Back to Journals » OncoTargets and Therapy » Volume 12
PRKD2 Promotes Progression and Chemoresistance of AML via Regulating Notch1 Pathway
Authors Liu Q, Li W, Zhou Y, Jian J, Han S, Liu C, Li W , Zhu X , Ma D, Ji M , Ji C
Received 2 October 2019
Accepted for publication 29 November 2019
Published 12 December 2019 Volume 2019:12 Pages 10931—10941
DOI https://doi.org/10.2147/OTT.S233234
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Qian Liu,1,2 Wei Li,1 Ying Zhou,1 Jimo Jian,1,3 Shijie Han,4 Chao Liu,5 Wei Li,1 Xunxun Zhu,6 Daoxin Ma,1,7 Min Ji,1 Chunyan Ji1
1Department of Hematology, Qilu Hospital, Shandong University, Jinan 250012, People’s Republic of China; 2Department of Pain, Qilu Hospital, Shandong University, Jinan 250012, People’s Republic of China; 3Department of Hematology, Qilu Hospital, Shandong University (Qingdao), Qingdao 266000, People’s Republic of China; 4Department of Orthopedics, Shandong Provincial Hospital Affiliated to Shandong University, Jinan 250021, People’s Republic of China; 5Department of Oral and Maxillofacial Surgery, Shandong Provincial Hospital Affiliated to Shandong University, Jinan 250012, People’s Republic of China; 6Department of Hematology, Tengzhou Central People’s Hospital, Tengzhou 277599, People’s Republic of China; 7Shandong Provincial Key Laboratory of Immunohematology, Qilu Hospital, Shandong University, Jinan 250012, People’s Republic of China
Correspondence: Min Ji
Department of Hematology, Qilu Hospital, Shandong University, West Wenhua Road, Jinan 250012, People’s Republic of China
Email [email protected]
Introduction: Progression and chemoresistance of acute myeloid leukemia (AML) contribute to most of the treatment failure. Notch pathway has been proven to be involved in many biological processes and diseases, especially AML. In this study, we aimed to explore genes correlated with Notch1 pathway in AML and determine their roles in the regulation of AML progression and chemoresistance.
Methods: TCGA database was used to explore Notch1 associated genes. Kaplan–Meier survival analysis was performed to evaluate the prognostic significance of genes. Quantitative RT-PCR (qRT-PCR) and Western blot were performed to examine the expression of genes. The expression of PRKD2 was up-regulated or knocked down in AML cell lines by lentivirus or siRNAs. CCK-8 and flow cytometry were used to analyze the effect of PRKD2 on cell proliferation and chemoresistance.
Results: Based on TCGA database, PRKD2 was found to be positively correlated with Notch1 expression, cytogenetic risk status and poorer prognosis in AML. Moreover, the expression level of PRKD2 was higher in AML chemo-resistant cells than in chemo-sensitive cells. Functionally, knockdown of PRKD2-induced apoptosis and increased chemosensitivity of AML cells. PRKD2 overexpression promoted proliferation and chemoresistance of AML cells. Furthermore, we found PRKD2 could regulate Notch1 pathway. Besides, high PRKD2 expression was correlated with higher risk group of AML patients which indicated that PRKD2 was an independent prognostic marker for AML.
Conclusion: Taken together, our results showed that PRKD2 could promote the proliferation and chemoresistance of AML cells by regulating Notch1 pathway. The study broadened our insights into the underlying mechanisms in chemoresistance and proliferation of AML, and provided a new prognostic marker and treatment target for AML.
