Back to Journals » Infection and Drug Resistance » Volume 13
Saliva as an Alternative Specimen for Molecular COVID-19 Testing in Community Settings and Population-Based Screening
Authors Senok A , Alsuwaidi H, Atrah Y , Al Ayedi O, Al Zahid J , Han A , Al Marzooqi A, Al Heialy S, Altrabulsi B, AbdelWareth L, Idaghdour Y, Ali R, Loney T , Alsheikh-Ali A
Received 2 August 2020
Accepted for publication 20 August 2020
Published 1 October 2020 Volume 2020:13 Pages 3393—3399
DOI https://doi.org/10.2147/IDR.S275152
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sahil Khanna
Abiola Senok,1,* Hanan Alsuwaidi,1,* Yusrah Atrah,2 Ola Al Ayedi,3 Janan Al Zahid,2 Aaron Han,1 Asma Al Marzooqi,3 Saba Al Heialy,1,4 Basel Altrabulsi,5 Laila AbdelWareth,5,6 Youssef Idaghdour,7 Raghib Ali,7 Tom Loney,1 Alawi Alsheikh-Ali1
1College of Medicine, Mohammed Bin Rashid University of Medicine and Health Sciences, Dubai, United Arab Emirates; 2Molecular Department, Unilabs UAE, Dubai, United Arab Emirates; 3Al Khawaneej Health Center, Dubai Health Authority, Dubai, United Arab Emirates; 4Meakins-Christie Laboratories, Research Institute of the McGill University Health Center, Montreal, QC, Canada; 5National Reference Laboratory, Abu Dhabi, United Arab Emirates; 6Pathology and Laboratory Medicine Institute, Cleveland Clinic Abu Dhabi, Abu Dhabi, United Arab Emirates; 7Public Health Research Center, New York University Abu Dhabi, Abu Dhabi, United Arab Emirates
*These authors contributed equally to this work
Correspondence: Abiola Senok
College of Medicine, Mohammed Bin Rashid University of Medicine and Health Sciences, Building 14, Dubai Healthcare City, Dubai 505055, United Arab Emirates
Tel +97143838717
Email [email protected]
Purpose: With the easing of restriction measures, repeated community-based sampling for tracking new COVID-19 infections is anticipated for the next 6 to 12 months. A non-invasive, self-collected specimen like saliva will be useful for such public health surveillance. Investigations on the use of saliva for SARS-CoV-2 RT-PCR have largely been among COVID-19 in-pa\tients and symptomatic ambulatory patients with limited work in a community-based screening setting. This study was carried out to address this paucity of data and reported discrepancies in diagnostic accuracy for saliva samples.
Patients and Methods: From 29th June to 14th July 2020, adults presenting for COVID-19 testing at a community-based screening facility in Dubai, United Arab Emirates were recruited. Clinical data, nasopharyngeal swab in universal transport media and drooling saliva in sterile containers were obtained. Reverse transcriptase PCR amplification of SARS-CoV-2 RdRp and N genes was used to detect the presence of the SARS-CoV-2 virus.
Results: Of the 401 participants, 35 (8.7%) had viral detection in at least one specimen type and the majority (n=20/35; 57.1%) were asymptomatic. Both swab and saliva were positive in 19 (54.2%) patients, while 7 (20.0%) patients had swab positive/saliva negative results. There were 9 (25.7%) patients with saliva positive/swab negative result and this included 5 asymptomatic COVID-19 patients undergoing repeat screening. Using the swab as the reference gold standard, the sensitivity and specificity of saliva were 73.1% (95% CI 52.2– 88.4%) and 97.6% (95% CI 95.5– 98.9%) while the positive and negative predictive values were 67.9% (95% CI 51.5– 80.8%) and 98.1% (95% CI 96.5– 99.0%), respectively.
Conclusion: The findings suggest good diagnostic accuracy for saliva and feasibility of utilization of specimen without transport media for SARS-CoV-2 RT-PCR. Saliva represents a potential specimen of choice in community settings and population-based screening.
