Back to Journals » International Journal of Nanomedicine » Volume 10 » Issue 1
Selective inhibition of MG-63 osteosarcoma cell proliferation induced by curcumin-loaded self-assembled arginine-rich-RGD nanospheres
Authors Chang R ![]() , Sun L, Webster T
, Sun L, Webster T ![]()
Received 5 December 2014
Accepted for publication 23 January 2015
Published 5 May 2015 Volume 2015:10(1) Pages 3351—3365
DOI https://doi.org/10.2147/IJN.S78756
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Lei Yang
Run Chang,1 Linlin Sun,1 Thomas J Webster1,2
1Department of Chemical Engineering, Northeastern University, Boston, MA, USA; 2Center of Excellence for Advanced Materials Research, King Abdulaziz University, Jeddah, Saudi Arabia
Abstract: Osteosarcoma is the most frequent primary malignant form of bone cancer, comprising 30% of all bone cancer cases. The objective of this in vitro study was to develop a treatment against osteosarcoma with higher selectivity toward osteosarcoma cells and lower cytotoxicity toward normal healthy osteoblast cells. Curcumin (or diferuloylmethane) has been found to have antioxidant and anticancer effects by multiple cellular pathways. However, it has lower water solubility and a higher degradation rate in alkaline conditions. In this study, the amphiphilic peptide C18GR7RGDS was used as a curcumin carrier in aqueous solution. This peptide contains a hydrophobic aliphatic tail group leading to their self-assembly by hydrophobic interactions, as well as a hydrophilic head group composed of an arginine-rich and an arginine-glycine-aspartic acid structure. Through characterization by transmission electron microscopy, self-assembled structures of spherical amphiphilic nanoparticles (APNPs) with diameters of 10–20 nm in water and phosphate-buffered saline were observed, but this structure dissociated when the pH value was reduced to 4. Using a method of codissolution with acetic acid and dialysis tubing, the solubility of curcumin was enhanced and a homogeneous solution was formed in the presence of APNPs. Successful encapsulation of curcumin in APNPs was then confirmed by Fourier transform infrared and X-ray diffraction analyses. The cytotoxicity and cellular uptake of the APNP/curcumin complexes on both osteosarcoma and normal osteoblast cell lines were also evaluated by methyl-thiazolyl-tetrazolium assays and confocal fluorescence microscopy. The results showed that the curcumin-loaded APNPs had significant selective cytotoxicity against MG-63 osteosarcoma cells when compared with normal osteoblasts. We have demonstrated for the first time that APNPs can encapsulate hydrophobic curcumin in their hydrophobic cores, and curcumin-loaded APNPs could be an innovative treatment for the selective inhibition of osteosarcoma cells.
Keywords: osteosarcoma, selective inhibition, curcumin, arginine-rich, arginine-glycine-aspartic acid, self-assembly
Introduction
Osteosarcoma is the most frequent primary malignant bone cancer, comprising 30% of all cases of bone cancer. It often arises in the metaphyses of long bones of adolescents, which have the highest growth potential.1 It is characterized by metastatic tumors that differentiate aggressively toward fibrous tissue, cartilage, and bone.2 The first peak of incidence of osteosarcoma is in 10–14-year-olds when pubertal growth starts and the second peak is in adults aged 65 years or older. In the USA, there are approximately 400 cases of osteosarcoma diagnosed every year.3
The occurrence of osteosarcoma could be related to dysfunction of the p53 tumor suppressor gene.3,4 Expression of the p53 tumor suppressor gene can upregulate apoptotic proteins (ie, Bax, Apaf-1, and CD95), initiating cell cycle arrest and apoptosis. When oncogene activation occurs, the inhibitor Mdm2 could be sequestered in the nucleolus, and p53 is then activated, promoting apoptotic cell death.5 Nonetheless, the first intron of the p53 gene is mutated in many osteosarcoma cell lines, resulting in the loss of p53 expression and abnormal DNA repair mechanisms.6 In addition, overexpression of nuclear factors can lead to irregular proliferation of many cancer cell lines, including osteosarcoma.3 Nuclear factor kappa B (NF-kB) is also an important transcription factor that could initiate and promote cancer. NF-kB is kept inactive in the cytoplasm when inhibited by IkB kinase. Stimulated by either oxidative stress or free radicals, the NF-kB inhibitor IkB is phosphorylated, causing a higher level of active NF-kB, thus fostering an inflammatory environment via various cytokines that favor malignant transformation.7 The active NF-kB transcription factor can translocate into the nucleus, and positively regulate different gene products related to antiapoptotic, proinflammatory, and pro-proliferation events. As a result, apoptosis is blocked under high levels of active NF-kB, this transcription factor could potentially benefit cancer activity. Increased activation of NF-kB is observed in many cancer cell lines, and is involved in antiapoptosis, tumor promotion, and metastasis.8,9
After adoption of multiagent chemotherapy, there was a significant increase in cancer survival rates reported for the years 1974–1994, during which there were lower rates of metastasis and local recurrence. However, there has been no significant improvement in the survival rate for osteosarcoma over the last two decades. According to a report by the National Cancer Institute, the survival rate of patients under the age of 45 years with osteosarcoma was steady at around 65% from 1975 to 2000. Thus, the development for safer, novel, and more targeted therapies is necessary.1,3,4
Curcumin (or diferuloylmethane) has been shown to have potential as a chemotherapeutic agent in many types of cancer since it possesses pleiotropic anticarcinogenesis efficacy. By affecting various molecular and cellular pathways, curcumin is able to target cellular processes, including gene expression, transcription, proliferation, and extracellular matrix synthesis.10 Curcumin not only shows antiproliferative effects towards many types of cancer by inhibiting NF-kB and its downstream gene products, but also affects various growth receptors and cell adhesion molecules in tumor growth.11–13 In addition, curcumin has been shown to upregulate p53 expression in various cancer cell lines, including osteosarcoma.14–16 However, with its polyphenol chemical structure, curcumin exists in an orange-yellow crystalline form and is insoluble in water.17 In basic conditions, curcumin is unstable and has a high degradation rate in phosphate buffers at pH 7.2 under different physiological conditions in vitro.18 Clearly, one solution to using curcumin as an anticancer drug is to develop a suitable carrier.
