Back to Journals » Infection and Drug Resistance » Volume 11
Prevalence of nontuberculous mycobacteria and high efficacy of D-cycloserine and its synergistic effect with clarithromycin against Mycobacterium fortuitum and Mycobacterium abscessus
Authors Khosravi AD , Mirsaeidi M, Farahani A , Tabandeh MR, Mohajeri P, Shoja S, Hoseini Lar KhosroShahi SR
Received 14 September 2018
Accepted for publication 30 October 2018
Published 7 December 2018 Volume 2018:11 Pages 2521—2532
DOI https://doi.org/10.2147/IDR.S187554
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Azar Dokht Khosravi,1,2 Mehdi Mirsaeidi,3 Abbas Farahani,1,2 Mohammad Reza Tabandeh,4 Parviz Mohajeri,5,6 Saeed Shoja,7 Seyedeh Roghayeh Hoseini Lar KhosroShahi8
1Infectious and Tropical Diseases Research Center, Health Research Institute, Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran; 2Department of Microbiology, School of Medicine, Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran; 3Division of Pulmonary, Critical Care, Sleep and Allergy, Department of Medicine, Miller School of Medicine, University of Miami, Miami, FL, USA; 4Department of Biochemistry and Molecular Biology, Faculty of Veterinary Medicine, Shahid Chamran University of Ahvaz, Ahvaz, Iran; 5Department of Microbiology, School of Medicine, Kermanshah University of Medical Sciences, Kermanshah, Iran; 6Nosocomial Infection Research Center, Kermanshah University of Medical Sciences, Kermanshah, Iran; 7Infectious and Tropical Diseases Research Center, Hormozgan Health Institute, Hormozgan University of Medical Sciences, Bandar Abbas, Iran; 8Tuberculosis Research Center, Hormozgan University of Medical Sciences, Bandar Abbas, Iran
Background: The prevalence of pulmonary disease caused by nontuberculous mycobacteria (NTM) is reportedly on the rise in the world. Some of the species are resistant to various antibiotics; hence, limited treatment options are available. The aims of this study were to investigate the prevalence of NTM and to determine the effect of D-cycloserine against Mycobacterium fortuitum and Mycobacterium abscessus isolated from clinical specimens to find out the synergistic effect of D-cycloserine and clarithromycin.
Methods: A total of 95 nonduplicate pulmonary isolates of NTM were collected from three major Regional Tuberculosis (TB) Centers. NTM isolates were identified by conventional tests and PCR sequence analysis of the rpoB gene. PCR sequencing of erm-41 was performed for detecting the inducible resistance to macrolides. In vitro susceptibilities and activities of D-cycloserine-clarithromycin combinations were accessed using the broth microdilution method.
Results: Among 714-positive acid-fast bacilli from TB-suspected cases, 95 isolates were identified as NTM (13.3%). The prevalence of identified isolates was as follows: M. fortuitum 46 (48.4%), Mycobacterium simiae 16 (16.8%), Mycobacterium kansasii 15 (15.7%), M. abscessus 7 (7.3%), Mycobacterium thermoresistibile 4 (4.2%), Mycobacterium elephantis 3 (3.2%), Mycobacterium porcinum 2 (2.1%), and Mycobacterium chimaera 2 (2.1%). In addition, rpoB sequence analysis could identify all NTM isolates. The effect of D-cycloserine was better than that of clarithromycin. The synergistic effect of D-cycloserine with clarithromycin was observed for six (100%) and five (71.5%) strains of M. fortuitum and M. abscessus, respectively.
Conclusion: In the present study, we demonstrated a wide range of NTM in processed samples from different provinces of Iran. Our observations indicated that D-cycloserine was very active against M. abscessus and M. fortuitum; hence, D-cycloserine, either alone or in combination with clarithromycin, may be promising for the treatment of M. abscessus- and M. fortuitum-associated diseases.