Keywords: SARS-CoV-2, nasopharyngeal swab, molecular test, population-based screening, saliva
Introduction
The impact of the ongoing COVID-19 pandemic on healthcare, social and economic structures across the world is unprecedented.1 With over 21 million laboratory-confirmed cases globally (as of 17th August 2020), widespread testing to rapidly identify SARS-CoV-2-infected individuals and ensure appropriate implementation of isolation measures and contact tracing has been a crucial element in the strategy to contain the pandemic. Nasopharyngeal swab (NPS) has been used as the gold standard respiratory specimen for SARS-CoV-2 reverse-transcriptase PCR (RT-PCR) tests. The collection of NPS is invasive and requires close contact between healthcare workers (HCWs) and patients. Therefore, to protect HCWs from the risk of viral transmission, the use of personal protective equipment (PPE) is mandated during specimen collection and this poses an additional strain on the resources of already stretched healthcare systems. Furthermore, patients experience a degree of discomfort during collection of the NPS specimens making it less acceptable particularly in children or when serial sampling is required.
As continued community screening is increasingly recognised as crucial for the rapid identification of new outbreak clusters, the need for a non-invasive specimen which can be self-collected by the patient without the need for supervision by trained personnel continues to be highlighted. In this respect, saliva represents an attractive choice of specimen especially as previous studies on the laboratory detection of respiratory viruses, including coronaviruses, in hospitalized patients have demonstrated a high concordance rate of over 90% in saliva and NPS specimens.2,3 Posterior oropharyngeal saliva which requires deep throat coughing has been shown to be sensitive in a cohort of hospitalized COVID-19 patients although the throat-clearing manoeuvre required is a limitation for the use of this approach for unsupervised self-collection.4 Neat saliva obtained by the drooling technique, which can be self-collected without supervision, has been reported to show greater sensitivity compared to NPS in hospitalized COVID-19 patients and at-risk HCW on COVID-19 wards.5–8 With the easing of lockdown restriction measures, repeated community-based sampling for rapid tracking of new infections in the population is anticipated and a non-invasive diagnostic specimen such as saliva will be useful for public health surveillance. Studies investigating the use of saliva for SARS-CoV-2 RT-PCR have largely been carried out among COVID-19 in-patients and symptomatic ambulatory patients with most of these studies being underpowered with small sample sizes. Furthermore, emerging data on the use of neat saliva samples for SARS-CoV-2 testing in the ambulatory setting have shown conflicting reports of diagnostic accuracy and the need for further adequately powered studies with larger sample sizes continues to be highlighted.9–11 Therefore, this study was carried out to assess the diagnostic accuracy of neat saliva sampling for SARS-CoV-2 screening in a community-based ambulatory setting compared to NPS.
Patients and Methods
Setting and Participants
This study has been reported according to the Standards for the Reporting of Diagnostic Accuracy Studies (STARD) guidelines 2015 (Supplementary Figure).12 The study was carried out from 29th June−14th July 2020 at the Al Khawaneej Health Center in Dubai, United Arab Emirates (UAE). This is a designated COVID-19 screening facility accessible to the general population. An average of 235 patients are screened daily at the center, including contacts of confirmed positive cases, those with presumptive symptoms with or without history of exposure as well as individuals concerned about possible infection. All adult patients undergoing COVID-19 testing at the center were eligible and informed consent was obtained from all participants. Ethical approval for the study was obtained from the Dubai Health Authority Research and Ethics Committee (DSREC-06/2020_15) and the Emirates Institutional Review Board for COVID-19 Research (DOH/CVDC/2020/1105).
Sample Size
We estimated a conservative infection prevalence of ~5.0% across our screening centers. Therefore, a minimum sample size of 400 participants (including 20 positive cases) was required to achieve a minimum power of at least 80% in order to detect a sensitivity of 80%, based on a target significance level of at least 0.05.13 We would also be able to detect a specificity of 95% as the minimum sample size to detect a specificity of 95% was 243 participants (including 12 positive cases).
