Back to Journals » Infection and Drug Resistance » Volume 11
Salmonella enterica serovars Typhimurium and Enteritidis causing mixed infections in febrile children in Mozambique
Authors García V , Mandomando I , Ruiz J , Herrera-León S, Alonso PL, Rodicio MR
Received 25 July 2017
Accepted for publication 27 September 2017
Published 31 January 2018 Volume 2018:11 Pages 195—204
DOI https://doi.org/10.2147/IDR.S147243
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Eric Nulens
Vanesa García,1 Inácio Mandomando,2,3 Joaquim Ruiz,4 Silvia Herrera-León,5 Pedro L Alonso,3,4 M Rosario Rodicio1
1Departamento de Biología Funcional, Área de Microbiología, Universidad de Oviedo, Oviedo, Spain; 2Centro de Investigação em Saúde de Manhiça, 3Instituto Nacional de Saúde, Ministério da Saúde, Maputo, Mozambique; 4ISGlobal, Barcelona Centre for International Health Research, Hospital Clínic, Universitat de Barcelona, Barcelona, 5Centro Nacional de Microbiología, Instituto de Salud Carlos III, Majadahonda, Madrid, Spain
Background and purpose: Invasive nontyphoidal salmonellosis, mostly caused by serovars Typhimurium and Enteritidis of Salmonella enterica, has emerged as a major public health problem in sub-Saharan Africa. The aim of this study was the clinical and microbiological characterization of nontyphoidal salmonellosis episodes affecting febrile children in Mozambique.
Patients and methods: The clinical records of the patients were evaluated, and S. enterica isolates were characterized with regard to serovar, phage type, antimicrobial resistance (phenotype/responsible genes), plasmid content, pulsed-field gel electrophoresis, and multilocus sequence typing.
Results: Fifteen S. Typhimurium and 21 S. Enteritidis isolates were recovered from blood samples of 25 children, the majority with underlying risk factors. With regard to phage typing, most isolates were either untypeable or reacted but did not conform, revealing that a number of previously unrecognized patterns are circulating in Mozambique. Most isolates were multidrug-resistant, with nearly all of the responsible genes located on derivatives of serovar-specific virulence plasmids. ST313 and ST11 were the predominant sequence types associated with S. Typhimurium and S. Enteritidis, respectively, and the uncommon ST1479 was also detected in S. Enteritidis. A distinct XbaI fragment of ~350 kb was associated with pulsed-field gel electrophoresis patterns of multidrug-resistant isolates of S. Enteritidis. Nearly half of the children were coinfected with both serovars, a fact expected to aggravate the disease and hamper the treatment. However, particularly poor outcomes were not observed for the coinfected patients.
Conclusion: Mixed Salmonella infections could frequently occur in febrile children in Mozambique. Additional studies are required to determine their actual impact and consequences, not only in this country, but also in other African countries.
Keywords: invasive nontyphoidal salmonellosis, bloodstream infection, multidrug resistance, virulence-resistance plasmid, ST313, ST1479
Introduction
Nontyphoidal serovars of Salmonella enterica are a prominent cause of bloodstream infections in sub-Saharan Africa, which mainly affect children with specific risk factors, including malnutrition, severe anemia, malaria, and HIV infection, as well as HIV-infected or immunocompromised adults.1–3 S. Typhimurium and S. Enteritidis are the most common serovars involved in invasive nontyphoidal salmonellosis (iNTS).4–6 Phylogenetic analysis based on single-nucleotide polymorphisms revealed that the majority of the invasive S. Typhimurium from sub-Saharan countries belong to sequence type (ST) 313 and are distributed into two closely related lineages designated I and II.7,8 Both lineages comprise multidrug-resistant (MDR) isolates carrying distinct derivatives of pSLT, the ~94 kb virulence plasmid specific of S. Typhimurium, where multiple resistance genes are located.7,8 Examples of such plasmids are pSLT-A130 (conferring resistance to ampicillin, kanamycin, sulfonamides, and trimethoprim), carried by the A130 isolate representative of lineage I, and pSLT-BT (responsible for resistance to ampicillin, chloramphenicol, kanamycin, streptomycin, sulfonamides, and trimethoprim), found in D23580 of lineage II. With regard to S. Enteritidis, it is of note that MDR isolates of this serovar were recovered