Back to Journals » International Journal of Chronic Obstructive Pulmonary Disease » Volume 14
Phenotyping and outcomes of hospitalized COPD patients using rapid molecular diagnostics on sputum samples
Authors Alotaibi NM, Chen V , Hollander Z , Leipsic JA, Hague CJ, Murphy DT , DeMarco ML, FitzGerald JM, McManus BM, Ng RT, Sin DD
Received 20 September 2018
Accepted for publication 28 December 2018
Published 23 January 2019 Volume 2019:14 Pages 311—319
DOI https://doi.org/10.2147/COPD.S188186
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Richard Russell
Nawaf M Alotaibi,1,2 Virginia Chen,1,3,4 Zsuzsanna Hollander,1,3,4 Jonathon A Leipsic,5 Cameron J Hague,5 Darra T Murphy,5 Mari L DeMarco,1,6 JM FitzGerald,3,7,8 Bruce M McManus,1,3,4,6 Raymond T Ng,4,9 Don D Sin1,3,7
1Centre for Heart Lung Innovation, James Hogg Research Centre, St Paul’s Hospital, Vancouver, BC, Canada; 2Department of Medicine, Division of Pulmonary Medicine, College of Medicine, King Saud University, Riyadh, Saudi Arabia; 3Institute for Heart and Lung Health, Vancouver, BC, Canada; 4PROOF Centre of Excellence, Vancouver, BC, Canada; 5Department of Radiology, St Paul’s Hospital, Vancouver, BC, Canada; 6Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, BC, Canada; 7Division of Respiratory Medicine, Department of Medicine, University of British Columbia, Vancouver, BC, Canada; 8The Lung Centre, Vancouver General Hospital, Vancouver, BC, Canada; 9Department of Computer Sciences, University of British Columbia, Vancouver, BC, Canada
Background: Etiologies of acute exacerbations of chronic obstructive pulmonary disease (AECOPD) are heterogeneous. We phenotyped severe AECOPD based on molecular pathogen detection of sputum samples collected at hospitalization of COPD patients and determined their outcomes.
Methods: We phenotyped 72 sputum samples of COPD patients who were hospitalized with a primary diagnosis of AECOPD using a molecular array that detected common bacterial and viral respiratory pathogens. Based on these results, the patients were classified into positive or negative pathogen groups. The pathogen-positive group was further divided into virus or bacteria subgroups. Admission day 1 blood samples were assayed for N-terminal prohormone brain natriuretic peptide, CRP, and complete blood counts.
Results: A total of 52 patients had a positive result on the array, while 20 patients had no pathogens detected. The most common bacterial pathogen detected was Haemophilus influenzae and the most common virus was rhinovirus. The pathogen-negative group had the worse outcomes with longer hospital stays (median 6.5 vs 5 days for bacteria-positive group, P=0.02) and a trend toward increased 1-year mortality (P=0.052). The bacteria-positive group had the best prognosis, whereas the virus-positive group had outcomes somewhere in between the bacteria-positive and pathogen-negative groups.
Conclusion: Molecular diagnostics on sputum can rapidly phenotype serious AECOPD into bacteria-, virus-, or pathogen-negative groups. The bacteria-positive group appears to have the best prognosis, while pathogen-negative group has the worst. These data suggest that AECOPD is a heterogeneous event and that accurate phenotyping of AECOPD may lead to novel management strategies that are personalized and more precise.
Keywords: COPD, molecular pathogen detection, exacerbation phenotypes
Introduction
Acute exacerbations of COPD (AECOPD) are caused by a variety of etiological factors.1 In AECOPD, the major drivers are respiratory tract infections; however, in roughly 30% of cases no clear inciting factor is found.2 An autopsy study of COPD patients who died within 24 hours of hospital admission due to AECOPD demonstrated that acute heart failure and pulmonary embolism were common primary causes of death.3 These data highlighted the importance of phenotyping AECOPD to target and treat underlying causes of AECOPD. According to the latest GOLD document,4 sputum cultures are generally not considered useful for guiding initial antibiotic choice or in phenotyping AECOPD. This is because sputum cultures have relatively poor sensitivity in identifying respiratory pathogens and determining therapeutic responsiveness to antimicrobials.5 Therefore, AECOPD are empirically treated with antibiotics and/or systemic corticosteroids, irrespective of the microbial or nonmicrobial drivers.4 The advent of molecular diagnostics in sputum has significantly increased the sensitivity of pathogen detection compared with traditional culture-based methods. However, its role in the prognosis or management of AECOPD has not been well defined.