Bottom-up nanotechnology using self-assembled amphiphilic molecules has recently been investigated in many drug delivery studies. These consist of one or more polar hydrophilic moieties and nonpolar hydrophobic moieties.19,20 Hydrophobic interaction of the hydrophobic moieties is the main driving force of self-assembly, so these amphiphilic molecules can self-assemble into micelles in aqueous solution, while the hydrophilic moieties can directly interact with water and govern the morphology of micelles by molecular interactions (ie, electrostatic interactions and hydrogen bonds).21
Surfactant-like amphiphilic peptides are amphiphiles that contain natural l-amino acids. In addition to improving biocompatibility, amphiphilic peptides can also be functionalized by a variety of peptide sequences for different applications.20 For instance, the arginine-glycine-aspartic acid (RGD) tripeptide can target overexpressed receptors, such as ανβ3 integrins on cancer cells,22–24 while cationic peptides with consecutive 5–11 arginine residues can facilitate the macropinocytosis-meditated pathway for cellular uptake.25–29 An amphiphilic peptide, C18GR7RGDS, was used as a gene delivery carrier,30 but its potential use for drug delivery has rarely been discussed. In the present study, this molecule was tested for the first time as a surfactant-like drug carrier encapsulating hydrophobic curcumin in its hydrophobic core. We hypothesized that curcumin-loaded amphiphilic nanoparticles (APNPs) could be an innovative strategy for the selective inhibition of osteosarcoma cells over healthy osteoblasts.
Materials and methods
Eagle’s Minimum Essential Medium was purchased from the American Type Culture Collection (Manassas, VA, USA), and osteoblast basal medium and osteoblast growth medium Supplemental Mix were purchased from PromoCell (Heidelberg, Germany). Curcumin (diferuloylmethane), acetic acid, dimethyl sulfoxide (DMSO), 4′,6-diamidino-2-phenylindole (DAPI), and Atto Rho6G phalloidin were supplied by Sigma-Aldrich (St Louis, MO, USA). The dry powder for the C18GR7RGDS (Figure 1) amphiphilic peptide was obtained from Biomatik (Wilmington, DE, USA). The PlusOne Mini Dialysis Kit (molecular weight cutoff 1 kDa) was purchased from GE Healthcare (Buckinghamshire, UK) and the methyl-thiazolyl-tetrazolium (MTT) dye solution was purchased from Promega (Madison, WI, USA).
  | Figure 1 Three-dimensional schematic structure of C18GR7RGDS amphiphilic peptides. |
Preparation of APNPs
The APNPs were prepared by dissolving dry powder of C18GR7RGDS (molecular weight 1,850.28 g/mole) in deionized water followed by sonication for 60 seconds. The amphiphilic peptide was also suspended in phosphate-buffered saline and acetic acid solutions at pH 2, 4, and 6. The self-assembly behavior of APNPs in these different solutions were then observed under a transmission electron microscope (TEM) as described in the next section.
Transmission electron microscopy
The morphologies of the APNPs in different solutions were determined using a JEM-1010 Transmission Electron Microscope (JEOL, Tokyo, Japan). Samples in different aqueous conditions were prepared by dissolving the amphiphilic peptides in deionized water, phosphate-buffered saline, and acetic acid solutions at pH 2, 4, and 6. Next, 5 μL of each sample was mounted on a 300-mesh copper grid (EM Sciences Ltd, North Vancouver, BC, Canada) and negatively stained by 5 μL of 1.5% aqueous phosphotungstic acid for 5 seconds. The excess liquid was removed carefully using filter paper. The images were captured by TEM at 40,000–50,000×, operating at an accelerating voltage of 80 kV.
Preparation of curcumin-loaded APNPs
The curcumin-loaded APNPs were prepared by codissolution and dialysis tubing methods. Firstly, curcumin was dissolved in 50% acetic acid and then codissolved with APNPs. In the mixture, the molar ratio of peptide to curcumin was equal to 1:2. The mixture was then transferred to a dialysis tube (molecular weight cutoff 1 kDa) against 800 mL of deionized water. The water was replaced by fresh deionized water every 4 hours in order to eliminate acetic acid and unloaded curcumin from the mixture in the dialysis tube. When the pH of the mixture was close to 7.0, the dialysis tubing was removed. The morphology of the curcumin-loaded APNPs in the final solution were characterized by TEM as previously described. The amount of curcumin encapsulated in the APNPs was characterized by a standard curve showing a linear correlation between the known concentrations of curcumin in DMSO and the corresponding absorbance measured by ultraviolet-visible spectroscopy (SpectraMax M3, Molecular Devices, Sunnyvale, CA, USA) at a wavelength of 430 nm (R2>0.98). Briefly, an aliquot of the curcumin-loaded APNP solution was lyophilized by a freeze-dryer (FreeZone 2.5 Plus, Labconco, Kansas City, MO, USA). The dry powder was then dissolved in DMSO, and the concentration of curcumin was evaluated by correlating the ultraviolet-visible absorption of this solution at a 430 nm wavelength with a standard curve. The concentration of curcumin was evaluated three times for each sample.