in the UK from blood and feces of patients who had an African name or had recently traveled to Africa.9 The UK isolates were phage type (PT) 42 and belonged to ST11, the most common ST in S. Enteritidis, or to the infrequent ST1479. MDR in these isolates was encoded by derivatives of pSEV, the ~60 kb virulence plasmid of S. Enteritidis, related to pUO-SeVR1, a plasmid first detected in a PT14b, ST11 isolate collected in Spain from feces of a young child with gastroenteritis.10 Recently, whole genome sequence analysis of S. Enteritidis from sub-Saharan Africa identified two related but phylogenetically and geographically distinct clades, the Western African clade and the Central/Eastern African clade. In both clades, MDR was encoded by pSEV derivatives highly similar to pUO-SeVR1.11
In rural Mozambique, previous studies revealed that nontyphoidal Salmonella is the most frequent cause of bacteremia in children younger than 15 years admitted to the Manhiça District Hospital (MDH).5,12,13 Like in other African countries, S. Typhimurium and S. Enteritidis were the most frequent serovars, accounting for 66% and 25% of the cases, respectively.13 To broad our knowledge on iNTS in Mozambique, the present study focused on a number of isolates causing bacteremia in children, which were characterized with regard to serovar, PT, resistance properties (including phenotypes, genes, integrons, and plasmids involved), and genomic types established by pulsed-field gel electrophoresis (PFGE) and multilocus sequence typing (MLST). The study disclosed a relatively high number of mixed infections with the simultaneous detection of S. Typhimurium and S. Enteritidis isolates in a single episode of bacteremia.
Material and methods
Setting, patients, and bacterial isolates
Twenty-five blood samples positive for Salmonella were recovered during the period 2002–2004 from different children admitted to the MDH (the referral health facility for a rural area located in Southern Mozambique), and the clinical histories of the patients were retrospectively revised. The identification of Salmonella was based on Gram staining and biochemical methods.14 Serotyping was performed at the Centro de Investigação em Saúde da Manhiça (CISM, Maputo, Moçambique) by traditional slide agglutination with commercially available antisera (polyvalent and monoclonal; Bio-Rad, Marnes-la-Coquette, France) and by a PCR-based method.15 Serotyping results were confirmed at the Centro Nacional de Microbiología (CNM, Madrid, Spain), where PTs were also determined. Isolates showing a pattern that did not conform to previously recognized PTs were designated as “reacted but did not conform” (RDNC). Isolates that did not react with any of the typing phages were designated as “untypeable” (UT).
Antimicrobial susceptibility testing and detection of resistance genes
Susceptibility to ampicillin, chloramphenicol, ciprofloxacin, cefotaxime, cefoxitin, cephalothin, gentamicin, kanamycin, nalidixic acid, sulfonamides, nitrofurantoin, streptomycin, tetracycline, and trimethoprim was tested by the disc diffusion method on Mueller–Hinton agar, using commercially available discs (Oxoid, Madrid, Spain). Results were interpreted according to the guidelines of the Clinical and Laboratory Standards Institute.16
Genes responsible for resistance to ampicillin (blaOXA-1-like, blaPSE-1, and blaTEM-1-like), chloramphenicol (catA1, catA2, cmlA1-like, and floR), gentamicin/kanamycin [aac(3)-II, aac(3)-IV, and aadB], streptomycin (aadA1-like, aadA2, strA, and strB), sulfonamides (sul1, sul2, and sul3), tetracycline [tet(A), tet(B), and tet(G)], and trimethoprim (dfrA1-like, dfrA7-dfrA17, and dfrA12) were screened by PCR amplification using previously described primers and conditions,17–19 except for catA2 (catA2-F/R: 5’-AGCTTACGGAGGACGGAATG-3’/5’-CACCAATCCTGCCGGATACG-3’). The genetic bases of nalidixic acid resistance were investigated by using PCR amplification and sequencing of the quinolone resistance-determining region of the gyrA gene.20 The presence of class 1 and 2 integrons was investigated by using the 5’-CS/3’-CS and Hep74/Hep51 primers, and gene cassettes located therein were identified by nested PCR amplification with primers selected according to the detected resistance genes.21,22 Gene cassettes were sequenced at Macrogen Europe (Amsterdam, The Netherlands), when required for allele identification.