Keywords: PRDK2, AML, progression, chemoresistance, Notch1
Introduction
Acute myeloid leukemia (AML) is the most common and aggressive type of acute leukemia, which is characterized by the uncontrolled proliferation of myeloid progenitor cells leading to abnormal accumulation of immature precursors and insufficient generation of normal mature blood cells.1,2 It has been reported that AML is the most common cause of leukemia-related mortality in adult patients with a 5-year overall survival (OS) around 40%.3 Despite the use of new target drugs such as FLT3 inhibitors and IDH1/2 inhibitors in the clinic, the mainstay of treatment approach is chemotherapy and several chemotherapeutic drugs including anthracycline are predominantly used.4–6 Although great prognostic improvement of AML has been made during the past few decades, 60–80% of patients who have initially achieved complete remission finally relapse while parts of them eventually die of this disease due to chemoresistance.7 Therefore, it is of great significance to elucidate the underlying mechanisms of chemoresistance in AML and develop novel therapeutic strategies for reversing chemoresistance.
Notch signaling pathway is a highly conserved cell signaling system present in most multicellular organisms.8 Recent studies show that Notch signaling pathway plays crucial roles in the regulation of cell-cell contacts, cell survival, proliferation, differentiation, as well as fate determination during multiple physiological processes.9,10 Because of its potential in maintaining the balance between cell survival and apoptosis, its roles in tumor initiation and progression have been evaluated previously. For instance, Baker et al reported Notch1-PTEN-ERK1/2 signaling axis promoted HER2+ breast cancer cell proliferation and supported cancer stem cell survival.11 Another study showed that patients with lung adenocarcinoma had higher Notch1 expression compared to normal controls and overexpression of Notch1 also suggested a higher rate of cancer recurrence.12 Moreover, we previously reported that Notch1/Dll4 pathway was activated in AML cells and could promote disease progression.13 Besides, we found that activation of Notch1 pathway may indicate an unfavorable prognosis in AML.14 Since Nothch1 pathway has played critical roles in AML pathogenesis, it is of great value to further explore its regulatory network.
In this study, we analyzed TCGA database and found that PRKD2 was positively correlated with Notch1 expression and its overexpression predicted poorer prognosis in AML patients. Further results showed that PRKD2 could promote cell proliferation and chemoresistance, as well as inhibit drug-induced apoptosis via activating Notch1 pathway in AML cells. In conclusion, our study demonstrated biological functions, molecular mechanisms and clinical significance of PRKD2/Notch1 regulation axis in AML, and PRKD2 may hopefully serve as a new prognostic marker and treatment target for AML.
Materials and Methods
TCGA RNA Sequencing and Clinical Data
The RNA-seq and clinical profiles of 134 newly diagnosed AML patients (non-APL) were downloaded from TCGA database, which contains RNA sequencing data for multiple types of cancer.15 The RNA-seq data were analyzed using R packages.
Cell Lines and Cell Culture
Human AML cell lines K562, K562/A02, KASUMI and U937 were purchased from the Institute of Hematology & Blood Diseases Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Tianjin, China. Cells were cultured in RPMI 1640 (Invitrogen, Carlsbad, CA) supplemented with 10% fetal bovine serum (FBS). The multidrug-resistant cell line K562/A02 was maintained in the medium supplemented with 1 mg/l adriamycin (ADM) which was removed from the medium 2 weeks before experiments. All cells were cultured in a 37°C humidified incubator with 5% CO2.
In vitro Transfection with Lentivirus or Synthetic siRNAs
Lentivirus that stably overexpressed PRDK2 was purchased from GeneChem Corporation (Shanghai, China). Cells infected with lentivirus were then cultured with puromycin to select the PRKD2 overexpression cell lines. siRNAs of PRKD2 were purchased from GenePharma Corporation (Shanghai GenePharma Co, Ltd, Shanghai, China). siRNAs were transfected into AML cells using Lipofectamine® RNAiMAX Reagent (Thermo Fisher Scientific, Inc.) according to the provided protocol. The siRNA sequences of PRKD2 were shown as follows:
siRNA-1: 5ʹ-AAUGACCUUAACUGCCACGUCCCGG-3ʹ;
siRNA-2: 5ʹ-CCUGAGUGUGGCUUCUACGGCCUUU-3ʹ.
Cell Viability Assay
Cell viability was determined by CCK8 assays. In brief, cells were seeded in 96-well plates in culture medium and incubated in 5% CO2 at 37°C. After incubation for the indicated time, 10 μL of CCK8 was added into each well and incubated for 4 hrs. The optical density (OD) was measured at a wavelength of 450 nm on a Microplate Reader (Bio-Rad).