Keywords: nontuberculous mycobacteria, in vitro activity, D-cycloserine, Mycobacterium fortuitum, Mycobacterium abscessus
Introduction
Nontuberculous mycobacteria (NTM) are generally found in the environmental sources including tap water, soil, and dust but in certain circumstances are able to cause disease in humans especially in immunosuppressed conditions.1 Many opportunistic NTM pathogens are associated with a wide spectrum of localized and systemic disorders such as skin and soft tissue infections2 and pulmonary and extrapulmonary infections. The prevalence of pulmonary disease caused by NTM is reportedly on the rise in the world.3
Based on the growth rate, NTM species are categorized into slowly growing mycobacteria (SGM) and rapidly growing mycobacteria (RGM).4 Mycobacterium chelonae, Mycobacterium peregrinum, Mycobacterium abscessus, Mycobacterium fortuitum, and Mycobacterium smegmatis group are among RGM that act as opportunistic pathogens of humans and produce various diseases.5,6 M. fortuitum is most often associated with skin and soft tissue infections and hospital-acquired postoperative infections.7 M. abscessus has emerged as an important human pathogen and is responsible for a wide variety of soft tissue and disseminated infections in immunocompromised patients and can complicate lung transplantation.3 NTM infections, especially M. abscessus infection, is particularly problematic, and often, its treatment is costly and lengthy and comprises side effects.8,9
The description of the mechanisms of resistance does not limit the therapeutic options.10 These organisms have not become resistant, and they are intrinsically resistant to most of the antibiotics that are currently available.11 Also, infections caused by M. fortuitum isolates need long-term antibiotic therapy, because of their high resistance to chemotherapeutic agents and disinfectants.12 The antibiotic of choice for infections caused by M. fortuitum is amikacin, but resistance to aminoglycosides and macrolides has also been reported.12,13 Therefore, there is a great need to develop new treatment regimens for RGM infections, especially infections due to M. fortuitum and M. abscessus.14 Combination therapy of oral macrolides and amikacin for 2–4 months has been recommended by other researchers and also by the American Thoracic Society (ATS).15 After initial therapy, macrolide administration with at least one other antimicrobial agent to which the organism is susceptible should be used for the treatment. In vitro activity of aminoglycosides, such as clofazimine, D-cycloserine, and dapsone, has also been studied for certain SGM species16; however, the combined activities of a macrolide with D-cycloserine against M. abscessus and M. fortuitum have not been evaluated so far. As M. abscessus is the most common cause of NTM infections after M. fortuitum and Mycobacterium simiae in Iran,17 the aims of this study were to investigate the prevalence of NTM isolated from clinical samples in Iran and to determine the effect of D-cycloserine against M. abscessus and M. fortuitum to find out the synergistic effect of D-cycloserine and clarithromycin on these organisms.
Methods
Sample collection
In this cross-sectional study, a total of 2,414 pulmonary samples were collected from three major Regional Tuberculosis (TB) Centers in Iran, including Khuzestan (south west), Kermanshah (west), and Hormozgan (south), from February 2016 to January 2018. Thirteen provinces of Iran are covered by these centers. As a part of the centers’ policy, referred patients were requested to sign the informed consent in case their samples are used for research purposes apart from routine clinical investigation.
The preliminary proposal of the work was approved in joined Institutional Review Board (IRB) and Ethics Committee (Code: IR.AJUMS.REC.1396.103) of the Ahvaz Jundishapur University of Medical Sciences, Iran, and the necessary permission was granted for sample collection.
Phenotypic identification
The isolates were inoculated into Lowenstein–Jensen (LJ) medium and incubated at 37°C. All LJ tubes were examined daily for 30 days and twice weekly for 4 weeks thereafter. The mycobacterial isolates were subjected to phenotypic identification tests including growth characteristics, colony morphology, pigment production, niacin production, Tween 80 hydrolysis, semiquantitative catalase test, salt tolerance, arylsulfatase test, growth on MacConkey agar, heat-stable catalase (68°C), iron uptake, urease production, and tellurite and nitrate reduction.18
Molecular identification
DNA extraction and PCR amplification
Chromosomal DNA was extracted from mycobacterial colonies grown on LJ medium using an extraction and purification QIAamp DNA Mini Kit (Qiagen NV, Venlo, the Netherlands) according to the manufacturer’s protocol. DNA concentrations and purity were determined using the NanoDrop One instrument (Thermo Fisher Scientific, Waltham, MA, USA) at 260 nm and used as template in PCR amplification.
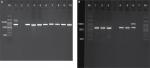
A 751 bp fragment of the rpoB gene was amplified by using two specific primers of MycoF (5′-GCAAGGTCACCCCGAAGGG-3′) and MycoR (5′-AGCGGCTGCTGGGTGATCATC-3′) as previously described.19 PCR amplification was performed in a final volume of 20 µL comprising 10 µL of Taq DNA Polymerase Master Mix RED (Ampliqon, Copenhagen, Denmark), 0.5 µL of each primer at 10 µM, 3 µL of extracted DNA (50 ng), and 6 µL of ddH2O. The amplification was performed in a thermocycler (C1000 Touch; Bio-Rad Laboratories Inc., Hercules, CA, USA), with the following program: initial denaturation at 95°C for 1 minute, followed by 35 cycles of denaturation at 94°C for 30 seconds, annealing 64°C for 30 seconds, and extension at 72°C for 90 seconds with a final extension at 72°C for 5 minutes. The PCR products were separated by electrophoresis on a 1.2% agarose gel (EMD Millipore, Billerica, MA, USA), stained with the SYBR Safe DNA Gel Stain (Thermo Fisher Scientific), and the DNA bands were visualized by a UV transilluminator (Uvidoc, gel documentation system; Jencons Scientific Inc., Cambridge, UK). The PCR Clean-Up Kit (Vivantis Technologies Sdn. Bhd., Subang Jaya, Malaysia) was used to purify PCR amplicons according to the manufacturer’s instructions.
