Back to Journals » International Journal of Nanomedicine » Volume 16
Platinum Nanoparticles Enhance Exosome Release in Human Lung Epithelial Adenocarcinoma Cancer Cells (A549): Oxidative Stress and the Ceramide Pathway are Key Players
Authors Gurunathan S , Kang MH , Jeyaraj M, Kim JH
Received 21 November 2020
Accepted for publication 31 December 2020
Published 22 January 2021 Volume 2021:16 Pages 515—538
DOI https://doi.org/10.2147/IJN.S291138
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Thomas Webster
Sangiliyandi Gurunathan, Min-Hee Kang, Muniyandi Jeyaraj, Jin-Hoi Kim
Department of Stem Cell and Regenerative Biotechnology, Konkuk University, Seoul 05029, Korea
Correspondence: Sangiliyandi Gurunathan; Jin-Hoi Kim
Department of Stem Cell and Regenerative Biotechnology, Konkuk University, Seoul 05029, Korea
Tel +82 2 450 3687
Fax + 82 2 544 4645
Email [email protected]; [email protected]
Background: Several studies have demonstrated various molecular mechanisms involved in the biogenesis and release of exosomes. However, how external stimuli, such as platinum nanoparticles (PtNPs), induces the biogenesis and release of exosomes remains unclear. To address this, PtNPs were synthesized using lutein to examine their effect on the biogenesis and release of exosomes in human lung epithelial adenocarcinoma cancer cells (A549).
Methods: The size and concentration of isolated exosomes were characterized by dynamic light scattering (DLS) and nanoparticle tracking analysis system (NTA). Morphology and structure of exosomes were examined using scanning electron microscopy and transmission electron microscopy (TEM), respectively. Quantification of exosomes were analyzed by EXOCETTM assay and fluorescence polarization (FP). The expression of typical markers of exosomes were analyzed by quantitative reverse transcription-polymerase chain reaction (qRT-PCR) and enzyme-linked immunosorbent assay (ELISA).
Results: A549 cells cultured with PtNPs enhance exosome secretion by altering various physiological processes. Interestingly, A549 cells treated with PtNPs increases total protein concentration, biogenesis and release of exosomes associated with PtNPs-induced oxidative stress. GW4869 inhibits PtNPs induced biogenesis and release of exosomes and also acetylcholinesterase (AChE), neutral sphingomyelinase activity (n-SMase), and exosome counts. A549 cells pre-treated with N-acetylcysteine (NAC) significantly inhibited PtNPs induced exosome biogenesis and release. These findings confirmed that PtNPs-induced exosome release was due to the induction of oxidative stress and the ceramide pathway. These factors enhanced exosome biogenesis and release and may be useful in understanding the mechanism of exosome formation, release, and function.
Conclusion: PtNPs provide a promising agent to increase exosome production in A549 cells. These findings offer novel strategies for enhancing exosome release, which can be applied in the treatment and prevention of cancer. Importantly, this is the first study, to our knowledge, showing that PtNPs stimulate exosome biogenesis by inducing oxidative stress and the ceramide pathway.
Keywords: exosome, platinum nanoparticle, cytotoxicity, oxidative stress, acetylcholinesterase activity, neutral sphingomyelinase activity
Introduction
Cancer is one of the complex disease and serious health issues arising from alterations in the cellular genome and affecting the expression and functions of oncogenes and tumor suppressor genes. It is widely accepted that the tumor microenvironment plays an essential role in tumor development and progression.1 Lung cancer is the most commonly diagnosed cancer in both males and females. However, the degree of mortality depends on the degree of economic development, social and lifestyle factors of individual countries.2 Although several types of therapies are available, such as radiation, stem cell, chemo, immuno, hormone, and targeted drug therapy, patients are getting undesired side effects and chemo-resistance. In addition, some natural products have shown potential in the prevention and treatment of cancers3. However, conventional cancer therapies cause chemoresistance and undesired side effects. Therefore, studies on cancer and the anticancer therapies are necessary, and great attention is required in the clinical arena. Recent studies have shown that cancer-derived exosomes can be beneficial in recruiting and reprogramming of constituents correlated with the tumor microenvironment. The tumor microenvironment is vital in detection, treatment, and prevention of cancer. Therefore, understanding the molecular mechanisms, detection at an early stage, and development of safe and highly efficient targeted therapeutic methods are crucial for lung cancer treatment.4 Recently, extracellular vehicles (EVs) have shown immense beneficial effects in assisting early detection and diagnosis and improving treatment outcomes in cancer.5 In a previous study, the combination of exosomes derived from gefitinib-treated lung cancer cells and cisplatin considerably reduced the cisplatin sensitivity of cancer cells, indicating a synergistic effect.6 These findings underline the importance of exosomes in mediating the antagonistic effects of cisplatin and gefitinib in lung cancer.
Exosomes are nanovesicles with an average size between 30 and 150 nm and are secreted by normal and diseased cells are playing significant role in intercellular communication to transfer various biomolecules to a recipient cell.7,8 Tumor cells have been shown to secrete significantly higher number of exosomes than normal cells.9 Several studies demonstrate that cell-released exosomes play a significant role in intercellular communication. Moreover, unique signatures of exosomes allow us to identify specific biomarkers for cancer identification and predict therapeutic outcomes through the transfer of bioactive molecules.10–12 Exosomes are serving as therapeutic tools and as diagnostic biomarkers for various diseases.10,13 The functions of exosomes depend on the origin of the cell type. Thakur et al reported that exosomes derived from lung tumor cells in culture reflected the mutational status of specific genes (EGFR) in the parental cell lines.14 Microvesicles/exosomes derived from various lung cancer cell lines transferred activated EGFR from cancer cells to endothelial cells and transferred microvesicles/exosomes induced the mitogen-activated protein kinase (MAPK) and AKT pathways.15 Exosomal miRNAs from lung cancer cells serve as diagnostic and prognostic biomarkers.16 Immune cells containing exosomes, which carry miR-21 and miR-29a, activate endosomal TLR7 and TLR8 are responsible for pro-inflammatory phenotype.17
Exosomes are secreted in almost all cell types including prokaryotes, eukaryotes and almost all body fluids. Due to its unique properties which are immensely used in biomedical applications.18 Therefore, exosomes are efficient delivery agents and it can across different biological barriers to target cells.19–21 Many different studies tried to increase the biogenesis and secretion of exosomes through various factors including intrinsic stresses, such as low pH-, hypoxia-, oxidative stress-, and thermal stress-induced alterations of melanoma exosomes, and their ability to transfer drug resistance.22,23 The biogenesis of exosomes depends on the percentage of confluency of approximately 60–90%, which influences the yield and functions of EVs.24 Exogenous stimulation, including Ca2+ ionophores25 and hypoxia,26,27 can influence the condition of the cells, including the phenotype, such as detachment of cells,28 as well as efficacy of secretion. Oxidative stress increases exosome secretion in retinal pigment epithelial cells.29 Oxidative stress increases autophagy in retinal astrocytes and promotes these cells to regulate endothelial cell function by releasing exosomes.30
Stress playing a vital role in exosomes release. Various types of stress conditions such as thermal and oxidative stress aggravate increased exosome release from cancer cells including leukemia/lymphoma T and B cell lines.31 Hypoxic conditions have been shown to be effective in enhancing Tex release from breast cancer cells.32 Sub-lethal doses of various chemotherapeutic agents stimulate exosome secretion in different tumor models.33,34 Similarly, 5-fluorouracil, cisplatin, and doxorubicin are known to induce an increased amount of HSP70+ exosome release from melanoma and colon cancer cell lines.35 However, nanoparticle-induced oxidative stress-elicited changes or release of exosomes have not been elucidated yet.
Due to its unique physical and chemical properties, nanoparticles represent an increasingly important material in the development of novel nano-devices, including silver nanoparticles, which are used as antimicrobial agents,36 gold nanoparticles as anticancer agents,37 and platinum nanoparticles used in various applications.38 Nanoparticles induce cell toxicity and cell death by generating oxidative stress. It was previously reported that ultra-small platinum nanoparticles induce cytotoxicity by inducing oxidative stress in the human monocytic THP-1 cell line.39 Platinum nanoparticles (PtNPs) increase the apoptotic potential of doxorubicin through the generation of reactive oxygen species in human bone OS epithelial cells (U2OS).40 Moreover, PtNPs have been shown to induce apoptosis in neuroblastoma cancer cells by triggering loss of cell viability, proliferation, endoplasmic reticulum-mediated stress, and apoptosis by activation of p53, Bax, and caspase-3 and downregulation of BCl-2.41 However, whether platinum nanoparticles influence exosome biogenesis and release by altering cell survival, cytotoxicity, and apoptosis in human lung epithelial adenocarcinoma cells (A549) remains unclear. In this study, we investigated the platinum nanoparticle-mediated effects on the cellular and exosomal functions of human lung epithelial adenocarcinoma cells (A549) cells cultured in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% EVs-depleted serum and in serum-free Opti-MEM. We first analyzed how cell viability, cell proliferation, total exosome protein concentration, acetylcholine esterase activity and exosomes secretion was affected by media and incubation time. Our findings show that serum-free conditions lead to an increase in EVs release, and the protein concentration of exosomes differ to those derived from cells grown in serum-containing media. Further, we evaluated the effect of platinum nanoparticles induced oxidative stress on exosomes release using series of parameters and released exosomes were characterized by nanoparticle tracking analysis and dynamic light scattering, scanning electron microscopy, transmission electron microscopy and also we analyzed the expression of typical biomarkers of exosomes such as TSG101, CD81, CD63 and CD9. Finally, for mechanistic investigation, we analyzed the possible links between oxidative stress and ceramide-dependent pathways for biogenesis and release of exosomes.
Materials and Methods
Synthesis and Characterization of PtNPs
PtNPs synthesis and characterization were performed as described previously.39 PtNPs were synthesized by the reduction of PtCl6 2- ions into PtNPs by mixing 10 mL of 1 mg/mL lutein with 90 mL of 1 mM aqueous H2PtCl6.6H2O (Sigma-Aldrich, St. Louis, MO, USA). The mixture was maintained at 100°C (on a hotplate) in a sealed flask to avoid evaporation, for 1 h, since temperature catalyzes the reduction process. The detailed methodology is given as supplementary file 1.