Cell Apoptosis Assay
Cell apoptosis was detected using an Annexin V-FITC/PI apoptosis detection kit (JingMei Biotech, Beijing, China) according to the manufacturer’s protocols. Briefly, cells were treated with EDTA-free trypsin, and resuspended with 400 μL Annexin binding buffer at a concentration of 106 cells/mL. Then, 5 μL FITC-conjugated Annexin V and 10 μL PI were added to the cells and incubated at room temperature for 15 mins in dark. Cell apoptosis assay was performed within 1 hr after staining by Becton Dickinson FACS Calibur flow cytometer.
Quantitative RT-PCR Analysis
Total RNA was extracted from cells using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). PrimeScript reverse transcriptase (RT) reagent kit (TaKaRa, Shiga, Japan) was used to synthesize cDNA from total RNA. Real-time PCR was performed on Applied Biosystems StepOne plus System, and results were analyzed as previously described.16
Western Blot Analysis
Cells were harvested and lysed in lysis buffer containing protease inhibitors. Subsequently, 50 μg of total cellular protein from each sample was separated by 10% SDS-PAGE and electro-transferred onto polyvinylidene fluoride (PVDF) membrane using a semi-dry blotting apparatus (Bio-Rad, Hercules, CA, USA). The membranes were blocked with 5% nonfat milk at room temperature for 1 hr, and then incubated with primary antibodies overnight at 4°C. After incubation with the appropriate secondary antibodies, the protein bands were detected using the Pro-lighting HRP agent. Expression of β-actin was used as a loading control. The antibodies used in this study were obtained from Abcam: PRKD2 (ab51251), NOTCH1 (ab8925), NICD (ab83232), HES1 (ab108937), β-actin (ab8226).
Statistical Analysis
All data were presented as means ± S.D. from at least three independent experiments. The software SPSS V18.0 was used for statistical analysis. Spearman correlation test was used to test the correlation between gene expression and Notch1. Statistical significance of differences between two groups was evaluated using Student’s t-test, and one-way ANOVA was used to determine the significance of differences among multiple groups. Pearson’s Chi-square test was used to compare the clinicopathological characteristics between groups. The mean expression value of each gene in our cohort had been calculated and patients were separated into two groups based on mean expression value. Survival curves were generated using the Kaplan–Meier method, and differences between curves were analyzed with log-rank test. Differences with P < 0.05 were considered statistically significant.
Results
TCGA RNA Sequencing and Clinical Data
TCGA database contains RNA sequencing data for multiple types of cancer. The RNA-seq and clinical profiles of 134 newly diagnosed AML patients (non-APL) were downloaded and analyzed. AML patients were classified into three groups according to cytogenetic risk status. Detailed clinical characteristics of AML patients are shown in Table 1.
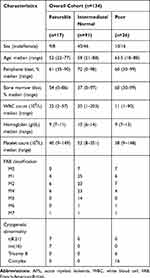 |
Table 1 Clinical Characteristics of AML Patients |
PRKD2 Expression Was Positively Correlated with Notch1 and Its Up-Regulation was Associated with Poorer Prognosis and Drug Resistance in AML
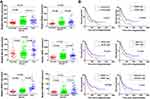
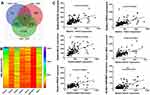
To identify Notch1 related genes, we firstly analyzed the TCGA database, from which the correlation between expression of Notch1 and all the other genes was studied. A total of 172 genes with correlation coefficient > 0.5 and P <0.05 were filtered. To further predict functional genes, we analyzed the association between gene expression and cytogenetic risk status. 5931 genes were found correlated with AML risk status. Then, we analyzed the association between gene expression and prognosis of AML patients, and 1584 genes were demonstrated prognostically significant. Finally, 3 groups of filtered genes mentioned above were intersected with each other and only 19 genes that simultaneously met three conditions showed up (Figure 1A). After reviewing articles, 6 genes (PACS2, PRKD2, BCL9L, SIPA1, PPM1F, RAPGEF3) which were previously published to be functional in cancers were selected. The relative mRNA expression of 6 genes in AML patients was shown in Figure 1B. Moreover, the gene expression correlation with Notch1, risk status and prognosis is shown in Figures 1C and 2A, B.
