age-related osteoporosis.2,3 Plasma antioxidant levels significantly decrease in women with age-related osteoporosis.4 Mesenchymal stem cells (MSCs) can induce repair processes at sites of injury in vivo by replenishing bone cells, including chondrocytes and osteoblasts.5,6 The infinite self-renewal capacity and greater differentiation potential of pluripotent embryonic stem cells (ESCs) have enabled their application in biology and medicine.7–9 For many clinical applications, in vitro expansion of MSCs is required to obtain high yields of functionally viable cells before transplantation.10 Unfortunately, the in vitro expansion of human mesenchymal stem cells (hMSCs) in long-term cell culture leads to a gradual decline in the viability and multipotency of MSCs, which is a major obstacle in cell-based therapy.11,12
In long-term cell culture, OS is a potential source of cellular damage, affecting the in vitro expansion and longevity of MSCs, with a decreased number of functionally viable cells for clinical applications.13,14 During cell differentiation, increased mitochondrial biogenesis and a metabolic shift toward oxidative phosphorylation change the redox status and spontaneously increase reactive oxygen species (ROS) generation.15 Many tissue regeneration–based studies have reported that the most transplanted MSCs undergo OS due to excessive ROS generated by either host tissues or the MSCs themselves.16 After several days, this increase in ROS levels leads to cell cycle arrest and the death of transplanted MSCs, and only a small percentage of transplanted MSCs survive, which is insufficient for replacing lost tissue.17 High ROS levels can lead to cellular damage and dysfunction, while physiological/moderate ROS levels can improve cell survival, proliferation, and differentiation.18 Therefore, ROS regulation by an antioxidant can improve the viability and differentiation of MSCs, which might augment the MSC treatment efficiency in reversing age-related bone loss by osteogenic progenitors.19,20
Selenium (Se) is an essential trace element that protects cells against oxidative damage. Se is also required for regulating bone cell proliferation and differentiation.21 Nanosized Se is a highly effective molecular compound with higher antioxidant activity and lower toxicity compared to ordinary Se.22 Although the potential use of nanomaterials in regulating MSC differentiation is well understood, the role of selenium nanoparticles (SeNPs) in governing MSC differentiation toward an osteogenic lineage is unclear. Therefore, this study investigated the effect of SeNPs on the cell viability and osteogenesis of human embryonic stem cell (hESC)-derived hMSCs. We also analyzed the effect of SeNPs on intracellular OS markers. In addition, we investigated the expression of the OS-regulated mitogen-activated protein kinases (MAPKs) p38, c-Jun N-terminal kinase (JNK), and extracellular signal-related kinase (ERK), in addition to Forkhead box O3a (FOXO3a), during osteogenesis.
Materials and Methods
Cell Culture
We purchased hESC-derived hMSCs (P3) from Merck Millipore (Burlington, MA, USA) and cultured them in stromal medium containing Dulbecco’s Modified Eagle Medium (DMEM; Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 10% fetal bovine serum (FBS; Thermo Fisher Scientific), 1x penicillin-streptomycin (Thermo Fisher Scientific), 110 mg/L of sodium pyruvate (Thermo Fisher Scientific), 4 mM L-glutamine (Thermo Fisher Scientific), and 4500 mg/L of D-glucose (Thermo Fisher Scientific) in a humidified tissue culture incubator at 37°C in a 5% CO2 atmosphere. The cultured hESCs were used for experiments after 80% confluence.
Characterization of SeNPs
SeNPs (5–50 nm) were purchased from Nanocs Inc. (Boston, MA, USA), separated by centrifugation, and redispersed in aqueous medium by sonication. Their shape and size were examined using transmission electron microscopy (TEM) at 200 kV (JEM-2100F; JEOL, Tokyo, Japan). In addition, their fingerprint or composition was measured by Fourier transform infrared (FTIR) spectroscopy in the wavenumber range of 4000–400 cm–1.
Uptake of SeNPs by MSCs Visualized Through Fluorescent Microscopy
To observe the cellular uptake of SeNPs, we incubated hMSCs with fluorescently labeled SeNPs for 4 h and then prefixed them with 2.5% glutaraldehyde in phosphate-buffered saline (PBS) at 4°C overnight. Subsequently, we washed the cells thrice with PBS, postfixed them for 45 min with 1% osmium dissolved in PBS, and washed them by dipping them in water for 10 min. Once the cells were dehydrated, we observed them under a fluorescent microscope to see in the internalization of SeNPs by MSCs.