A549 Exposure to PtNPs, C6-Cer, CSP and GW4869
A 100 μL aliquot of cells at a density of 1 × 105 cells/mL was then plated in each well of the 96-well plates. After 24 h, the culture medium was replaced with a medium containing concentrations of PtNPs (10 µM) or C6-cer (10 µM), or CSP (10 µM) or GW4869 (20 µM). A dose-dependent effect was observed upon using various concentrations of PtNPs, C6-cer, CSP and GW4869 for 24 h. Cells were pre-incubated with GW4869 (20 µM) or 1 mM NAC for 12 h prior to PtNPs or C6-cer or CSP exposure.
Cell Viability
Cell viability was measured in A549 cells exposed to PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) or various concentrations of PtNPs (2.5–40 µM) or C6-cer (2.5–40 µM) or CSP (2.5–40 µM) or GW4869 (2.5–40 µM). After incubation for 24 h at 37°C and 5% CO2 in a humidified incubator, 10 μL of the CCK-8 solution was added to each well. The plates were then incubated for another 2 h at 37°C. Absorbance was measured at 450 nm using a microplate reader (Multiskan FC; Thermo Fisher Scientific Inc., Waltham, MA, USA).
Cell Proliferation
Cell viability was measured in A549 cells exposed to PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) or various concentrations of PtNPs (2.5–40 µM) or C6-cer (2.5–40 µM) or CSP (2.5–40 µM) or GW4869 (2.5–40 µM). After incubation for 24 h at 37°C and 5% CO2 in a humidified incubator and then cell proliferation was measured according to the manufacturer’s instructions.
Cellular Uptake of PtNPs and Quantification of Pt Ions
Cellular uptake of PtNPs was determined as described previously.42 Pt ion concentration was determined by inductively coupled plasma mass spectrometry (ICP-MS) ICP-MS was performed to measure the Pt contents in the PtNPs exposed cells. Cells were treated with 10 µM of PtNPs for 24 h, and then cells were collected by Trypsin/EDTA treatment. Silver ions in the cells by ICP-MS. Briefly, 1 x 106 A549 cells were cultured on the each 6-well cell culture plate, after treating PtNPs, the cells were collected by trypsin/EDTA, and silver ions were measured.
Membrane Integrity
We evaluated the membrane integrity of A549 cells using the LDH Cytotoxicity Detection Kit. Briefly, the cells were exposed to PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) for 24 h. After 3 h of incubation under standard conditions, the optical density of the final solution was determined at a wavelength of 490 nm, using a microplate reader.
Estimation of ROS
ROS production was estimated as described previously.41 A549 cells were exposed to PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) and the cells were incubated for 24 h. The cells were supplemented with 20 μM of DCFH2-DA and incubation continued for 30 min at 37°C. The fluorescence intensity was determined using a Gemini EM spectrofluorometer.
Estimation of Caspase-3 Activity
Caspase-3 activity was measured as described previously.43 The cells were treated with PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) for 24 h, and subsequently the activity of caspase-3 was measured using a kit from Sigma-Aldrich Co., according to the manufacturer’s instructions.
Exosome Isolation
The cells were treated with PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM) for 24 h. Exosomes were prepared from culture supernatants of A549 cells by differential centrifugation according to a previously described method.44,45 Exosomes were also isolated and purified using ExoQuick (EXOQ5TM-1, System Biosciences, Palo Alto, CA, USA) according to the manufacturer’s instructions.
Scanning Electron Microscopy (SEM) and Transmission Electron Microscopy (TEM)
The SEM analysis was carried out according to a previously described method.46 Briefly, the pellets containing exosomes were vortexed and resuspended in 0.21 mL of PBS. SEM images were obtained using a field emission scanning electron microscope (HITACHI SU8010, Hitachi Corporation, Japan). TEM analysis were performed according to a previously described method.47
Determination of Total Protein Concentration of Exosomes
Total protein concentration of exosomes was determined using the bicinchoninic acid (BCA) assay kit (Thermo Scientific, Waltham, MA, USA) according to the manufacturer’s instructions. In addition, we followed a previously described protocol.47
Quantitation of Exosomes Using a Colorimetric Assay
Exosome concentration was estimated using the EXOCETTM assay (System Biosciences), performed as described previously.48 Quantification of exosomes by fluorescence polarization (FP) was performed as described previously.49
Particle Size and Concentration Analyses
The particle size and concentration were measured using the Nanoparticle Tracking Analysis System 300 (NTA300, Malvern Instruments, UK). The exosome size/diameter was estimated by dynamic light scattering (DLS) as described previously.50
Measurement of Acetylcholinesterase (AChE) Activity
AChE activity was determined according to a previously described method.51 To measure the quantity of exosomes in culture supernatants, cells were treated with PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM) or GW4869 (20 µM).
Sphingomyelinase Activity Assay
The sphingomyelinase activity assay was performed as described previously,52 using a kit and following the manufacturer’s protocol. The Amplex Red sphingomyelinase assay kit (Molecular Probes Inc., Eugene, OR, USA) was used to quantify neutral sphingomyelinase activity.
Quantitative Reverse Transcription-Polymerase Chain Reaction (qRT-PCR)
Total RNA was isolated using the TRI reagent protocol, and cDNA synthesis was performed using oligo (dT) primers. qRT-PCR was performed using the SYBR Green Master Mix (Life Technologies) according to the manufacturer’s instructions on Applied Biosystems 7300 Sequence Detection System. The expression levels of target genes were quantified and normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH). The list of primers used in this study mentioned in supplementary Table 1.
ELISA Assay
The expression of TSG101, CD9, CD63, and CD81 was determined with ELISA, as described previously47 and also we followed according to manufacturer instructions (Thermo Fisher Scientific and system biosciences).
Statistical Analysis
All experiments were repeated at least thrice (triplicate). Data were analyzed with the Student’s t-test or ANOVA, as required. To assess the differences between two groups, Student’s t-test (paired or unpaired, 2-tailed) was performed, as applicable. Differences were considered as significant when the p value was less than 0.05 (p < 0.05).
Results and Discussion
Synthesis and Characterization of PtNPs Using Lutein
Bio-reduction of Pt ions to PtNPs was carried out using a biomolecule called lutein as a reducing and stabilizing agent. PtNPs synthesis was monitored by tracking the color change from pale yellow to brown. Upon completion of the reaction, the color changed from brown to black. The conversion rate was rapid at high temperatures, such as 100°C. The conversion of Pt (IV) into Pt (0) indicates formation of PtNPs.39,53,54 UV/VIS spectra exhibited a peak at 300 nm, which is attributed to the excitation of surface plasmon vibrations, indicating the synthesis of PtNPs (Figure 1A). The disappearance of the original peak corresponding to H2PtCl6 and appearance of typical peak at 300 nm indicated the reduction of Pt (IV) ions into Pt (0) nanoparticles.
The crystal nature of PtNPs corresponding XRD peaks were shown in Figure 1B. The XRD spectrum of the prepared PtNPs exhibited five different intense peaks at 2θ = 39.9°, 46.7° and 67.7° corresponding to the indexed planes (111), (200) and (220) respectively, which were consistent with the fcc structure of platinum.55 The crystallographic plane of platinum is face-centered cubic (fcc) (JCPDS #87-0644). The broadening of the Bragg peaks indicated the purity and formation of nanoparticles. Our results are in good agreement with those of biologically synthesized PtNPs using Ocimum sanctum53 and are consistent with our previous reports.39,41
The FTIR spectrum of lutein-functionalized PtNPs showed strong peaks at 3,370 cm-1, which is attributed to the hydroxyl group in polyphenolic compounds (Figure 1C). In addition, other characteristic peaks were observed in the spectrum of PtNPs, in agreement with our previous reports.39,41 Sharp peaks at 2,140, 1,640, and 1,025 cm-1, associated with C–H stretching, N–O stretching in the nitro group, and C–C stretching, respectively, have been reported.39,53 The presence of almost all the characteristic peaks of PtNPs indicated the stabilization of nanoparticles by lutein due to the coating of biomolecules. Overall, the data suggest that lutein plays a key role in reducing metal ions to their corresponding metallic nanoparticles.
TGA was performed to determine the weight loss between lutein and lutein functionalized PtNPs. As shown in supplementary Figure 1, The TGA spectra of the PtNPs show a weight loss of 10% between 100°C and 200°C due to the loss of adsorbed moisture. The weight loss of 20% between 400°C and 500°C and 40–70% weight loss was observed between 500°C and 800°C, it shows the stability of lutein functionalized PtNPs. The lutein curve exhibits an 80%–90% weight loss between 100°C and 400°C, which is the characteristic decomposition temperature of the organic molecules. While comparing the thermogram of PtNPs-lutein with those of lutein provides the formulation’s composition and chemical modifications. The weight loss of PtNPs-lutein significantly lower than lutein, which suggests reduction of the lutein during the functionalization. The lower percentage of weight loss between 200°C and 400°C indicates the presence of the PtNPs-lutein complex, and implies that PtNPs-lutein contains at 20% by weight.