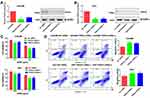
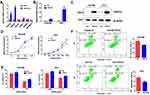
In addition, we detected the expression of the 6 genes in a drug-resistant cell line (K562/A02) and its parental cell line (K562). As shown in Figure 3A, PACS2, PRKD2 and SIPA1 were significantly up-regulated in K562/A02 cells with PRKD2 most remarkably changed, which indicated that PRKD2 might also play crucial roles in the drug resistance of AML.
Overexpression of PRKD2 Promoted Proliferation and Chemoresistance of AML Cells
It was well known that chemoresistance resulted in fast progression of neoplasms. As we had explored that PRKD2 was highly expressed in K562/A02 cells, we wondered whether PRKD2 was involved in the chemoresistance of AML cells. To better understand the roles and mechanisms of PRKD2 in AML, we infected KASUMI and U937 cell lines with lentivirus that stably overexpressed PRDK2 (Figure 3B and C). We firstly verified the effect of PRKD2 overexpression on cell proliferation, and we found the up-regulation of PRKD2-promoted proliferation in both KASUMI and U937 cell lines (Figure 3D). Moreover, we confirmed that PRKD2 overexpression significantly increased the chemoresistance of AML cells to ADM (Figure 3E). What is more, we found ADM induced apoptosis was decreased in AML cell lines with PRKD2 up-regulation (Figure 3F), which indicated that PRKD2-regulated proliferation and chemoresistance via induction of apoptosis.
PRKD2 Knockdown Suppressed Chemoresistance via Induction of Apoptosis in AML Cells
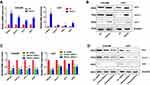
Based on the above results, we have showed that overexpression of PRKD2 contributed to chemoresistance of AML cells by suppressing cell apoptosis. To further validate our results, we transfected both KASUMI and U937 cells with PRKD2 siRNAs, and the expression of PRKD2 was downregulated on both mRNA and protein levels (Figure 4A and B). We found knockdown of PRKD2 sensitized both cell lines to ADM (Figure 4C). Apoptosis assay was also conducted and the ADM induced apoptosis was remarkably increased after PRKD2 knockdown (Figure 4D). In brief, our findings further demonstrated that inhibition of PRKD2 could suppress the chemoresistance of AML cells via induction of apoptosis.
PRKD2 Expression Was Correlated with Notch1 Pathway in AML Cells
To further clarify the association between PRKD2 and Notch1 signaling pathway in AML cells, key genes in Notch1 pathway were detected by qRT-PCR and Western blot after modulating PRKD2 expression. As shown in Figure 5A and C, we presented that PRKD2 expression was positively correlated with Notch1, NICD and HES1 on mRNA levels. Similarly, PRDK2 could also regulate the activity of Notch1 pathway on protein levels (Figure 5B and D). Based on these results, we proposed that PRKD2 may exert effects on AML drug resistance via manipulating the Notch1 pathway.
PRKD2 Overexpression Was Associated with Older Age and Higher Cytogenetic Risk in Newly Diagnosed AML Patients
As we have shown the crucial roles and underlying mechanisms of PRKD2 in AML drug resistance, we further divided patients into two groups based on PRKD2 mean expression level. As shown in Table 2, characteristics of 47 patients with low PRKD2 expression and 87 patients with high PRKD2 expression were analyzed. High expression of PRKD2 was associated with older age and higher cytogenetic risk, which might lead to poorer prognosis in AML patients.