Analysis of sequence data
The sequence analyses based on the rpoB gene were performed in both directions (Bioneer Corporation, Daejeon, South Korea). The sequences were trimmed at both the 5′ and 3′ ends to include the most corresponding gene fragment sequences of mycobacteria collected in NCBI Nucleotide database. All the obtained sequences were analyzed by the Chromas software ver 2.6.
Phylogenetic analysis
The sequences were aligned using the ClustalW format in the MEGA 6.0 program Mega software (version 5.2).20 The methodology for phylogenetic trees was constructed using the neighbor-joining method21 and verified by the maximum likelihood method with 1,000 bootstrap replications. The phylogenetic tree was rooted with Mycobacterium tuberculosis and used Nocardia asteroids as an outgroup.
PCR assay for erm-41 gene
PCR amplification for the erm-41 gene was performed on extracted DNA by using a set of primers of ermF (5′-GAC CGG GGC CTT CTT CGT GAT-3′) and ermR1 (5′-GAC TTC CCCGCA CCG ATT CC-3′), described by Shallom et al,22 and the amplification protocol as described earlier for the rpoB gene. The target fragments were amplified using a thermal cycler (C1000 Touch), with following conditions: 95°C for 2 minutes, followed by 35 cycles of 95°C for 1 minute, 62°C for 1 minute, and 72°C for 90 seconds and a final extension at 72°C for 10 minutes. The electrophoresis of PCR products was performed on 1.5% agarose gel, and the gels were stained with the SYBR Safe DNA Gel Stain and were then visualized by the UV transilluminator. The PCR products were sent to Bioneer Co. for sequencing. The sequences obtained were analyzed using the BLAST algorithm at the NCBI website (http://blast.ncbi.nlm.nih.gov/).
Antimicrobial agents
D-cycloserine and clarithromycin were purchased from Sigma-Aldrich Co. (St Louis, MO, USA). For stock solution’s preparation, potency of each antibiotic was calculated according to the CLSI guideline.23 In brief, D-cycloserine and clarithromycin were dissolved in water and dimethyl sulfoxide (DMSO) at a concentration of 1,024 and 256 mg/L, respectively, to serve as stock solutions. Each stock solution was filtered using a sterilized 0.22 mm syringe filters (Sartorius, Melbourne, VIC, Australia). Stock solutions were stored at –80°C prior to each experiment. Staphylococcus aureus ATCC 29213 and M. abscessus ATCC 19977 were used as the reference strains for the determination of minimum inhibitory concentrations (MICs).
Drug susceptibility testing
Broth microdilution method was performed according to Clinical and Laboratory Standards Institute (CLSI) guidelines,23 using 96-well round bottom microtiter plates. Briefly, the colonies of M. fortuitum and M. abscessus clinical isolates were harvested from growth on Luria-Bertani (LB) agar and suspended in 2 mL of sterile saline in tubes containing glass beads and mixed by vortexing. The mycobacterial suspensions were adjusted to McFarland 0.5 and inoculated into the cation-adjusted Mueller-Hinton broth (CAMHB) medium (Sigma-Aldrich Co.), containing twofold dilutions of antibiotics. The final drug concentration was from 2 to 256 mg/L for each antibiotic. The prepared MIC plates were incubated aerobically at 30°C. MICs were measured at day 3 or 4 in comparison to control sample well without antibiotic showing sufficient bacterial growth. The MIC was defined as the lowest drug concentration at which bacterial growth was not visualized. To detect the inducible macrolide-resistant M. abscessus isolates, MIC measurements were carried out at days 3 and 14 for clarithromycin.
Since to date, there are no reference MIC data for D-cycloserine, M. fortuitum, and M. abscessus isolates to determine the MIC breakpoints (susceptible, intermediate, and resistant), we referred to the MIC breakpoints of D-cycloserine (≤16, 32, and ≥64 µg/mL) for Mycobacterium avium complex proposed by Huang et al.16 The MIC breakpoints (susceptible, intermediate, and resistant) of ≤2, 4, and ≥8 µg/mL, respectively, for clarithromycin were interpreted according to the CLSI guideline.23
The synergistic effects testing between clarithromycin and D-cycloserine
The synergistic effects of clarithromycin and D-cycloserine against six randomly selected M. fortuitum strains, and all of the seven M. abscessus strains were determined by the checkerboard approach using the broth microdilution method described elsewhere.24,25
Briefly, colonies were harvested from the LB agar and transferred to a CAMHB with glass beads for vigorous mixing on a vortex for 1 minute. After mixing, the mycobacterial suspensions were adjusted to the McFarland standard 5.0 (further diluted 1:200 in CAMHB). Clarithromycin and D-cycloserine were prepared as working solutions in 2.5 mL of CAMHB. For each isolate, 11 concentrations of clarithromycin (0.062–64 µg/mL) in the horizontal wells were tested for synergy against eight concentrations of D-cycloserine (0.25–32 µg/mL) in the vertical wells, using 96-well round bottom microtiter plates. After adding drug concentrations in wells, 100 µL of mycobacterial suspensions were added to each well of the microplate containing twofold diluted antimicrobials and, then, the plates were sealed by parafilm and incubated at 37°C for 3 days. Also, each drug was tested alone for two species within the range of concentrations from 0.062 to 64 µg/mL, at twofold serial dilutions. MIC measurements were carried out at day 3 of incubation.