Plasmid analysis, conjugation, and transformation experiments
Plasmid DNA was extracted by using an alkaline lysis method.23 Electrophoresis was carried out on 0.6% agarose gels for 3–4 hours at 100 V. Plasmids obtained from Escherichia coli V517 (54, 7.3, 5.6, 5.1, 3.9, 3, 2.7, and 2.1 kb), E. coli 39R861 (150, 65, 37, and 7 kb), S. Typhimurium ATCC 14028 (pSLT; 94 kb), S. Enteritidis ATCC 13076 (pSEV; 60 kb), and S. Enteritidis CNM 4839/03 (which contains two plasmids, pUO-SeVR1 of 110 kb and a cryptic plasmid of ~80 kb) were used as size standards.
All isolates were screened for the IncFIIA and IncFIB replicons and the spvC gene of serovar-specific virulence plasmids, by PCR amplification using previously described primers and conditions.10,24 pSLT and pSEV were included as positive controls. Isolates harboring small plasmids were also tested for the ColE replicon.25
Triparental conjugation experiments were performed on Luria–Bertani liquid medium at 37°C for 20 hours using the invasive S. Typhimurium and S. Enteritidis isolates as donors, a derivative of E. coli K12 J53 resistant to rifampicin as the recipient, and E. coli MT1694, resistant to kanamycin and containing the conjugative pRK2013 plasmid, as a helper strain. Transconjugants were selected on Eosin Methylene Blue agar (Oxoid) supplemented with rifampicin (50 mg/L) plus either ampicillin (100 mg/L), chloramphenicol (30 mg/L), streptomycin (30 mg/L), sulfonamides (300 mg/L), tetracycline (30 mg/L), or trimethoprim (10 mg/L). For transformation experiments, plasmid DNA obtained from selected isolates was purified by the High Pure Plasmid Isolation Kit (Roche, Madrid, Spain) and transformed into a nalidixic acid-resistant derivative of E. coli DH5α. The preparation of competent cells and transformation were as in Sambrook and Russell.26 Transformants were selected in Luria–Bertani agar medium supplemented with nalidixic acid and ampicillin (30 and 100 mg/L, respectively). Transfer of additional resistance genes was tested by the disc diffusion method and PCR amplification.
PFGE analysis and MLST
PFGE was performed with the XbaI endonuclease (Thermo Fisher Scientific, Waltham, MA, USA), following the PulseNet protocol for S. enterica (www.pulsenetinternational.org). XbaI-digested DNA of S. Braenderup H9812 was used as a size marker. The resulting profiles were visually analyzed by recording the presence or absence of fragments >33 kb, and those with differences in two or more bands were designated using serial numbers (eg, X1 and X2). MLST was carried out by PCR amplification and sequencing, using the specific primers for seven housekeeping genes (aroC, dnaN, hemD, hisD, purE, sucA, and thrA) described in the MLST website (http://mlst.warwick.ac.uk/mlst/dbs/Senterica). The STs were assigned according to the database in the same website.
Hybridization experiments
PFGE and plasmid profiles were transferred into nylon membranes (Hybridon™ N+; GE Healthcare Life Sciences, Barcelona, Spain) and hybridized with probes for IncFIIA, IncFIB, and spvC selected as markers of serovar-specific virulence plasmids and for the identified resistance genes and integrons [blaOXA-1-like, blaTEM-1-like, catA1, catA2, aac(3)-IV, aadB, aadA1-like, strA, strB, sul1, sul2, tet(A), dfrA7, dfrA12, and intI1] to establish their location.26 The probes were labeled using the PCR DIG labeling mix (Roche Applied Science, Barcelona, Spain), followed by gel extraction with the GFX™ DNA and Gel Band Purification Kit (GE Healthcare Life Sciences).
