Back to Journals » Infection and Drug Resistance » Volume 13
Predictors of Severity and Co-Infection Resistance Profile in COVID-19 Patients: First Report from Upper Egypt
Authors Ramadan HKA, Mahmoud MA, Aburahma MZ , Elkhawaga AA , El-Mokhtar MA , Sayed IM , Hosni A , Hassany SM , Medhat MA
Received 18 July 2020
Accepted for publication 11 September 2020
Published 5 October 2020 Volume 2020:13 Pages 3409—3422
DOI https://doi.org/10.2147/IDR.S272605
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Haidi Karam-Allah Ramadan,1 Manal A Mahmoud,2 Mohamed Zakaria Aburahma,1 Amal A Elkhawaga,3 Mohamed A El-Mokhtar,3 Ibrahim M Sayed,3,4 Amal Hosni,5 Sahar M Hassany,1 Mohammed A Medhat1
1Department of Tropical Medicine and Gastroenterology, Faculty of Medicine, Assiut University, Assiut, Egypt; 2Department of Chest Diseases and Tuberculosis, Faculty of Medicine, Assiut University, Assiut, Egypt; 3Department of Medical Microbiology and Immunology, Faculty of Medicine, Assiut University, Assiut, Egypt; 4Department of Pathology, School of Medicine, University of California, San Diego, CA, USA; 5Department of Clinical Pathology, Faculty of Medicine, Assiut University, Assiut, Egypt
Correspondence: Haidi Karam-Allah Ramadan
Department of Tropical Medicine and Gastroenterology, Faculty of Medicine, Assiut University, Assiut, Egypt
Email [email protected]
Background: The emerging severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) resulted in a worldwide devastating effect with a diagnostic challenge. Identifying risk factors of severity aids in assessment for the need of early hospitalization. We aimed to demonstrate, for the first time, the clinical, laboratory and radiological characteristics of coronavirus disease 2019 (COVID-19) patients, to identify the predictors of severity and to describe the antimicrobial resistance profile in patients from Upper Egypt.
Materials and Methods: Demographic characters, clinical presentations, laboratory, and radiological data were recorded and analyzed. Presence of other respiratory microorganisms and their sensitivity patterns were identified using the VITEK2 system. Resistance-associated genes were tested by PCR.
Results: The study included 260 COVID-19 patients. The majority were males (55.4%) aged between 51 and 70 years. Hypertension, diabetes, and ischemic heart disease were common comorbidities. Main clinical manifestations were fever (63.8%), cough (57.7%), dyspnea (40%) and fatigue (30%). According to severity, 51.5% were moderate, 25.4% mild and 23% severe/critical. Lymphopenia, elevated CRP, ferritin, and D-dimer occurred in all patients with significantly higher value in the severe group. Age > 53 years and elevated ferritin ≥ 484 ng/mL were significant risk factors for severity. About 10.7% of the COVID-19 patients showed bacterial and/or fungal infections. Klebsiella pneumoniae, Acinetobacter baumannii, and Staphylococcus aureus were the predominant isolated bacteria while Candida albicans and Candida glabrata were the predominant isolated fungi. All Staphylococci were methicillin-resistant and carried the mecA gene. Gram-negative isolates were multidrug-resistant and carried different resistance-associated genes, including NDM-1, KPC, TEM, CTX-M, and SHV.
Conclusion: Older age and elevated serum ferritin were significant risk factors for severe COVID-19. Bacterial co-infection and multidrug resistance among patients with COVID-19 in Upper Egypt is common. Testing for presence of other co-infecting agents should be considered, and prompt treatment should be carried out according to the antimicrobial sensitivity reports.
Keywords: COVID-19, clinical characteristics, ESBL, KPC, NDM-1, risk factors, disease severity, Upper Egypt
Introduction
The novel coronavirus (COVID-19), causing acute respiratory illness, initially spread from China and resulted in a pandemic.1 The World Health Organization (WHO) declared COVID-19 as a public health emergency.2 To the date of writing this study, there have been more than 25 million confirmed cases of COVID-19, including about 845.000 deaths, according to WHO reports.3 COVID-19 infection in Egypt showed a rapid increase in incidence over a short period of time (Figure 1).4 Coronavirus belongs to the family of Coronavirinae, the phylogenetic analysis of COVID-19 genome shows that the virus is closely related to the Bat-SL-CoV and without any divergence at the genomic level.5
It mainly infects the respiratory system and is transmitted via respiratory droplets primarily through close contact. The virus could result in mild illness up to a critical care condition, which requires specialized treatment at intensive care units.6–8 Recovery after 1 week was usually reported in patients with mild symptoms. Meanwhile, severe cases could develop progressive respiratory failure by the alveolar damage with rapid progression to acute respiratory distress syndrome (ARDS), which could lead to death.9
Clinical symptoms of COVID-19 resemble those of other viruses or bacteria, causing respiratory illnesses.10 Therefore, the overlapping presentation makes the clinical diagnosis problematic, particularly during seasonal flu.11 The diagnosis of COVID-19 using laboratory panels was recommended. These panels include complete blood count (CBC) and coagulation tests such as fibrinogen or D-dimer, prothrombin time or partial thromboplastin time.12 Other biochemical tests are recommended such as C-reactive protein (CRP), ferritin, liver, renal tests and cardiac troponin reflecting the development of viral sepsis, systemic inflammatory response syndrome (SIRS), and/or multiple organ failure.12
Although reverse transcription polymerase chain reaction (RT-PCR) is considered the standard test to diagnose COVID-19, yet it may yield a false-negative result in some cases. The role of radiological imaging has been recognized in most cases as a crucial step for the diagnosis of suspected COVID-19 cases. Computed tomography (CT) seems to be feasible, available, and rapid method with higher sensitivity when compared to RT-PCR.11,13 As there is no specific curative treatment for COVID-19 till now, the WHO guidelines recommend the management according to each patient’s condition with particular caution to pediatric patients, pregnancy, and patients with comorbidities.14
Cases of COVID-19 were announced in Egypt by the end of February 2020. Meanwhile, the number of cases increased to more than 98,000 cases by the end of August 2020.4 The Egyptian government has made considerable efforts to contain the outbreak, and the fight against coronavirus is still ongoing. The Egyptian government has put several strategies including social distance and home management to reduce the spread of the virus. Identifying risk factors of severe disease will aid in assessment for early hospitalization and reduce mortality.