In this study, our primary aim was to phenotype severe AECOPD by using a molecular pathogen-detection method. We hypothesized that a nucleic acid-based assay for detecting pathogens in sputum would enable classification of AECOPD into viral, bacterial, and uninfectious groups, which in turn would be associated with different health outcomes.
Methods
Study patients
This study consisted of patients who were able to provide adequate sputum samples in the COPD Rapid Transition Program. This program’s cohort has been described in detail previously.6,7 In brief, all patients included in this study were hospitalized with a confirmed primary diagnosis of AECOPD by board-certified general internists or pulmonologists who cared for these patients. All diagnoses were validated through a detailed chart review by at least one additional pulmonologist using the criteria recommended by the GOLD committee.4 Samples were classified as being adequate sputum samples based on color, transparency, and viscosity by study personnel, who were blinded to characteristics of the study patients. All patients received standard antiexacerbation therapy, including prednisone and antibiotics. The study is registered with ClinicalTrials.gov (NCT02050022, registered January 28, 2014). The study was approved by the University of British Columbia Providence Health Care Research Ethics Board (certificate H11-00786) for patients enrolled at St Paul’s Hospital, Vancouver, Canada and the University of British Columbia Clinical Research Ethics Board (certificate H13-00790) for patients enrolled at Vancouver General Hospital, Vancouver, Canada.
Specimens and measurement technique
Following receipt of written informed consent from patients, blood samples were collected in PaxGene, EDTA, and serum tubes on days 1 and 3 of hospitalization, at discharge, and on days 30 and 90 postdischarge. Blood components were processed as per standardized protocol and stored at −80°C until analysis. N-terminal prohormone brain natriuretic peptide (NT-proBNP), CRP, and complete blood count and differentials were measured using standard techniques (Supplementary material).
Spontaneously expectorated sputum samples were collected on day 1 in OmniGene oral (OM505) tubes and stored in −80°C freezers until measurement. The tubes were thawed in a hot bath at 50°C for 1 hour and then placed in an air incubator at 24°C for 30 minutes. Next, Sputolysin was added in a 1:1 ratio to liquefy the samples, after which samples were subaliquoted into 500 μL volumes and stored at −80°C. Bacterial load was quantified (Supplementary material), and for detecting pathogenic microorganisms, the Randox Respiratory Multiplex Array II was used. This array detects 22 common bacterial and viral pathogens within 6 hours using nucleic acids extracted from sputum samples (Supplementary material). In brief, the assay combines multiplex PCR and biochip-array hybridization. Baseline lung-function measurements were performed at the time of convalescence (ie, at day 30 or day 90) for AECOPD patients. Spirometry was used to obtain lung-function parameters after bronchodilator administration during clinical stability according to recommendations from American Thoracic Society–European Respiratory Society guidelines.8
Statistical analysis
Continuous variables that were normally distributed are reported as mean ± SD, abnormally distributed variables as medians and IQR, and categorical variables as percentages. Continuous variables that were not normally distributed were log10-transformed prior to application of a parametric test where appropriate. Student’s t-test and Mann–Whitney–Wilcoxon tests were used to determine differences between the pathogen-negative and pathogen-positive groups, and ANOVA and Kruskal–Wallis tests were used to determine differences between pathogen-negative, virus, and bacteria groups. Fisher’s exact test was used to test for differences in categorical variables between the groups. Comparisons of 1-year mortality rate across groups were analyzed by Kaplan–Meier survival curves using a log-rank test, and a Cox proportional-hazard model adjusted for age and sex was used to calculate HRs. Statistical tests were two-sided, and significance was assigned to results with P<0.05.
Ethics approval and consent to participate
The study is registered with ClinicalTrials.gov (NCT02050022, registered January 28, 2014). The study was approved by the University of British Columbia Providence Health Care Research Ethics Board (certificate H11-00786) for patients enrolled at St Paul’s Hospital, Vancouver, Canada and the University of British Columbia Clinical Research Ethics Board (certificate H13-00790) for patients enrolled at Vancouver General Hospital, Vancouver, Canada. Written informed consent was obtained from all enrolled patients. The study was conducted in accordance with the Declaration of Helsinki.
Results
Patient characteristics
Demographic and clinical data for the 72 patients studied are displayed in Table 1. Patients had a mean age of 65.8±11.5 years, 63.9% were male, 80.6% were Caucasian, and 61.1% were current smokers. All patients had airflow limitation, with mean FEV1 of 46.6%±16.7% predicted, and 38.9% had a history of cardiac comorbidities (heart failure, coronary artery disease, myocardial infarction, and arrhythmia).