Zeta potential
The zeta potentials of pure APNPs and curcumin-loaded APNPs were determined using a ZS90 Nanosizer (Malvern Instruments, Malvern, UK). Next, 0.4 mg/mL of pure APNPs and curcumin-loaded APNPs were prepared in deionized water followed by sonication for 60 seconds at room temperature. The zeta potential of 1 mL of each sample was then measured over ten runs in triplicate.
Fourier transform infrared spectroscopy
Fourier transform infrared (FT-IR) spectra of plain curcumin, the C18GR7RGDS peptide powder, and the lyophilized curcumin-loaded APNPs were collected in order to analyze the chemical structure of these compounds and possible changes therein after drug loading. Samples were analyzed by a FT-IR spectrometer (Vertex 70, Bruker Corporation, Billerica, MA, USA) using the attenuated total reflectance method. The FT-IR spectra were collected in the wavelength range of 550–4,000 cm−1 with a resolution of 2 cm−1.
X-ray diffraction
An X-ray diffraction (XRD) study was conducted to analyze the crystallographic structure of curcumin, pure APNPs, and lyophilized curcumin-loaded APNPs. Samples were analyzed using an X-ray diffractometer (Ultima IV, Rigaku Corporation, Tokyo, Japan) at a voltage of 40 kV, 44 mA, and 1.76 kW. The scanned angle was in the range of 5° ≤ 2θ° ≤ 40° and the scan rate was 3° per minute.
Cell culture
MG-63 osteosarcoma (CRL-1427) cells (American Type Culture Collection) were cultured in Eagle’s Minimum Essential Medium supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin, while healthy human osteoblasts (C-12760, PromoCell) were cultured in complete growth medium composed of osteoblast basal medium and osteoblast growth medium Supplement Mix. Both cell lines were incubated at 37°C in a humidified incubator with an atmosphere of 95% oxygen and 5% CO2. Cells were used at population numbers less than 3.
In vitro cytotoxicity
MG-63 osteosarcoma and noncancerous human healthy osteoblast cell lines were used to evaluate the cytotoxicity of plain curcumin suspended in phosphate-buffered saline, curcumin dissolved in DMSO, a solution of pure APNPs, and a curcumin-loaded APNP solution by the colorimetric MTT assay. First, 100 μL of the osteosarcoma and healthy osteoblast cell solutions were seeded on a 96-well plate at 2×103 cells/well (cell density 6,154 cells/cm2). After 24 hours of incubation in 5% CO2 at 37°C for attachment, the cells were treated with plain curcumin in phosphate-buffered saline, curcumin dissolved in DMSO, and a curcumin-loaded APNP solution with different curcumin concentrations (3, 5, 10, 20, and 30 μM). For the cells treated with a solution of pure APNPs, the solution was prepared by the same codissolution and dialysis tubing method as that used for the preparation of curcumin-loaded APNPs mentioned previously. Cells treated with medium only were used as a positive control. For the samples treated with curcumin dissolved in DMSO, cells treated with the same amount of DMSO (less than 0.5% v/v) were regarded as control samples. Serum-free medium was used in all samples to avoid interactions between the arginine-rich peptides and serum albumin.
The cells were then treated for 24 hours. The medium was then removed from each sample and the cells were washed three times with phosphate-buffered saline. Next, 100 μL of cell medium and 15 μL of the MTT dye solution were added to each well, and the cells were incubated for 4 hours to allow the formation of formazan crystals. At the end of incubation, 100 μL of the MTT stop solution were added to each well. The 96-well plates were then tested using a spectrophotometer (SpectraMax M3, Molecular Devices) at a wavelength of 570 nm to obtain the optical density. Cell density was obtained from a standard curve expressing the linear correlation between different cell densities and optical densities (R2>0.98). Cell viability was expressed as the ratio of cell density in each sample to the cell density in the control sample.
Qualitative cellular uptake study
A confocal laser scanning microscope and a bright field microscope were used for the qualitative cellular uptake study. First, 1 mL each of the osteosarcoma cell line and the healthy human osteoblast cell line were seeded on a 24-well plate at a density of 2×104 cells/mL. After 24 hours of incubation in 5% CO2 and at 37°C, the cells were treated for 2 hours with 20 μM of curcumin encapsulated in APNPs or plain curcumin suspended in phosphate-buffered saline. The cells were then rinsed with phosphate-buffered saline three times to remove the unabsorbed curcumin. The qualitative uptake of curcumin was then monitored by bright field microscopy.
After 10 minutes of fixation by 10% formaldehyde solution and subsequent treatment with a 0.1% Triton X-100 solution for 10 minutes, the nuclei of the cells were stained by DAPI and F-actin was stained by Atto Rho6G phalloidin for observation by confocal laser scanning microscopy (LSM710, Zeiss, Dublin, CA, USA). The stained cells were then viewed under a confocal microscope for DAPI (excitation at 358 nm, emission at 461 nm), Atto Rho6G phalloidin (excitation at 525 nm, emission at 560 nm), and curcumin uptake was observed using a fluorescein isothiocyanate filter (excitation at 495 nm, emission at 519 nm).
Statistical analysis
All experiments were conducted at least three times for each cell line, with five samples for each group. The data are expressed as the mean ± standard error of the mean and unpaired Student’s t-tests were used to test for significant differences between the two treatment groups. P<0.01 indicated significant differences and P<0.005 indicated highly significant differences.