Bacterial and fungal co-infection in hospitalized patients with COVID-19 is poorly understood and the differentiation between viral infection by COVID-19 and possible presence of secondary bacterial or fungal infection on clinical basis is challenging. In Wuhan, secondary bacterial infections were reported in 15% of COVID-19 hospitalized patients, with higher prevalence in the non-survivor group compared to survivors (50% versus 1%).15,16 During influenza viral pneumonitis epidemics and pandemics, increased secondary bacterial and fungal infections were reported,16–18 poor patient outcomes were noticed with S. pneumoniae, H. influenzae, S. aureus and Aspergillus spp.19 As the COVID-19 pandemic continues, a considerable increase in antimicrobial resistance is foreseen through the heavy use of antibiotics in COVID-19 patients. Multidrug-resistant bacteria are leading to increased morbidity and mortality. Therefore, the need for antibiotic treatment should be promptly evaluated and discontinued when not appropriate.20
Nevertheless, there is a lack of studies conducted in Egypt to characterize the disease and predict its severity. Therefore, we aimed in this study to demonstrate the clinical, laboratory, radiological characteristics, and the predictors of severity of COVID-19 infection at the initial diagnosis of the disease. In addition, we aimed to evaluate the antibiotic resistance profile for co-infections associated with COVID-19 infection.
Materials and Methods
Study Population and Ethical Statement
This was a prospective study conducted on COVID-19 patients admitted to Alrajhi Liver University hospital and Assiut University hospital as tertiary hospitals in the period between the 3rd of May and the 30th of June. Diagnosis of the cases was confirmed using RT-PCR for detection of the viral RNA (Cobas 6800 system-Roche), according to the WHO and Egyptian Ministry of Health and Population (MOH) definitions.14 Written informed consent was provided by adult participants or parent or legal guardian of patients under the age of 18 years. The study protocol was approved by the ethical committee of the Faculty of Medicine, Assiut University (IRB no 17,300,381). It was conducted in accordance with the provisions of the Declaration of Helsinki. Informed written consent was obtained from each eligible patient who participated in the study. The inclusion criteria included hospitalized COVID-19 patients of different age groups including children (confirmed by RT-PCR test). Patients were divided according to disease severity into three groups: mild, moderate, severe and critical cases. Pregnant women were excluded from the study.
Demographic and Clinical Characteristics
Demographic characteristics, including sex, age, smoking status, underlying comorbidities, history of contact with a confirmed case were recorded. We also reported clinical features including; fever, headache, fatigue or myalgia, respiratory manifestations such as dry cough, expectoration, sore throat, shortness of breath, rhinorrhea or chest pain, gastrointestinal symptoms, e.g., nausea, vomiting or diarrhea and assessment for eye congestion and signs of respiratory distress.
Classification of Patients
Patients were categorized according to the severity of the disease based on the Egyptian MOH protocol (version 1.4, 30 May) into mild, moderate, severe or critical groups. Cases were considered mild if they were asymptomatic or symptomatic with leucopenia or lymphopenia and no evidence of pneumonia in the radiological image. Moderate cases were symptomatic with pneumonia on radiology and/or leucopenia or lymphopenia. Severe cases were presented with either; respiratory rate >30 breaths/min; SpO2 ≤92% on room air, PaO2/FiO2 ratio <300 or CT was showing more than 50% lesion or progressive lesion within 24–48 hours. Critical cases were defined as cases who had respiratory rate >30 breaths/min, SpO2 ≤92% on room air or PaO2/FiO2 ratio <300 despite O2 therapy.
Laboratory Characteristics
Laboratory investigations included CBC (by Pentra 80 Horiba blood counter), D-dimer (on sysmex 1500) and serum ferritin (by Advia centaur). In addition, CRP, serum urea, creatinine, liver enzymes (ALT and AST), troponin (Dimension RXL automated blood chemistry analyser), and oxygen saturation by blood gas analysis (Gym 3000).
Radiological Characteristics
Radiological features were described as regarding chest X-ray in the postero-anterior erect position and/or chest multislice CT (MSCT) in the supine position. Lesions were defined according to the radiological pattern, the affected side, and the involved lung lobes.
Microbiological Examination
Microbiological examination was only done for patients with suspected co-infection. Under proper infection control measures, sputum or endotracheal aspirates were collected from COVID-19 patients throughout the course of illness. Blood samples were inoculated directly into BacT/ALERT blood culture bottles (BioMérieux, Marcy l’Etoile, France) and monitored for bacterial growth. Samples were streaked out on Blood agar (Oxoid, CM0271) with 5% mammalian blood, initial identification of the bacterial isolates was done by Gram staining and growth characteristic on Mannitolsalt agar (Oxoid, PO0151) and MacConkey agar (Oxoid, CM0007). Microorganisms and their antimicrobial sensitivity patterns were identified using the VITEK2 Microbiology automated system (BioMerieux’s). To distinguish between colonization and infection, 10 μL of the respiratory specimen were inoculated on appropriate bacteriological media and incubated at 37◦C for 24–48 h. Then, the number of CFU per mL of the sample was determined. Presence of ≥105 CFU/mL in sputum or endotracheal aspirates indicated bacterial co-infection based on the American Thoracic Society definitions.21,22
Detection of Antibiotic Resistance Genes by Monoplex PCR Technique
Bacterial DNA was extracted from the clinical isolates obtained from COVID-19 co-infected patients using QIAamp® DNA Mini Kit (QIAGEN GmbH, Hilden, Germany) in accordance with manufacturer’s instructions and stored at −20 °C. Monoplex PCR assay was performed in order to identify and amplify the following antimicrobial resistance genes (mecA, NDM-1, KPC, TEM, CTX-M, and SHV) obtained from multidrug-resistant isolated strains. Supplementary Table 1 summarizes the specific primers used for each resistance gene. In each run, non-template control was used as a negative control and the following standard control strains; MRSA (ATCC 43,300, mecA), K. pneumoniae ATCC strain BAA-2146 (NDM-1), Klebsiella pneumoniae ATCC BAA-1705 (KPC-TEM-SHV), E. coli CCUG62975 (CTX-M) were used as positive controls. Amplified products were visualized by electrophoresis in 2% agarose gels stained with ethidium bromide.