Cell Viability Assay
We seeded hMSCs in 96-well culture plates at a density of 2 × 104 cells/well and incubated them for 9 days. After incubation, we added different concentrations (25, 50, 100, and 300 ng/mL) of SeNPs to the wells and further incubated the cells for 3, 7, and 9 days. Untreated cells were used as the control group. Next, we measured the cell viability on days 0, 3, 7, and 9 using alamarBlue assay (Bio-Rad AbD Serotec, USA). Briefly, we added 10 μL of alamarBlue substrate to the cultured hMSCs and incubated the plates in the dark at 37°C for 2 h. We measured fluorescence levels at excitation and emission wavelengths of 530 and 590 nm, respectively. The fluorescence intensity was proportional to the number of viable cells.
Differentiation of hMSCs into an Osteogenic Lineage
We performed alkaline phosphatase (ALP) activity and alizarin red (AR)-staining assays to assess the osteogenic differentiation of hMSCs. To induce osteogenesis, we cultured ~1 × 104 hMSCs in osteogenic induction supplement (OIS) containing 10% FB, 1% penicillin-streptomycin, 50 μg/mL of L-ascorbic acid (Wako Chemicals, Germany), 10 mM β-glycerophosphate, and 10 nM dexamethasone (Sigma-Aldrich, St. Louis, MO, USA). We assayed five experimental groups for ALP staining: (i) hMSCs cultured in OIS only (control group); (ii) hMSCs cultured in OIS + 25 ng/mL of SeNPs (group 1); (iii) hMSCs cultured in OIS + 50 ng/mL of SeNPs (group 2); (iv) hMSCs cultured in OIS + 100 ng/mL of SeNPs (group 3); and (v) hMSCs cultured in OIS + 300 ng/mL of SeNPs (group 4). In groups 1–4, we quantified the ALP activity after 7 days of cell differentiation using the ALP activity colorimetric assay kit (BioVision, Inc, CA, USA) according to the manufacturer’s instructions. Finally, absorbance was measured at a wavelength of 405 nm.
We detected mineralized matrix nodule formation in hMSCs cultured in OIS with or without different concentrations of SeNPs by performing AR staining; hMSCs cultured in OIS without SeNPs were the control group. Briefly, hMSCs were seeded in a 48-well tissue culture plate at a density of 5 × 106 cells/well, containing OIS and OIS + SeNPs (25, 50, 100, and 300 ng/mL) for 14 days. After 14 days, the cells were washed with PBS, fixed in 70% ice-cold ethanol at room temperature for 1 h, and stained with the filtered Alizarin red S (ARS) Staining Kit (ScienCell Research Laboratories, Carlsbad, CA, USA) at room temperature for 20 min. Next, the excess dye was washed away with distilled water, and images of stained mineral deposits were taken using an inverted light microscope. To quantify the ARS staining, we eluted the dye with 10% (w/v) cetylpyridinium chloride solution (Sigma-Aldrich) at room temperature for 20 min and measured the optical density (OD) at a wavelength of 570 nm. Finally, the mineralization rate was calculated as a percentage using the following equation:
[(ODsample – ODcontrol)/ODcontrol] × 100
Detection and Quantification of Adipogenesis
We assessed the adipogenic differentiation of hMSCs in the absence and presence of SeNPs using Nile red staining, which is used to detect intracellular lipid droplets. We added adipogenetic induction supplements, including 10% horse serum, 10% FBS (Sigma-Aldrich), 1% penicillin-streptomycin, 100 nM dexamethasone, 0.45 mM isobutyl methyl xanthine (Sigma-Aldrich), 3 μg/mL of insulin (Sigma-Aldrich), and SeNPs (25, 50, 100, or 300 ng/mL), to the culture medium. After 14 days of adipogenic induction, we washed the hMSCs with PBS, added Nile red directly to the cells (5 μg/mL in acetone), and incubated the cells at room temperature for 10 min. After staining, the excess dye was washed away with PBS, and images of stain-detecting intracellular lipid droplets were taken using a fluorescent microscope. To quantify Nile red fluorescent staining, we measured the signal using a Spectra Max/M5 fluorescence spectrophotometer plate reader (Molecular Devices, Sunnyvale, CA, USA) at excitation and emission wavelengths of 485 and 572 nm, respectively. Finally, the adipogenic differentiation was calculated as a percentage using the following equation:
[(ODsample – ODcontrol)/ODcontrol] × 100
Quantitative, Real-Time Reverse Transcription Polymerase Chain Reaction
After 18 days of osteogenic and adipogenic induction of hMSCs incubation with or without 50 ng/mL of SeNPs, the cells were lysed and total RNA was extracted using the innuPREP RNA Mini Kit (Analytik Jena AG, Jena, Germany). Next, we reverse-transcribed 1 μg of RNA into complementary DNA (cDNA) using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) and a Labnet Multigene thermal cycler (Labnet International Inc., Edison, NJ, USA) according to the manufacturer’s instructions. To measure the expression of different osteogenic pathway–focused genes, including ALP, Runt-related transcription factor 2 (RUNX2), type I collagen (COL1), osteocalcin (OCN), osteopontin (OPN), cadherin-11 (CDH11), and osteonectin (SPARC), the samples were then subjected to quantitative, real-time reverse transcription polymerase chain reaction (RT-PCR) analysis using the Power SYBR® Green RNA-to-CT™ 1-Step Kit (Applied Biosystems). In addition, we detected the gene expression of the master regulator for adipogenesis, peroxisome proliferator-activated receptor-gamma (PPAR-γ), and CCAAT/enhancer-binding protein α (C/EBPα). We also analyzed the messenger RNA (mRNA) expression of MAPK and FOXO3 to check its consistency with protein expression. Finally, we expressed the relative expression of the genes of interest normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) as a fold-change. Supplementary Table 1 lists the primer sequences used in this study.