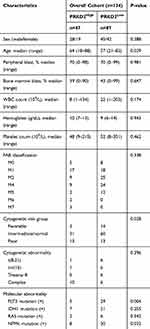 |
Table 2 Association of PRKD2 Expression with Clinical Pathologic Factors of AML Patients |
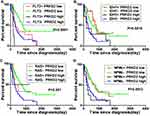
AML is characterized by clonal evolution and genetic heterogeneity.17 Genetic alterations including amplifications, deletions, rearrangements, and point mutations are recurrent.18 Here, we also analyzed the association between PRKD2 expression and key genomic mutations including FLT3, IDH1, RAS and NPMc mutations in AML patients. As shown in Figure 6, high expression of PRKD2 predicted poorer prognosis in both FLT3-mutated (P=0.002) and wild-type (P<0.001) AML patients. FLT3 mutation was an unfavorable factor in PRDK2 low expression group (P=0.019) but not in PRDK2 high expression patients (P=0.051). For IDH1 and RAS, high expression of PRKD2 only predicted poorer prognosis in wild-type IDH1 or RAS AML patients. For NPMc, our results showed PRKD2 expression was an independent prognostic marker as high expression of PRKD2 contributed to poorer prognosis in both NPMc-mutated and non-mutated AML patients.
Discussion
Chemotherapy resistance still remains a major barrier to the clinical treatment and prognosis of AML patients. Therefore, it is of great significance to explore the molecular mechanisms of chemoresistance and identify novel targets for clinical therapy. In this study, we found PRKD2 expression was positively correlated with the expression of key genes in Notch1 pathway, cytogenetic risk status and poorer prognosis in AML patients. Moreover, PRKD2 was up-regulated in chemo-resistant AML cells. Knockdown of PRKD2 led to increased sensitivity of AML cells to ADM via induction of apoptosis. Overexpression of PRKD2 further promoted ADM resistance and proliferation of AML cells. Moreover, we found PRKD2 could regulate Notch1 and HES1 expression which were the key genes of Notch1 pathway. Finally, we demonstrated that PRKD2 expression was associated with several AML patient characteristics. Based on our results, PRKD2 might serve as a promising therapeutic target for AML in clinic.
Exploring the molecular mechanisms underlying AML chemoresistance could contribute to developing reasonable and effective therapeutic strategies to overcome chemoresistance. Notch1 pathway has been proven to serve as key regulators of chemoresistance in numerous cancers.19–21 For instance, previous studies reported that Notch1 controlled cell chemoresistance in small cell lung carcinoma cells,19 enhanced prostate cancer chemoresistance,20 regulated chemoresistance of gastric cancer stem cells via induction of autophagy.21 To further discover Notch1 related genes, we downloaded and analyzed data of AML patients from TCGA database, which indicated PRKD2 was positively correlated with Notch1 expression, higher risk status and poorer prognosis in AML patients. Moreover, PRKD2 were also up-regulated in drug-resistant AML cells. The protein kinase D (PRKD) family of serine/threonine kinases belongs to the calcium/calmodulin-dependent protein kinase superfamily and comprises 3 isoforms, PRKD1, PRKD2, and PRKD3.22 Previous studies have shown that PRKD2 plays important oncogenic roles in the regulation of cancer progression, such as in gastrointestinal cancers,23 triple-negative breast cancer,24 glioma and so on.25 However, its functions in chemoresistance of AML have not been well studied.
To demonstrate the functions of PRKD2 in AML and its effects on Notch1 pathway, knockdown and overexpression assays were conducted. As shown above, we unveiled that PRKD2-enhanced proliferation and ADM-resistance in AML cells, which was the consequence of decreased apoptosis rate. Moreover, we demonstrated that PRKD2 expression was positively correlated with the activity of Notch1 pathway. Notch1, NICD and HES1 expression were remarkably changed after PRKD2 knockdown or overexpression. Then, we further explored the clinical significance of high PRKD2 expression in AML patients by analyzing data from TCGA. Apart from characteristics such as older age and higher risk group, high PRKD2 expression also indicated the poorer prognosis of AML patients with specific genomic mutations. Based on previous work, FLT3,26 IDH1,27 NPM128 and RAS29 mutations were important prognostic markers for AML patients, and were all regarded as unfavorable mutations. Our results showed PRKD2 could also act as an independent prognostic marker, because its high expression indicated poorer prognosis in AML patients. Moreover, our results indicated that low PRKD2 expression could alleviate the poor prognostic effects of FLT3 mutation in AML patients.