Hospital Management and Complications
We also recorded care provided in the hospitals regarding the medical treatment, the received drugs, the requirement for oxygen therapy or mechanical ventilation, duration of hospitalization, and the patients’ mortality. Development of complications such as ARDS, sepsis, and disseminated intravascular coagulopathy (DIC) and renal failure were recorded.
Statistical Analysis
Statistical analyses were carried out using SPSS for Windows version 19 (SPSS, Chicago, IL, USA), and GraphPad Prism 8.4 (GraphPad, La Jolla, CA, USA). Categorical variables were expressed as a number and percentages and compared using chi-squared (χ2) or Fisher’s exact probability test. Continuous normally distributed measurements were presented as means ± standard deviations. Results of patient laboratory parameters were compared among clinical groups (mild, moderate, and severe groups) and statistically analyzed, and significance of differences was determined on the ranks with a one-way analysis of variance (ANOVA) and Tukey’s multiple comparison tests. A p-value of <0.05 was considered statistically significant. To analyze the risk factors associated with severity, a univariate logistic regression model for all variables was carried out, and variables that achieved a p <0.05 were included in the multivariate logistic regression model. Continuous variables were dichotomized, according to the normal reference values. Odds ratios (OR) and 95% confidence intervals (CI) for the tested variables were calculated in univariate and multivariate logistic regression analysis. Using DeLong test, significant variables on the multivariate regression analysis were evaluated using receiver-operating-characteristic (ROC) curve analysis and is expressed as the area under the ROC curve and the estimated sensitivity and specificity.
Results
Demographic and Clinical Characteristics
Among the COVID-19 suspected cases, 260 patients were confirmed to have COVID-19 infection by RT-PCR. Demographic and clinical characteristics are shown in Table 1. The majority of the included cases were males (55.4%); the most common age range was between 51 and 70 years (36.2%). Hypertension and diabetes mellitus were the most common reported comorbidities (29.2% and 23.7%, respectively). More than half of the patients did not have a history of contact with confirmed cases (54%). Regrading the clinical manifestations, about two-thirds of the included patients (63.8%) presented with fever, more than half had a dry cough (57.7%), 40% had shortness of breath, and about one-third (30%) had fatigue. However, 6.9% were asymptomatic contacts to positive cases. When categorizing the patients according to severity, it was shown that most cases were of moderate severity 51.5% (n=134), 25.4% were mild cases (n=66), and 23% were severe or critical cases (n=60).
 |
Table 1 Demographic and Clinical Characteristics of the Included Patients |
Laboratory Characteristics
As demonstrated in Table 2, the mean lymphocytic count was at the lower limit of normal (1.5±0.85), but a significantly lower percent of lymphocytes was observed in the severe group relative to moderate and mild groups. There was an elevation of the level of CRP, ferritin, and D-dimer in all patients. Their mean level was higher in the severe group with a statistical significance for CRP and ferritin. ALT and AST were significantly higher in the severe group compared to mild group (p=0.0038 and p<0.0001, respectively). AST level also showed a significant difference among all groups. Serum creatinine was significantly different among all groups, with the highest level in the severe group (2±1.3 mg/dL). Significantly low oxygen saturation was found in the moderate and severe groups compared to the mild group (p <0.0001).
 |
Table 2 The Laboratory Data of the Included Patients According to the Groups of Severity |
Radiological Characteristics
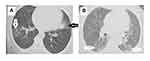
As imaging in asymptomatic and mild cases with no risk factors for disease progression are not routinely indicated according to recent recommendations by the American College of Chest Physicians, not all cases in these categories had radiological data.23 However, chest x-ray in those who did not perform MSCT was normal. Meanwhile, patients in moderate and severe or critical groups usually develop MSCT abnormalities, as shown in Table 3. All patients in the severe group had CT examination except for 16 cases who were in a critical condition on admission and required immediate management without delay. The most frequent findings were scattered irregular peripheral patches of ground-glass opacities in 72.2% (n=140), followed by consolidation in 22.7% (n=44), then crazy paving in 17.5% (n=34), while the least presentations were tree in bud appearance in 5.2% (n=10) and mediastinal lymphadenopathy in 3% (n=6) (Figure 2A and B). CT lesions were bilateral in the majority of cases (70%). Pulmonary involvement was detected in two or more lung lobes in the majority of cases (n=114) followed by the affection of lower lung lobes alone (n=44), suggesting that lower lobes were predominantly infected while the upper lobes were the least to be infected.
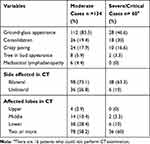 |
Table 3 Radiological Characteristics of Lung Lesions in the CT in Moderate and Severe Cases |
Hospital Management and Complications
Patients in the mild group were younger (mean 23.3± 18.9 years) relative to the moderate group (46.6±14.9) and severe group (57.7± 16.2). As demonstrated in Table 4, a higher percentage of males were observed in moderate (56.7%) and severe groups (66.6%). Comorbidities and smoking were more in severe groups relative to moderate group (63.3%, 26.7%, respectively). All symptomatic patients received treatment in the form of either paracetamol as antipyretic, cough suppressants, fluid therapy, multivitamins, e.g.: vitamin C or zinc, etc. According to the Egyptian protocol for management of COVID-19, patients could receive antivirals as azithromycin, or hydroxychloroquine, while favipiravir was recently introduced in Egypt and was received by only 14 cases. Anticoagulants were used in 8.9% of moderate cases and in 13.3% of severe cases. Oxygen therapy was used in moderate and severe cases (1.5% and 86.7%, respectively). Mechanical ventilation was required by 13.3% in the severe group. Complications such as ARDS occurred in 20% (n=12) of the severe and critical group. After the exclusion of patients with chronic kidney disease, acute renal injury was observed in moderate and severe groups (10.4% and 6.7%, respectively); however, it was mild and transient in moderate cases. Both ARDS and renal injury complicated 13.3% of the severe group.