Results
Characterization of self-assembly behavior of APNPs
The C18GR7RGDS amphiphilic peptide could be dissolved easily in deionized water, phosphate-buffered saline (pH 7.4), and acetic acid (pH 2, 4, and 6). A 10 mg/mL stock solution of APNPs was prepared in autoclaved deionized water and then preserved at −80°C before use. The morphology of the self-assembled C18GR7RGDS APNPs in the different solutions was characterized by TEM.
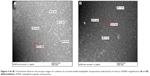
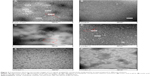
The TEM images showed that the peptide could self-assemble into nanospheres in deionized water and phosphate-buffered saline, with a mean diameter of 15.6 (range 10–20) nm at a concentration of 1.5 mg/mL (Figure 2A and B). The C18 aliphatic tail group could serve as the driving force for the self-assembly behavior of APNPs. The hydrophilic head group of the peptide functionalized by positively charged arginine-rich groups could result in strong electrostatic interactions between each adjacent molecule. APNPs with a spherical morphology were thus formed by the equilibrium of the hydrophobic interaction led by the tail group and the electrostatic interaction led by the head group.31–33 Interestingly, the APNPs could aggregate (Figure 2C) with each other when the peptide was dissolved in deionized water without sonication before characterization.
Self-assembled nanospherical aggregates could still be observed at pH 6 in an acetic acid solution (Figure 2D). However, at pH 2 and 4 (Figure 2E and F), only random cloud-like layers were observed, and the amphiphilic peptides could not self-assemble into nanospheres. Therefore, APNPs could be opened in acidic conditions at pH values <4, and due to this pH-sensitive property, the external material can be encapsulated.
Since curcumin is soluble in acetic acid, it is possible to encapsulate curcumin into APNP aggregates by codissolution with acetic acid disrupting the self-assembled structure, and to reform the nanoparticles by removing acetic acid using a dialysis-tubing purification method. Arginine deprotonation may not be the main factor for this pH-sensitive activity since the pKa of arginine is 12.48, indicating that the guanidinium groups on the arginine-rich structure could be positively charged in a physiological environment. A possible reason for the dissociation of APNPs at low pH might be that the electrostatic interaction is so strong that the repulsive force among molecules can disrupt the self-assembled structure.
This pH-sensitive property might be beneficial for cellular uptake of encapsulated bioactive molecules in the inner core. For example, endosomes, in which the pH is 5–6, are membrane-bound compartments that can transport extracellular molecules from the plasma membrane to the lysosome. The lysosomes can then engulf the molecules by digestive enzymes at a working pH around 4–5. Therefore, this low pH environment could cause dissociation of micelles and release bioactive molecules into the cytosol, followed by digestion by the lysosome.
Formation of curcumin-loaded APNPs
APNPs can be disrupted by acetic acid when the pH is less than 4, so a method for codissolution with curcumin in 50% acetic acid was developed to dissociate the aggregates. Due to their low molecular weights, the acetic acid and the free curcumin molecules could be purified with dialysis tubing. Only APNPs and the encapsulated curcumin could be retained in the solution. Using this method, a homogeneous solution of curcumin was successfully prepared with the help of APNPs. Compared with the same amount of a solid curcumin suspension in water (solubility less than 0.1 mg/mL), the resulting solution showed significantly increased solubility and homogeneity (Figure 3A). Moreover, the lyophilized powder of the solution could be redissolved in water easily (Figure 3B and C).
Subsequently, nanospheres were observed in the TEM images of the resulting solution after drug loading. These nanoparticles had a morphology similar to that of the pure APNPs but with comparably larger diameters at around 18–30 nm (average diameter 22.8 nm, Figure 4). This result indicates that the self-assembly behavior was not altered during the drug preparation procedure, and the pH-sensitive nanoparticles were able to reform by a reversible process after removal of acetic acid. Meanwhile, hydrophobic molecules, like curcumin, could be entrapped and solubilized in the stearyl C18 aliphatic cores of the micelles by energetically favorable hydrophobic interactions,31,34 suggesting successful drug encapsulation in the aqueous surfactant solution of APNPs.
Zeta potential
The measured average zeta potential of pure APNPs was +59±3.15 mV, while that of curcumin-loaded APNPs was +70.63±3.02 mV (Figure 5). This result indicates that both pure and curcumin-loaded APNPs were stable in aqueous solution. The curcumin-loaded micelles have a higher zeta-potential, possibly resulting from the increased number of free peptide monomers aggregated to form stable micelles after drug loading, and these positively charged micelles could facilitate cellular uptake mediated by the membrane potential.26
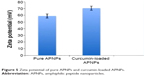  | Figure 5 Zeta potential of pure APNPs and curcumin-loaded APNPs. |
Fourier transform infrared spectroscopy
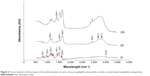
To further investigate the chemical and molecular interactions after drug loading, FT-IR analysis was used to show the spectra of solid-state plain curcumin, pure APNPs, and curcumin-loaded APNPs (Figure 6).