Ethical clearance
The strains characterized here were isolated from the ongoing invasive bacterial surveillance system that included several research protocols reviewed and approved by the Mozambican National Bioethics Committee for Health (IR00002657) and by Institutional Review Boards of Hospital Clinic of Barcelona, Spain; the US Centers for Disease Control and Prevention; and the School of Medicine, University of Maryland. Written informed consent was obtained from parents or caretakers of the eligible children.
Results
Serovars and PTs of the bloodstream isolates
From blood samples of 25 febrile children admitted to the MDH, 36 Salmonella isolates were recovered. Table 1 shows the relevant features of the patients, including sex and age, risk factors, symptoms and outcome. Fifteen isolates were assigned to S. Typhimurium and 21 isolates to S. Enteritidis. Both serovars were found in eleven samples, indicating a high frequency of mixed infections (Table 1). PT32 and U302 (shown by single isolates of S. Typhimurium), and PT1 (shown by two isolates of S. Enteritidis) were the only recognizable PTs associated with the detected isolates. In contrast, seven S. Typhimurium and 16 S. Enteritidis isolates displayed five and two RDNC patterns, respectively. The remaining isolates were UT (Tables 2 and 3).
Resistance properties of S. Typhimurium
As shown in Table 2, only three of the 15 invasive isolates of S. Typhimurium were susceptible to all antimicrobials tested. The remaining twelve were MDR, ie, resistant to three, five, or six unrelated antimicrobials, including ampicillin (blaTEM-1 ± blaOXA-1), chloramphenicol (catA1), gentamicin/kanamycin [aac(3)-IV and aadB], streptomycin (aadA1 ± strAB), sulfonamides (sul1 ± sul2), and trimethoprim (dfrA1 or not identified). Resistance to cephalosporins, nalidixic acid, ciprofloxacin, nitrofurantoin, or tetracycline was not detected. All MDR isolates carried one or two class 1 integrons, with three different variable regions: 2000 bp/blaOXA-1- aadA1, 700 bp/aadB, and 1600 bp/dfrA1-aadA1, but class 2 integrons were not found.
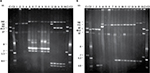
On the basis of plasmid content, the isolates could be distributed in three groups, characterized by the presence of pSLT (found in susceptible isolates), pSLT-A130-like plasmids or pSLT-BT-like plasmids (carried by seven and five MDR isolates, respectively; Figure 1A; Table 2). In agreement with this, all isolates were positive for the IncFIIA and IncFIB replicons and the spvC gene.
All except one of the pSLT-A130-like plasmids had a size of about 120 kb. These plasmids were found in S. Typhimurium isolates resistant to ampicillin, gentamicin/kanamycin, streptomycin, sulfonamides, and trimethoprim and carried the blaOXA-1, blaTEM-1, aadA1, and sul1 genes as well as the 2000 bp/blaOXA-1-aadA1 integron, as demonstrated by hybridization experiments of plasmid profiles (Table 2). In contrast, aac(3)-IV and the 700 bp/aadB integron, also present in these isolates, failed to hybridized with plasmid DNA. In a single isolate of the pSLT-A130-like group (resistant to ampicillin, streptomycin, and sulfonamides, but susceptible to gentamicin/kanamycin and trimethoprim), the plasmid was slightly smaller (~115 kb) but carried the same resistance genes and the 2000 bp/blaOXA-1-aadA1 integron of the 120 kb plasmids (Table 2). The pSLT derivatives were always accompanied by small ColE1 plasmids (~7 kb), which harbored a second copy of the blaTEM-1 gene and could be transferred into E. coli by transformation. Biparental and triparental conjugations, using four representative isolates as donors (Table 2), did not yield E. coli transconjugants under the applied conditions. This supports that the pSLT-A130-like plasmids are neither conjugative nor mobilizable.
pSLT-BT-like plasmids, of about 117 kb, were found in five isolates resistant to ampicillin, chloramphenicol, streptomycin, sulfonamides, and trimethoprim (Table 2). All resistance genes (blaTEM-1, catA1, aadA1, strAB, sul1, sul2, and dfrA1) as well as the 1600 bp/dfrA1-aadA1 integron found in these isolates were mapped on the plasmids. Additional cryptic plasmids of variable sizes (~84 and 50 kb, of undetermined incompatibility group, and ColE1 of ~3 kb) were also detected. Although the pSLT-BT-like plasmids failed to conjugate through biparental mattings, they could be mobilized from representative isolates (Table 2) into E. coli by triparental mattings. The obtained transconjugants had the same resistance phenotype and genes as the donor isolates.