The size and shape of the PtNPs were evaluated using scanning electron microscopy (SEM). SEM images of the PtNPs prepared using lutein are shown in Figure 1D. The size of the platinum nanoparticles was approximately 40 nm. The morphology of the synthesized particles showed a flower-like structure, which is similar to that in previously published reports.39,41
We next determined the size of the PtNPs, using DLS. DLS is a simple and feasible technique for understanding the diffusion behavior, which is affected by the size and shape of macromolecules.56 The size of the particles was found to be 50 nm (Figure 1E), slightly larger than the size of the particles analyzed with SEM. The size of the particles is slightly larger than that obtained with other biomolecule-assisted synthesis of PtNPs, such as Ocimum sanctum-, apigenin-, and beta carotene-functionalized PtNPs.39,41
Furthermore, we determined the size, shape, and morphologies of the lutein-reduced nanoparticles using TEM analysis. The TEM micrograph image showed various morphologies, such as spherical, triangular, cubic, oval, hexagonal, and rod-shaped (Figure 1F). The lutein-functionalized particles were not agglomerated and were well scattered, indicating that PtNPs were stabilized in the matrix of the biomaterial. The average size of the particles was found to be 40 nm. Pal et al57 reported that PtNPs obtained by microwave irradiation-mediated synthesis were spherical in shape. We have previously reported obtaining similar shaped platinum nanoparticles, which were synthesized using apigenin.39 Conversely, Sheny et al58 reported the synthesis of crystalline and irregular rod-shaped nanoparticles using dried leaf powder of Anacardium occidentale. We also obtained similar shaped platinum nanoparticles, which were synthesized using beta-carotene.41 It is well known that the concentration of reducing agents plays a significant role in the size and morphology of metal nanoparticles. The size and morphology of the synthesized PtNPs can be correlated with the interactions between lutein and platinum atoms. The merits of these multifaceted shapes of nanoparticles are that they can easily enter into tissues and cells rapidly.59 The various shapes of nanoparticles play critical roles in biotechnology and biomedical applications, including therapeutic delivery processes, such as particle adhesion, distribution, and cell internalization.39,41
Effect of Serum on the Viability and Proliferation of A549 Cells
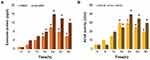
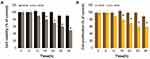
Understanding the cellular responses to serum starvation conditions is critical for biogenesis and secretion of exosomes. Therefore, we first investigated how these cells reacted to changes in culture media, by culturing them in DMEM containing 10% serum and Opti-MEM with low serum. Since A549 cells are normally grown in a medium supplemented with low serum, cell viability is low and rate of cell proliferation is generally slow in medium containing low serum concentration compared to those in medium containing 10% serum. Opti-MEM is a well-accepted reduced-serum medium used for transfection experiments. To understand the impact of Opti-MEM on cell viability and cell proliferation, A549 cells were grown in Opti-MEM for 36 h and cell viability and proliferation were monitored. As a control of optimal growth conditions, A549 cells were also grown in DMEM with 10% FBS. There was no remarkable change observed in cells cultured in DMEM for 24 h; however, after 24 h, a slight reduction in cell viability was observed, but it was not significant. Conversely, a significant change in cell viability was observed in Opti-MEM cultured cells compared to those cultured in DMEM. The cell viability started to decrease in Opti-MEM at 12 h and continuously decreased up to 36 h, indicating that it was significantly reduced in Opti-MEM compared to DMEM (Figure 2A). A significant effect of Opti-MEM on cell viability was observed between 18 and 36 h.
Next, we examined the effect of serum concentration on the proliferation of A549 cells. The cell proliferation assay results showed that the proliferation rate of cells growing in Opti-MEM was significantly lower throughout the experimental period from 12 to 36 h, than that of the cells grown in DMEM (Figure 2B). Serum starvation significantly influences the growth of cells and causes apoptosis-induced cell death in different human cell lines, including human neuroblastoma cells60 and human adenocarcinoma lung cancer cells.61 Serum starvation arrests the growth of A549 cells in the G1 phase without inducing apoptosis.62 Our findings suggested that cell death was increased after growing the cells in reduced-serum medium. As the use of low serum medium can reduce the undesired effects of serum in cell culture experiments and also determine the specific effect of PtNPs on exosome biogenesis and secretion, we selected the reduced-serum medium for further experiments.
Effect of Serum on Exosome Protein Expression and AChE Activity
Before studying PtNPs-induced exosome secretion in detail, selection of media is crucial, as it plays a critical role in different cellular signaling events. Thus, we next investigated the effect of DMEM and Opti-MEM on exosome protein expression and AChE activity. A549 cells were grown in DMEM containing 10% serum and Opti-MEM for 36 h, and exosome protein expression and AChE activity were determined with colorimetric method, after harvesting the cells. In addition, the cells were grown in DMEM supplemented with 10% FBS as a control, to determine the normal expression patterns of proteins in A549 cells. Expression of exosome proteins in DMEM-cultured cells increased until 24 h, and high expression levels were observed at 24 h, which subsequently started to decrease. This suggested that the expression of exosome proteins was time-dependent (Figure 3A). In contrast, cells treated with Opti-MEM showed higher expression levels of exosome proteins than those treated with DMEM. Although the pattern of expression of exosome proteins seemed to be similar, the protein content was higher in Opti-MEM-grown cells than in DMEM-cultured cells. Rashid and Coombs61 reported serum-dependent effect of HSPA5 expression in A549 cells, and found that HSPA5 expression was significantly increased after 2 days of incubation in Opti-MEM. Li et al60 compared the effect of pre-spun medium (high serum) and Opti-MEM (low serum) on the concentration of exosome particles in human neuroblastoma cells. The findings from this study suggested that substantially increased number of particles were released in Opti-MEM than in pre-spun medium. Our findings also confirmed that the total protein concentration of exosomes was higher in Opti-MEM than in DMEM, which is suitable for producing increased number of exosomes in A549 cells.
Next, we examined AChE activity, which is a parameter used to indirectly determine the quantity of exosomes in culture supernatants, since acetylcholinesterase are localized in the membrane of exosomes.51,63 The results indicated that AChE activity increased until 24 h under both conditions (Figure 3B). Subsequently, the AChE activity began to decrease at 30 h in both DMEM- and Opti-MEM-cultured cells. However, AChE activity was significantly higher in Opti-MEM throughout the incubation period than that in DMEM. Essandoh et al51 reported that a significantly high AChE activity was observed in the supernatants of LPS-treated macrophages compared to that in the supernatants of the controls. Similarly, the protein concentration of exosomes isolated from LPS-treated macrophages (LPS exosomes) was significantly higher than the protein quantities in ‘non-LPS exosomes’. Our data showed that cells cultured in Opti-MEM expressed more enzymes, indicating that enzyme activity is directly proportional to the quantity of exosomes. Pérez-Aguilar et al64 demonstrated a clear relationship between AChE expression and cell cycle progression. Inhibition of AChE activity led to an increase in cell proliferation, which was associated with the downregulation of p27 and cyclins in hepatocellular carcinoma cells. Therefore, Opti-MEM is more suitable than DMEM to produce more yield of exosomes.
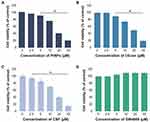
Dose-Dependent Effect of PtNPs, C6-Cer, CSP and GW4869 on the Viability of A549 Cells
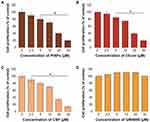
To determine the effects of PtNPs, C6-cer, CSP and GW4869 on cell viability, A549 cells were treated with PtNPs or C6-cer or CSP or GW4869 at increasing concentrations (0, 2.5, 5, 10, 20, and 40 μmol/L) for 24 h and then cell viability was assessed using the CCK-8 assay. Herein, CSP and C6-cer served as a positive control and GW4869 served as a negative control. As shown in Figure 4, the viability of A549 cells was dose-dependently inhibited by PtNPs, C6-cer and CSP (Figure 4A–C). In contrast, the cell viability rate did not change after GW4869 treatment (Figure 4D). Consistent with this finding, Cheng et al65 have previously reported that the cell proliferation rate was increased after GW4869 treatment. The inhibition of cell viability and proliferation was significant only when the concentration of GW4869 was 40 μmol/L or higher. Previously, several studies have reported that PtNPs dose-dependently inhibit the viability of several types of cancer cells, including human ovarian cancer cells,66 osteosarcoma cells40 and human neuroblastoma cancer cells.41 C6-cer inhibited the viability of human multiple myeloma cells in a dose-dependent manner.65 CSP dose-dependently inhibited cell viability of A549 cells.67 In our study, we observed that GW4869 did not alter both viability as well as proliferation of A549 cells. PtNPs were functionalized with lutein, the lutein bound with PtNPs could have effect on cell viability, and therefore we analyzed dose-dependent effect of lutein on cell viability of A549 cells. The cells were incubated with various concentrations of lutein (0.2 −1.0 mg/mL) for 24 h. The results show that there is no significant effect on cell viability of A549 cells at tested concentrations (supplementary Figure 2). Hence, the concentration (1mg/mL) taken for synthesis of PtNPs is non-toxic to the cells and also there is no further cytotoxic effects to the cells, the effect to the cells is due to PtNPs.
Dose-Dependent Effect of PtNPs, C6-Cer, CSP and GW4869 on the Proliferation of A549 Cells
To determine the effect of PtNPs, C6-cer, CSP and GW4869 on cell proliferation, A549 cells were treated with PtNPs or C6-cer, or CSP or GW4869 at increasing concentrations (0, 2.5, 5, 10, 20, and 40 μmol/L) for 24 h and then the cell proliferation rate was assessed using the BrdU incorporation assay. As shown in Figure 5A–C, the rate of proliferation of A549 cells was dose-dependently inhibited by PtNPs, C6-cer and CSP. We found that PtNPs dose-dependently (2.5–40 μmol/L) inhibited cell proliferation (10%–80%). Similar trend was observed with both CSP and C6-cer treated cells. In contrast, the cell proliferation rate did not change after GW4869 treatment (Figure 5D). Previous studies have demonstrated that PtNPs dose-dependently inhibit the proliferation of several types of cancer cells, including human ovarian cancer cells,66 osteosarcoma cells,40 and human neuroblastoma cancer cells.41 C6-cer inhibited the viability of human multiple myeloma cells in a dose-dependent manner.65 We found that C6-cer dose-dependently (2.5–40 μmol/L) inhibited cell proliferation (15%–80%), whereas GW4869 did not affect cell proliferation at the same concentration range. Similarly, CSP dose-dependently inhibits proliferation of A549 cells. These results are consistent with previous reports, which showed the effects of C6-cer and GW4869 on the proliferation of breast cancer cells.68,69 Interestingly, GW4869 increased the proliferation of human multiple myeloma cells.65 Taken together, data from the above experiments suggest that PtNPs, C6-cer and CSP potentially inhibit cell viability and proliferation in a dose-dependent manner.
Cellular Uptake of PtNPs and Quantification of Pt Ions
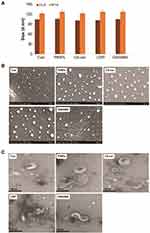
Cellular uptake studies were carried out to determine the fate of PtNPs in terms of internalization and cell attachment. The A549 cells were exposed to the PtNPs dose (10 μM) for 24 h, fixed, and prepared for TEM analysis. The A549 controls appeared normal (Figure 6A). After 24 h exposure, PtNPs were internalized and it induces cellular morphological changes which suggest that oxidative stress induced autophagy had occurred in the treated cells. PtNPs were localized within membrane-bound cytoplasmic vacuoles and in enlarged lysosomes (Figure 6B).