Sample Collection
Paired swab and saliva samples were obtained from all participants. The swab specimens were obtained by trained healthcare personnel using the standard technique for NPS collection as per COVID-19 screening protocol already in place at the facility. Swabs were placed in the Greiner Bio-One universal transport system (Greiner Bio-One, Kremsmünster, Austria) for transport to the diagnostic laboratory. To ensure consistency in saliva collection, the healthcare personnel were trained in giving instructions to the patients for saliva collection and a unified script was used. Saliva was collected using sterile containers without transport medium and samples were obtained at least one hour after the patient last consumed food, fluid, or smoked tobacco. Patients were asked to pool saliva in their mouth for 1–2 minutes and then gently spit 2–4 mL of saliva into the provided sterile container. Demographic data and clinical information including indication for screening, history of co-morbidities, COVID-19 symptoms and vital signs were recorded for all participants. Paired swab and saliva samples were stored at room temperature and transferred to the diagnostic laboratory within three hours of collection.
SARS-CoV-2 Detection
Samples were processed for SARS-CoV-2 detection at Unilabs, Dubai, UAE which is a diagnostic laboratory approved for COVID-19 testing by the health authority and accredited by the EIAC accreditation body for COVID-19 testing under the ISO 15189:2012 standards. A record of the time of specimen collection and the time of arrival at the diagnostic laboratory was kept. All samples were processed immediately on arrival at the diagnostic laboratory. Viral RNA was extracted from 300 µL of each sample using the Chemagic viral RNA extraction kit on the automated Chemagic™ 360 Nucleic Acid Extractor (PerkinElmer, Baesweiler, Germany) according to the manufacturer’s instructions. Detection of SARS-CoV-2 was carried out using the NeoPlex COVID-19 kit (GeneMatrix, Seoul, South Korea) for the RT-PCR amplification of the SARS-CoV-2 RdRp and N gene targets. The kit contains specific primers and dual-labelled probes for the amplification and simultaneous differentiation of SARS-CoV-2 and other beta-coronaviruses. The preparation of master mix and RT-PCR was carried out in accordance with manufacturer-provided instructions. Briefly, the PCR master mix was prepared according to manufacturer-provided protocol to give a final volume per test of COVID-19 PPM 5 µL, One-step master mix 5 µL and 5 µL DW (RNase-free water). The master mix was vortexed and 15 µL aliquots were placed into 0.2 mL PCR tubes followed by addition of 5 µL of extracted patient sample nucleic acid. The cycling protocol comprised of a first segment of one cycle of 50°C for 30 minutes followed by the second segment of a single cycle of 95°C for 15 minutes. This was followed by 40 cycles of 95°C for 15 seconds and 60°C for 60 seconds. Cycle threshold of ≤40 was taken as cut off for positive result for both target genes. The results were interpreted in accordance with the UAE Federal guidelines (NCEMA UAE Expert Panel-May 2020) which are in line with manufacturer guidelines, and a “detected” result for SARS-CoV-2 was rendered if two gene targets or only the RdRp gene target was amplified and “non-detected” result if all gene targets were not amplified, while the manufacturer provided internal, negative and positive controls as well as third party negative and positive controls included in every run remained validated.
Statistical Analysis
Data were entered and analysed using SPSS statistical software version 24. Descriptive statistics for categorical variables are presented as number (percent) and for continuous variables as mean ±standard deviation (SD) or median (interquartile range; IQR). Comparison of means was carried out using Student’s t-test with statistical significance at 0.05. The sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV) and a 95% confidence interval (CI) were calculated. Kappa coefficient was used to estimate agreement between swab and saliva RT-PCR test results.