Estimation of Oxidative Stress Markers
After 3 days of osteogenic differentiation, we measured the intracellular ROS levels using flow cytometry (FACSCalibur; Becton Dickinson, San Jose, CA, USA) on the basis of the fluorescence-associated oxidation of 2′,7′-dichlorodihydrofluorescein (DCFH) to 2′,7′-dichlorofluorescein. Briefly, we added 1.0 μM 2′,7′-dicholorodihydrofluorescein diacetate (DCF-DA) to harvested hMSCs and incubated them at 37°C for 15 min. Next, the cells were washed with PBS, and the ROS levels in the incubated samples were estimated using flow cytometry (BD Biosciences, San Jose, CA, USA). Data were acquired and analyzed using CellQuest pro ver 6.0 (excitation at 488 nm and detection at 535 nm). We also detected reduced glutathione (GSH) levels using the Glutathione Colorimetric Assay Kit (BioVision Inc., CA, USA). Finally, we analyzed superoxide dismutase (SOD) and catalase activities using the Superoxide Dismutase Activity Assay Kit and the Catalase Activity Colorimetric/Fluorometric Assay Kit (Abcam, MA, USA), respectively, according to the manufacturer’s instructions.
Detection of MAPK and FOXO3a Expression by Western Blotting
We performed Western blotting to study the protein expression levels of p-p38, p-ERK1/2, p-JNK, and p-FOXO3a in hMSCs after incubation with or without 50 ng/mL of SeNPs during osteogenic differentiation. After 3 days of osteogenic induction, the hMSCs were harvested and washed twice with PBS. Next, cell lysates were prepared in radioimmunoprecipitation assay (RIPA) lysis buffer (Boston Bioproducts, Ashland, MA, USA), the mixture was centrifuged at 14,000 rpm for 15 min, and the supernatant was collected. Subsequently, 10–20 μg of protein estimated using Bradford protein assay was loaded on electrophoresis gels, the separated proteins were transferred to a 0.2 μm polyvinylidene difluoride (PVDF) membrane using a turbo protein transfer system (Bio-Rad Laboratories, Hercules, CA, USA), and the blots were blocked using Sea Block blocking buffer (Thermo Fisher Scientific) at room temperature for 1 h. The PVDF membrane was washed with PBS containing 0.1% Tween-20 (PBST) and incubated overnight at 4°C with primary antibodies p-p38, p-ERK1/2, p-JNK, p-FOXO3a (Abcam, MA, USA), and β-actin (Santa Cruz Biotechnology, CA, USA). Next, the blots were washed and then incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies at 25°C for 1 h. Finally, the signals were detected using the Pierce enhanced chemiluminescence (ECL) detection system (Thermo Fisher Scientific), and image analysis was performed using ImageLab software (Bio-Rad Laboratories).
Statistical Analysis
All data were expressed as means ± standard deviation (SD). We performed analysis of variance (ANOVA) to test differences between control and experimental groups using GraphPad Prism7 (GraphPad Software Inc., CA, USA). P < 0.05 was considered statistically significant.
Results
Characterization and Cellular Uptake of SeNPs
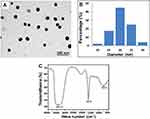
The SeNPs were characterized to check their stability in terms of their shape, size, and composition. The TEM image (Figure 1A) showed a clear, spherical morphology of the SeNPs, with an average diameter of 20 ± 5.6 nm (Figure 1B). The FTIR spectrum, which represented symmetric and asymmetric vibration bands of the SeNPs, showed a broad peak at 3200–3500 cm–1, indicating the presence of the amide (–NH) group, while a strong peak at 1637 cm–1 corresponded to the C=O group.23 The peak at 767.5 cm–1 was related to the stretching mode of the metal peaks of Se (Figure 1C).