Pathogens detected
Pathogens that were detected in our 72-patient cohort are presented in Table 2. The most common pathogen was Haemophilus influenzae, accounting for 33.7% of all pathogens detected. Rhinovirus was the most common virus detected, accounting for 13.3% of all pathogens detected. Details on the pathogens detected for each patient are provided in the Supplementary material.
  | Table 2 Pathogens detected on the panel |
Phenotyping COPD exacerbations
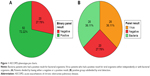
Twenty of 72 patients (27.8%) had a negative result on the array. Nucleic acid concentrations were similar between samples that did and did not have a positive result on the array (P=0.71, Supplementary material). Patients with a negative result on the array were considered to have had an uninfectious exacerbation, whereas those who had a positive result on the array were considered to have had an infectious exacerbation. We further subdivided the patients with an infectious exacerbation into either a bacteria- or a virus-associated exacerbation based on the results. The virus group consisted of those who had a virus identified on the array, with or without detection of bacteria (Figure 1).
Demographics and clinical data for AECOPD exacerbations
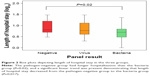
Demographic and clinical data for the groups are listed in the Supplementary material. There were no statistically significant differences between the pathogen-negative and pathogen-positive groups in terms of age, sex, cardiac comorbidities, inhaled corticosteroid use, or baseline FEV1% predicted. Of note, the pathogen-positive group was less likely to be on home (domiciliary) oxygen therapy (35% in the negative group and 14% in the positive group), but this comparison did not reach statistical significance (P=0.094). All hospitalizations were right-censored at 30 days, as hospital stays beyond this time frame were likely driven by factors other than AECOPD. There were no significant differences in length of hospital stay between the pathogen-negative and the pathogen-positive groups (P=0.096). However, a subgroup analysis revealed that there were significant differences in length of hospital stay across the bacteria, virus, and pathogen-negative groups (P=0.046 for overall ANOVA). These differences were largely driven by the comparison between the bacteria-associated AECOPD group and the pathogen-negative group (P=0.02 on post hoc analysis with Fisher’s least-significant-difference test). Consistent with this analysis, there was a significant trend in length of hospitalization across the three groups, with the pathogen-negative group having the longest stay and the bacteria group having the shortest (Ptrend=0.017, Figure 2).
NT-proBNP, CRP, complete blood count, and bacterial load
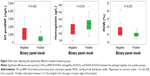
Concentrations of NT-proBNP, CRP, and complete blood counts for each of the groups are provided in the Supplementary material. We examined NT-proBNP, CRP, and complete blood counts at the date of hospital admission for all patients. The pathogen-negative group had significantly higher NT-proBNP concentrations (P=0.042), lower concentrations of hemoglobin (P=0.031), and higher red-blood-cell distribution width (P=0.025) compared with the pathogen-positive group (Figure 3). Subgroup analyses demonstrated a statistically significant difference between the pathogen-negative and virus groups for red-blood-cell distribution width (P=0.046 on overall Kruskal–Wallis test, Bonferroni-adjusted P=0.04 on post hoc pairwise comparison between the pathogen-negative and virus groups). There was no statistically significant difference among the three groups for NT-proBNP concentrations (P=0.081), but there was a significant trend in NT-proBNP concentrations across the three groups (P=0.029 on Jonckheere–Terpstra test for trend), with the pathogen-negative group having the highest concentrations and the bacteria group having the lowest (Supplementary material). Bacterial load measured by droplet digital PCR is shown in the Supplementary material. There was no statistically significant difference between the pathogen-negative and -positive groups (P=0.503) or the virus and other groups (P=0.625). Of note, we had day 30 follow-up NT-proBNP levels in 23 patients, which showed that subjects in the negative group had a median decrease in NT-proBNP levels of 1,102 (61−1,768) ng/L, while the positive group had a median decrease of 127.5 (30−458) ng/L. However, the results did not reach statistical significance, most likely due to the small sample.