In the spectra of plain curcumin, the bands that appeared in the ranges of 1,225–1,175 cm−1 and 1,125–1,090 cm−1 together with two additional weak bands in the ranges around 1,070–1,000 cm−1 could represent the 1:2:4-substitution of the aromatic rings. The two C=C bonds conjugated with the neighborhood aromatic rings and C=O bonds could be characterized at 1,629 cm−1 and 1,606 cm−1, respectively. The hydroxyl group with intramolecular hydrogen bonds in the phenol groups could be characterized by the relatively weak absorption at 3,519 cm−1.35
In the spectra of pure APNPs, the absorption at 1,654 cm−1 could represent the amide I group,36 while the band at 1,560 cm−1 could indicate the COOH group in the amino acid sequence. In addition, the two wide bands at 3,400–3,300 cm−1 could characterize the amine group of the arginine-rich structure. For the spectra of lyophilized curcumin-loaded APNPs, the bands appeared at a wavelength similar to that for pure APNPs, but the band at 1,409 cm−1 could represent the OH deformation vibration on phenols.35 The FT-IR spectra may suggest that the chemical structure of the amphiphilic peptide was not altered after drug loading since no significant band shifts were observed. Furthermore, most of the absorbance bands for curcumin could not be observed, except for the OH deformation vibration on the phenols. This result indicates successful encapsulation of curcumin by APNPs, as curcumin molecules were shielded in the inner core of micelles, and the infrared radiation could not be transmitted through the encapsulated molecules.
X-ray diffraction
XRD was conducted to further analyze the crystalline and molecular structure of curcumin before and after encapsulation by APNPs (Figure 7). In the XRD pattern for curcumin, a series of characteristic peaks could be observed in the range of 15° ≤ 2θ° ≤ 30°, representing the distinct crystalline structure of curcumin molecules. In contrast, pure APNPs may not have a characteristic crystalline structure since no peaks were evident in its XRD pattern. More importantly, the curcumin-loaded APNPs showed an XRD pattern similar to that of pure APNPs and did not show an observable crystalline structure. Disappearance of peaks characteristic of the crystalline structure of curcumin can result from encapsulation by APNPs. The XRD pattern for the pure APNPs demonstrated that these molecules could exist in a disordered crystalline structure or be amorphous. APNPs could shield the curcumin molecules from diffracting the X-ray beams, causing the XRD of the curcumin crystalline structure to be undetectable. Thus, the XRD pattern of curcumin-loaded APNPs further confirmed successful drug encapsulation.
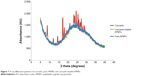  | Figure 7 X-ray diffraction patterns of curcumin, pure APNPs, and curcumin-loaded APNPs. |
Qualitative cellular uptake of curcumin-loaded APNPs
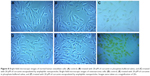
Internalization of curcumin-loaded APNPs by the osteosarcoma and human osteoblast cell lines was tracked by bright field microscopy and confocal laser scanning microscopy. After incubation with curcumin-loaded APNPs at a 20 μM curcumin concentration for 2 hours, the osteosarcoma cells showed significantly higher uptake of curcumin than the normal human osteoblast cells in bright field microscopy images (Figure 8). In the samples treated only with plain curcumin suspended in phosphate-buffered saline, very small amounts of crystalline curcumin could be observed in the cell membranes, but curcumin did not accumulate in the cytosol by endocytosis and internalization was inefficient.
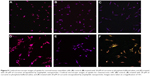
The nuclei of the cells were tracked by blue fluorescent DAPI staining using confocal microscopy (Figure 9), and the F-actin filaments of cells were stained with red fluorescent Rhodamine 6G. The cellular uptake of curcumin was monitored using a green fluorescent fluorescein isothiocyanate filter.25,37 Similar to the images taken by bright field microscopy, both cell lines did not show detectable fluorescence of curcumin in the samples treated by plain curcumin, which indicated that APNPs could enhance cellular uptake. Osteosarcoma cells treated with curcumin-loaded APNPs showed a strong green fluorescent signal, indicating that these cells accumulated significant amounts of curcumin into the cytosol. However, normal human osteoblast cells showed only a weak green fluorescent signal in the cytosol.
These results demonstrated that curcumin-loaded APNPs could penetrate the membrane of osteosarcoma cells more efficiently and induce significantly higher cellular uptake than in human osteoblast cells. With an RGD-functionalized head group, the curcumin-loaded micelles might selectively attach to the receptors of the overexpressed integrins on osteosarcoma cells, leading to more drug accumulation on the surface of the osteosarcoma cells than on the normal human osteoblast cells. Meanwhile, the micelles could stably attach to carboxylate, sulfate, and phosphate groups on the cell membrane by two hydrogen bonds of each arginine group in the peptide sequence. This would favor macropinocytosis-meditated internalization of arginine-rich peptides.26 Hence, curcumin molecules could internalize into the cytosol efficiently via the endosomal pathway from the cell membrane to the lysosome, although the exact mechanisms involved will need to be elucidated in future studies.
In vitro cytotoxicity
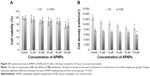
The cytotoxicity of the curcumin-loaded APNP solution, the solution containing pure APNP, curcumin dissolved in DMSO, and plain curcumin suspended in phosphate-buffered saline was investigated (expressed as cell viability and cell density). The pure APNPs showed minor cytotoxicity in both the osteosarcoma cell line and the human osteoblast cell line at the highest concentration investigated (Figure 10), which might have resulted from the increased positive charge along with increased peptide concentrations.26 The cytotoxicity of plain curcumin suspended in phosphate-buffered saline was insignificant for both cell lines (Figures 11 and 12A and B), possibly reflecting low cellular uptake due to the low solubility of curcumin in phosphate-buffered saline. When dissolved in DMSO, curcumin was more cytotoxic to osteosarcoma cells at all concentrations investigated, which is in accordance with previous studies.38,39 More importantly, the curcumin-loaded APNPs showed significant selective inhibition of viability of osteosarcoma cells. Compared with the curcumin/DMSO sample, the cytotoxicity of curcumin-loaded APNPs was more selective for osteosarcoma cells in the concentration range of 20–30 μM. At a curcumin concentration of 30 μM, the viability of osteosarcoma cells was as low as 15% after treatment with curcumin-loaded APNPs, whereas over 50% of human osteoblast cells were viable. The curcumin-loaded APNPs with a 20 μM curcumin concentration could be the optimal concentration, given that the viability of osteosarcoma was the minimum value at this concentration. This result quantitatively confirms the targeting effects of the RGD peptide sequence on ανβ3 integrins, which are overexpressed on cancer cells, bringing about more uptake of encapsulated drug by osteosarcoma cells than by healthy human osteoblast cells.