Next, we quantified the internalized PtNPs into the cells by ICP-MS. ICP-MS is a conventional approach to quantify NPs uptake. Generally, cellular uptake of NPs is size dependent. The amount of Pt associated with the A549 cells was determined by ICP-MS. After incubation with 10 µM PtNPs for 24 h, the Pt–ion concentration after 24 h of incubation with PtNPs was about 100 nM/106 cells, whereas the cells untreated shows there is no Pt ion concentration (Figure 6C).
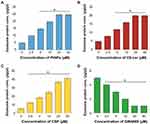
Dose-Dependent Effect of PtNPs, C6-Cer, CSP and GW4869 on the Expression of Exosome Proteins
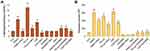
To validate the data obtained from cell viability and proliferation assays of A549 cells and to determine whether PtNPs can stimulate the secretion of exosomes in Opti-MEM, the cells were treated with various concentrations of PtNPs or C6-cer or CSP (0, 2.5, 5, 10, 20, and 40 μmol/L) for 24 h, and then total protein concentration of PtNPs, C6-cer and CSP-treated samples was determined. Treatment with PtNPs, C6-cer and CSP at the tested concentrations increased exosome release significantly in a dose-dependent manner (Figure 7A–C), which could possibly be due to the oxidative stress induced by PtNPs, C6-Cer and CSP. Conversely, the cells treated with GW4869 significantly decreased exosome production in a dose-dependent manner (Figure 7D). Furthermore, increasing concentrations of PtNPs or C6-cer or CSP increased exosome protein concentration, thereby implying that at higher concentrations of PtNPs or C6-cer or CSP, ROS production was significantly higher than that at lower concentrations. Therefore, at least a minimum level of oxidative stress is required to induce biogenesis and release of exosomes. Emam et al70 reported the enhancement of exosome yield by incubating various types of liposomes, including neutral, cationic-bare, or PEGylated liposomes, with four different tumor cell lines, including Colon 26 (C26) murine colorectal cancer cell line, B16BL6 murine melanoma cell line, MKN45 human gastric cancer cell line, and DLD-1 human colorectal cancer cell line, and subsequently estimating the total protein concentration of isolated exosomes. The results showed that both neutral and cationic-bare liposomes enhanced exosome secretion in a dose-dependent manner. Among the tested liposomes, fluid cationic liposomes exhibited the strongest stimulation. Unexpectedly, the PEGylation of bare liposomes diminished exosome secretion. Collectively, our findings suggest that increasing the concentration of PtNPs potentially induces exosome release in a dose-dependent manner. Based on all the above optimization parameters, in order to reduce the undesired effects of serum in cell culture experiments and minimize the toxicity of high concentration of PtNPs, C6-cer and CSP, we selected Opti-MEM media, 24-h incubation time, and concentrations of PtNPs, C6-cer, CSP and GW4869 as 10 µM, 10 µM, 10 µM and 20 µM, respectively, for further experiments.
PtNPs Enhance Exosome Protein Concentration and Exosome Counts
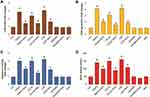
To determine the effect of PtNPs on exosome protein concentration and exosome counts, cells were treated with concentration of PtNPs (10 µM) or C6-cer (10 µM) or CSP or GW4869 (20 µM) for 24 h. Then, we isolated the exosomes from PtNPs-, C6-cer-, CSP and GW4869-treated cells. The total protein concentration of exosomes was determined. The quantity of exosomes was indirectly determined by measuring the protein concentration of the isolated exosomes. Among the four tested compounds, PtNPs, C6-cer and CSP significantly induced exosome protein concentration, whereas GW4869 significantly lowered the level of protein concentration (Figure 8A). These data suggest that the number of exosomes secreted by PtNPs is directly related to concentration of protein and it is slightly higher to that secreted by C6-cer-treated cells. CSP treated cells show equivalent to PtNPs treated cells. It is well known that ceramide plays a critical role in exosome secretion and function in cancer cells. The results confirmed that PtNPs, C6-cer and CSP significantly increased protein concentrations. To further demonstrate the consistency and interpretation of results, we determined the quantity of exosomes using various methods, including fluorescence polarization (Figure 8B), NanoTrack (Figure 8C), and EXOCET (Figure 8D). Analysis using all these techniques consistently indicated that there was a significant increase in exosome number in the PtNPs, C6-cer and CSP treated cells compared to that in the control cells.
All these methods confirmed that the PtNPs-treated cultures produced slightly higher number of exosomes than C6-cer-treated cells, which is due to high level of oxidative stress produced by PtNPs. Interestingly, GW4869-treated cells showed significantly decreased exosome counts compared to the control, PtNPs-, or C6-cer or CSP-treated cells. Sinha et al71 reported that the actin cytoskeletal regulatory protein cortactin promotes exosome secretion. Among all the tested techniques, such as fluorescence polarization, NanoTrack and EXOCET, for quantification of exosomes, EXOCET displayed more sensitivity and exhibited a high number of exosome counts, and seems to be a better method compared to other techniques. Previous studies have reported that various types of stresses, such as hypoxia,27 acidosis,72 thermal and oxidative stress,31 radiation,73 and cytotoxic drugs74–76 release a higher amount of exosomes, including tumor cells. Our findings consistent with previous report suggest that CSP induces extracellular vesicles in human ovarian cancer cells.77 Previously several studies were reported increased level of EVs by various types of stress such as radiation, oxidative stress, ionizing radiation, senescence and CSP29,77–80 Li et al81 found that EV markers, such as PDC6I/Alix, CD9, and TSG101, were consistently detected a day earlier in Opti-MEM-cultured cells than in serum-rich medium. In addition, the expression of calnexin was observed in serum-free medium earlier than in serum-rich medium. Phuyal et al82 reported the involvement of glycosphingolipids and flotillins in the release of exosomes from PC-3 cells and indicated that the role of ceramide in exosome formation may be cell-dependent. Altogether, PtNPs-induced oxidative stress could be the possible mechanism for the increased release of exosomes in PtNPs-treated cells.
Size, Size Distribution, Zeta Potential and Surface Morphology Analysis of Exosomes
After the confirmation of exosome isolation by determining protein concentration and exosome counts, we set out to determine the size of the particles using the NTA and DLS. Both techniques are simple, feasible, and dependable, and can detect the overall nanoparticle concentration and size using the Stokes–Einstein equation.83,84 The size distribution of the isolated exosomes was determined using the NTA and DLS, as shown in Figure 9A.
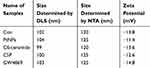
The exosomes released from the control, PtNPs-, C6-cer, CSP and GW4869-treated cells exhibited average sizes of 120, 125, 120, 120 and 125 nm, respectively, as ascertained with NTA, whereas the exosomes released from control, PtNPs-, C6-cer-, CSP and GW4869-treated cells exhibited average sizes of 102, 104, 99, 100 and 103 nm, respectively, as determined by DLS. The sizes of majority of the particles fit between 100 −120 nm (Figure 9A). Interestingly, the same pattern was observed in all the samples tested exclusively with PtNPs-, C6-cer-, CSP and GW4869-treated A549 cells; there was no significant size change between the treated samples and the control. The NTA measures slightly larger mean sizes of exosomes than the DLS. However, the sizes obtained using both NTA and DLS were slightly larger than those obtained using TEM, because both techniques measure the size in solution. The sizes of the particles are consistent with a previous report suggesting that CSP-treated A549 cells release round vesicles measuring 30–100 nm in diameter.85 However, the sizes of exosomes were larger than those determined by SEM and TEM. The reason for the larger sizes of the particles is that the NTA and DLS measure Brownian motion. Serrano-Pertierra et al suggested that EVs isolated from plasma contain a range of particle populations with different average sizes, and NTA measurement indicated larger mean sizes of EVs than those obtained using DLS.81 As shown in Table 1 A549 cell-derived exosomes exhibited size distribution profile with a peak at 102 nm for control, at 104 nm for PtNPs treated cells, at 99 nm for C6-ceramide treated cells, at 100 nm for CSP treated cells and at 103 nm for GW4869 treated cells. Further, to obtain information about the stability of the particles in terms of dispersion, aggregation or flocculation, we measured the zeta potential of exosomes isolated from the cells treated with PtNPs, C6-ceramide, CSP and GW4869. The zeta potential ranged from 14.8 to 11.4 mV (Table 1), suggesting a good and significant nanoparticle stability.
Exosomes are released by cells and can be distinguished based on their morphology and size distribution. Electron microscopy (EM) is a vital technique to characterize the morphology of exosomes, which are smaller than 300 nm.86 SEM images showed that PtNPs, C6-cer and CSP-treated cells exhibited a round and uniform morphology, spherical with unimodal size distribution that was consistent (Figure 9B). The size of exosomes released from control cells was mostly found to be 50–100 nm. The cells treated with PtNPs or C6-cer or CSP released an increased number of exosomes compared to the control cells under the same conditions, with sizes ranging from 50 to 100 nm. In contrast, cells treated with GW4869 released significantly reduced number of exosomes compared to either PtNPs- or C6-cer-or CSP treated cells. We observed that the SEM images of exosomes derived from all treated samples were consistently spheroidal, similar to exosomes isolated from human adenocarcinoma cells.85 Based on SEM observations, PtNPs release significantly increased numbers of exosomes compared to the control. Interestingly, GW4869 significantly inhibited the release of exosomes from A549 cells.