Resistance properties of S. Enteritidis
With regard to S. Enteritidis (Table 3), only two of 21 invasive isolates were susceptible to all antimicrobials tested, while the remaining 19 were resistant to five, six, or seven unrelated antimicrobials. Resistance to ampicillin (blaTEM-1), chloramphenicol (catA2 ± catA1), nalidixic acid (due to a single mutation in gyrA, changing TCC/Ser83 to TAC/Tyr, in three isolates tested), streptomycin (strAB ± aadA1), sulfonamides (sul1 and sul2), and trimethoprim (dfrA7) was found in all MDR isolates, and all except two were also resistant to tetracycline [tet(A)]. In contrast, resistance to cephalosporins, ciprofloxacin, or nitrofurantoin was not detected. All isolates were negative for the integrase gene of class 1 and 2 integrons, but positive for the 700 bp/dfrA7 variable region, consistent with the presence of a defective class 1 integron.
pSEV was present in the two susceptible isolates, together with a small ColE1 plasmid of ~5 kb (Figure 1B; Table 3). Larger plasmids, ranging in size from ~95 to 130 kb, were found in all MDR isolates, and all except three carried a small ColE1 plasmid of about 6 kb. Through hybridization experiments, the blaTEM-1, catA2, strAB, sul1, sul2, tet(A), and dfrA7, and so the defective class 1 integron, were located on the larger plasmids. These plasmids also were hybridized with probes specific for the IncFIIA and IncFIB replicons, and for spvC, identifying them as derivatives of pSEV which, according to their resistance properties, appear to be related to pUO-SeVR1.9,10,27 Also coinciding with pUO-SeVR1, the pSEV derivatives from Mozambique could not be transferred into E. coli by biparental conjugation. However, 17 of the 21 pUO-SeVR1-like plasmids could be mobilized through triparental mattings. The exceptions were the two plasmids lacking tet(A). The obtained transconjugants displayed the resistance phenotype of the corresponding donor strain and carried all resistance genes except catA1, aadA1, and dfrA1, which did not alter the resistance phenotype since they are redundant with the catA2, strAB, and dfrA7 genes of plasmid origin.
PFGE and MLST analysis of the isolates
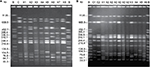
Eight XbaI-PFGE profiles were identified among invasive S. Typhimurium isolates (Figure 2A), differentially connected with susceptible (X1 and X2), pSLT-A130-like (X3, X4, and X5), and pSLT-BT-like (X6, X7, and X8) isolates. S. Enteritidis isolates showed six profiles (Figure 2B), with X1 associated with the susceptible isolates and X2–X6 with the MDR group. The latter profiles can readily be distinguished by a ~350 kb fragment, absent in susceptible isolates from Mozambique as well as in the control ATCC 13076. By MLST, the three susceptible S. Typhimurium isolates were assigned to ST19, while all MDR isolates belonged to ST313. In the case of S. Enteritidis, two susceptible and 15 MDR isolates were ST11. The remaining four MDR isolates were ST1479.
Discussion
Here we present the resistance properties and molecular types of S. Typhimurium and S. Enteritidis isolates recovered from blood samples taken from febrile children in rural Mozambique. As previously reported for iNTS, diarrhea, vomiting, and dehydration were infrequent, and most children presented risk factors, particularly anemia, severe malnutrition, and/or malaria.3 Like in other sub-Saharan countries, HIV infection is a well-recognized risk factor for iNTS in Mozambique, being highly prevalent in the population, with a high rate of mother-to-child transmission in the community of Manhiça.28–30 Unfortunately, no HIV data were available for the study period (2002–2004), since HIV counseling and testing in Manhiça District (southern Mozambique) were not established until 2005.