The fluorescence microscopic image of hMSCs cultured in the absence of SeNPs did not show internalized SeNPs (Figure 2A). In contrast, in hMSCs cultured in the presence of 50 ng of SeNPs, the SeNPs were localized inside the hMSCs and not merely attached to their surface (Figure 2B and C). We also visualized agglomerated SeNPs in a few vesicles in the cytoplasm (Figure 2C).
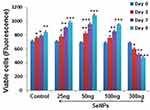
SeNPs Increase Cell Viability
Figure 3 shows the fluorescence intensities of untreated (control) and SeNP-treated hMSCs. We observed higher cell viability in hMSCs treated with 25, 50, and 100 ng of SeNPs, while the cell viability significantly decreased at a concentration of 300 ng compared to the control group (P < 0.001). In addition, hMSCs treated with 50 ng of SeNPs showed the highest cell viability (P < 0.001) compared to hMSCs treated with 25 and 100 ng of SeNPs.
SeNPs Induce Osteogenic Differentiation
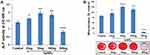
We determined the effect of SeNPs on osteogenic differentiation of hMSCs cultured for 7 days by measuring their ALP activity in order to evaluate the expression of an early phenotypic marker for osteogenesis (Figure 4A). MSCs treated with SeNPs 25, 50, 100 ng had higher ALP activities than the control group. The results revealed a considerable (p<0.001) increase of ALP activity in MSCs treated with SeNPs at a concentration of 50 ng. In contrast, ALP activity in MSCs treated with 300 ng of SeNPs significantly decreased compared to the control group.
Mineralized matrix nodule formation, a phenotypic marker of the last stage of mature osteoblasts, was analyzed to assess the progression of hMSCs to an osteogenic lineage. Calcium deposits can specifically be stained a bright orange-red by AR staining. After 14 days of osteogenic induction, hMSCs treated with 25, 50, 100 ng showed increased mineralization nodule formation compared to the control group. In contrast, the cells treated with 300 ng significantly suppressed mineralization nodule formation in hMSCs undergoing osteogenesis (Figure 4B). This indicates that lower concentrations of SeNPs promoted successful hMSC differentiation into osteoblasts. Treatment with SeNPs at a dosage of 50 ng showed maximum (p<0.001) mineralized nodule formation, compared to the control group.
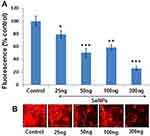
SeNPs Inhibit Adipogenic Differentiation
Figure 5A and B show the concentration-dependent effect of SeNPs on adipogenic differentiation of hMSCs in the presence of adipogenic induction medium, as determined by staining intracytoplasmic lipids with Nile red. SeNPs significantly inhibited adipogenic differentiation of hMSCs compared to the control group. We observed significant inhibition of adipogenic differentiation at a SeNP concentration of 50 ng on day 14 and more decreased adipocyte formation at a SeNP concentration of 300 ng compared to the untreated control group.
SeNPs Lower Oxidative Stress During Osteogenesis
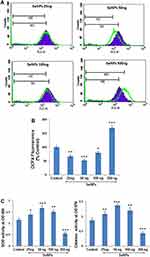
ROS levels can be used to evaluate the concentration-dependent free-radical-scavenging effect of SeNPs on hMSCs undergoing osteogenic differentiation. In this assay, we measured the intracellular levels of ROS using DCFH-DA in hMSCs with or without SeNP treatment. DCF fluorescence significantly decreased at SeNP concentration of 25, 50, and 100 ng, indicating that ROS generation decreases compared to the control group (Figure 6B). The shift in the histogram (green line) toward the right indicated enhanced ROS scavenging at a SeNP concentration of 50 ng (Figure 6A). In contrast, ROS levels significantly increased at a SeNP concentration of 300 ng compared to control group (P < 0.001) (Figure 6A and B).
In addition, lower SeNP concentrations significantly enhanced the level of enzymatic antioxidants, such as SOD and catalase. MSCs treated with 50 ng of SeNPs showed the maximum increase in antioxidant activity compared to the control group (P < 0.001), while hMSCs treated with a SeNP concentration of 300 ng significantly inhibited the activity of the enzymatic antioxidants (P < 0.001) (Figure 6C).