One-year mortality
Of the 72 patients included in our study, 12 died within 1 year of follow-up (Table 3). On Kaplan–Meier survival analysis, there were no statistically significant differences between the pathogen-negative and -positive groups (P=0.065, Figure 4A). There were no statistically significant differences in survival between the virus and pathogen-negative groups (P=0.262), or between the virus and bacteria groups (P=0.376). However, there was moderate evidence of a difference in survival between the pathogen-negative and bacteria groups, which did not reach the predefined level of statistical significance (P=0.053, Figure 4B). Across the three groups, there was a trend toward increased mortality in the pathogen-negative group compared with the bacteria group (P=0.052). On the Cox proportional-hazard model adjusting for age and sex (Table 4), the pathogen-negative group was approximately 4.7 times more likely to die than the bacteria group during the 1-year follow-up period (HR 4.69, 95% CI 0.92–23.8).
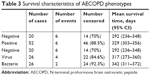  | Table 3 Survival characteristics of AECOPD phenotypes |
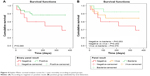  | Figure 4 Kaplan–Meier survival-analysis curves for 1-year mortality according to panel groups. |
  | Table 4 Cox proportional-hazard model comparing the pathogen-negative, virus, and bacteria groups |
Discussion
To our knowledge this is the first study to examine the utility of a molecular method for pathogen detection in AECOPD for the purpose of phenotyping exacerbations. Here, we showed that one in four patients hospitalized for AECOPD did not have detectable (common) respiratory pathogens in their sputum, one in three demonstrated a potential viral pathogen, and the rest had potential bacterial pathogens detected in their sputum. Most importantly, we showed that patients with a negative sputum-pathogen-array test had worse outcomes, including longer stays in hospital and a trend toward increased 1-year mortality. Interestingly, the group that demonstrated only bacteria in their sputum had the best outcomes, including the shortest hospital stay.
In this study, we demonstrated the feasibility of using molecular pathogen-detection methods that have been proven to be more sensitive in detecting viral pathogens than culture and serology methods in patients with serious AECOPD.9 Multiple studies have shown that molecular methods have higher sensitivity in detecting pathogens compared to traditional culture-based methods.10–12 Moreover, molecular methods enable simultaneous detection of multiple microorganisms, including both viral and bacterial pathogens, from a single clinical specimen. This approach, known as multiplexing, is increasingly being utilized for the diagnosis of a variety of different infectious diseases, and currently, there are multiple US Food and Drug Administration-approved panels designed to aid in the diagnosis of respiratory, gastrointestinal, and central nervous system infections.13 A strength of our study is that we chose a clinically approved panel that detects common respiratory pathogens. This enabled us to use a single clinical specimen, in contrast to other studies, which have used different tests and different sampling sites (ie, nasopharyngeal swabs for viruses, blood and urine samples for atypical bacteria, and sputum for typical bacteria).14–17 The pathogens detected in our study are consistent with the published literature on AECOPD in terms of type and prevalence.18
The current paradigm of AECOPD pathogenesis suggests that roughly 80% of cases are infectious in origin,15,19 with a third being caused by viruses20 and the remainder being attributed to multiple uninfectious etiologies. Uninfectious exacerbations in COPD are frequently attributed to heart failure,21 atrial fibrillation,22 gastroesophageal reflux disease,23 and acute pulmonary embolism.24 Here, we showed that patients with a negative result on the array had higher NT-proBNP concentrations, which suggests that these patients may have experienced acute cardiac dysfunction.25,26 This raises the possibility that at least in a subset of patients with serious AECOPD, cardiac dysfunction may play a significant role in their AECOPD.
We did not observe any significant differences in CRP concentrations across groups. Several explanations are possible for this observation. First, we had a relatively small sample, which may have limited our ability to detect statistically significant differences in CRP concentrations. Second, the array does not cover all the bacterial organisms that are associated with AECOPD. Most notably, Staphylococcus aureus and Pseudomonas aeruginosa are not on this array. Therefore, patients with these infectious organisms may have been overlooked and included in the pathogen-negative group. We did not find any significant differences in CRP concentrations between viral and bacterial groups either, which have been previously noted by other groups.27,28
In AECOPD, there are specific treatments for bacterial infections that have very high cure rates, while in the case of viral infections, influenza is the only respiratory virus that has an available treatment.29 We showed here that patients who had only bacteria detected in their sputum had the shortest hospitalization, while interestingly patients who had a negative result on the array had the longest hospitalization. Previous studies have shown that patients with a high burden of comorbidities, such as anemia, have longer hospitalizations for AECOPD, independently of age, sex, or FEV1.30 Similarly, those who have elevated NT-proBNP concentrations on admission also experience longer hospitalizations for their AECOPD.6
Interestingly, we observed a trend toward a higher 1-year mortality rate in patients who were part of the pathogen-negative group compared to the bacteria group. The pathogen-negative group was fourfold more likely to die within 1 year of follow-up than the bacteria group. The most common causes of death in COPD patients according to death-certificate data are cardiac diseases,31 and elevated NT proBNP concentrations are strongly associated with mortality in AECOPD,6,32 which might explain the increased mortality rate observed in patients who had a negative result on the array.