Results
A total of 401 participants who provided both swab and saliva specimens were enrolled. The mean age (± SD) of participants was 35.5 ± 9.5 years and the majority were males (n/N=329/401; 82.0%). History of contact with confirmed COVID-19 patients was obtained in 176 (43.9%) participants and 48.6% (n=195) reported that they were asymptomatic. Majority of samples arrived in the laboratory within 2–3 hours of collection with median (IQR) of 2 hours 48 minutes (1 hr 6 mins – 2 hrs 45 mins). The volume of saliva produced by the majority of patients was 2–3 mL and viral detection in both swab and saliva was recorded for three patients who had a saliva volume of 1 mL.
A total of 35 (8.7%) patients showed viral detection by SARS-CoV-2 RT-PCR from at least one specimen type and both the RdRp and N gene targets were detectable in all positive samples. The overall prevalence for COVID-19 diagnosis by swab RT-PCR was 6.5% (n/N=26/401) and 7.0% by saliva RT-PCR (n/N=28/401). Among the 35 patients that had a positive test by either specimen, 20 (57.1%) were asymptomatic and whilst 12 patients self-reported having fever, only four were febrile with temperature ≥37.5°C (mean ± SD 38.0 ± 0.3°C). Other self-reported symptoms were sore-throat (n=12), muscle pain (n=6) and two patients each reported shortness of breath, anosmia, and loss of taste. The median (IQR) onset of symptoms prior to the test was 2.5 (1–14) days. Table 1 shows the demographic and clinical characteristics of all patients who tested positive for SARS-CoV-2.
 |
Table 1 Profile of Patients with SARS-CoV-2 RT-PCR Positive Finding |
Among the 35 patients with positive viral detection, both swab and saliva were positive in 19 (54.3%), while 7 (20.0%) had swab positive/saliva negative results. There were 9 (25.7%) patients with saliva positive/swab negative result and five of these were confirmed asymptomatic COVID-19 patients undergoing repeat screening. The mean (± SD) duration since the last test result in these 5 patients was 14 ± 7 days. Table 2 shows the distribution of SARS-CoV-2 detection in swab and saliva specimens. Using the swab as the reference gold standard, the sensitivity and specificity of saliva was 73.1 % (95% CI 52.2–88.4%) and 97.6% (95% CI 95.5–98.9%), respectively. The PPV and NPV were 67.9% (95% CI 51.5–80.8%) and 98.1% (95% CI 96.5–99.0%), respectively. The accuracy was 96.0% (95% CI 93.6–97.7%) and Kappa coefficient was 0.68 (95% CI 0.53–0.82).
 |
Table 2 Viral Detection in Saliva and Swab Specimens |
Among the 19 patients with paired positive samples, the median cycle threshold (Ct) values for RdRp and N gene targets were 29.6 (IQR 24.8–35.0) and 28.9 (IQR 24.5–35.0), respectively, in saliva; while the swab Ct values were RdRp 28.1 (IQR 22.6–31.8) and 28.0 (IQR 23.1–32.3) N gene target (Figure 1 shows the Ct values for paired samples). RdRp and N gene median Ct of 36.1 (IQR 33.9–37.1) and 36.3 (IQR 34.0–37.5), respectively, were found in saliva samples (n=9) which had a negative paired swab. Similarly, median RdRp and N gene Ct was 36.4 (IQR 35.1–38.6) and 37.5 (IQR 32.5–38.2), respectively, for swab samples (n=7) which had paired negative saliva. Comparison of Ct values in patients with both saliva and swab positive samples versus those with either saliva or swab only positive showed significantly higher Ct values for both target genes in the latter (p<0.05; Figure 2).