SeNPs Enhance Osteogenic- and Inhibit Adipogenic-Specific Gene Expression
On the basis of the concentrated-dependent protective effect of SeNPs, we selected a SeNP concentration of 50 ng/mL for subsequent experiments. Specific transcription factors regulate the interconversion of hMSCs to a particular lineage with repression of alternative phenotypes. ALP, RUNX2, and COL1a1 determine the osteoblast lineage from pluripotent MSCs, while PPAR-γ and C/EBPα induce hMSCs to undergo adipocyte differentiation. RT-PCR results showed significantly increased ALP, RUNX2, and COL1a1 expression and significantly attenuated PPAR-γ and C/EBPα expression compared to the control group (Figure 7), which is consistent with our observations in cellular assays, indicating that SeNPs preferentially favor hMSC differentiation into an osteogenic over an adipogenic lineage. We also performed RT-PCR to test the transcriptional profile of proteins, such as OCN, OPN, CDH11, and SPARC that significantly contribute to osteoblast differentiation and bone formation. SeNPs markedly upregulated all osteogenesis-promoting genes examined. The transcriptional profile of osteogenic marker genes clearly indicated the effect of SeNPs on osteogenic differentiation, as observed by increased ALP activity (Figure 4A) and mineralized matrix nodule formation (Figure 4B).
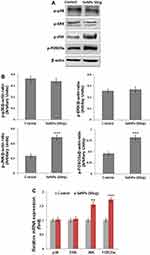
SeNPs Modulate JNK and FOXO3a Expression
We performed Western blotting to evaluate the effect of SeNP treatment on the key proteins of the MAPK pathway and FOXO3a expression during osteogenesis. We analyzed the expression MAPK signaling proteins, including p38, JNK, and ERK (Figure 8A). SeNP treatment significantly increased p-JNK and p-FOXO3a expression, while p-ERK and p-p38 expression did not substantially change compared to the control group (Figure 8B).
In addition, we performed RT-PCR to study the effect of SeNPs on MAPK and FOXO3a gene expression. The transcriptional profile was consistent with protein expression levels (Figure 8C).
Discussion
In this study, we investigated the effect of supplementation of SeNPs as a potential means to promote osteogenesis of hESC-derived hMSCs in vitro. The hMSCs have a controlled redox status to manage OS. However, in vitro expansion and differentiation disturb the redox status of proliferating hMSCs.24–26 Antioxidants, such as SOD and glutathione peroxidase, which are expressed in high levels in hESCs, are downregulated during osteogenic differentiation.24 Therefore, in addition to the metabolic shift toward oxidative phosphorylation, antioxidant downregulation might also be associated with increased intracellular ROS levels during osteogenic differentiation.
Culturing hMSCs with a low SeNP concentration significantly inhibits ROS levels and increases the antioxidant activity within the cells. Decreased OS is associated with improved cell viability and enhanced differentiation of hMSCs to an osteogenic over an adipogenic lineage. There is an inverse relationship between osteogenic and adipogenic differentiation.27 This inverse relationship is indicated by the concurrent upregulation of RUNX-2 (a master regulator of osteogenesis) and downregulation of the adipogenic marker PPAR-γ (which induces cells to differentiate into adipocytes).28–30 In addition, supplementation with a high (300 ng/mL) SeNP concentration results in excessive ROS generation with compromised antioxidant defense and decreased cell viability. These findings indicate that SeNPs at supraphysiological levels might change from antioxidants to pro-oxidants, which might induce cytotoxicity.31 Therefore, increased cell viability and osteogenesis at low SeNP concentrations could be due to regulation of ROS to controlled or moderate levels in cell culture and the commitment to an osteogenic lineage, with inhibited alternative phenotypes. In addition, increased OS favors hMSC differentiation to adipocytes, indicating that the OS level determines the fate of the cell.32
The positive effect of low SeNPs concentrations on osteogenesis is indicated by a significant increase in the activity and gene expression of ALP (an early osteogenic marker) and mineralized matrix nodule formation (a phenotypic marker of mature osteoblasts).33 Enhanced gene expression of bone matrix proteins, such as COL1a1, OPN, OCN, and CDH11, which mediate cell–cell adhesion, further indicates osteoblast lineage commitment and differentiation.34 Overall, the transcriptional profile of osteogenic marker genes clearly illustrates the positive effect of SeNPs in enhancing osteogenesis, also supported by observations of enhanced ALP activity and mineralized matrix nodule formation (Figure 4A and B).