Blood infections with simultaneous detection of two Salmonella serovars have rarely been reported and mainly concerned S. Typhi and S. Paratyphi A coinfecting Asian patients or patients from developed countries who had traveled to Asia.31–35 Thus, a surprising result of the present study was the high number of patients coinfected with both S. Typhimurium and S. Enteritidis. However, a systematic assessment of mixed infections was not attempted, so well-designed comprehensive studies are required to determine the actual frequency and impact of coinfections, not only in Mozambique, but also in other African countries. Coinfections could increase the severity of the disease and led to treatment failure. The latter is of special relevance in low-income countries, due to the limited number of cost-effective therapeutic alternatives. Nonetheless, the two deaths reported in our series were not related with this type of infections.
MDR isolates of S. Typhimurium were ST313 and contained two kinds of pSLT derivatives that probably correspond to the pSLT-A130-like and pSLT-BT-like plasmids previously reported in lineages I and II of this invasive clone.7,8 According to Okoro et al8, chloramphenicol resistance conferred by pSLT-BT-like plasmids together with the use of this antibiotic as a drug of choice for the treatment of iNTS in Africa could have led to the replacement of lineage I by lineage II, which took place around 2003–2005. As indicated above, the isolates from Mozambique were collected along the period 2002–2004, which overlaps the time proposed for the replacement; therefore, temporal coincidence of the two kinds of plasmids was to be expected. Interestingly, despite being invasive, the three susceptible S. Typhimurium isolates belonged to ST19 and carried pSLT.
In contrast to S. Typhimurium, information on the STs of invasive S. Enteritidis from African countries is still limited. ST11, the most prevalent ST in this serovar, was detected in both susceptible and MDR isolates from Mozambique. Interestingly, four MDR isolates were assigned to ST1479, an uncommon ST that has only been reported in invasive UK isolates from patients linked to Africa.9 All MDR isolates carried pSEV derivatives that appear to be related to pUO-SeVR1, like those found in the UK isolates, as well as in those belonging to the Central/Eastern African clade, which characteristically carry the catA2 and tet(A) genes for chloramphenicol and tetracycline resistance.9–11,27 Interestingly, XbaI profiles of all S. Enteritidis isolates positive for pUO-SeVR1-like plasmids that were analyzed thus far included a distinct ~350 kb fragment, absent in susceptible isolates.10,36
Up till now little information exists on the PTs of S. Typhimurium and S. Enteritidis causing iNTS in Africa. In the present study, most isolates were UT or showed RDNC patterns, which did not coincide with any of those previously recognized in S. Typhimurium or S. Enteritidis. However, data available on PTs mainly were from European countries. In Kenya, UT and RDNC isolates have also been recovered from patients with gastroenteritis.37 Therefore, distinctive PTs appear to be circulating in Africa. PT42, the PT shown by invasive UK S. Enteritidis related to Africa,9 was not represented among the limited number of isolates tested in this study, although it has been reported previously in the study community.12
Conclusion
Results of the present study have revealed that 1) serovar-specific virulence plasmids were responsible for the MDR phenotype of S. Typhimurium and S. Enteritidis recovered from febrile children in Mozambique; 2) ST313 (S. Typhimurium), ST11, and the uncommon ST1479 (S. Enteritidis), as well as previously unrecognized PTs (UT and RDNC) were associated with these isolates; and 3) nearly half of the children were coinfected with both serovars, although there were no particularly poor outcomes in the affected patients.
Acknowledgments
The authors thank all the study participants (children and caretakers) and all workers of the CISM for their help in obtaining the data. Special thanks to Nélio Nobela, Marcelino Garrine, and all laboratory technicians from the CISM for their support on the laboratory-related activities and the clinicians for seeing the patients. The authors also thank the Manhiça district health authorities for their collaboration in the research activities ongoing in the Manhiça district.
This work was supported by projects UO-15-INVES-09 (Consejería de Educación, Cultura y Deporte, Principado de Asturias, Spain) and FIS PI11-00808 and FIS PI14CIII/00051 (Fondo de Investigación Sanitaria, Instituto de Salud Carlos III, Ministerio de Economía y Competitividad, Spain, cofunded by European Regional Development Fund of the European Union). CISM received core funding from the Spanish Agency for International Cooperation and Development. JR was supported by a fellowship from the program I3, of the ISCIII (Grant Number: CES11/012).