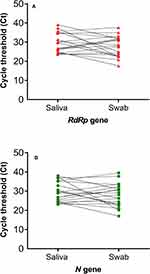 |
Figure 1 SARS-CoV-2 RT-PCR cycle threshold (Ct) values for the target genes in patients with paired positive samples (i.e. both swab and saliva samples positive). |
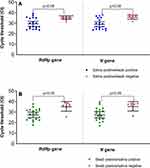 |
Figure 2 Comparison of RT-PCR cycle threshold (Ct) in paired positive swab and saliva versus samples with only one specimen type positive. |
Discussion
The findings from this study indicate that neat saliva has good diagnostic accuracy when compared to the gold standard NPS and it has the potential to be a specimen of choice in community settings and population-based screening, especially for children. As communities re-open and restriction measures are eased, repeated sampling of individuals might become necessary for the rapid identification of new infection clusters. Hence, an understanding of the usefulness of saliva in a community-based screening setting is crucial. This study was carried out in the context of a busy community-based screening facility which caters to a population with diverse levels of risk potential and absent or mild symptoms. Our findings show that in such a setting, neat saliva is useful for SARS-CoV-2 RT-PCR, particularly in asymptomatic patients. These findings are in agreement with previous work investigating ambulatory patients in a busy screening clinic in Melbourne, Australia.11 Furthermore, we also demonstrate the feasibility of using saliva without the addition of transport media. During the ongoing pandemic, the cost of sample collection devices with transport media and challenges with supply logistics significantly impacted screening capability of healthcare systems particularly in resource-limited settings. The ability to use neat saliva will be crucial in addressing such challenges especially given the potential future scenario of repeated population screening. The timeframe from sample collection to arrival in the laboratory for processing we have shown in this study is envisaged to be feasible for high-volume community-based screening centers. In similar work by Williams et al,11 samples were transported to the laboratory (median time of three hours) without transport media with the addition of Amies media in the laboratory.
In previous comparative studies of upper and lower respiratory samples for SARS-CoV-2 RT-PCR, the sensitivity for NPS specimen was 63–73% which is similar to our reported sensitivity for saliva.8,14,15 Although most of the indices for diagnostic accuracy in this study are aligned with those reported in similar studies in cohorts of hospitalized COVID-19 patients and symptomatic ambulatory individuals, our sensitivity and positive predictive values are on the lower end.5,7,9,10 This difference is probably due to clinical heterogeneity in patients (48.6% asymptomatic in our sample). As this study was carried out in a screening center, patients with severe COVID-19 did not form part of our study population. It has been shown that higher sensitivity is consistently seen in those with severe disease probably reflecting the higher viral load associated with COVID-19 severity.16 Indeed, our finding that patients with lower Ct values (indicative of higher viral load) are more likely to have combined swab and saliva positive results is consistent with reported literature on the association between test sensitivity and high viral load. Diagnostic accuracy is crucial for the use of saliva in the community setting wherein lower prevalence with a predominance of asymptomatic and mildly symptomatic cases are expected. In light of these, it is interesting that our findings also demonstrate that half of the patients who were identified only by their saliva RT-PCR findings were confirmed COVID-19 patients undergoing retesting and they would have been missed based on their negative swab results. However, in keeping with previous literature, our data also indicate that neither saliva or swab specimen are 100% sensitive for the detection of SARS-CoV-2 as we also had seven patients with swab positive/saliva negative results.7,8 Although the use of combined saliva and swab specimens for increased diagnostic accuracy has been suggested,7 in a scenario wherein repeated population sampling might be expected, our findings support the notion that saliva represents an alternative low-cost diagnostic specimen for SARS-CoV-2 screening particularly with the associated ease of collection and minimal risk for HCWs. A limitation of this study was that we only used one type of SARS-CoV-2 RT-PCR kit; hence, we suggest further investigations using kits for detection of other viral gene targets. As there are limited studies carried out in the context of community-based screening, we recommend additional studies for further evaluation of saliva for SARS-CoV-2 screening in these settings possibly including children and the elderly as well as multiple sampling for temporal profiling.
Conclusion
The findings suggest good diagnostic accuracy for neat saliva and indicate the feasibility of utilization of saliva specimen without transport media for SARS-CoV-2 RT-PCR. Saliva represents a potential specimen of choice in community settings and population-based screening, especially as a non-invasive alternative for children and the elderly.
Data Sharing Statement
The data supporting the findings of this study are available within the article and are available from the corresponding author on reasonable request.
Funding
There is no funding to report.