Incubating cells with nanomaterials in the culture medium enables serum proteins to adsorb to the nanomaterial surface, which induces nanomaterial entry into cells by receptor-mediated endocytosis.35 Light microscopy image shows the presence of fluorescence-labeled SeNPs in the cytoplasm. Once inside a cell, the SeNPs might interact with cytoplasmic proteins, which can regulate the fate of hMSCs through mechanical stimuli–activated signal transduction.9 On the basis of these findings, we hypothesized that the MAPK pathway, which is activated by mechanical stresses, might be affected during hMSC–SeNP interaction. To test this hypothesis, we assessed the involvement of the key proteins of the MAPK pathway by Western blotting and RT-PCR during hMSC–SeNP interaction. Increased expression of p-JNK, a stress-activated kinase, and no substantial change in p-ERK and p-p38 expression indicates that activation of the JNK signaling pathway might be involved in the preferential differentiation of hMSCs to osteoblasts. The JNK-MAPK pathway plays an important role in the OS-coordinated cellular response to different stresses, including ROS, which also play a positive role in modulating osteogenic differentiation.36 A mild oxidative environment can activate the pro-survival cellular mechanism of JNK-MAPKs to repair the damage before committing cells to death.37 The pro-survival function of JNK depends on the regulation of the antioxidant response. In response to low OS, the activated JNK survival signaling pathway mediates FOXO phosphorylation and translocation from the cytoplasm into the nucleus.38 As a transcription factor, FOXO not only enhances the synthesis of antioxidant enzymes Mn‐SOD and catalase but also binds to the promoters of RUNX2, ALP, and OCN and assists in commitment to an osteogenic lineage.39,40 Therefore, increased gene and protein expression of p-JNK and FOXO3a after SeNP treatment might lead to decreased ROS levels via induction of antioxidant enzymes. Consequently, activation of the JNK/FOXO3a pathway by enhancing cellular antioxidants might improve cell viability and enhance the osteogenic potential of hMSCs, as observed by the increased expression of bone marker proteins in SeNP-treated hMSCs compared to untreated hMSCs.
Conclusion
A low SeNP concentration (50 ng/mL) by decreasing OS plays an important role in enhancing the differentiation of hMSCs to an osteogenic lineage. Increased gene and protein expression of JNK and FOXO3a indicates the role of the JNK/FOXO3a pathway in SeNP-induced osteogenesis. This, in turn, indicates that SeNPs due to its antioxidative and osteoinductive properties can be used as potential therapeutic agents in treating osteoporosis through cell-based therapy.
Abbreviations
MSCs, mesenchymal stem cells; mRNA, messenger RNA; SeNPs, selenium nanoparticles; ESCs, embryonic stem cells; MAPK, mitogen-activated protein kinase; FOXO, Forkhead box O3; ROS, reactive oxygen species; ERK, extracellular signal-related kinase; JNK, c-Jun N-terminal kinase; OS, oxidative stress; hMSCs, human mesenchymal stem cells; DMEM, Dulbecco’s Modified Eagle Medium; FBS, fetal bovine serum; TEM, transmission electron microscopy; FTIR, Fourier transform infrared; PBS, phosphate-buffered saline; OIS, osteogenic induction supplement; ALP, alkaline phosphatase; AR, alizarin red; ARS, Alizarin red S; OD, optical density; cDNA, complementary DNA; RT-PCR, reverse transcription polymerase chain reaction; RUNX2, Runt-related transcription factor 2; COL1, type I collagen; OCN, osteocalcin; OPN, osteopontin; CDH11, cadherin-11; SPARC, osteonectin; C/EBPα, CCAAT/enhancer-binding protein α; PPAR-G, peroxisome proliferator-activated receptor-gamma; GAPDH, glyceraldehyde 3-phosphate dehydrogenase; DCFH, 2′,7′-dichlorodihydrofluorescein; DCF-DA, 2′,7′-dicholorodihydrofluorescein diacetate; GSH, glutathione; SOD, superoxide dismutase; SD, standard deviation; ANOVA, analysis of variance; RIPA, radioimmunoprecipitation assay; PVDF, polyvinylidene difluoride; HRP, horseradish peroxidase; ECL, enhanced chemiluminescence.
Acknowledgment
This research project was supported by a grant from the “Research Center of the Female Scientific and Medical Colleges”, Deanship of Scientific Research, King Saud University.
Author Contributions
All authors contributed to data analysis, drafting or revising the article, have agreed on the journal to which the article will be submitted, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Sözen T, Özışık L, Başaran NÇ. An overview and management of osteoporosis. Eur J Rheumatol. 2017;4(1):46–56. doi:10.5152/eurjrheum.2016.048
2. Domazetovic V, Marcucci G, Iantomasi T, Brandi ML, Vincenzini MT. Oxidative stress in bone remodeling: role of antioxidants. Clin Cases Miner Bone Metab. 2017;14(2):209–216. doi:10.11138/ccmbm/2017.14.1.209
3. Geng Q, Gao H, Yang R, Guo K, Miao D. Pyrroloquinoline quinone prevents estrogen deficiency-induced osteoporosis by inhibiting oxidative stress and osteocyte senescence. Int J Biol Sci. 2019;15(1):58–68. doi:10.7150/ijbs.25783
4. Maggio D, Barabani M, Pierandrei M, et al. Marked decrease in plasma antioxidants in aged osteoporotic women: results of a cross-sectional study. J Clin Endocrinol Metab. 2003;88(4):1523–1527. doi:10.1210/jc.2002-021496
5. Shao J, Zhang W, Yang T. Using mesenchymal stem cells as a therapy for bone regeneration and repairing. Biol Res. 2015;48:62. doi:10.1186/s40659-015-0053-4
6. Lin S, Lee WYW, Feng Q, et al. Synergistic effects on mesenchymal stem cell-based cartilage regeneration by chondrogenic preconditioning and mechanical stimulation. Stem Cell Res Ther. 2017;8(1):221.