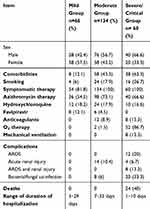 |
Table 4 Clinical Features, Treatment, Complications, Mortality, and Duration of Hospitalization Among the Groups of Patients According to Severity |
The mortality rate in severe and critical cases was 40%. The mean age of death was 62.8±14 years, the majority were complicated by ARDS (n=20); out of them, eight cases had both ARDS with renal injury. Six cases were complicated with co-infections. Longer duration of hospitalization was observed in the moderate group (7–33 days) relative to the other groups. Patients were discharged from the hospital after having two negative RT-PCR 48 hours apart after clinical and/or radiological improvement.
Microbiological Examination
Among the studied COVID-19 patients, 28/260 cases (10.7%) showed bacterial and/or fungal co-infections. Only respiratory samples showed bacterial growth while blood cultures were all negative. Total number of clinical isolates obtained was 42, of which 37 were bacteria and 5 were fungi. Gram-negative isolates were more predominant 30/42 (71.4%) than Gram-positive bacteria. There were 5 (11.9%) Staphylococcus aureus, 2 (4.7%) Streptococcus pneumoniae, 1 (2.3%) E. faecalis, 12 (28.5%) Klebsiella pneumoniae, 7 (16.6%) Acinetobacter baumannii, 4 (9.5%) Escherichia coli, 4 (9.5%) Pseudomonas aeruginosa and 2 (4.7%) E.cloacae. In addition, five cases (11.9%) showed fungal infections, which were all mixed with bacteria. Isolated fungi belonged to Candida spp. and included three isolates of Candida albicans and two isolates of Candida glabrata. Candida albicans isolates were sensitive to amphotericin B, flucytosine, caspofungin and micafungin, but resistant to fluconazole and voriconazole. Interestingly, no co-infections were observed in patients of the mild group. Eight patients who had co-infections were moderate cases, while 20 were detected in severe COVID-19 patients. Presence of co-infection was linked to 6/24 (25%) of the cases of mortality. These cases were also associated with increased respiratory severity and complications.
According to the results of the antibiogram (Table 5), most isolated strains had a multidrug-resistant pattern. In total, Gram-positive clinical isolates showed 100% resistance to amoxicillin, gentamicin and piperacillin/tazobactam and moderate resistance against amikacin (3/8) 37.5% and levofloxacin (6/8) 75%. On the other hand, all Gram-positive isolates were sensitive to vancomycin, linezolid and tigecycline. Regarding Gram-negative clinical isolates, they were mostly ESBL and carbapenemase producers. One hundred percent resistance was observed against amoxicillin, amoxicillin–clavulanic acid, cefoxitin, erythromycin, while variable resistance was observed against levofloxacin (20/29) 69%, ciprofloxacin (22/29) 75.9%, cefepime and meropenem (24/29) 82.8%, clindamycin (25/29) 86.3%, both imipenem and piperacillin/tazobactam (26/29) 89.7%. The highest sensitivity was observed for tigecycline (24/29) 82.7% and amikacin (23/29) 79.3%.
 |
Table 5 Antimicrobial Resistance Pattern and Genes Profiling in COVID-19 Co-Infections |
In addition, the isolated strains were evaluated for the presence of different antimicrobial resistance genes. All Staphylococcus aureus carried the mecA gene. On the other hand, NDM-1 was the most predominant gene (16/29) 55.1% among the Gram-negative clinical isolates, followed by CTX-M (15/29) 51.7%, TEM (12/29) 41.3%, KPC (10/29) 34.4% and SHV (2/29) 6.8%.
Risk Factors for Severity of COVID-19
The univariate regression analysis for predicting the severity of COVID-19 patients (Table 6) showed that age >53 years, diabetes mellitus, ischemic heart disease, CT involvement of two or more lung lobes, bilateral lung lesion, lymphopenia, elevated CRP, raised ferritin, increased liver enzymes (ALT and AST), urea, creatinine and presence of co-infection were significant risk factors for severe disease. CT involvement of two or more lung lobes had the highest odds ratio (OR: 7.342, 95% CI: 2.519 to 26.89, p=0.0007) followed by bilateral lesion in CT (OR: 6.592, 95% CI: 2.082 to 29.31, p=0.0039), ischemic heart disease (OR: 4.800, 95% CI: 1.188 to 20.67, p=0.0266) and diabetes mellitus (OR: 3.734, 95% CI: 1.527 to 9.175, p=0.0038).
 |
Table 6 Logistic Regression Analysis Evaluating Risk Factors for Severity of COVID-19 in the Studied Patients |
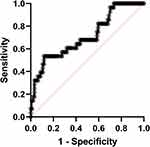
On multivariate regression analysis, only age >53 years and elevated ferritin were significant risk factors for severe COVID-19 (OR: 1.074 and 1.004, respectively). The ROC curve analysis of ferritin level discriminates severe COVID-19 patients from mild or moderate severity showed the sensitivity, specificity, and cut-off point were (64.3%, 60% and ≥484 ng/mL) for severe COVID-19 (Figure 3).
Discussion
The spread of COVID-19 infection caused a major devastating effect worldwide by its rapid transmission and high rate of infection. Similarly, COVID-19 infection in Egypt showed a rapid increase in incidence over a short period of time (Figure 1).4
Knowledge of the different characteristics of COVID-19 infection would help in the management of infected patients. However, the diagnosis of COVID-19 infection may be challenging due to the diversity of clinical picture and radiological findings in the suspected cases. In this report, we studied the clinical, radiological, laboratory characteristics, and risk factors for severe COVID-19 infection in Egypt. To the best of our knowledge, this is the first study on the risk factors of severity of COVID-19 infection in Upper Egypt.