There are several limitations to our study and studies utilizing molecular methods of pathogen detection in general. First, we used a qualitative diagnostic method, and colonization could have led to false-positive results.33 Colonization rates may also be increased in those with concomitant bronchiectasis, though in our study all patients had chest imaging and those with significant bronchiectasis were excluded from the study (data not shown). Second, we used sputum samples to phenotype exacerbations. However, a sputum sample does not necessarily represent the whole lung. Moreover, there are regional differences in detection rates of bacterial pathogens within the same lung.34,35 Third, our study was a retrospective study in which stored sputum samples in −80°C were tested for respiratory pathogens, and the effects of prolonged storage at low temperatures on microbial pathogen detection have not been systematically studied.36 However, in a study that examined the ability of the Xpert Mycobacterium tuberculosis–resistance to rifampicin assay to detect M. tuberculosis in sputum samples that had been stored in −80°C freezers for up to 4 years, the assay showed sensitivity of 95.7%, which was within the range reported in fresh samples.37 Another study that examined microbial communities in stored bronchoalveolar lavage samples for cystic fibrosis patients in −80°C for >5 years showed results that were consistent with historical culturing results.38 Fourth, a limitation of the array is that it does not cover all pathogenic organisms implicated in AECOPD, and of these, P. aeruginosa and S. aureus are the most notable. For Staphylococcus, as with other organisms that possess thick cell walls, specialized DNA-extraction methods are required.39–42 If these methods are used, it has been shown that increased detection of Staphylococcus comes at the expense of microbial organisms with a fragile cell wall and viruses.43,44 Currently, there is no commercial nucleic acid-extraction kit that can simultaneously extract Gram-positive, Gram-negative, and viruses from the same sample. Kajiura et al44 proposed a method for achieving this goal through techniques that prevent the loss of smaller viral particles while still extracting Gram-positive bacteria. This has been successfully used on 300 archived clinical samples of respiratory origin. This method appears promising, and if externally and prospectively validated, it could become the gold standard for nucleic acid extraction for molecular pathogen detection. Fifth, we did not have data on whether the subjects had received antibiotics or systemic steroids prior to presenting to our emergency department. Sixth, the small sample of the study may be a potential reason for not uncovering statistically significant outcomes. Lastly, we did not have echocardiography data to confirm with the elevated NT-proBNP concentrations the evidence of cardiac dysfunction in the pathogen-negative group.
Conclusion
Comprehensive molecular methods of pathogen detection are soon to be considered a cornerstone for diagnosing respiratory infectious diseases. Molecular diagnostic methods are significantly faster and more sensitive than culture methods, and not affected by prior antibiotic use to the same extent as culture methods. One important aspect is that these methods are not only capable of simultaneously detecting a wide gamut of different pathogens (bacteria, viruses, and fungi) but also are capable of simultaneously and accurately detecting antibiotic-resistance genes.45 We show here that a commercially available respiratory multiplex array could aid in phenotyping AECOPD into infectious or uninfectious exacerbations. These results are encouraging to explore further and develop better molecular pathogen-detection panels that possess broader pathogen coverage and address the issue of colonization. Prospectively examining respiratory multiplex arrays in AECOPD that requires hospital admission and assessing the point-of-care advantages that these panels might possess would be the logical next step to validate their value in the utilization of hospital resources and clinical outcomes.
Availability of data and materials
Data sets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Acknowledgments
We would like to thank all patients for their participation and Sara Assadian, Roxanne Rousseau, and Breanne Crouch for their roles in patient recruitment for the study and collecting and entering clinical data, and Sheena Tam, Grace Lee, Yeni Oh, and David Ngan for processing samples, as well as Meghan McLennan and Jackson Wu for performing clinical laboratory analyses. An abstract of this paper was presented at the 2018 American Thoracic Society International Conference as a poster discussion with interim findings. The poster’s abstract was published in the 2018 abstract issue of the American Journal of Respiratory and Critical Care Medicine (https://www.atsjournals.org/doi/abs/10.1164/ajrccm-conference.2018.197.1_MeetingAbstracts.A2749). The study was funded by Genome Canada, Genome British Columbia, Genome Quebec, Canadian Institutes of Health Research, Providence Health Care, St Paul’s Hospital Foundation, and the PROOF Center.