In conclusion, our results showed that PRKD2 could promote the proliferation and chemoresistance of AML cells by regulating the Notch1 signaling pathway. The study broadened our insights into the underlying mechanisms in chemoresistance and proliferation of AML, and provided a new prognostic marker and treatment target for AML.
Ethics Statement
Our study used public data and cell lines and no human or animal subjects were involved. There were no ethical concerns.
Acknowledgments
This work was supported by grants from Taishan Scholars Program, National Natural Science Foundation of China (81770159), Natural Science Foundation of Shandong Province (ZR2017BH088, 2018CXGC1215, ZR2018PH012), and the Fundamental Research Funds of Shandong University (2018JC003).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Zheng Z, Zheng X, Zhu Y, et al. miR-183-5p inhibits occurrence and progression of acute myeloid leukemia via targeting erbin. Mol Ther. 2019;27(3):542–558. doi:10.1016/j.ymthe.2019.01.016
2. Khasawneh MK, Abdel-Wahab O. Recent discoveries in molecular characterization of acute myeloid leukemia. Curr Hematol Malig Rep. 2014;9(2):93–99. doi:10.1007/s11899-014-0200-y
3. Medinger M, Lengerke C, Passweg J. Novel prognostic and therapeutic mutations in acute myeloid leukemia. Cancer Genomics Proteomics. 2016;13(5):317–329.
4. Lan QY, Zhi YL, Heng H, et al. Chemical space of FLT3 inhibitors as potential anti-AML drugs. Recent Pat Anticancer Drug Discov. 2017;12(4):296–322. doi:10.2174/1574892812666170727154643
5. Molenaar RJ, Radivoyevitch T, Nagata Y, et al. IDH1/2 mutations sensitize acute myeloid leukemia to PARP inhibition and this is reversed by IDH1/2-mutant inhibitors. Clin Cancer Res. 2018;24(7):1705–1715. doi:10.1158/1078-0432.CCR-17-2796
6. de Rooij JD, Zwaan CM, van den Heuvel-eibrink M, Pediatric AML. From biology to clinical management. J Clin Med. 2015;4(1):127–149. doi:10.3390/jcm4010127
7. Feldman EJ. Novel therapeutics for therapy-related acute myeloid leukemia: 2014. Clin Lymphoma Myeloma Leuk. 2015;15 Suppl:S91–S93. doi:10.1016/j.clml.2015.02.031
8. Aster JC, Pear WS, Blacklow SC. The varied roles of notch in cancer. Annu Rev Pathol. 2017;12:245–275. doi:10.1146/annurev-pathol-052016-100127
9. Previs RA, Coleman RL, Harris AL, Sood AK. Molecular pathways: translational and therapeutic implications of the Notch signaling pathway in cancer. Clin Cancer Res. 2015;21(5):955–961. doi:10.1158/1078-0432.CCR-14-0809
10. Yen WC, Fischer MM, Axelrod F, et al. Targeting Notch signaling with a Notch2/Notch3 antagonist (tarextumab) inhibits tumor growth and decreases tumor-initiating cell frequency. Clin Cancer Res. 2015;21(9):2084–2095. doi:10.1158/1078-0432.CCR-14-2808
11. Baker A, Wyatt D, Bocchetta M, et al. Notch-1-PTEN-ERK1/2 signaling axis promotes HER2+ breast cancer cell proliferation and stem cell survival. Oncogene. 2018;37(33):4489–4504. doi:10.1038/s41388-018-0251-y
12. Chen CY, Chen YY, Hsieh MS, et al. Expression of Notch gene and its impact on survival of patients with resectable non-small cell lung cancer. J Cancer. 2017;8(7):1292–1300. doi:10.7150/jca.17741