The fractional inhibitory concentration index (FICI) method was calculated as previously described,26
FICI = FICA + FICB = (MICA combination/MIC alone) + (MICB combination/MIC alone)
where A represents D-cycloserine and B represents clarithromycin. An FICI of ≤0.5 was interpreted as synergism, an FICI of 0.5 < FICI ≤ 4 was interpreted as indifference, and an FICI of >4 was interpreted as antagonism between the two drugs.
Results
A total of 2,414 pulmonary samples were examined during the period of study. Of these, 714 samples were positive for acid-fast bacilli by the Ziehl–Neelsen (ZN) staining and culture method. Among 714 isolates, 619 (86.69%) isolates were identified as M. tuberculosis complex (MTC) based on biochemical and molecular tests and 95 isolates were identified as NTM (13.3%). The samples were obtained from TB-suspected patients who were referred to the three major Regional TB Centers covering 13 provinces of Iran.
Of the 95 NTM-positive isolates, 55 (57.9%) isolates were belonged to male patients and 40 (42.1%) isolates were from females, with the mean age of 47.4±19.9 years. Coughing, fever, phlegm, and weight loss were the most common signs and symptoms of NTM-positive patients (Table 1).
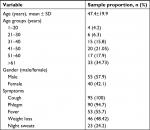  | Table 1 Demographic characteristics of nontuberculous mycobacteria-positive patients |
Phenotypic identification
According to the growth rate, the isolates were classified into two groups (RGM and SGM). On the basis of growth characteristics, morphological, and biochemical properties, which are presented in Table 2, M. fortuitum (38 isolates) was the most frequent isolate, followed by M. simiae (10 isolates), Mycobacterium kansasii like (nine isolates), and M. chelonae (nine isolates). Among the 95 NTM isolates studied, only 76 (80%) strains were detected by phenotypic tests and the remaining isolates were unidentifiable.
  | Table 2 Phenotypic and molecular characteristics of clinical isolates Abbreviations: Sc, scotochromogen; SQ, semiquantitative; Y, yellow. |
Molecular identification
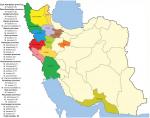
According to the sequencing of the rpoB gene, all 95 isolates belonged to eight different species and were clearly delineated (Figure 1A), including five RGM and three SGM. The most prevalent NTM species isolated from different provinces were M. fortuitum 46 isolates (48.4%), M. simiae 16 isolates (16.8%), and M. kansasii 15 isolates (15.7%). These were followed by M. abscessus seven isolates (7.3%), Mycobacterium thermoresistibile four isolates (4.2%), Mycobacterium elephantis three isolates (3.2%), Mycobacterium porcinum two isolates (2.1%), and Mycobacterium chimaera two isolates (2.1%) (Figures 2 and 3). As indicated in Table 3 and Figure 2, the majority of NTM were isolated from Kermanshah, Hormozgan and Khuzestan provinces. There was no significant correlation in the distribution of NTM in different regions.
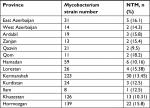  | Table 3 Distribution of clinical NTM strains throughout 13 Iranian provinces Abbreviation: NTM, nontuberculous mycobacteria. |
In addition, all the M. abscessus isolates confirmed as M. abscessus subsp. abscessus by the replication and sequencing of the erm-41 gene. Additionally, sequence analysis revealed that all M. abscessus strains showed mutation at position 28 in the erm-41 gene (Figure 1B). Sequencing of erm-41 and rpoB genes could identify all M. abscessus subsp. abscessus and showed valuable sensitivity and specificity, but erm-41 was limited to the identification of M. abscessus species. M. fortuitum and M. abscessus isolates were then selected for further characterization.