7. Hass R, Kasper C, Bohm S, Jacobs R. Different populations and sources of human mesenchymal stem cells (MSC): a comparison of adult and neonatal tissue-derived MSC. Cell Commun Signal. 2011;9:12. doi:10.1186/1478-811X-9-12
8. Yoon SW, Kim DK, Kim KP, Park KS. Rad51 regulates cell cycle progression by preserving G2/M transition in mouse embryonic stem cells. Stem Cells Dev. 2014;23(22):2700–2711. doi:10.1089/scd.2014.0129
9. Wei M, Li S, Le W. Nanomaterials modulate stem cell differentiation: biological interaction and underlying mechanisms. J Nanobiotechnol. 2017;15(1):75. doi:10.1186/s12951-017-0310-5
10. Drela K, Stanaszek L, Nowakowski A, Kuczynska Z, Lukomska B. Experimental strategies of mesenchymal stem cell propagation: adverse events and potential risk of functional changes. Stem Cells Int. 2019;2019:7012692. doi:10.1155/2019/7012692
11. Wei X, Yang X, Han ZP, Qu FF, Shao L, Shi YF. Mesenchymal stem cells: a new trend for cell therapy. Acta Pharmacol Sin. 2013;34(6):747–754. doi:10.1038/aps.2013.50
12. Chen G, Yue A, Ruan Z, et al. Monitoring the biology stability of human umbilical cord-derived mesenchymal stem cells during long-term culture in serum-free medium. Cell Tissue Bank. 2014;15(4):513–521. doi:10.1007/s10561-014-9420-6
13. Alves H, Mentink A, Le B, van Blitterswijk CA, de Boer J. Effect of antioxidant supplementation on the total yield, oxidative stress levels, and multipotency of bone marrow-derived human mesenchymal stromal cells. Tissue Eng Part A. 2013;19:928–937. doi:10.1089/ten.tea.2011.0700
14. Denu RA, Hematti P. Effects of oxidative stress on mesenchymal stem cell biology. Oxid Med Cell Longev. 2016;2016:2989076. doi:10.1155/2016/2989076
15. Li Q, Gao Z, Chen Y, Guan MX. The role of mitochondria in osteogenic, adipogenic and chondrogenic differentiation of mesenchymal stem cells. Protein Cell. 2017;8(6):439–445. doi:10.1007/s13238-017-0385-7
16. Liu SP, Ding DC, Wang HJ, et al. Nonsenescent Hsp27-upregulated MSCs implantation promotes neuroplasticity in stroke model. Cell Transplant. 2010;19(10):1261–1279. doi:10.3727/096368910X507204
17. Yang SR, Park JR, Kang KS. Reactive oxygen species in mesenchymal stem cell aging: implication to lung diseases. Oxid Med Cell Longev. 2015;2015:486263. doi:10.1155/2015/486263
18. Lee J, Cho YS, Jung H, Choi I. Pharmacological regulation of oxidative stress in stem cells. Oxid Med Cell Longev. 2018;2018:4081890. doi:10.1155/2018/4081890
19. Muthusami S, Ramachandran I, Muthusamy B, et al. Ovariectomy induces oxidative stress and impairs bone antioxidant system in adult rats. Clin Chim Acta. 2005;360(1–2):81–86. doi:10.1016/j.cccn.2005.04.014
20. Shaban S, El-Husseny MW, Abushouk AI, Salem AMA, Mamdouh M, Abdel-Daim MM. Effects of antioxidant supplements on the survival and differentiation of stem cells. Oxid Med Cell Longev. 2017;2017:5032102. doi:10.1155/2017/5032102
21. Hosnedlova B, Kepinska M, Skalickova S, et al. Nano-selenium and its nanomedicine applications: a critical review. Int J Nanomedicine. 2018;13:2107–2128. doi:10.2147/IJN.S157541
22. Khurana A, Tekula S, Saifi MA, Venkatesh P, Godugu C. Therapeutic applications of selenium nanoparticles. Biomed Pharmacother. 2019;111:802–812. doi:10.1016/j.biopha.2018.12.146
23. Alkhudhayri A, Al-Shaebi EM, Qasem MAA, et al. Antioxidant and anti-apoptotic effects of selenium nanoparticles against murine eimeriosis. An Acad Bras Cienc. 2020;92(2):e20191107. doi:10.1590/0001-3765202020191107