Disclosure
YA, JAZ are employees of Unilabs UAE but this did not influence the study design, and they have no competing interests to declare. None of the other authors has any financial or other relationships that may constitute a conflict of interest.
References
1. Uddin M, Mustafa F, Rizvi TA, et al. SARS-CoV-2/COVID-19: viral genomics, epidemiology, vaccines, and therapeutic interventions. Viruses. 2020;12(5):526. doi:10.3390/v12050526
2. To KK, Lu L, Yip CC, et al. Additional molecular testing of saliva specimens improves the detection of respiratory viruses. Emerg Microbes Infect. 2017;6(6):e49. doi:10.1038/emi.2017.35
3. To KKW, Yip CCY, Lai CYW, et al. Saliva as a diagnostic specimen for testing respiratory virus by a point-of-care molecular assay: a diagnostic validity study. Clin Microbiol Infect. 2019;25(3):372–378. doi:10.1016/j.cmi.2018.06.009
4. To KK-W, Tsang OT-Y, Leung W-S, et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20(5):565–574. doi:10.1016/S1473-3099(20)30196-1
5. Azzi L, Carcano G, Gianfagna F, et al. Saliva is a reliable tool to detect SARS-CoV-2. J Infect. 2020;81(1):e45–e50. doi:10.1016/j.jinf.2020.04.005
6. Wyllie AL, Fournier J, Casanovas-Massana A, et al. Saliva is more sensitive for SARS-CoV-2 detection in COVID-19 patients than nasopharyngeal swabs. medRxiv. 2020;2020:
7. Jamal AJ, Mozafarihashjin M, Coomes E, et al. Sensitivity of nasopharyngeal swabs and saliva for the detection of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Clin Infect Dis. 2020. doi:10.1093/cid/ciaa848
8. Wang W, Xu Y, Gao R, et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA. 2020;323(18):1843–1844.
9. Pasomsub E, Watcharananan SP, Boonyawat K, et al. Saliva sample as a non-invasive specimen for the diagnosis of coronavirus disease-2019 (COVID-19): a cross-sectional study. Clin Microbiol Infect. 2020. doi:10.1016/j.cmi.2020.05.001
10. Skolimowska K, Rayment M, Jones R, Madona P, Moore LS, Randell P. Non-invasive saliva specimens for the diagnosis of COVID-19: caution in mild outpatient cohorts with low prevalence. Clin Microbiol Infect. 2020. doi:10.1016/j.cmi.2020.07.015
11. Williams E, Bond K, Zhang B, Putland M, Williamson DA, McAdam AJ. Saliva as a non-invasive specimen for detection of SARS-CoV-2. J Clin Microbiol. 2020;58(8):e00776–e00720. doi:10.1128/JCM.00776-20
12. Bossuyt PM, Reitsma JB, Bruns DE, et al. STARD 2015: an updated list of essential items for reporting diagnostic accuracy studies. BMJ. 2015;351:h5527. doi:10.1136/bmj.h5527
13. Bujang MA, Adnan TH. Requirements for minimum sample size for sensitivity and specificity analysis. J Clin Diagn Res. 2016;10(10):YE01–YE06. doi:10.7860/JCDR/2016/18129.8744
14. Chan JF, Yip CC, To KK, et al. Improved molecular diagnosis of COVID-19 by the novel, highly sensitive and specific COVID-19-RdRp/Hel real-time reverse transcription-PCR assay validated in vitro and with clinical specimens. J Clin Microbiol. 2020;58(5):e00310–e00320. doi:10.1128/JCM.00310-20
15. Yang Y, Yang M, Shen C, et al. Evaluating the accuracy of different respiratory specimens in the laboratory diagnosis and monitoring the viral shedding of 2019-nCoV infections. medRxiv. 2020;2020:
16. Liu Y, Yan LM, Wan L, et al. Viral dynamics in mild and severe cases of COVID-19. Lancet Infect Dis. 2020;20(6):656–657. doi:10.1016/S1473-3099(20)30232-2
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