In vitro susceptibilities to D-cycloserine and clarithromycin
For M. abscessus and M. fortuitum, MICs were determined for two antibiotics and their susceptibility patterns are shown in Table 4. From total M. fortuitum strains, 10 (21.7%), 10 (21.7%), and 26 (56.5%) isolates were susceptible, intermediate, and resistant, respectively, to clarithromycin. All M. abscessus isolates in this study were resistant to clarithromycin.
For D-cycloserine, 41 (89.1) of the M. fortuitum isolates were susceptible to D-cycloserine and all M. abscessus isolates were susceptible to this antibiotic. Thus, D-cycloserine showed good activity against both M. fortuitum and M. abscessus, while clarithromycin showed lower activity, inhibiting only 21.7% (n=10) of the M. fortuitum isolates and none of M. abscessus isolates. Moreover, two (4.4%) of M. fortuitum strains were co-resistant to both antibiotics.
In vitro activities of D-cycloserine–clarithromycin combinations
The combination of clarithromycin + D-cycloserine showed good activity against M. fortuitum isolates. So, the well in which more than 90% of isolates were killed after colony-forming unit (CFU) enumeration, was determined as MIC value (Table 5). The synergistic effect of D-cycloserine with clarithromycin was observed for six (100%) M. fortuitum, and combination results showed at least a 2.4-fold decrease in the concentration of each drug compared to either drug alone. These results suggested that this novel combination is active against clarithromycin-resistant M. fortuitum with FICI values ranging from 0.375 to 0.5.
Among inducible macrolide-resistant M. abscessus isolates, the synergistic effect of D-cycloserine with clarithromycin was observed for five (71.5%) isolates, by FICI values ranging from 0.375 to 0.5. The other two (28.5%) M. abscessus strains showed indifferent interaction with the drugs’ combination by FICI value of 0.75 (Table 5). Nevertheless, antagonism was not observed for both species. Our combination results showed at least a 2.6-fold decrease in the concentration of each drug when needed to achieve >90% inhibition as compared to the concentration needed to achieve the same level of inhibition when used alone.
Discussion
Infections due to NTM are a serious problem for immunocompromised patients, and its prevalence among clinical isolates is greatly on the increase in recent years worldwide.27 The antibiotic choices for the treatment of NTM infections are different from those for TB, as NTM is naturally resistant to the classical antituberculous drugs. Additionally, among NTM, RGM shows resistance to most of the currently available antibiotics in different geographical regions of the world. By the improvement in microbiological and laboratory techniques, more RGM has been identified28; therefore, identification of NTM species and determination of drug susceptibility pattern are crucial.29–31 In the current study, only 76 (80%) strains were assigned to a species or complex by phenotypic tests. By the rpoB sequence analysis, all NTM isolates (100%) were identified and clearly delineated (Figure 3). According to the sequence analysis, the rpoB gene showed high discriminatory power for species’ identification and all isolates were clustered with their reference strains. In concordance with our study, other previous reports demonstrated the efficacy of molecular tests such as rpoB sequencing. They introduced the technique as one of the most common techniques currently used for NTM identification, which is significantly more accurate than the phenotypic tests.32,33
Our findings indicated that the phenotypic tests alone cannot be applied as an efficient method for the identification of NTM species. The phenotypic method is time consuming and costly, and its results are very variable compared to those of molecular tests.33
Based on the sequences’ analyses of erm-41 and rpoB genes, all M. abscessus isolates were identified more correctly and this indicated that the erm-41 and rpoB genes had valuable sensitivity and specificity for M. abscessus group identification. Further studies with more diverse selection of Mycobacterium abscessus group should be conducted to verify the discriminatory power of these genes for identification.34
In the present study, we demonstrated that M. fortuitum, M. abscessus, M. simiae, and M. kansasii are the most common RGM and SGM species in samples of patients from different provinces investigated in this study; this finding is in line with other reports from Iran.17,35,36 Moreover, M. fortuitum was reported as the most common NTM, isolated from environmental and clinical sources in the distinctive part of Iran.37,38
Several studies indicate a high incidence of M. simiae in Iran,39,40 while according to our results, M. fortuitum (48.4%) was the most prevalent NTM in pulmonary samples. We indicated that, in 12 provinces of Iran, some patients with TB misdiagnosis and relevant treatment were actually infected by M. fortuitum. In contrast, M. simiae isolates (16.8%) were the second dominant NTM species. These differences may be the result of different geographical distributions of NTM throughout Iran. Umrao et al41 reported that the predominant species were M. abscessus and M. fortuitum in Northern Indian population.
Infection with NTM is often misdiagnosed for TB and may be treated as drug-resistant TB. NTM species are resistant to the first and some second lines, resulting in treatment failure that may lead to an increase in mortality and morbidity; therefore, accurate and timely diagnosis is essential. Our study showed that most of the isolates of NTM were obtained from pulmonary samples and thus, there is a need to develop prevention strategies against them.