Discussion
The present TEM results show that the amphiphilic peptide C18GR7RGDS can self-assemble into spherical nanoparticles with diameters of 10–20 nm in water and phosphate-buffered saline. This self-assembly behavior might be driven by the hydrophobic interactions of the aliphatic tail group and the electrostatic interaction attributed to the positive charge in the head group. A three-dimensional schematic molecular structure of the amphiphilic peptide was created and analyzed using ChemBio3D Ultra 14 software (Figure 1). The estimated length of an amphiphilic peptide molecule is about 6.74 nm. Comparing the diameters of micelles measured in the TEM images and those in the schematic measurement, the micelle structures in APNPs could be monolayer aggregates with solid hydrophobic cores, although the critical packing parameter should be precisely evaluated.19 Furthermore, given that micelles can form when the concentration of amphiphilic molecules is above the critical micelle concentration whereas amphiphilic molecules behave differently when their concentration is lower than that of the critical micelle concentration,19 the morphology of APNPs with lower peptide concentrations need to be observed in order to determine the stability of APNPs upon dilution.
The self-assembly properties of APNPs were also found to be pH-sensitive. Under acidic conditions, in which the pH was lower than 4, the APNP aggregates could dissociate, allowing for entry of external compounds. When the samples were negatively stained by uranyl acetate, the self-assembled APNP structures could not be observed by TEM (images not shown), possibly because the nanoparticles were disrupted in the acidic uranyl acetate solution (pH 4.2–4.5).
By codissolution with a curcumin solution in 50% acetic acid and dialysis tubing, APNPs could reform into spherical nanostructures but with larger diameters. More importantly, the solubility of curcumin increased significantly and the orange-yellow solution showed more stability and homogeneity with the assistance of APNPs. Even after freeze-drying, the powder of the solute of this solution could be dissolved easily and rapidly in water. Therefore, it could be hypothesized that curcumin can be successfully encapsulated in the inner hydrophobic core of micelles of APNPs.33,34 The encapsulation of curcumin was further studied by FT-IR and XRD analysis. After drug loading, no significant band shift occurred in the FT-IR spectra for pure APNPs, but bands representing most of the chemical groups in curcumin molecules could not be detected. This result might provide evidence for successful encapsulation of curcumin since the encapsulated molecules could be shielded from the infrared signal. XRD analysis further confirmed the successful encapsulation of curcumin by APNPs, since the distinct peaks for the crystalline structure of curcumin disappeared and the curcumin-loaded APNPs had an XRD pattern similar to that of pure APNPs after drug loading.
In vitro experiments were conducted using both MG-63 osteosarcoma cells and normal healthy human osteoblast cells to investigate the ability of APNPs to target bone cancer cells. At curcumin concentrations of 3–30 μM, the cytotoxicity of curcumin-loaded APNPs against osteosarcoma cells was significantly greater than against healthy osteoblast cells. The viability of healthy osteoblast cells and osteosarcoma cells was 54% and 15%, respectively, after 24 hours of treatment with 30 μM curcumin concentration in APNPs, while the same amount of pure APNPs and plain curcumin suspended in phosphate-buffered saline had low toxicity in both cell lines. The 20 μM curcumin concentration in APNPs could be optimal to inhibit the proliferation of osteosarcoma cells since their viability reached its minimum value at this point. The selective cytotoxicity of curcumin-loaded APNPs was also greater than that of curcumin dissolved in the organic solvent DMSO, which may be because the RGD sequence targets overexpressed integrins on cancerous cells.34,40–42 Since DMSO has been shown to enhance the penetration of both hydrophobic and hydrophilic agents into the cell membrane,43 curcumin could be delivered through the lipid membrane with the aid of DMSO. By becoming incorporated below the hydrophilic head group in a nonspecific manner, molecules of DMSO increase the average head group area and the amenability to bending in the lipid membranes, the entry of water molecules is then facilitated, inducing non-specific water pore formation.44 However, although the dermal application of DMSO did not show significant toxicity in earlier studies, it was reported that DMSO caused eye damage, including lens abnormalities, in animals.45 Moreover, Hanslick et al reported that DMSO could induce widespread apoptotic neurodegeneration in the central nervous system of juvenile mice, and possibly caused similar damage in children. Even at a concentration as low as 0.5%, DMSO could lead to apoptosis in the central nervous system, and at the postnatal age of 7 and 30 which are equivalent to human childhood ages, severe damage was observed after DMSO exposure.46 Therefore, as the peak incidence of osteosarcoma in adolescents is in 10–14-year-olds, when pubertal growth starts,4 DMSO might cause damage in the central nervous system when used as a solvent for hydrophobic anticancer drugs. In this study, APNPs did not show significant toxicity, so the curcumin-loaded APNPs could accelerate the internalization of curcumin with low toxicity induced by short polypeptides, although further in vivo studies are required to investigate this. Moreover, the drug delivery performance of APNPs should be compared with traditional drug carriers like liposomes.