In the current study, males were more frequently infected than females, which keeps in line with a recent meta-analysis.24–26 Females often have lower susceptibility to viral infections because they develop stronger immune responses than males, which could be attributed to sex hormones that play an essential role in innate and adaptive immunity.27
In this study, hypertension, diabetes mellitus, and ischemic heart disease were the most common reported comorbidities. Similarly, previous reports showed that hypertension was the most common comorbidity in the recruited patients, followed by diabetes mellitus.24,28 In a parallel line to the previous reports, we found that fever, dry cough, shortness of breath, and fatigue were the most common presenting symptoms, while sore throat, eye congestion, rhinorrhea, and diarrhea were less predominant.25,29 Asymptomatic patients were 6.9%, similar to a meta-analysis that reported a 5.6% proportion were asymptomatic COVID-19.30
Regarding COVID-19 severity, a Chinese study reported that the majority of patients were of mild severity (59.79%), 21.13%were moderate, 12.89% were severe, and 6.19% were critically ill.31 On the other side, another study showed that the majority had severe disease (47.5%), while37.8% had a moderate type, 14.2% were in critical conditions, and 0.5% had mild infection.32 Meanwhile, in our study, moderate severity of COVID-19 was observed in the majority of patients, which is consistent with a meta-analysis that reported 25.6% of patients presented with severe illness.30 These changes in the proportion of disease severity among studies could be attributed to different criteria used for the classification or the time of presentation of the patients to the hospital, patient criteria and geographical distribution. Late hospitalization of patients increases the probability of having a severe or critical stage on admission, while early medical evaluation reduces the severity.
The most frequently reported laboratory abnormalities were lymphopenia, elevated CRP, serum ferritin, and D-dimers in all patients. However, all these laboratory markers are very non-specific, causing limited clinical utility, and they are not indistinguishable from patients with other coronavirus infections, such as severe acute respiratory syndrome (SARS) and Middle-East respiratory syndrome (MERS).33,34 The predominance of lymphopenia in COVID-19 suggests that the virus might act on lymphocytes, particularly T lymphocytes, inducing a reduction of CD4.35
In the current study, we observed higher frequency of comorbidities, marked abnormalities in laboratory or radiological findings and complications were higher in patients with severe or critical disease compared to patients without severe illness. This is in concordance with other studies.30 Negative correlation was reported between lymphocytes count and the severity of the disease combined with increased ferritin and D-dimer in severe and extremely severe patients.36
In this study, the predominance of bilateral peripheral multilobar ground-glass opacities was consistent with a systematic review that reported the characteristic CT patterns in COVID-19 infection. It showed that ground-glass opacification occurs in 88%, consolidation in 32%, bilateral involvement in 87.5% with peripheral distribution in 76.0%, and multilobar involvement in 78.8%.37 Possible findings such as CT halo sign, pleural effusion, cavitation, lymphadenopathy, and pneumothorax were uncommon but can be seen with the disease progression.38,39 Patients of moderate severity in this study showed that CT findings were suggestive of the disease progression. This could be explained by Jin et al study that showed five temporal stages of the CT changes; ultra-early, early, rapid progression, consolidation, and dissipation stages. These stages depend on the duration of illness.40 Similarly, Pan et al41 on an assessment of follow-up CT within 3–14 days after an initial CT examination found that >85% of the cases showed radiological signs associated with disease progressions such as an increase in ground-glass appearance, consolidative opacities, and interstitial septal thickening.41 Moreover, in patients who have typical clinical manifestations but false-negative RT-PCR, chest CT will be valuable in assessment showing the characteristic features of the disease.42 The correlation between CT findings and the disease severity and mortality can be concluded from several studies. In severely ill patients, the most common CT imaging was bilateral, multilobar involvement, and subsegmental consolidation.37
Microbial co-infection notably increases the risk of disease severity and mortality among humans.43,44 In the present study, most co-infections occurred within 1–7 days of COVID-19 disease onset and these microorganisms showed MDR phenotype. Among large pandemics of the viral respiratory infections, the role of co-infection is understudied.44 Our results showed that co-infections could affect COVID-19 disease progression and mortality. The most frequently detected pathogens in COVID-19 patients were Klebsiella pneumoniae (28.5%) followed by Acinetobacter baumannii (16.6%), Staphylococcus aureus (MRSA) (11.9%), Escherichia coli and Pseudomonas aeruginosa were (9.5%), Candida albicans (7.1%). Lansbury, Lim45 reported a different co-pathogen profile, as the most common bacteria were Mycoplasma pneumoniae (42%) followed by Pseudomonas aeruginosa (12%) and Haemophilus influenzae (12%), while fungi were Aspergillus flavus (two cases), Aspergillus fumigatus and Candida glabrata (one case). Many viral respiratory pathogens are associated with other bacterial co-infections. Importantly, viral agents can reduce the mucociliary clearance of bacteria from the lungs.46 Therefore, different investigators have described the bacterial/fungal co-infection in patients with COVID-19. However, the burden of co-infections in COVID-19 patients varies significantly between studies.8,45,47–49 The discrepancy in results could be attributed to many reasons. It seems that the geographical location, the presence of antimicrobial policy, appropriate stewardship interventions, robust surveillance for healthcare-associated infections and antimicrobial resistance have a great impact on the presence of bacterial/fungal co-infection in patients with coronavirus infection.45 Of particular importance, is the application of strict infection control measures since most of the detected pathogens are important causes of nosocomial infections within the hospital environment.50–53
We noticed the predominance of Gram-negative co-infections in COVID-19 patients, which may be attributed to the administration of azithromycin in the regimen of COVID-19 therapy that acts primarily against Gram-positive bacteria.54 Besides, in line with our results, a study performed at Assiut University Hospitals identified Gram-negative bacteria (Klebsiella pneumoniae and H. influenzae) as the main bacteria isolated from hospitalized patients with chronic obstructive pulmonary disease.55 In addition, an outbreak caused by Acinetobacter baumanii was recorded in Assiut University Hospitals.56,57 This was also comparable to observations of Lin, Kuo58 who reported Klebsiella pneumoniae and P. aeruginosa, followed by A. baumannii as the most common agents. Prevalence of Gram-negative resistant bacteria has increased because only few new antibiotic agents attacking this group of pathogens have been produced successfully.59