This study evaluated the activity of D-cycloserine and clarithromycin and the combinations of both antibiotics against M. abscessus and M. fortuitum clinical isolates. The findings demonstrated that the effect of D-cycloserine alone against M. abscessus and M. fortuitum was better than that of clarithromycin. Furthermore, we observed the synergistic effect of D-cycloserine–clarithromycin combination against M. abscessus and M. fortuitum. These results indicate that both D-cycloserine and clarithromycin may serve as an alternative treatment for M. abscessus- and M. fortuitum-associated infections. However, the D-cycloserine–clarithromycin combination was not effective against the minority of tested M. abscessus and 28.5% of the isolates showed an indifferent interaction.
Cycloserine acts by inhibiting cell wall biosynthesis in bacteria. In fact, cycloserine acts against two crucial enzymes important in the peptidoglycan synthesis, including d-alanine:d-alanine ligase (Ddl) and alanine racemase (Alr).42 Cycloserine also has good capability of penetrating into liver and tissues. Cycloserine was found to have low resistance rate in MTC and also high susceptibility rates against M. intracellulare and M. avium.16
In combination, cycloserine may interrupt cell wall synthesis, thus increasing the penetration of clarithromycin into the bacteria.
Previous studies have demonstrated an in vitro synergistic effect of combination of clofazimine with amikacin or clarithromycin against M. abscessus.43 The only oral drugs used in the treatment of M. abscessus are the macrolides and clofazimine,44 but there is less supportive clinical evidence of its efficacy.45 We found that two isolates of M. fortuitum were resistant to D-cycloserine, while all M. abscessus strains were susceptible. The mechanism of resistance to D-cycloserine is unknown in M. fortuitum, but overexpression of ddlA and alr genes has been reported in D-cycloserine-resistant Mycobacterium smegmatis.46 Another report demonstrated that those mutations in alr and ald genes and the loss of function in the ald gene conferred resistance to D-cycloserine in M. tuberculosis.47 In concordance with our findings, Huang et al recently reported the suitability of D-cycloserine for the treatment of both M. avium- and M. intracellulare-associated diseases,16 in addition, Cowman et al reported that cycloserine, tigecycline, and clofazimine might be useful in the treatment of the most resistant SGM.48 These reports emphasize on the D-cycloserine efficacy against other NTM and M. tuberculosis clinical isolates as well. A limitation of two drugs’ combination, which was studied for synergy in this work, was small sample size; hence, we suggest to use the method with a larger sample size in future studies to achieve a better understanding of the synergistic effect of present antibiotics.
Conclusion
The present study showed that the pulmonary samples comprise a wide range of NTM in different provinces of Iran. Based on our observations, we came to the conclusion that D-cycloserine is very active against both M. abscessus and M. fortuitum. D-cycloserine, either alone or in combination with clarithromycin, may be promising for the treatment of M. abscessus- and M. fortuitum-associated diseases. Future clinical trials are needed to confirm our findings.
Acknowledgments
This work is part of PhD thesis of Abbas Farahani, which was approved in Infectious and Tropical Diseases Research Center, and was financially supported by a grant (OG-96104) from Research Affairs, Ahvaz Jundishapur University of Medical Sciences, Ahvaz, Iran. We are greatly thankful to the staff of Regional TB Reference Laboratories for their collaboration in sample collection.
Disclosure
The authors report no conflicts of interest in this work.