Qualitative cellular uptake studies further confirmed the selectivity of curcumin-loaded APNPs. In both bright field microscopy and confocal laser scanning microscopy images, significant amounts of internalized APNPs containing curcumin could be observed in the cytosol of osteosarcoma cells, whereas limited amounts of curcumin were observed in healthy human osteoblast cells. Additionally, when compared with cells treated using a plain curcumin suspension in phosphate-buffered saline, the arginine-rich sequence of APNPs induced higher internalization efficiency, causing macropinocytosis-meditated internalization of curcumin from the plasma membrane into the cytosol.26 Since the self-assembled APNPs are pH-sensitive, the micelles of APNPs loaded with curcumin may be stable in early endosomes at pH 6, but disassemble in the lysosome at a lower pH, releasing the drug in a controllable manner. This assumption should be confirmed by studies of the cellular uptake mechanism. It is still unclear if these APNPs can protect curcumin from degradation in an alkaline environment, so the stability of pure APNPs and curcumin-loaded APNPs should be studied in alkaline conditions. Also, in vivo studies are necessary to investigate the therapeutic efficacy of curcumin-loaded APNPs in the human body.
Conclusion
This study demonstrates for the first time that arginine-rich-RGD amphiphilic peptides can serve as a drug delivery vehicle for curcumin and selectively inhibit the proliferation of osteosarcoma cells over healthy osteoblast cells. After loading with curcumin, micelles of APNPs could selectively inhibit MG-63 osteosarcoma cells rather than normal human osteoblasts. Further studies for enhancing curcumin loading efficiency, the stability of micelles, combined delivery of curcumin and genes via APNPs, quantitative polymerase chain reaction assays, and in vivo studies are necessary to continue this promising line of research.
Acknowledgments
The authors would like to thank Northeastern University for funding, William Fowle for TEM training, and Wenwen (Vivian) Liu for her assistance with confocal microscopy.
Disclosure
The authors have no relevant financial or nonfinancial relationships to disclose in relation to this study.
References
Broadhead ML, Clark JC, Myers DE, Dass CR, Choong PF. The molecular pathogenesis of osteosarcoma: a review. Sarcoma. 2011;2011:959248. | ||
He JP, Hao Y, Wang XL, et al. Review of the molecular pathogenesis of osteosarcoma. Asian Pac J Cancer Prev. 2014;15(15):5967–5976. | ||
Cotterill SJ, Wright CM, Pearce MS, Craft AW. Stature of young people with malignant bone tumors. Pediatr Blood Cancer. 2004;42(1):59–63. | ||
Ottaviani G, Jaffe N. The Etiology of Osteosarcoma. In: Jaffe N, Bruland ØS, Bielack SS, editors. Pediatric and Adolescent Osteosarcoma. Houston, TX, USA; Springer; 2009:15–32. | ||
Dutra FR, Largent EJ. Osteosarcoma induced by beryllium oxide. Am J Pathol. 1950;26(2):197–209. | ||
Pollard TD, Earnshaw WC. Cell Biology. 2nd ed. Philadelphia, PA, USA: Saunders Elsevier; 2008. | ||
Wang ZQ, Liang J, Schellander K, Wagner EF, Grigoriadis AE. c-Fos-induced osteosarcoma formation in transgenic mice: cooperativity with c-Jun and the role of endogenous c-Fos. Cancer Res. 1995;55(24):6244–6251. | ||
Staudt LM. Oncogenic activation of NF-kB. Cold Spring Harb Perspect Biol. 2010;2:a000109. | ||
Barnes PJ, Karin M. Nuclear factor-kappa B: a pivotal transcription factor in chronic inflammatory diseases. N Engl J Med. 1997;336(15):1066–1071. | ||
Strimpakos AS, Sharma RA. Curcumin: preventive and therapeutic properties in laboratory studies and clinical trials. Antioxid Redox Signal. 2008;10(3):511–545. | ||
Wilken R, Veena MS, Wang MB, Srivatsan ES. Curcumin: a review of anti-cancer properties and therapeutic activity in head and neck squamous cell carcinoma. Mol Cancer. 2011;10:12. | ||
Brennan P, O’Neill LA. Inhibition of nuclear factor κB by direct modification in whole cells – mechanism of action of nordihydroguaiaritic acid, curcumin and thiol modifiers. Biochem Pharmacol. 1998;55(7):965–973. | ||
Jobin C, Bradham CA, Russo MP, et al. Curcumin blocks cytokine-mediated NF-kappa B activation and proinflammatory gene expression by inhibiting inhibitory factor I-kappa B kinase activity. J Immunol. 1999;163(6):3474–3483. | ||
Choudhuri T, Pal S, Das T, Sa G. Curcumin selectively induces apoptosis in deregulated cyclin D1-expressed cells at G2 phase of cell cycle in a p53-dependent manner. J Biol Chem. 2005;280(20):20059–20068. | ||
Liu E, Wu J, Cao W, et al. Curcumin induces G2/M cell cycle arrest in a p53-dependent manner and upregulates ING4 expression in human glioma. J Neurooncol. 2007;85(3):263–270. | ||
Collins HM, Abdelghany MK, Messmer M, et al. Differential effects of garcinol and curcumin on histone and p53 modifications in tumour cells. BMC Cancer. 2013;13:37. | ||