TEM was performed to determine the size as well as for the qualitative assessment of the morphology of isolated exosomes. TEM images showed a typical cup-shaped morphology of exosomes, whereas SEM images showed exosomes as round spheroids. TEM analysis indicated that both PtNPs, C6-cer and CSP-treated A549 cells released exosomes that are spherical, membrane-encapsulated particles with a cup-shaped morphology, characteristic of exosomes (Figure 9C). The majority of exosomes released from PtNPs- or C6-cer-treated cells were in the size range of 80–100 nm. This finding was confirmed by the NTA, which showed similar size distribution profiles for PtNPs- or C6-cer or CSP-treated A549 cell-released exosomes, indicating that PtNPs, C6-cer and CSP can produce exosomes that exhibit homogenous size distribution. Altogether, these results suggested that, while SEM and TEM showed similarity regarding the size of the exosomes, they slightly differed with respect to the morphology of exosomes. Exosome particle counts from TEM were found to be lower than those measured with the NTA, which is due to the incomplete adhesion of vesicles to the surface of EM grids.87
mRNA and Protein Expression of TSG101, CD9, CD63 and CD81
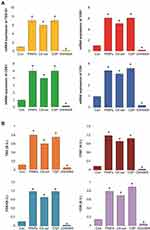
To corroborate the results obtained from protein quantification, exosome counts, and exosome quantification, we determined the mRNA and protein expression levels of TSG101, CD9, CD63, and CD81. A549 cells were treated with PtNPs (10 µM), C6-cer (10 µM), CSP (10 µM) or GW4869 (20 µM) for 24 h and the expression of TSG101, CD9, CD63, and CD81 was analyzed with qRT-PCR (Figure 10A). The A549 cells treated with PtNPs or C6-cer or CSP exhibited significantly increased levels of exosomes than the untreated cells. These results suggest that the exosomes released from A549 cells also carry information at the mRNA level, which could be helpful in understanding cancer prevention and therapy.88 TSG101, CD63, CD81, and CD9 are typical markers for exosomes.89
We next measured the expression of TSG101 and tetraspanin (CD9, CD63, and CD81) in the exosomes released by the control as well as PtNPs-, C6-cer or CSP or GW4869-treated cells. The results showed increased expression of TSG101, CD9, CD63, and CD81 in the exosomes released from PtNPs- C6-cer and CSP treated A549 cells compared to that in the exosomes of untreated control cells (Figure 10B). The expression of CD9, CD63, and CD81 proteins confirmed the release of exosomes by A549 cells. These observations support that PtNPs potentially induce exosome secretion. Interestingly, high optical density was observed in the expression of all tested proteins, including TSG101, CD9, CD63, and CD81. Among the four proteins, CD81 and CD63 showed significantly greater optical density compared to that of TSG101 and CD9. The results indicated that ELISA-based approach offers an accessible, versatile, and flexible method for exosome characterization. Willms et al reported that proteomic analysis showed enrichment of the transmembrane tetraspanin proteins CD63, CD9, and CD81 in both low-density and high-density fractions of exosomes compared to microvesicles (MVs).90 Li et al81 found that EV markers, such as PDC6I/Alix, CD9, and TSG101, were consistently detected a day earlier in Opti-MEM than in pre-spun medium, and the expression of calnexin was significantly higher in Opti-MEM-cultured cells than in serum-rich medium-cultured cells.
NAC Inhibits Oxidative Stress-Induced Exosome Release
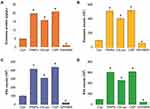
Oxidative stress is involved in nanoparticle-mediated cytotoxicity and apoptosis. We aimed to address the role of oxidative stress and the mechanism involved in the release of exosomes in A549 cells. To determine the effects of cytotoxicity, oxidative stress, apoptosis, and AChE activity on exosome release from A549 cells, the cells were treated with PtNPs (10 µM) or C6-cer (10 µM) or CSP (10 µM), GW4869 (20 µM) for 24 h, and subsequently LDH, ROS, caspase-9, and AChE activity were measured (Figure 11A–D). First, to test the possible toxicity of PtNPs, C6-cer, CSP and GW4869, we measured the levels of lactate dehydrogenase (LDH), a marker of cell injury. Our results showed that LDH levels were threefold higher in PtNPs- or C6-cer- or CSP-treated cells than in the untreated control (Figure 11A). Our findings are in agreement with a previous report, which showed that PtNPs increase the leakage of LDH and enhance toxicity in various types of cancer cells, including osteosarcoma cells40 and human neuroblastoma cancer cells.41 Conversely, no significant effect was observed in either the GW4869-treated cells or NAC-treated cells compared to the control. Interestingly, cells pre-treated with NAC showed that the LDH levels were equal to those of the control, indicating that NAC inhibits the PtNPs- or C6-cer-or CSP induced toxicity. Essandoh et al51 observed that LDH levels were similar in 0.005% DMSO, 10 μM GW4869, 20 μM GW4869, and non-treated cells. This suggests that 0.005% DMSO and GW4869 do not have toxic effects on macrophages. Similarly, we found that GW4869 treatment was not toxic to the cells.
Next, we determined the level of ROS in cells treated with PtNPs (10 µM), C6-cer (10 µM), CSP (10 µM), or GW4869 (20 µM) for 24 h, by measuring the fluorescence intensity. Our results showed that ROS levels were fourfold higher in PtNPs- or C6-cer-or CSP-treated cells than those in the untreated control (Figure 11B). Surprisingly, no significant difference was observed between either GW4869-treated cells or NAC-treated cells compared to the control. Interestingly, cells pre-treated with NAC showed similar ROS levels as those of the control, indicating that NAC inhibits the PtNPs- C6-cer and CSP-induced oxidative stress. For instance, NAC potentially mitigates AgNPs-induced ROS generation in male Leydig cells (TM3) and Sertoli cells (TM4).91 Carver and Yang92 reported that N-acetylcysteine amide (NACA) protects against H2O2 oxidative stress-induced microparticle release from human retinal pigment epithelial cells. NACA protects ARPE-19 cells against oxidative stress-induced cell death and decreases cellular GSH levels. A previous study demonstrated that NACA serves as a protective agent against oxidative stress under numerous pathological conditions both in vitro and in vivo, including in a mouse model of retinal degeneration.93
To determine whether NAC can protect cultured A549 cells against the PtNPs- or C6-cer-or CSP induced caspase-3 activity and prevent caspase-mediated cell death, we pre-treated A549 cells with NAC and analyzed cell apoptosis by estimating the caspase-3 activity. Pre-treatment of A549 cells with NAC significantly reduced the PtNPs- or C6-cer-or CSP induced caspase-3 activity. Our results showed that caspase-3 levels were threefold higher in PtNPs- or C6-cer-or CSP treated cells than in the untreated control (Figure 11C). Surprisingly, no significant difference was observed between either GW4869-treated cells or NAC-treated cells compared to the control. Interestingly, cells pre-treated with NAC showed that caspase-3 levels were equal to those in the control, indicating that NAC inhibits the PtNPs- or C6-cer-or CSP induced apoptosis. Previous studies have demonstrated that PtNPs or platinum nanocomposites induce apoptosis through the activation of caspase-3 in various types of cancer cells, including prostate cancer cells,39 human acute monocytic leukemia cells94 human hepatic cancer cells,95 and human neuroblastoma cancer cells.41 Ceramide triggers caspase-3 activation and induces apoptosis in A549 lung cancer cells.96 Cheng et al65 reported that C6 ceramide inhibited proliferation and promoted apoptosis in human multiple myeloma cells, which were associated with elevated caspase 3/9 and PARP cleavage.
Further, to confirm that PtNPs induced exosome quantity and release, acetylcholinesterase activity assay was performed. Acetylcholinesterase enzyme is enriched in exosomes. Our findings suggest that AChE activity levels were 3-fold higher in PtNPs- or C6-cer-or CSP treated cells than in the untreated control. Furthermore, NAC significantly suppressed the PtNPs, C6-cer and CSP induced AChE activity compared to the control (Figure 11D). No significant difference was observed in AChE activity between the exosomes derived from untreated A549 cells and the exosomes derived from A549 cells treated with either GW4869 or NAC. Altogether, these results demonstrate that NAC inhibits the PtNPs- or C6-cer-or CSP induced LDH activity, ROS generation, caspase-3 activity, and AChE activity, indicating that the PtNPs-induced oxidative stress could be a possible reason for increased exosome release in A549 cells. These findings were corroborated by a previous report showing that NAC inhibits ceramide-induced cell death by decreasing ROS generation from mitochondria.97 Exposure of bovine granulosa cells to H2O2 induced the accumulation of ROS, reduced mitochondrial activity, increased Nrf2 expression, altered cell cycle transitions, and induced cellular apoptosis. Conversely, granulosa cells exposed to oxidative stress released exosomes enriched with Nrf2 mRNA and candidate antioxidants, such as CAT, PRDX1, and TXN1.98 Our studies good agreement with previous report suggest that CSP induces expression of AChE during apoptosis in a time- and concentration-dependent manner in human breast cancer cells. CSP induced apoptosis play significant role in for the upregulation of AChE and p53.99 Collectively, these results suggest that NAC may function upstream of PtNPs and ceramide.
NAC Suppresses PtNPs-Induced Neutral Sphingomyelinase Activation
We next examined whether NAC inhibits neutral SMase activity directly or indirectly, which is mainly involved in ceramide formation. Ceramide plays a vital role in exosome secretion and function in cancer cells. Hence, we investigated the effects of NAC on neutral sphingomyelinase activity in the exosomes isolated from PtNPs- or C6-cer-or CSP treated A549 cells. When the cells were pre-incubated with NAC, neutral SMase activity was inhibited. Our findings suggest that sphingomyelinase activity levels were threefold, fivefold and 3 fold higher in PtNPs, C6-cer and CSP treated cells, respectively, than those in the untreated control. Furthermore, NAC significantly suppressed the PtNPs, C6-cer and CSP induced sphingomyelinase activity compared to the control. No significant difference was observed in sphingomyelinase activity between the exosomes derived from untreated A549 cells and those derived from A549 cells treated with either GW4869 or NAC (Figure 12A). Furthermore, inhibition of PtNPs, C6-cer and CSP activated the nSMase pathway, and GW4869 treatment revealed the involvement of the nSMase pathway in both exosome formation as well as release. Sphingomyelinase (SMase) is responsible for the conversion of SM to ceramide, and key signaling pathways of ceramide-induced apoptosis have been well documented.100,101 In this study, PtNPs could activate sphingomyelinase either directly or indirectly, via ceramide, which plays a major role in apoptosis. Cisplatin is known to induce apoptosis by targeting DNA. Studies have suggested that cisplatin induces apoptosis by signaling through plasma membrane lipid rafts that contain abundant sphingolipids, which play critical roles in maintaining the structural integrity of cell membranes and in modulating apoptosis via gene regulation and signal transduction.102–104 Maurmann et al105 reported that CSP mediated apoptosis pathway is associated with acid sphingomyelinase and FAS pro-apoptotic protein activation in ovarian cancer by alteration of various lipid profiles including increased level of phosphocholine, and glycerophosphocholine; elevated cellular energetics; and phosphocreatine and nucleoside triphosphate concentrations. Similarly, the possible mechanism of PtNPs could alter lipid profile of the membrane and it can influence n-sphingomyelinase activity. Collectively, these findings suggest that PtNPs may activate sphingomyelinase; however, further studies are required to elucidate the molecular mechanism of sphingomyelinase activation by PtNPs.