References
Hruska K, Kaevska M. Mycobacteria in water, soil, plants and air: a review. Vet Med. 2013;57(12):623–679. | ||
Gonzalez-Santiago TM, Drage LA. Nontuberculous mycobacteria: skin and soft tissue infections. Dermatol Clin. 2015;33(3):563–577. | ||
Henkle E, Winthrop KL. Nontuberculous mycobacteria infections in immunosuppressed hosts. Clin Chest Med. 2015;36(1):91–99. | ||
Kim CJ, Kim NH, Song KH, et al. Differentiating rapid- and slow-growing mycobacteria by difference in time to growth detection in liquid media. Diagn Microbiol Infect Dis. 2013;75(1):73–76. | ||
Brown-Elliott BA, Wallace RJ. Clinical and taxonomic status of pathogenic nonpigmented or late-pigmenting rapidly growing mycobacteria. Clin Microbiol Rev. 2002;15(4):716–746. | ||
Lavollay M, Dubée V, Heym B, et al. In vitro activity of cefoxitin and imipenem against Mycobacterium abscessus complex. Clin Microbiol Infect. 2014;20(5):O297–O300. | ||
Yu JR, Heo ST, Lee KH, et al. Skin and soft tissue infection due to rapidly growing mycobacteria: Case series and literature review. Infect Chemother. 2013;45(1):85–93. | ||
Ferro BE, van Ingen J, Wattenberg M, van Soolingen D, Mouton JW. Time-kill kinetics of antibiotics active against rapidly growing mycobacteria. J Antimicrob Chemother. 2015;70(3):811–817. | ||
Luthra S, Rominski A, Sander P. The role of antibiotic-target-modifying and antibiotic-modifying enzymes in Mycobacterium abscessus drug resistance. Front Microbiol. 2018;9:2179. | ||
Nessar R, Cambau E, Reyrat JM, Murray A, Gicquel B. Mycobacterium abscessus: a new antibiotic nightmare. J Antimicrob Chemother. 2012;67(4):810–818. | ||
Chopra S, Matsuyama K, Hutson C, Madrid P. Identification of antimicrobial activity among FDA-approved drugs for combating Mycobacterium abscessus and Mycobacterium chelonae. J Antimicrob Chemother. 2011;66(7):1533–1536. | ||
Santos DR, Lourenço MC, Coelho FS, Mello FC, Duarte RS. Resistance profile of strains of Mycobacterium fortuitum isolated from clinical specimens. J Bras Pneumol. 2016;42(4):299–301. | ||
Jayasingam SD, Zin T, Ngeow YF. Antibiotic resistance in Mycobacterium abscessus and Mycobacterium fortuitum isolates from Malaysian patients. Int J Mycobacteriol. 2017;6(4):387–390. | ||
Huang CW, Chen JH, Hu ST, et al. Synergistic activities of tigecycline with clarithromycin or amikacin against rapidly growing mycobacteria in Taiwan. Int J Antimicrob Agents. 2013;41(3):218–223. | ||
Griffith DE, Aksamit T, Brown-Elliott BA, et al. An official ATS/IDSA statement: diagnosis, treatment, and prevention of nontuberculous mycobacterial diseases. Am J Respir Crit Care Med. 2007;175(4):367–416. | ||
Huang CC, Wu MF, Chen HC, Huang WC. In vitro activity of aminoglycosides, clofazimine, D-cycloserine and dapsone against 83 Mycobacterium avium complex clinical isolates. J Microbiol Immunol Infect. 2018;51(5):636–643. | ||
Velayati AA, Farnia P, Mozafari M, Mirsaeidi M. Nontuberculous mycobacteria isolation from clinical and environmental samples in Iran: Twenty years of surveillance. Biomed Res Int. 2015;2015:254285. | ||
Procop G, Church D, Hall G, Janda W, Koneman E, Schrekenberger P. Koneman’s Color Atlas and Textbook of Diagnostic Microbiology. Philadelphia: Wolters Kluwer Health; 2017. | ||
Adékambi T, Colson P, Drancourt M. rpoB-based identification of nonpigmented and late-pigmenting rapidly growing mycobacteria. J Clin Microbiol. 2003;41(12):5699–5708. | ||
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28(10):2731–2739. | ||
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406–425. | ||
Shallom SJ, Moura NS, Olivier KN, Sampaio EP, Holland SM, Zelazny AM. New real-time PCR assays for detection of inducible and acquired clarithromycin resistance in the Mycobacterium abscessus Group. J Clin Microbiol. 2015;53(11):3430–3437. | ||
Woods GL. Susceptibility testing of mycobacteria, nocardiae, and other aerobic actinomycetes. Approved Standard M24-A2. 2011;31(5). | ||
Kohno Y, Ohno H, Miyazaki Y, et al. In vitro and in vivo activities of novel fluoroquinolones alone and in combination with clarithromycin against clinically isolated Mycobacterium avium complex strains in Japan. Antimicrob Agents Chemother. 2007;51(11):4071–4076. | ||
Kaushik A, Makkar N, Pandey P, Parrish N, Singh U, Lamichhane G. Carbapenems and Rifampin Exhibit Synergy against Mycobacterium tuberculosis and Mycobacterium abscessus. Antimicrob Agents Chemother. 2015;59(10):6561–6567. | ||