Antimicrobial resistance is a growing crisis that affects the global health and demands urgent action. Notably, most isolated strains in our study had a multidrug-resistant pattern. Among the isolated Gram-positive strains, all Staphylococcus aureus were MRSA and 100% carried mecA gene. Methicillin resistance in a growing problem in coagulates positive and negative staphylococci.60,61 Interestingly, about half of Staphylococci aureus isolated from ambulance vehicles were MRSA.52 A recent study from our university hospital also reported that all staphylococci isolated from patients with nosocomial pneumonia were methicillin resistant.62 Regarding Gram-negative clinical isolates, mostly they were ESBL and/or carbapenemase producers. The highest sensitivity was observed for tigecycline and Amikacin. NDM1 was the predominant detected gene followed by CTX-M, TEM, KPC and SHV. Coexistence of several resistance genes was markedly observed in the Gram-negative isolates which is in consistent with observations of Lincopan and McCulloch,63 who detected the coexistence of MBL IMP-1 and ESBL CTX-M in Klebsiella pneumoniae. The production of these enzymes (ESBL and MBL) together allows bacteria to resist all antibiotics, including carbapenems, and is the main cause for treatment failure.63 Therefore, for COVID-19 cases who show co-infections with other respiratory pathogens, prompt administration of antibiotics according to the antimicrobial sensitivity reports as well as strict application of infection control strategies are essential to reduce severity, complications, mortality and nosocomial spread achieved by these superbugs.64–66
Analysis of risk factors of severe COVID-19 showed that age >53 years, and high level of ferritin were significant risk factors for severe disease. In one study, predictors of severity were age ≥65 years, hypertension, and D-dimer >1 mg/L were significantly associated with severe cases.67
In this study, a ferritin level ≥484 ng/mL was a significant risk factor for severe disease. A recent meta-analysis showed a ferritin level 408.3 ng/mL could be used to differentiate between severe and non-severe cases, indicating the development of a SIRS in patients with severe illness. The severe disease in this study was defined as patients with ARDS, need mechanical ventilation, need vital life support, or need intensive care management.68 However, in this study analysis of the cutoff level of ferritin showed low sensitivity and low accuracy. Therefore, further studies on variable risk factors and laboratory parameters in severe illness should be conducted, considering differences in the methodology used to define the severe disease.
One of the limitations of our study is the diversity of the pattern of bacterial profile and drug resistance, which could be attributed to geographical difference of the patients referred to the hospital from various regions of Upper Egypt. Another limitation is the lack of analysis of other respiratory viruses including influenza viruses.
The Egyptian MOH guideline, by the end of May, recommended that mild and some moderate cases will be managed by home isolation. We are providing an overview of the disease in Egypt and the risk factors of the severe disease so that cases managed at home will be alerted if they have any of these factors for early hospitalization.
Conclusions
Older age and elevated serum ferritin were significant risk factors for severe COVID-19. Bacterial co-infection and multidrug resistance among patients with COVID-19 in Upper Egypt is common. Our results highlighted the importance of prompt administration of antibiotics according to the antimicrobial sensitivity reports.
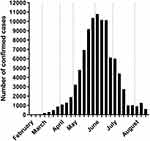 |
Figure 1 WHO report of the COVID-19 cases in Egypt. |
Abbreviations
SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; COVID-19, coronavirus disease 2019; ESBL, extended-spectrum β-lactamase; CRP, C-reactive protein.
Funding
The authors have not received external funding for this study.
Disclosure
The authors declare no conflicts of interest for this work.
References
1. Guo Y-R, Cao QD, Hong ZS, et al. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak–an update on the status. Mil Med Res. 2020;7(1):1–10.
2. Sohrabi C, Alsafi Z, O’Neill N, et al. World Health Organization declares global emergency: a review of the 2019 novel coronavirus (COVID-19). Int J Surg. 2020.
3. WHO. Coronavirus Disease (COVID-19) Dashboard; 2020.
4. WHO. 2020. Available from: https://covid19.who.int/region/emro/country/eg.
5. Kumar Y, Singh H, Patel CN. In silico prediction of potential inhibitors for the main protease of SARS-CoV-2 using molecular docking and dynamics simulation based drug-repurposing. J Infect Public Health. 2020;13(9):1210–1223. doi:10.1016/j.jiph.2020.06.016
6. Zhu N, Zhang D, Wang W, et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med. 2020.
7. Chan JF, Yuan S, Kok KH, et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395(10223):514–523.
8. Chen N, Zhou M, Dong X, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–513. doi:10.1016/S0140-6736(20)30211-7
9. Li T, Wei C, Li W, Hongwei F, Shi J. Beijing Union Medical College Hospital on “pneumonia of novel coronavirus infection” diagnosis and treatment proposal (V2. 0). Med J Peking Union Med Coll Hosp. 2020.
10. Li Q, Guan X, Wu P, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus–infected pneumonia. N Engl J Med. 2020;382(13):1199–1207. doi:10.1056/NEJMoa2001316
11. Ai T, Yang Z, Hou H, et al. Correlation of chest CT and RT-PCR testing in coronavirus disease 2019 (COVID-19) in China: a report of 1014 cases. Radiology. 2020;200642.
12. Favaloro EJ, Lippi G. Recommendations for Minimal Laboratory Testing Panels in Patients with COVID-19: Potential for Prognostic Monitoring. In Seminars in Thrombosis and Hemostasis. Thieme Medical Publishers; 2020.
13. Mahdavi A, Khalili N, Davarpanah AH, et al. Radiologic management of COVID-19: preliminary experience of the Iranian Society of Radiology COVID-19 Consultant Group (ISRCC). Iran J Radiol. 2020;17(2). doi:10.5812/iranjradiol.102324
14. Organization, W.H. Clinical Management of Severe Acute Respiratory Infection (SARI) When COVID-19 Disease is Suspected: Interim Guidance, 13 March 2020. World Health Organization; 2020.
15. Ruan Q, Yang K, Wang W, et al. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 2020;46(5):846–848. doi:10.1007/s00134-020-05991-x
16. Rice TW, Rubinson L, Uyeki TM, et al. Critical illness from 2009 pandemic influenza A (H1N1) virus and bacterial co-infection in the United States. Crit Care Med. 2012;40(5):1487.