The present address of Vanesa García is Laboratorio de Referencia de E. coli, Departamento de Microbioloxía e Parasitoloxía, Facultade de Veterinaria, Universidade de Santiago de Compostela, 27002, Lugo, Spain.
Disclosure
The authors report no conflicts of interest in this work.
References
Morpeth SC, Ramadhani HO, Crump JA. Invasive non-Typhi Salmonella disease in Africa. Clin Infect Dis. 2009;49(4):606–611. | ||
Graham SM. Nontyphoidal salmonellosis in Africa. Curr Opin Infect Dis. 2010;23(5):409–414. | ||
Feasey NA, Dougan G, Kingsley RA, Heyderman RS, Gordon MA. Invasive non-typhoidal salmonella disease: an emerging and neglected tropical disease in Africa. Lancet. 2012;379(9835):2489–2499. | ||
Gordon MA, Graham SM, Walsh AL, et al. Epidemics of invasive Salmonella enterica serovar enteritidis and S. enterica Serovar typhimurium infection associated with multidrug resistance among adults and children in Malawi. Clin Infect Dis. 2008;46(7):963–969. | ||
Sigaúque B, Roca A, Bassat Q, et al. Severe pneumonia in Mozambican young children: clinical and radiological characteristics and risk factors. J Trop Pediatr. 2009;55(6):379–387. | ||
Reddy EA, Shaw AV, Crump JA. Community-acquired bloodstream infections in Africa: a systematic review and meta-analysis. Lancet Infect Dis. 2010;10(6):417–432. | ||
Kingsley RA, Msefula CL, Thomson NR, et al. Epidemic multiple drug resistant Salmonella Typhimurium causing invasive disease in sub-Saharan Africa have a distinct genotype. Genome Res. 2009;19(12):2279–2287. | ||
Okoro CK, Kingsley RA, Connor TR, et al. Intracontinental spread of human invasive Salmonella Typhimurium pathovariants in sub-Saharan Africa. Nat Genet. 2012;44(11):1215–1221. | ||
Rodríguez I, Rodicio MR, Guerra B, Hopkins KL. Potential international spread of multidrug-resistant invasive Salmonella enterica serovar enteritidis. Emerg Infect Dis. 2012;18(7):1173–1176. | ||
Rodríguez I, Guerra B, Mendoza MC, Rodicio MR. pUO-SeVR1 is an emergent virulence-resistance complex plasmid of Salmonella enterica serovar Enteritidis. J Antimicrob Chemother. 2011;66::218–220. | ||
Feasey NA, Hadfield J, Keddy KH, et al. Distinct Salmonella Enteritidis lineages associated with enterocolitis in high-income settings and invasive disease in low-income settings. Nat Genet. 2016;48(10):1211–1217. | ||
Mandomando I, Bassat Q, Sigauque B, et al. Invasive Salmonella infections among children from rural Mozambique, 2001-2014. Clin Infect Dis. 2015;61(Suppl 4):S339–S345. | ||
Mandomando I, Macete E, Sigauque B, et al. Invasive non-typhoidal Salmonella in Mozambican children. Trop Med Int Health. 2009;14(12):1467–1474. | ||
Murray PR, Baron E, Pfaller MA, Tenover FC, Yolken RH. Manual of Clinical Microbiology. 6th ed. Washington: American Society for Microbiology (ASM); 1995. | ||
Tennant SM, Diallo S, Levy H, et al. Identification by PCR of non-typhoidal Salmonella enterica serovars associated with invasive infections among febrile patients in Mali. PLoS Negl Trop Dis. 2010;4(3):e621. | ||
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing. 25rd informational supplement document M100-S25. Wayne, PA; CLSI, 2015. | ||
Guerra B, Junker E, Miko A, Helmuth R, Mendoza MC. Characterization and localization of drug resistance determinants in multidrug-resistant, integron-carrying Salmonella enterica serotype Typhimurium strains. Microb Drug Resist. 2004;10(2):83–91. | ||
García P, Guerra B, Bances M, Mendoza MC, Rodicio MR. IncA/C plasmids mediate antimicrobial resistance linked to virulence genes in the Spanish clone of the emerging Salmonella enterica serotype 4,[5],12:i:-. J Antimicrob Chemother. 2011;66(3):543–549. | ||