13. Zhang J, Ye J, Ma D, et al. Cross-talk between leukemic and endothelial cells promotes angiogenesis by VEGF activation of the Notch/Dll4 pathway. Carcinogenesis. 2013;34(3):667–677. doi:10.1093/carcin/bgs386
14. Zhang J, Ma D, Ye J, et al. Prognostic impact of delta-like ligand 4 and Notch1 in acute myeloid leukemia. Oncol Rep. 2012;28(4):1503–1511. doi:10.3892/or.2012.1943
15. Tomczak K, Czerwinska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol. 2015;19(1A):A68–A77. doi:10.5114/wo.2014.47136
16. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25(4):402–408. doi:10.1006/meth.2001.1262
17. Dohner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373(12):1136–1152. doi:10.1056/NEJMra1406184
18. Walter MJ, Payton JE, Ries RE, et al. Acquired copy number alterations in adult acute myeloid leukemia genomes. Proc Natl Acad Sci U S A. 2009;106(31):12950–12955. doi:10.1073/pnas.0903091106
19. Hassan WA, Yoshida R, Kudoh S, et al. Notch1 controls cell chemoresistance in small cell lung carcinoma cells. Thorac Cancer. 2016;7(1):123–128. doi:10.1111/1759-7714.12297
20. Xiao YS, Zeng LYK, Wu Y, et al. Major vault protein is a direct target of Notch1 signaling and contributes to chemoresistance in triple-negative breast cancer cells. Cancer Lett. 2019;440–441:156–167. doi:10.1016/j.canlet.2018.09.031
21. Li LQ, Pan D, Zhang SW, Xie D, XL Z, Chen H. Autophagy regulates chemoresistance of gastric cancer stem cells via the Notch signaling pathway. Eur Rev Med Pharmacol Sci. 2018;22(11):3402–3407. doi:10.26355/eurrev_201806_15162
22. Azoitei N, Frohling S, Scholl C, Seufferlein T. PRKD2: a two-pronged kinase crucial for the tumor-supporting activity of HSP90. Mol Cell Oncol. 2015;2(2):e981444. doi:10.4161/23723556.2014.981444
23. Azoitei N, Pusapati GV, Kleger A, et al. Protein kinase D2 is a crucial regulator of tumour cell-endothelial cell communication in gastrointestinal tumours. Gut. 2010;59(10):1316–1330. doi:10.1136/gut.2009.206813
24. Liu Y, Wang Y, Yu S, et al. The role and mechanism of CRT0066101 as an effective drug for treatment of triple-negative breast cancer. Cell Physiol Biochem. 2019;52(3):382–396.
25. Bernhart E, Damm S, Heffeter P, et al. Silencing of protein kinase D2 induces glioma cell senescence via p53-dependent and -independent pathways. Neuro-Oncology. 2014;16(7):933–945. doi:10.1093/neuonc/not303
26. Schlenk RF, Dohner K, Krauter J, et al. Mutations and treatment outcome in cytogenetically normal acute myeloid leukemia. N Engl J Med. 2008;358(18):1909–1918. doi:10.1056/NEJMoa074306
27. Salem D, El-Aziz SA, El-Menshawy N, Abouzeid T, Ebrahim M. Prevalence and prognostic value of IDH1 R132 mutation in newly diagnosed AML Egyptian patients with normal karyotype. Indian J Hematol Blood Transfusion. 2017;33(1):49–55. doi:10.1007/s12288-016-0649-z
28. Heath EM, Chan SM, Minden MD, Murphy T, Shlush LI, Schimmer AD. Biological and clinical consequences of NPM1 mutations in AML. Leukemia. 2017;31(4):798–807. doi:10.1038/leu.2017.30
29. Liu X, Ye Q, Zhao XP, et al. RAS mutations in acute myeloid leukaemia patients: a review and meta-analysis. Clin Chim Acta. 2019;489:254–260. doi:10.1016/j.cca.2018.08.040
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.