Kumar A, Ahuja A, Ali J, Baboota S. Conundrum and therapeutic potential of curcumin in drug delivery. Crit Rev Ther Drug Carrier Syst. 2010;27(4):279–312. | ||
Tønnesen HH. Solubility, chemical and photochemical stability of curcumin in surfactant solutions. Pharmazie. 2002;57(12):820–824. | ||
Ramanathan M, Shrestha LK, Mori T, Ji Q, Hill JP, Ariga K. Amphiphile nanoarchitectonics: from basic physical chemistry to advanced applications. Phys Chem Chem Phys. 2013;15(26):10580–10611. | ||
Tu RS, Tirrell M, Bottom-up design of biomimetic assemblies. Adv Drug Deliv Rev. 2004;56(11):1537–1563. | ||
Zhao X, Pan F, Xu H, et al. Molecular self-assembly and applications of designer peptide amphiphilies. Chem Soc Rev. 2010;39(9):3480–3498. | ||
Raj A, Saraf P, Javali NM, Li X, Jasti B. Binding and uptake of novel RGD micelles to the αvβ3 integrin receptor for targeted drug delivery. J Drug Target. 2014;22(6):518–527. | ||
Javali NM, Raj A, Saraf P, Li X, Jasti B. Fatty acid-RGD peptide amphiphilic micelles as potential paclitaxel delivery carriers to αvβ3 integrin overexpressing tumors. Pharm Res. 2012;29(12):3347–3361. | ||
Giancotti FG, Ruoslahti E. Integrin signaling. Science. 1999;285(5430):1028–1032. | ||
Mohanty C, Sahoo SK. The in vitro stability and in vivo pharmacokinetics of curcumin prepared as an aqueous nanoparticulate formulation. Biomaterials. 2010;31(25):6597–6611. | ||
Futaki S, Hirose H, Nakase I. Arginine-rich peptides: methods of translocation through biological membranes. Curr Pharm Des. 2013;19(16):2863–2868. | ||
Nakase I, Tadokoro A, Kawabata N, et al. Interaction of arginine-rich peptides with membrane-associated proteoglycans is crucial for induction of actin organization and macropinocytosis. Biochemistry. 2007;46(2):492–501. | ||
Nakase I, Niwa M, Takeuchi T, et al. Cellular uptake of arginine-rich peptides: roles for macropinocytosis and actin rearrangement. Mol Ther. 2004;10(6):1011–1022. | ||
Rothbard JB, Kreider E, VanDeusen CL, Wright L, Wylie BL, Wender PA. Arginine-rich molecular transporters for drug delivery: role of backbone spacing in cellular uptake. J Med Chem. 2002;45(17):3612–3618. | ||
Chen JX, Wang HY, Quan CY, Xu XD, Zhang XZ, Zhuo RX. Amphiphilic cationic lipopeptides with RGD sequences as gene vectors. Org Biomol Chem. 2010;8(14):3142–3148. | ||
Southall NT, Dill KA, Haymet AJ. A view of the hydrophobic effect. J Phys Chem B. 2002;106(3):521–533. | ||
Chandler D. Interfaces and the driving force of hydrophobic assembly. Nature. 2005;437(7059):640–647. | ||
Nagarajan R, Ruckenstein E. Theory of surfactant self-assembly: a predictive molecular thermodynamic approach. Langmuir. 1991;7(12):2934–2969. | ||
Fahr A, Liu X. Drug delivery strategies for poorly water-soluble drugs. Expert Opin Drug Deliv. 2007;4(4):403–416. | ||
Bellamy LJ. The Infra-Red Spectra of Complex Molecules. 2nd ed. London, UK: Chapman and Hall Ltd; 1958. | ||
Xu XD, Chen JX, Cheng H, Zhang XZ, Zhuo RX. Controlled peptide coated nanostructures via the self-assembly of functional peptide building blocks. Polym Chem. 2012;3:2479–2486. | ||
Khaw AK, Hande MP, Kalthur G, Hande MP. Curcumin inhibits telomerase and induces telomere shortening and apoptosis in brain tumour cells. J Cell Biochem. 2013;114(6):1257–1270. | ||
Chang R, Sun L, Webster TJ. Short communication: selective cytotoxicity of curcumin against osteosarcoma cell lines compared to healthy osteoblasts. Int J Nanomedicine. 2014;9:461–465. | ||
Jin S, Xu HG, Shen JN, Chen XW, Wang H, Zhou JG. Apoptotic effects of curcumin on human osteosarcoma U2OS cells. Orthop Surg. 2009;1(2):144–152. | ||
Jin H, Varner J. Integrins: roles in cancer development and as treatment targets. Br J Cancer. 2004;90(3):561–565. | ||
Liu S. Radiolabeled multimeric cyclic RGD peptides as integrin αvβ3 targeted radiotracers for tumor imaging. Mol Pharm. 2006;3(5):472–487. | ||
Hassenpflug WA, Tschan MP, Torbett BE, Habermann BF. Integrin alpha(v)beta(3) and osteopontin promote hematogenous metastasis of osteosarcoma cells to the lung. Thromb Res. 2007;120:S172. | ||
Williams AC, Barry BW. Penetration enhancers. Adv Drug Deliv Rev. 2004;56(5):603–618. | ||
Notman R, Noro M, O’Malley B, Anwar J. Molecular basis for dimethylsulfoxide (DMSO) action on lipid membranes. J Am Chem Soc. 2006;128(43):13982–13983. | ||
David NA. The pharmacology of dimethyl sulfoxide 6544. Annu Rev Pharmacol. 1972;12:353–374. | ||
Hanslick JL, Lau K, Noguchi KK, et al. Dimethyl sulfoxide (DMSO) produces widespread apoptosis in the developing central nervous system. Neurobiol Dis. 2009;34(1):1–10. |
 © 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2015 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms
and incorporate the Creative Commons Attribution
- Non Commercial (unported, 3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.