Next, to determine whether NAC inhibits the PtNPs- or C6-cer-or CSP induced oxidative stress-mediated release of exosomes through the activation of sphingomyelinase, the cells were pre-treated with either NAC or GW4869 and exosome counts were analyzed. As expected, PtNPs- or C6-cer- or CSP treated cells induced exosome release. However, pre-treatment of cells with NAC in the presence of either PtNPs or C6-cer or CSP reduced the level of exosome counts compared to those in the untreated control. Furthermore, the inhibition of sphingomyelinase activity by NAC significantly suppressed the PtNPs- or C6-cer- or CSP mediated exosome release compared to the control. No significant difference was observed in exosome counts between the control and A549 cells treated with either GW4869 or NAC (Figure 12B). GW4869 is a specific inhibitor of nSMases that inhibits exosome release from cells.106 Furthermore, inhibition of the nSMase pathway using GW4869, which is activated by PtNPs or C6-cer or CSP, released reduced number of exosomes, which clearly indicates that exosome formation and release is mediated by sphingomyelinase (SMase) activation. Gills et al107 reported that exogenous ceramide increases nanovesicle release by stimulating the sphingomyelinase activity and generating ceramide. Interestingly, NAC inhibited PtNPs- or C6-cer- or CSP mediated exosome release. Altogether, these findings suggest that PtNPs induce exosome biogenesis and release by generating oxidative stress and activating the ceramide pathway.
Conclusions
Exosomes are nanovesicles, with sizes ranging between 30 and 150 nm in diameter that are secreted by numerous cell types, including cancer cells, and play a vital role in intercellular communication between the cells. A limitation in exosome application is limited secretion from cells, which is a major bottleneck for efficient exosome production and application. Here, we examined the ability of PtNPs in enhancing exosome secretion from human adenocarcinoma cells (A549). Treatment of A549 cells with PtNPs significantly increased exosome production by sixfold, which was achieved by altering cell viability, proliferation, oxidative stress, and apoptosis. The PtNPs-induced exosomes exhibited sizes ranging between 90 and 100 nm. The size, shape, and quality of exosomes were characterized with various analytical techniques, including dynamic light scattering, scanning electron microscopy, and transmission electron microscopy. The quantity of exosomes was determined with fluorescence polarization, Nanotrack analyzer, and EXOCET. The results from these studies indicate that PtNPs promote exosome secretion from cancer cells by modulating the oxidative stress. The increased level of exosome release was confirmed by adapting various methodologies, including examining the concentration of exosome proteins, numbers, size, and shape and expression levels of TSG101, CD63, CD81, and CD9. The expression of exosome markers was significantly elevated in the presence of PtNPs or C6-cer. In contrast, cells treated with GW4869 remarkably decreased the expression levels of genes and proteins of exosome markers. Our data suggest that the modulation of oxidative stress by PtNPs is a critical control point for exosome secretion. In addition, we describe a novel connection between the ceramide pathway and exosome biogenesis, where PtNPs-induced exosome release was inhibited by GW4869, a specific inhibitor of nSMases, which suppressed exosome release from A549 cells. PtNPs stimulation may be a useful strategy to increase exosome yield, by promoting the secretion of exosomes. Importantly, this is the first study, to our knowledge, showing that PtNPs stimulate exosome biogenesis by inducing oxidative stress and the ceramide pathway.
Acknowledgments
This study was supported by the KU-Research Professor Program of Konkuk University.
Funding
This work was supported by a grant from the Science Research Center (2015R1A5A1009701) of the National Research Foundation of Korea.
Disclosure
The authors declare no conflict of interest.
References
1. Balkwill F, Charles KA, Mantovani A. Smoldering and polarized inflammation in the initiation and promotion of malignant disease. Cancer Cell. 2005;7(3):211–217. doi:10.1016/j.ccr.2005.02.013
2. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
3. Zhou Y, Li Y, Zhou T, Zheng J, Li S, Li HB. Dietary natural products for prevention and treatment of liver cancer. Nutrients. 2016;8(3):156. doi:10.3390/nu8030156
4. Srivastava A, Amreddy N, Razaq M, et al. Exosomes as theranostics for lung cancer. Adv Cancer Res. 2018;139:1–33. doi:10.1016/bs.acr.2018.04.001
5. Vanni I, Alama A, Grossi F, Dal Bello MG, Coco S. Exosomes: a new horizon in lung cancer. Drug Discov Today. 2017;22(6):927–936. doi:10.1016/j.drudis.2017.03.004
6. Li XQ, Liu JT, Fan LL, et al. Exosomes derived from gefitinib-treated EGFR-mutant lung cancer cells alter cisplatin sensitivity via up-regulating autophagy. Oncotarget. 2016;7(17):24585–24595. doi:10.18632/oncotarget.8358
7. Tian T, Wang Y, Wang H, Zhu Z, Xiao Z. Visualizing of the cellular uptake and intracellular trafficking of exosomes by live-cell microscopy. J Cell Biochem. 2010;111(2):488–496. doi:10.1002/jcb.22733
8. Henderson MC, Azorsa DO. The genomic and proteomic content of cancer cell-derived exosomes. Front Oncol. 2012;2:38. doi:10.3389/fonc.2012.00038
9. Taylor DD, Gercel-Taylor C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol Oncol. 2008;110(1):13–21. doi:10.1016/j.ygyno.2008.04.033
10. Wang Z, Chen JQ, Liu JL, Tian L. Exosomes in tumor microenvironment: novel transporters and biomarkers. J Transl Med. 2016;14(1):297. doi:10.1186/s12967-016-1056-9
11. Munson P, Shukla A. Exosomes: potential in cancer diagnosis and therapy. Medicines (Basel). 2015;2(4):310–327. doi:10.3390/medicines2040310
12. Kadota T, Yoshioka Y, Fujita Y, Kuwano K, Ochiya T. Extracellular vesicles in lung cancer-From bench to bedside. Semin Cell Dev Biol. 2017;67:39–47. doi:10.1016/j.semcdb.2017.03.001
13. Wu X, Zheng T, Zhang B. Exosomes in parkinson’s disease. Neurosci Bull. 2017;33(3):331–338. doi:10.1007/s12264-016-0092-z
14. Thakur BK, Zhang H, Becker A, et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 2014;24(6):766–769. doi:10.1038/cr.2014.44
15. Al-Nedawi K, Meehan B, Kerbel RS, Allison AC, Rak J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proc Natl Acad Sci U S A. 2009;106(10):3794–3799. doi:10.1073/pnas.0804543106
16. Hu C, Meiners S, Lukas C, Stathopoulos GT, Chen J. Role of exosomal microRNAs in lung cancer biology and clinical applications. Cell Prolif. 2020;53(6):e12828. doi:10.1111/cpr.12828
17. Fabbri M, Paone A, Calore F, et al. MicroRNAs bind to Toll-like receptors to induce prometastatic inflammatory response. Proc Natl Acad Sci U S A. 2012;109(31):E2110–2116. doi:10.1073/pnas.1209414109
18. Zhang Y, Liu Y, Liu H, Tang WH. Exosomes: biogenesis, biologic function and clinical potential. Cell Biosci. 2019;9:19. doi:10.1186/s13578-019-0282-2
19. Pascucci L, Coccè V, Bonomi A, et al. Paclitaxel is incorporated by mesenchymal stromal cells and released in exosomes that inhibit in vitro tumor growth: a new approach for drug delivery. J Control Release. 2014;192:262–270. doi:10.1016/j.jconrel.2014.07.042
20. Haney MJ, Klyachko NL, Zhao Y, et al. Exosomes as drug delivery vehicles for Parkinson’s disease therapy. J Control Release. 2015;207:18–30. doi:10.1016/j.jconrel.2015.03.033
21. Minciacchi VR, You S, Spinelli C, et al. Large oncosomes contain distinct protein cargo and represent a separate functional class of tumor-derived extracellular vesicles. Oncotarget. 2015;6(13):11327–11341. doi:10.18632/oncotarget.3598
22. Wozniak DJ, Kajdacsy-Balla A, Macias V, et al. PTEN is a protein phosphatase that targets active PTK6 and inhibits PTK6 oncogenic signaling in prostate cancer. Nat Commun. 2017;8(1):1508. doi:10.1038/s41467-017-01574-5
23. Cesi G, Philippidou D, Kozar I, et al. A new ALK isoform transported by extracellular vesicles confers drug resistance to melanoma cells. Mol Cancer. 2018;17(1):145. doi:10.1186/s12943-018-0886-x
24. Gudbergsson JM, Johnsen KB, Skov MN, Duroux M. Systematic review of factors influencing extracellular vesicle yield from cell cultures. Cytotechnology. 2016;68(4):579–592. doi:10.1007/s10616-015-9913-6
25. Savina A, Furlán M, Vidal M, Colombo MI. Exosome release is regulated by a calcium-dependent mechanism in K562 cells. J Biol Chem. 2003;278(22):20083–20090. doi:10.1074/jbc.M301642200