Frana TS, Carlson SA, Griffith RW. Relative distribution and conservation of genes encoding aminoglycoside-modifying enzymes in Salmonella enterica serotype typhimurium phage type DT104. Appl Environ Microbiol. 2001;67(1):445–448. | ||
Vila J, Ruiz J, Marco F, et al. Association between double mutation in gyrA gene of ciprofloxacin-resistant clinical isolates of Escherichia coli and MICs. Antimicrob Agents Chemother. 1994;38(10):2477–2479. | ||
Lévesque C, Piché L, Larose C, Roy PH. PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob Agents Chemother. 1995;39(1):185–191. | ||
White PA, McIver CJ, Rawlinson WD. Integrons and gene cassettes in the enterobacteriaceae. Antimicrob Agents Chemother. 2001;45(9):2658–2661. | ||
Kado CI, Liu ST. Rapid procedure for detection and isolation of large and small plasmids. J Bacteriol. 1981;145(3):1365–1373. | ||
Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63(3):219–228. | ||
García-Fernández A, Fortini D, Veldman K, Mevius D, Carattoli A. Characterization of plasmids harbouring qnrS1, qnrB2 and qnrB19 genes in Salmonella. J Antimicrob Chemother. 2009;63(2):274–281. | ||
Sambrook J, Russell DW. Molecular Cloning. A Laboratory Manual. 3rd ed. New York: Cold Spring Harbor Laboratory; 2001. | ||
García V, García P, Rodríguez I, Rodicio R, Rodicio MR. The role of IS26 in evolution of a derivative of the virulence plasmid of Salmonella enterica serovar Enteritidis which confers multiple drug resistance. Infect Genet Evol. 2016;45:246–249. | ||
Moon TD, Silva WP, Buene M, et al. Bacteremia as a cause of fever in ambulatory, HIV-infected Mozambican adults: results and policy implications from a prospective observational study. PLoS One. 2013;8(12):e83591. | ||
González R, Munguambe K, Aponte J, et al. High HIV prevalence in a southern semi-rural area of Mozambique: a community-based survey. HIV Med. 2012;13(10):581–588. | ||
Naniche D, Bardají A, Lahuerta M, et al. Impact of maternal human immunodeficiency virus infection on birth outcomes and infant survival in rural Mozambique. Am J Trop Med Hyg. 2009;80(5):870–876. | ||
Joshi S, Wattal C, Sharma A, Prasad KJ. Mixed Salmonella infection – a case report. Indian J Med Microbiol. 2002;20(2):113–114. | ||
Perera N, Geary C, Wiselka M, Rajakumar K, Andrew Swann R. Mixed Salmonella infection: case report and review of the literature. J Travel Med. 2007;14(2):134–135. | ||
Ochiai RL, Acosta CJ, Danovaro-Holliday MC, et al. A study of typhoid fever in five Asian countries: disease burden and implications for controls. Bull World Health Organ. 2008;86(4):260–268. | ||
Humphries RM, Yeganeh N, Ward KW, Lewinski MA, Ching N. Enteric fever in a 6-year-old traveler caused by Salmonella enterica serotypes Typhi and Paratyphi A: laboratory detection strategies and treatment options. J Clin Microbiol. 2011;49(1):452–454. | ||
Sultan BA, Fasih N, Qaiser S, Khan E, Zafar A, Irfan S. Mixed Salmonella infection: a case series from Pakistan. J Pak Med Assoc. 2013;63(4):524–526. | ||
Rodríguez I, Rodicio MR, Herrera-León S, Echeita A, Mendoza MC. Class 1 integrons in multidrug-resistant non-typhoidal Salmonella enterica isolated in Spain between 2002 and 2004. Int J Antimicrob Agents. 2008;32(2):158–164. | ||
Akinyemi KO, Philipp W, Beyer W, Bohm R. Application of phage typing and pulsed-field gel electrophoresis to analyse Salmonella enterica isolates from a suspected outbreak in Lagos, Nigeria. J Infect Dev Ctries. 2010;4(12):828–833. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.