24. Sart S, Song L, Li Y. Controlling redox status for stem cell survival, expansion, and differentiation. Oxid Med Cell Longev. 2015;2015:105135. doi:10.1155/2015/105135
25. Hu C, Zhao L, Peng C, Li L. Regulation of the mitochondrial reactive oxygen species: strategies to control mesenchymal stem cell fates ex vivo and in vivo. J Cell Mol Med. 2018;22(11):5196–5207. doi:10.1111/jcmm.13835
26. Dai X, Yan X, Wintergerst KA, Cai L, Keller BB, Tan Y. Nrf2: redox and metabolic regulator of stem cell state and function. Trends Mol Med. 2020;26(2):185–200. doi:10.1016/j.molmed.2019.09.007
27. Li J, Zuo B, Zhang L, Dai L, Zhang X. Osteoblast versus adipocyte: bone marrow microenvironment-guided epigenetic control. Case Rep Orthop Res. 2018;1(1):2–18. doi:10.1159/000489053
28. James AW. Review of signaling pathways governing MSC osteogenic and adipogenic differentiation. Scientifica. 2013;2013:684736. doi:10.1155/2013/684736
29. Atashi F, Modarressi A, Pepper MS. The role of reactive oxygen species in mesenchymal stem cell adipogenic and osteogenic differentiation: a review. Stem Cells Dev. 2015;24(10):1150–1163. doi:10.1089/scd.2014.0484
30. Ge C, Cawthorn WP, Li Y, Zhao G, Macdougald OA, Franceschi RT. Reciprocal control of osteogenic and adipogenic differentiation by ERK/MAP kinase phosphorylation of Runx2 and PPARγ Transcription Factors. J Cell Physiol. 2016;231(3):587–596. doi:10.1002/jcp.25102
31. Kuršvietienė L, Mongirdienė A, Bernatonienė J, Šulinskienė J, Stanevičienė I. Selenium anticancer properties and impact on cellular redox status. Antioxidants. 2020;9(1):80. doi:10.3390/antiox9010080
32. Lin CH, Li NT, Cheng HS, Yen ML. Oxidative stress induces imbalance of adipogenic/osteoblastic lineage commitment in mesenchymal stem cells through decreasing SIRT1 functions. J Cell Mol Med. 2018;22(2):786–796.
33. Liu B, Lu Y, Wang Y, Ge L, Zhai N, Han J. A protocol for isolation and identification and comparative characterization of primary osteoblasts from mouse and rat calvaria. Cell Tissue Bank. 2019;20(2):173–182. doi:10.1007/s10561-019-09751-0
34. Yi C, Liu D, Fong CC, Zhang J, Yang M. Gold nanoparticles promote osteogenic differentiation of mesenchymal stem cells through p38 MAPK pathway. ACS Nano. 2010;4(11):6439–6448. doi:10.1021/nn101373r
35. Behzadi S, Serpooshan V, Tao W, et al. Cellular uptake of nanoparticles: journey inside the cell. Chem Soc Rev. 2017;46(14):4218–4244.
36. Gu H, Huang Z, Yin X, et al. Role of c-Jun N-terminal kinase in the osteogenic and adipogenic differentiation of human adipose-derived mesenchymal stem cells. Exp Cell Res. 2015;339(1):112–121. doi:10.1016/j.yexcr.2015.08.005
37. Courtial L, Picco V, Grover R, et al. The c-Jun N-terminal kinase prevents oxidative stress induced by UV and thermal stresses in corals and human cells. Sci Rep. 2017;7:45713. doi:10.1038/srep45713
38. Almeida M. Unraveling the role of FoxOs in bone–insights from mouse models. Bone. 2011;49(3):319–327. doi:10.1016/j.bone.2011.05.023
39. Dhoke NR, Geesala R, Das A. Low oxidative stress-mediated proliferation Via JNK-FOXO3a-Catalase signaling in transplanted adult stem cells promotes wound tissue regeneration. Antioxid Redox Signal. 2018;28(11):1047–1065. doi:10.1089/ars.2016.6974
40. Chen D, Gong Y, Xu L, Zhou M, Li J, Song J. Bidirectional regulation of osteogenic differentiation by the FOXO subfamily of Forkhead transcription factors in mammalian MSCs. Cell Prolif. 2019;52(2):e12540. doi:10.1111/cpr.12540
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.