26. Svensson KJ, Kucharzewska P, Christianson HC, et al. Hypoxia triggers a proangiogenic pathway involving cancer cell microvesicles and PAR-2-mediated heparin-binding EGF signaling in endothelial cells. Proc Natl Acad Sci U S A. 2011;108(32):13147–13152. doi:10.1073/pnas.1104261108
27. King HW, Michael MZ, Gleadle JM. Hypoxic enhancement of exosome release by breast cancer cells. BMC Cancer. 2012;12:421. doi:10.1186/1471-2407-12-421
28. Koumangoye RB, Sakwe AM, Goodwin JS, Patel T, Ochieng J. Detachment of breast tumor cells induces rapid secretion of exosomes which subsequently mediate cellular adhesion and spreading. PLoS One. 2011;6(9):e24234. doi:10.1371/journal.pone.0024234
29. Atienzar-Aroca S, Flores-Bellver M, Serrano-Heras G, et al. Oxidative stress in retinal pigment epithelium cells increases exosome secretion and promotes angiogenesis in endothelial cells. J Cell Mol Med. 2016;20(8):1457–1466. doi:10.1111/jcmm.12834
30. Zhu L, Zang J, Liu B, et al. Oxidative stress-induced RAC autophagy can improve the HUVEC functions by releasing exosomes. J Cell Physiol. 2020;235(10):7392–7409. doi:10.1002/jcp.29641
31. Hedlund M, Nagaeva O, Kargl D, Baranov V, Mincheva-Nilsson L. Thermal- and oxidative stress causes enhanced release of NKG2D ligand-bearing immunosuppressive exosomes in leukemia/lymphoma T and B cells. PLoS One. 2011;6(2):e16899. doi:10.1371/journal.pone.0016899
32. Wang T, Gilkes DM, Takano N, et al. Hypoxia-inducible factors and RAB22A mediate formation of microvesicles that stimulate breast cancer invasion and metastasis. Proc Natl Acad Sci U S A. 2014;111(31):E3234–3242. doi:10.1073/pnas.1410041111
33. Vulpis E, Soriani A, Cerboni C, Santoni A, Zingoni A. Cancer exosomes as conveyors of stress-induced molecules: new players in the modulation of NK cell response. Int J Mol Sci. 2019;20(3):611. doi:10.3390/ijms20030611
34. Bandari SK, Purushothaman A, Ramani VC, et al. Chemotherapy induces secretion of exosomes loaded with heparanase that degrades extracellular matrix and impacts tumor and host cell behavior. Matrix Biol. 2018;65:104–118. doi:10.1016/j.matbio.2017.09.001
35. Gobbo J, Marcion G, Cordonnier M, et al. Restoring anticancer immune response by targeting tumor-derived exosomes with a HSP70 peptide aptamer. J Natl Cancer Inst. 2016;108(3):djv330. doi:10.1093/jnci/djv330
36. Gurunathan S, Han JW, Kwon DN, Kim JH. Enhanced antibacterial and anti-biofilm activities of silver nanoparticles against Gram-negative and Gram-positive bacteria. Nanoscale Res Lett. 2014;9(1):373. doi:10.1186/1556-276x-9-373
37. Yang C, Bromma K, Di Ciano-Oliveira C, Zafarana G, van Prooijen M, Chithrani DB. Gold nanoparticle mediated combined cancer therapy. Cancer Nanotechnol. 2018;9(1):4. doi:10.1186/s12645-018-0039-3
38. Jeyaraj M, Gurunathan S, Qasim M, Kang MH, Kim JH. A comprehensive review on the synthesis, characterization, and biomedical application of platinum nanoparticles. Nanomaterials (Basel). 2019;9(12):1719. doi:10.3390/nano9121719
39. Gurunathan S, Jeyaraj M, Kang MH, Kim JH. The effects of apigenin-biosynthesized ultra-small platinum nanoparticles on the human monocytic THP-1 cell line. Cells. 2019;8(5):444. doi:10.3390/cells8050444
40. Gurunathan S, Jeyaraj M, Kang MH, Kim JH. Tangeretin-assisted platinum nanoparticles enhance the apoptotic properties of doxorubicin: combination therapy for osteosarcoma treatment. Nanomaterials (Basel). 2019;9(8):1089. doi:10.3390/nano9081089
41. Gurunathan S, Jeyaraj M, Kang MH, Kim JH. Anticancer properties of platinum nanoparticles and retinoic acid: combination therapy for the treatment of human neuroblastoma cancer. Int J Mol Sci. 2020;21(18):6792. doi:10.3390/ijms21186792
42. Park JH, Gurunathan S, Choi Y-J, Han JW, Song H, Kim J-H. Silver nanoparticles suppresses brain-derived neurotrophic factor-induced cell survival in the human neuroblastoma cell line SH-SY5Y. J Ind Eng Chem. 2017;47:62–73. doi:10.1016/j.jiec.2016.11.015
43. Gurunathan S, Kang MH, Kim JH. Combination effect of silver nanoparticles and histone deacetylases inhibitor in human alveolar basal epithelial cells. Molecules. 2018;23(8):2046. doi:10.3390/molecules23082046
44. Gurunathan S, Marash M, Weinberger A, Gerst JE. t-SNARE phosphorylation regulates endocytosis in yeast. Mol Biol Cell. 2002;13(5):1594–1607. doi:10.1091/mbc.01-11-0541
45. Cheng Y, Dai X, Yang T, Zhang N, Liu Z, Jiang Y. Low long noncoding RNA growth arrest-specific transcript 5 expression in the exosomes of lung cancer cells promotes tumor angiogenesis. J Oncol. 2019;2019:2476175. doi:10.1155/2019/2476175
46. Wu M, Ouyang Y, Wang Z, et al. Isolation of exosomes from whole blood by integrating acoustics and microfluidics. Proc Natl Acad Sci U S A. 2017;114(40):10584–10589. doi:10.1073/pnas.1709210114
47. Lim J, Choi M, Lee H, et al. Direct isolation and characterization of circulating exosomes from biological samples using magnetic nanowires. J Nanobiotechnology. 2019;17(1):1. doi:10.1186/s12951-018-0433-3
48. Soares Martins T, Catita J, Martins Rosa I, et al. Exosome isolation from distinct biofluids using precipitation and column-based approaches. PLoS One. 2018;13(6):e0198820. doi:10.1371/journal.pone.0198820
49. Kalimuthu K, Kwon WY, Park KS. A simple approach for rapid and cost-effective quantification of extracellular vesicles using a fluorescence polarization technique. J Biol Eng. 2019;13:31. doi:10.1186/s13036-019-0160-9
50. Gennebäck N, Hellman U, Malm L, et al. Growth factor stimulation of cardiomyocytes induces changes in the transcriptional contents of secreted exosomes. J Extracell Vesicles. 2013;2(1):20167. doi:10.3402/jev.v2i0.20167
51. Essandoh K, Yang L, Wang X, et al. Blockade of exosome generation with GW4869 dampens the sepsis-induced inflammation and cardiac dysfunction. Biochim Biophys Acta. 2015;1852(11):2362–2371. doi:10.1016/j.bbadis.2015.08.010
52. Tabatadze N, Savonenko A, Song H, Bandaru VV, Chu M, Haughey NJ. Inhibition of neutral sphingomyelinase-2 perturbs brain sphingolipid balance and spatial memory in mice. J Neurosci Res. 2010;88(13):2940–2951. doi:10.1002/jnr.22438
53. Soundarrajan C, Sankari A, Dhandapani P, et al. Rapid biological synthesis of platinum nanoparticles using Ocimum sanctum for water electrolysis applications. Bioprocess Biosyst Eng. 2012;35(5):827–833. doi:10.1007/s00449-011-0666-0
54. Ghosh S, Nitnavare R, Dewle A, et al. Novel platinum-palladium bimetallic nanoparticles synthesized by Dioscorea bulbifera: anticancer and antioxidant activities. Int J Nanomedicine. 2015;10:7477–7490. doi:10.2147/ijn.S91579
55. Venu R, Ramulu TS, Anandakumar S, Rani VS, Kim CG. Bio-directed synthesis of platinum nanoparticles using aqueous honey solutions and their catalytic applications. Colloids Surf a Physicochem Eng Asp. 2011;384(1):733–738. doi:10.1016/j.colsurfa.2011.05.045
56. Stetefeld J, McKenna SA, Patel TR. Dynamic light scattering: a practical guide and applications in biomedical sciences. Biophys Rev. 2016;8(4):409–427. doi:10.1007/s12551-016-0218-6
57. Pal J, Deb MK, Deshmukh DK, Sen BK. Microwave-assisted synthesis of platinum nanoparticles and their catalytic degradation of methyl violet in aqueous solution. Appl Nanosci. 2014;4(1):61–65. doi:10.1007/s13204-012-0170-0
58. Sheny DS, Philip D, Mathew J. Synthesis of platinum nanoparticles using dried Anacardium occidentale leaf and its catalytic and thermal applications. Spectrochim Acta a Mol Biomol Spectrosc. 2013;114:267–271. doi:10.1016/j.saa.2013.05.028
59. Hoshyar N, Gray S, Han H, Bao G. The effect of nanoparticle size on in vivo pharmacokinetics and cellular interaction. Nanomedicine (Lond). 2016;11(6):673–692. doi:10.2217/nnm.16.5
60. Li J, Lee Y, Johansson HJ, et al. Serum-free culture alters the quantity and protein composition of neuroblastoma-derived extra- 1425 cellular vesicles. J Extracell Vesicles. 2015;4:26883. doi:10.3402/jev.v4.26883
61. Rashid MU, Coombs KM. Serum-reduced media impacts on cell viability and protein expression in human lung epithelial cells. J Cell Physiol. 2019;234(6):7718–7724. doi:10.1002/jcp.27890
62. Nakhjavani M, Nikounezhad N, Ashtarinezhad A, Shirazi FH. Human lung carcinoma reaction against metabolic serum deficiency stress. Iran J Pharm Res. 2016;15(4):817–823.
63. Savina A, Vidal M, Colombo MI. The exosome pathway in K562 cells is regulated by Rab11. J Cell Sci. 2002;115(Pt 12):2505–2515.