De Logu A, Onnis V, Saddi B, Congiu C, Schivo ML, Cocco MT. Activity of a new class of isonicotinoylhydrazones used alone and in combination with isoniazid, rifampicin, ethambutol, para-aminosalicylic acid and clofazimine against Mycobacterium tuberculosis. J Antimicrob Chemother. 2002;49(2):275–282. | ||
Moore JE, Kruijshaar ME, Ormerod LP, Drobniewski F, Abubakar I. Increasing reports of non-tuberculous mycobacteria in England, Wales and Northern Ireland, 1995–2006. BMC Public Health. 2010;10:612. | ||
Ballarino GJ, Olivier KN, Claypool RJ, Holland SM, Prevots DR. Pulmonary nontuberculous mycobacterial infections: antibiotic treatment and associated costs. Respir Med. 2009;103(10):1448–1455. | ||
Kang YA, Koh WJ. Antibiotic treatment for nontuberculous mycobacterial lung disease. Expert Rev Respir Med. 2016;10(5):557–568. | ||
Mcgrath EE, Anderson PB. The therapeutic approach to non-tuberculous mycobacterial infection of the lung. Pulm Pharmacol Ther. 2010;23(5):389–396. | ||
Kothavade RJ, Dhurat RS, Mishra SN, Kothavade UR. Clinical and laboratory aspects of the diagnosis and management of cutaneous and subcutaneous infections caused by rapidly growing mycobacteria. Eur J Clin Microbiol Infect Dis. 2013;32(2):161–188. | ||
de Zwaan R, van Ingen J, van Soolingen D. Utility of rpoB gene sequencing for identification of nontuberculous mycobacteria in the Netherlands. J Clin Microbiol. 2014;52(7):2544–2551. | ||
Tortoli E. Phylogeny of the genus Mycobacterium: many doubts, few certainties. Infect Genet Evol. 2012;12(4):827–831. | ||
Yoshida S, Tsuyuguchi K, Suzuki K, et al. Rapid identification of strains belonging to the Mycobacterium abscessus group through erm(41) gene pyrosequencing. Diagn Microbiol Infect Dis. 2014;79(3):331–336. | ||
Nasiri MJ, Dabiri H, Fooladi AAI, Amini S, Hamzehloo G, Feizabadi MM. High rates of nontuberculous mycobacteria isolation from patients with presumptive tuberculosis in Iran. New Microbes New Infect. 2018;21:12–17. | ||
Dastranj M, Farahani A, Hashemi Shahraki A, Atashi S, Mohajeri P. Molecular identification and distribution of non-tuberculous mycobacteria isolated from clinical specimens by PCR-sequencing method in West of Iran. Clin Respir J. 2018;12(3):996–1002. | ||
Nasr Esfahani B, Zarkesh Esfahani FS, Bahador N, et al. Analysis of DNA gyrA gene mutation in clinical and environmental ciprofloxacin-resistant isolates of non-tuberculous mycobacteria using molecular methods. Jundishapur J Microbiol. 2016;9(3):e30018. | ||
Velayati AA, Farnia P, Mozafari M, et al. Molecular epidemiology of nontuberculous mycobacteria isolates from clinical and environmental sources of a metropolitan city. PLoS One. 2014;9(12):e114428. | ||
Hashemi-Shahraki A, Darban-Sarokhalil D, Heidarieh P, et al. Mycobacterium simiae: a possible emerging pathogen in Iran. Jpn J Infect Dis. 2013;66(6):475–479. | ||
Baghaei P, Tabarsi P, Farnia P, et al. Pulmonary disease caused by Mycobacterium simiae in Iran’s national referral center for tuberculosis. J Infect Dev Ctries. 2012;6(1):23–28. | ||
Umrao J, Singh D, Zia A, et al. Prevalence and species spectrum of both pulmonary and extrapulmonary nontuberculous mycobacteria isolates at a tertiary care center. Int J Mycobacteriol. 2016;5(3):288–293. | ||
Prosser GA, de Carvalho LP. Kinetic mechanism and inhibition of Mycobacterium tuberculosis D-alanine:D-alanine ligase by the antibiotic D-cycloserine. FEBS J. 2013;280(4):1150–1166. | ||
Ferro BE, Meletiadis J, Wattenberg M, et al. Clofazimine prevents the regrowth of Mycobacterium abscessus and Mycobacterium avium type strains exposed to amikacin and clarithromycin. Antimicrob Agents Chemother. 2016;60(2):1097–1105. | ||
Philley JV, Degroote MA, Honda JR, et al. Treatment of non-tuberculous mycobacterial lung disease. Curr Treat Options Infect Dis. 2016;8(4):275–296. | ||
Jarand J, Levin A, Zhang L, Huitt G, Mitchell JD, Daley CL. Clinical and microbiologic outcomes in patients receiving treatment for Mycobacterium abscessus pulmonary disease. Clin Infect Dis. 2011;52(5):565–571. | ||
Cáceres NE, Harris NB, Wellehan JF, Feng Z, Kapur V, Barletta RG. Overexpression of the D-alanine racemase gene confers resistance to D-cycloserine in Mycobacterium smegmatis. J Bacteriol. 1997;179(16):5046–5055. | ||
Desjardins CA, Cohen KA, Munsamy V, et al. Genomic and functional analyses of Mycobacterium tuberculosis strains implicate ald in D-cycloserine resistance. Nat Genet. 2016;48(5):544–551. | ||
Cowman S, Burns K, Benson S, Wilson R, Loebinger MR. The antimicrobial susceptibility of non-tuberculous mycobacteria. Journal of Infection. 2016;72(3):324–31. |
 © 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2018 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.