17. Morris DE, Cleary DW, Clarke SC. Secondary bacterial infections associated with influenza pandemics. Front Microbiol. 2017;8:1041.
18. Canning B, Senanayake V, Burns D, et al. Post-influenza Aspergillus ventriculitis. Clin Infect Pract. 2020;7–8:100026. doi:10.1016/j.clinpr.2020.100026
19. Mulcahy ME, McLoughlin RMJM. Staphylococcus aureus and influenza A virus: partners in coinfection. MBio. 2016;7(6).
20. Bengoechea JA, Bamford CG. SARS-CoV-2, bacterial co-infections, and AMR: the deadly trio in COVID-19? EMBO Mol Med. 2020;12(7):e12560. doi:10.15252/emmm.202012560
21. American Thoracic Society, Infectious Diseases Society of America. Guidelines for the management of adults with hospital-acquired, ventilator-associated, and healthcare-associated pneumonia. Am J Respir Crit Care Med. 2005;171(4):388.
22. Wiegers HMG, van Nijen L, van Woensel JBM, et al. Bacterial co-infection of the respiratory tract in ventilated children with bronchiolitis; a retrospective cohort study. BMC Infect Dis. 2019;19(1):938. doi:10.1186/s12879-019-4468-3
23. Rubin GD, Ryerson CJ, Haramati LB, et al. The role of chest imaging in patient management during the COVID-19 pandemic: a multinational consensus statement from the Fleischner Society. Chest. 2020.
24. Guan W-J, Ni Z-Y, Hu Y, et al. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020;382(18):1708–1720. doi:10.1056/NEJMoa2002032
25. Li L-Q, Huang T, Wang Y-Q, et al. COVID-19 patients’ clinical characteristics, discharge rate, and fatality rate of meta-analysis. J Med Virol. 2020;92(6):577–583. doi:10.1002/jmv.25757
26. Qian K, Deng Y, Tai Y, Peng J, Peng H, Jiang L. Clinical characteristics of 2019 novel infected coronavirus pneumonia: a systemic review and meta-analysis. medRxiv. 2020.
27. Klein SL, Huber S. Sex Differences in Susceptibility to Viral Infection, in Sex Hormones and Immunity to Infection. Springer; 2010:93–122.
28. Yang J, Zheng Y, Gou X, et al. Prevalence of comorbidities and its effects in patients infected with SARS-CoV-2: a systematic review and meta-analysis. Int J Infect Dis. 2020;94:91–95. doi:10.1016/j.ijid.2020.03.017
29. Chew NWS, Lee GKH, Tan BYQ, et al. A multinational, multicentre study on the psychological outcomes and associated physical symptoms amongst healthcare workers during COVID-19 outbreak. Brain Behav Immun. 2020;88:559–565. doi:10.1016/j.bbi.2020.04.049
30. Fu L, Wang B, Yuan T, et al. Clinical characteristics of coronavirus disease 2019 (COVID-19) in China: a systematic review and meta-analysis. J Infect. 2020;80(6):656–665. doi:10.1016/j.jinf.2020.03.041
31. Zhang H, Shang W, Liu Q, Zhang X, Zheng M, Yue M. Clinical characteristics of 194 cases of COVID-19 in Huanggang and Taian, China. Infection. 2020;1–8.
32. Zhang J, Wang X, Jia X, et al. Risk factors for disease severity, unimprovement, and mortality in COVID-19 patients in Wuhan, China. Clin Microbiol Infect. 2020;26(6):767–772. doi:10.1016/j.cmi.2020.04.012
33. Peiris JSM, Chu CM, Cheng V, et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: a prospective study. Lancet. 2003;361(9371):1767–1772. doi:10.1016/S0140-6736(03)13412-5
34. Assiri A, McGeer A, Perl TM, et al. Hospital outbreak of Middle East respiratory syndrome coronavirus. N Engl J Med. 2013;369(5):407–416. doi:10.1056/NEJMoa1306742
35. Chen G, Wu D, Guo W, et al. Clinical and immunologic features in severe and moderate forms of coronavirus disease. J Clin Invest. 2019;137244.
36. Wang F, Hou H, Luo Y, et al. The laboratory tests and host immunity of COVID-19 patients with different severity of illness. JCI Insight. 2020;5(10). doi:10.1172/jci.insight.137799
37. Salehi S, Abedi A, Balakrishnan S, Gholamrezanezhad A, et al. Coronavirus disease 2019 (COVID-19): a systematic review of imaging findings in 919 patients. Am J Roentgenol. 2020:1–7. doi:10.2214/AJR.20.24425
38. Song F, Shi N, Shan F, et al. Emerging 2019 novel coronavirus (2019-nCoV) pneumonia. Radiology. 2020;295(1):210–217.
39. Li X, Zeng X, Liu B, Yu Y. COVID-19 infection presenting with CT halo sign. Radiology. 2020;2(1):e200026.
40. Jin Y-H, Cai L, Cheng Z-S, et al. A rapid advice guideline for the diagnosis and treatment of 2019 novel coronavirus (2019-nCoV) infected pneumonia (standard version). Mil Med Res. 2020;7(1):4. doi:10.1186/s40779-020-0233-6
41. Pan Y, Guan H, Zhou S, et al. Initial CT findings and temporal changes in patients with the novel coronavirus pneumonia (2019-nCoV): a study of 63 patients in Wuhan, China. Eur Radiol. 2020;1–4.
42. Diao K, Han P, Pang T, et al. HRCT imaging features in representative imported cases of 2019 novel coronavirus pneumonia. Precis Clin Med. 2020;3(1):9–13. doi:10.1093/pcmedi/pbaa004
43. Zhu X, Ge Y, Wu T, et al. Co-infection with respiratory pathogens among COVID-2019 cases. Virus Res. 2020;285:198005. doi:10.1016/j.virusres.2020.198005
44. Cox MJ, Loman N, Bogaert D, et al. Co-infections: potentially lethal and unexplored in COVID-19. Lancet Microbe. 2020;1(1):e11. doi:10.1016/S2666-5247(20)30009-4
45. Lansbury L, Lim B, Baskaran V, et al. Co-infections in people with COVID-19: a systematic review and meta-analysis. J Infect. 2020;81(2):266–275. doi:10.1016/j.jinf.2020.05.046
46. Noviello S, Huang DB. The basics and the advancements in diagnosis of bacterial lower respiratory tract infections. Diagnostics. 2019;9(2):37.