64. Pérez-Aguilar B, Vidal CJ, Palomec G, et al. Acetylcholinesterase is associated with a decrease in cell proliferation of hepatocellular carcinoma cells. Biochim Biophys Acta. 2015;1852(7):1380–1387. doi:10.1016/j.bbadis.2015.04.003
65. Cheng Q, Li X, Wang Y, Dong M, Zhan FH, Liu J. The ceramide pathway is involved in the survival, apoptosis and exosome functions of human multiple myeloma cells in vitro. Acta Pharmacol Sin. 2018;39(4):561–568. doi:10.1038/aps.2017.118
66. Alshatwi AA, Athinarayanan J, Vaiyapuri Subbarayan P. Green synthesis of platinum nanoparticles that induce cell death and G2/M-phase cell cycle arrest in human cervical cancer cells. J Mater Sci Mater Med. 2015;26(1):5330. doi:10.1007/s10856-014-5330-1
67. Zhang X, He C, Yan R, et al. HIF-1 dependent reversal of cisplatin resistance via anti-oxidative nano selenium for effective cancer therapy. Chem Eng J. 2020;380:122540. doi:10.1016/j.cej.2019.122540
68. Wu M, Harvey KA, Ruzmetov N, et al. Omega-3 polyunsaturated fatty acids attenuate breast cancer growth through activation of a neutral sphingomyelinase-mediated pathway. Int J Cancer. 2005;117(3):340–348. doi:10.1002/ijc.21238
69. Flowers M, Fabriás G, Delgado A, Casas J, Abad JL, Cabot MC. C6-Ceramide and targeted inhibition of acid ceramidase induce synergistic decreases in breast cancer cell growth. Breast Cancer Res Treat. 2012;133(2):447–458. doi:10.1007/s10549-011-1768-8
70. Emam SE, Ando H, Lila ASA, et al. Liposome co-incubation with cancer cells secreted exosomes (extracellular vesicles) with different proteins expressions and different uptake pathways. Sci Rep. 2018;8(1):14493. doi:10.1038/s41598-018-32861-w
71. Sinha S, Hoshino D, Hong NH, et al. Cortactin promotes exosome secretion by controlling branched actin dynamics. J Cell Biol. 2016;214(2):197–213. doi:10.1083/jcb.201601025
72. Parolini I, Federici C, Raggi C, et al. Microenvironmental pH is a key factor for exosome traffic in tumor cells. J Biol Chem. 2009;284(49):34211–34222. doi:10.1074/jbc.M109.041152
73. Jelonek K, Widlak P, Pietrowska M. The influence of ionizing radiation on exosome composition, secretion and intercellular communication. Protein Pept Lett. 2016;23(7):656–663. doi:10.2174/0929866523666160427105138
74. Lv LH, Wan YL, Lin Y, et al. Anticancer drugs cause release of exosomes with heat shock proteins from human hepatocellular carcinoma cells that elicit effective natural killer cell antitumor responses in vitro. J Biol Chem. 2012;287(19):15874–15885. doi:10.1074/jbc.M112.340588
75. Yang Y, Chen Y, Zhang F, Zhao Q, Zhong H. Increased anti-tumour activity by exosomes derived from doxorubicin-treated tumour cells via heat stress. Int J Hyperthermia. 2015;31(5):498–506. doi:10.3109/02656736.2015.1036384
76. Harmati M, Tarnai Z, Decsi G, et al. Stressors alter intercellular communication and exosome profile of nasopharyngeal carcinoma cells. J Oral Pathol Med. 2017;46(4):259–266. doi:10.1111/jop.12486
77. Samuel P, Mulcahy LA, Furlong F, et al. Cisplatin induces the release of extracellular vesicles from ovarian cancer cells that can induce invasiveness and drug resistance in bystander cells. Philos Trans R Soc Lond B Biol Sci. 2018;373(1737):20170065. doi:10.1098/rstb.2017.0065
78. Jella KK, Rani S, O’Driscoll L, McClean B, Byrne HJ, Lyng FM. Exosomes are involved in mediating radiation induced bystander signaling in human keratinocyte cells. Radiat Res. 2014;181(2):138–145. doi:10.1667/rr13337.1
79. Arscott WT, Tandle AT, Zhao S, et al. Ionizing radiation and glioblastoma exosomes: implications in tumor biology and cell migration. Transl Oncol. 2013;6(6):638–648. doi:10.1593/tlo.13640
80. Lehmann BD, Paine MS, Brooks AM, et al. Senescence-associated exosome release from human prostate cancer cells. Cancer Res. 2008;68(19):7864–7871. doi:10.1158/0008-5472.Can-07-6538
81. Serrano-Pertierra E, Oliveira-Rodríguez M, Rivas M, et al. Characterization of Plasma-Derived Extracellular Vesicles Isolated by Different Methods: A Comparison Study. Bioengineering. 2019;6:8. doi:10.3390/bioengineering6010008
82. Phuyal S, Hessvik NP, Skotland T, Sandvig K, Llorente A. Regulation of exosome release by glycosphingolipids and flotillins. FEBS J. 2014;281(9):2214–2227. doi:10.1111/febs.12775
83. Goldburg W. Dynamic light scattering. Am J Phys. 1999;67(12):1152–1160. doi:10.1119/1.19101
84. Soo CY, Song Y, Zheng Y, et al. Nanoparticle tracking analysis monitors microvesicle and exosome secretion from immune cells. Immunology. 2012;136(2):192–197. doi:10.1111/j.1365-2567.2012.03569.x
85. Xiao X, Yu S, Li S, et al. Exosomes: decreased sensitivity of lung cancer A549 cells to cisplatin. PLoS One. 2014;9(2):e89534. doi:10.1371/journal.pone.0089534
86. van der Pol E, Hoekstra AG, Sturk A, Otto C, van Leeuwen TG, Nieuwland R. Optical and non-optical methods for detection and characterization of microparticles and exosomes. J Thromb Haemost. 2010;8(12):2596–2607. doi:10.1111/j.1538-7836.2010.04074.x
87. Akers JC, Ramakrishnan V, Nolan JP, et al. Comparative analysis of technologies for quantifying extracellular vesicles (EVs) in clinical cerebrospinal fluids (CSF). PLoS One. 2016;11(2):e0149866. doi:10.1371/journal.pone.0149866
88. Kumar D, Gupta D, Shankar S, Srivastava RK. Biomolecular characterization of exosomes released from cancer stem cells: possible implications for biomarker and treatment of cancer. Oncotarget. 2015;6(5):3280–3291. doi:10.18632/oncotarget.2462
89. McAndrews KM, Kalluri R. Mechanisms associated with biogenesis of exosomes in cancer. Mol Cancer. 2019;18(1):52. doi:10.1186/s12943-019-0963-9
90. Willms E, Cabañas C, Mäger I, Wood MJA, Vader P. Extracellular vesicle heterogeneity: subpopulations, isolation techniques, and diverse functions in cancer progression. Front Immunol. 2018;9:738. doi:10.3389/fimmu.2018.00738
91. Zhang XF, Choi YJ, Han JW, et al. Differential nanoreprotoxicity of silver nanoparticles in male somatic cells and spermatogonial stem cells. Int J Nanomedicine. 2015;10:1335–1357. doi:10.2147/ijn.S76062
92. Carver KA, Yang D. N-acetylcysteine amide protects against oxidative stress-induced microparticle release from human retinal pigment epithelial cells. Invest Ophthalmol Vis Sci. 2016;57(2):360–371. doi:10.1167/iovs.15-17117
93. Schimel AM, Abraham L, Cox D, et al. N-acetylcysteine amide (NACA) prevents retinal degeneration by up-regulating reduced glutathione production and reversing lipid peroxidation. Am J Pathol. 2011;178(5):2032–2043. doi:10.1016/j.ajpath.2011.01.036
94. Gurunathan S, Jeyaraj M, La H, et al. Anisotropic platinum nanoparticle-induced cytotoxicity, apoptosis, inflammatory response, and transcriptomic and molecular pathways in human acute monocytic leukemia cells. Int J Mol Sci. 2020;21(2):440. doi:10.3390/ijms21020440
95. Almarzoug MHA, Ali D, Alarifi S, Alkahtani S, Alhadheq AM. Platinum nanoparticles induced genotoxicity and apoptotic activity in human normal and cancer hepatic cells via oxidative stress-mediated Bax/Bcl-2 and caspase-3 expression. Environ Toxicol. 2020;35(9):930–941. doi:10.1002/tox.22929
96. Ravid T, Tsaba A, Gee P, Rasooly R, Medina EA, Goldkorn T. Ceramide accumulation precedes caspase-3 activation during apoptosis of A549 human lung adenocarcinoma cells. Am J Physiol Lung Cell Mol Physiol. 2003;284(6):L1082–1092. doi:10.1152/ajplung.00172.2002
97. Quillet-Mary A, Jaffrézou JP, Mansat V, Bordier C, Naval J, Laurent G. Implication of mitochondrial hydrogen peroxide generation in ceramide-induced apoptosis. J Biol Chem. 1997;272(34):21388–21395. doi:10.1074/jbc.272.34.21388
98. Saeed-Zidane M, Linden L, Salilew-Wondim D, et al. Cellular and exosome mediated molecular defense mechanism in bovine granulosa cells exposed to oxidative stress. PLoS One. 2017;12(11):e0187569. doi:10.1371/journal.pone.0187569
99. Ye X, Zhang C, Chen Y, Zhou T. Upregulation of acetylcholinesterase mediated by p53 contributes to cisplatin-induced apoptosis in Human breast cancer cell. J Cancer. 2015;6(1):48–53. doi:10.7150/jca.10521
100. Okazaki T, Bell RM, Hannun YA. Sphingomyelin turnover induced by vitamin D3 in HL-60 cells. Role in cell differentiation. J Biol Chem. 1989;264(32):19076–19080. doi:10.1016/S0021-9258(19)47268-2
101. Hannun YA. Functions of ceramide in coordinating cellular responses to stress. Science. 1996;274(5294):1855–1859. doi:10.1126/science.274.5294.1855
102. Simons K, Ikonen E. Functional rafts in cell membranes. Nature. 1997;387(6633):569–572. doi:10.1038/42408
103. Dimanche-Boitrel MT, Meurette O, Rebillard A, Lacour S. Role of early plasma membrane events in chemotherapy-induced cell death. Drug Resist Updat. 2005;8(1–2):5–14. doi:10.1016/j.drup.2005.02.003
104. Patwardhan GA, Liu YY. Sphingolipids and expression regulation of genes in cancer. Prog Lipid Res. 2011;50(1):104–114. doi:10.1016/j.plipres.2010.10.003
105. Maurmann L, Belkacemi L, Adams NR, Majmudar PM, Moghaddas S, Bose RN. A novel cisplatin mediated apoptosis pathway is associated with acid sphingomyelinase and FAS proapoptotic protein activation in ovarian cancer. Apoptosis. 2015;20(7):960–974. doi:10.1007/s10495-015-1124-2
106. Trajkovic K, Hsu C, Chiantia S, et al. Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science. 2008;319(5867):1244–1247. doi:10.1126/science.1153124
107. Gills JJ, Zhang C, Abu-Asab MS, et al. Ceramide mediates nanovesicle shedding and cell death in response to phosphatidylinositol ether lipid analogs and perifosine. Cell Death Dis. 2012;3(7):e340. doi:10.1038/cddis.2012.72
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.