47. Wu C, Chen X, Cai Y, et al. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern Med. 2020;180(7):934. doi:10.1001/jamainternmed.2020.0994
48. Wang Z, Yang B, Li Q, et al. Clinical features of 69 cases with coronavirus disease 2019 in Wuhan, China. Clin Infect Dis. 2020;71(15):769–777. doi:10.1093/cid/ciaa272
49. Arentz M, Yim E, Klaff L, et al. Characteristics and outcomes of 21 critically Ill patients with COVID-19 in Washington State. JAMA. 2020;323(16):1612. doi:10.1001/jama.2020.4326
50. El-Mokhtar MA, Hassanein KM, Ahmed AS, et al. Antagonistic activities of cell-free supernatants of lactobacilli against extended-spectrum β-lactamase producing Klebsiella pneumoniae and Pseudomonas aeruginosa. Infect Drug Resist. 2020;13:543–552. doi:10.2147/IDR.S235603
51. Elgendy SG, Abdel Hameed MR, El-Mokhtar MA. Tigecycline resistance among Klebsiella pneumoniae isolated from febrile neutropenic patients. J Med Microbiol. 2018;67(7):972–975. doi:10.1099/jmm.0.000770
52. El-Mokhtar MA, Hetta HF. Ambulance vehicles as a source of multidrug-resistant infections: a multicenter study in Assiut City, Egypt. Infect Drug Resist. 2018;11:587–594. doi:10.2147/IDR.S151783
53. Ahmed SH, Daef EA, Badary MS, et al. Nosocomial blood stream infection in intensive care units at Assiut University Hospitals (Upper Egypt) with special reference to extended spectrum β-lactamase producing organisms. BMC Res Notes. 2009;2(1):76. doi:10.1186/1756-0500-2-76
54. Drew RH, Gallis HA. Therapy, azithromycin—spectrum of activity, pharmacokinetics, and clinical applications. Pharmacotherapy. 1992;12(3):161–173.
55. Hassan AT, Mohamed SA, Mohamed MS, El-Mokhtar MA. Acute exacerbations of chronic obstructive pulmonary disease: etiological bacterial pathogens and antibiotic resistance in Upper Egypt. Egypt J Bronchol. 2016;10(3):283.
56. Daef EA, Mohamad IS, Ahmad AS, El-Gendy SG, Ahmed EH, Sayed IM. Relationship between clinical and environmental isolates of Acinetobacter baumannii in Assiut University hospitals. J Am Sci. 2013;9(11s):67–73.
57. Enas AD, Ismail SM, Ahmed SA, et al. Detection of outbreak caused by multi-drug resistant Acinetobacter baumannii in Assiut University Hospitals. Afr J Microbiol Res. 2014;8(22):2238–2244. doi:10.5897/AJMR2014.6795
58. Lin S-H, Kuo P-H, Hsueh P-R, et al. Sputum bacteriology in hospitalized patients with acute exacerbation of chronic obstructive pulmonary disease in Taiwan with an emphasis on Klebsiella pneumoniae and Pseudomonas aeruginosa. Respirology. 2007;12(1):81–87. doi:10.1111/j.1440-1843.2006.00999.x
59. Kollef MH, Golan Y, Micek ST, et al. Appraising contemporary strategies to combat multidrug resistant gram-negative bacterial infections–proceedings and data from the gram-negative resistance summit. Clin Infect Dis. 2011;53(Suppl suppl_2):SS33–SS38. doi:10.1093/cid/cir475
60. Martins A, de Lourdes RS Cunha M. Methicillin resistance in Staphylococcus aureus and coagulase-negative staphylococci: epidemiological and molecular aspects. Microbiol Immunol. 2007;51(9):787–795.
61. Eltwisy HO, Abdel-Fattah M, Elsisi AM, et al. Pathogenesis of Staphylococcus haemolyticus on primary human skin fibroblast cells. Virulence. 2020;11(1):1142–1157. doi:10.1080/21505594.2020.1809962
62. Abdel-Rahim MH, Mohammed AH, Ghandour AM, Shaaban LH, Hassan HA. Bacterial nosocomial pneumonia in respiratory intensive care unit of Assiut University Hospital. Egypt J Med Microbiol. 2019;28(4):113–139.
63. Lincopan N, McCulloch JA, Reinert C, Cassettari VC, Gales AC, Mamizuka EM. First isolation of metallo-beta-lactamase-producing multiresistant Klebsiella pneumoniae from a patient in Brazil. J Clin Microbiol. 2005;43(1):516–519. doi:10.1128/JCM.43.1.516-519.2005
64. Rawson TM, Moore LS, Zhu N, et al. Bacterial and fungal co-infection in individuals with coronavirus: a rapid review to support COVID-19 antimicrobial prescribing. Clin Infect Dis. 2020. doi:10.1093/cid/ciaa530
65. Bills CB, Newberry JA, Rao GR, et al. Acute respiratory illness among a prospective cohort of pediatric patients using emergency medical services in India: demographic and prehospital clinical predictors of mortality. PLoS One. 2020;15(4):e0230911. doi:10.1371/journal.pone.0230911
66. Ruan Q, Yang K, Wang W, Jiang L, Song J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 2020;1–3.
67. Fann RJ, Rani VR, Vaithlingam I. Clinical presentations and predictors of mortality for leptospirosis - a study from suburban area in Malaysia. Med J Malaysia. 2020;75(1):52–56.
68. Henry BM, De Oliveira MH, Benoit S, Plebani M, Lippi G. Hematologic, biochemical and immune biomarker abnormalities associated with severe illness and mortality in coronavirus disease 2019 (COVID-19): a meta-analysis. Clin Chem Lab Med. 2020;58(7):1021–1028.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.