Author contributions
NMA had full access to all the data in the study, takes responsibility for the integrity of the data and the accuracy of the data analysis, and contributed to drafting the manuscript. DDS contributed substantially to the study design, data analysis and interpretation, and had authority over manuscript preparation and the decision to submit the manuscript for publication. CJH interpreted the radiological studies. All authors contributed to data analysis, drafting and revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
DDS reports grants and personal fees from AstraZeneca and Boehringer Ingelheim, grants from Merck Frosst, and personal fees from Novartis, Regeneron, and Sanofi Aventis outside the submitted work. JAL reports grants from GE Healthcare outside the submitted work. The authors report no other conflicts of interest in this work.
References
Pavord ID, Jones PW, Burgel PR, Rabe KF. Exacerbations of COPD. Int J Chron Obstruct Pulmon Dis. 2016;11 Spec Iss:21–30. | ||
Connors AF, Dawson NV, Thomas C, et al. Outcomes following acute exacerbation of severe chronic obstructive lung disease. The support Investigators (study to understand prognoses and preferences for outcomes and risks of treatments). Am J Respir Crit Care Med. 1996;154(4 Pt 1):959–967. | ||
Zvezdin B, Milutinov S, Kojicic M, et al. A postmortem analysis of major causes of early death in patients hospitalized with COPD exacerbation. Chest. 2009;136(2):376–380. | ||
Vogelmeier CF, Criner GJ, Martinez FJ, et al. Global strategy for the diagnosis, management, and prevention of chronic obstructive lung disease 2017 report. Gold executive summary. Am J Respir Crit Care Med. 2017;195(5):557–582. | ||
Sethi S. Molecular diagnosis of respiratory tract infection in acute exacerbations of chronic obstructive pulmonary disease. Clin Infect Dis. 2011;52 Suppl 4:S290–S295. | ||
Chen YR, Chen V, Hollander Z, et al. C-reactive protein and N-terminal prohormone brain natriuretic peptide as biomarkers in acute exacerbations of COPD leading to hospitalizations. PLoS One. 2017;12(3):e0174063. | ||
Alotaibi NM, Chen V, Hollander Z, et al. Phenotyping COPD exacerbations using imaging and blood-based biomarkers. Int J Chron Obstruct Pulmon Dis. 2018;13:217–229. | ||
Miller MR, Hankinson J, Brusasco V, et al. Standardisation of spirometry. Eur Respir J. 2005;26(2):319–338. | ||
Beckham JD, Cadena A, Lin J, et al. Respiratory viral infections in patients with chronic, obstructive pulmonary disease. J Infect. 2005;50(4):322–330. | ||
Simpson JL, Baines KJ, Horvat JC, et al. COPD is characterized by increased detection of Haemophilus influenzae, Streptococcus pneumoniae and a deficiency of Bacillus species. Respirology. 2016;21(4):697–704. | ||
Murphy TF, Brauer AL, Schiffmacher AT, Sethi S. Persistent colonization by Haemophilus influenzae in chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2004;170(3):266–272. | ||
Garcha DS, Thurston SJ, Patel AR, et al. Changes in prevalence and load of airway bacteria using quantitative PCR in stable and exacerbated COPD. Thorax. 2012;67(12):1075–1080. | ||
Hanson KE, Couturier MR. Multiplexed molecular diagnostics for respiratory, gastrointestinal, and central nervous system infections. Clin Infect Dis. 2016;63(10):1361–1367. | ||
Kim HC, Choi SH, Huh JW, et al. Different pattern of viral infections and clinical outcomes in patient with acute exacerbation of chronic obstructive pulmonary disease and chronic obstructive pulmonary disease with pneumonia. J Med Virol. 2016;88(12):2092–2099. | ||
Papi A, Bellettato CM, Braccioni F, et al. Infections and airway inflammation in chronic obstructive pulmonary disease severe exacerbations. Am J Respir Crit Care Med. 2006;173(10):1114–1121. | ||
Bafadhel M, McKenna S, Terry S, et al. Acute exacerbations of chronic obstructive pulmonary disease: identification of biologic clusters and their biomarkers. Am J Respir Crit Care Med. 2011;184(6):662–671. | ||
Shimizu K, Yoshii Y, Morozumi M, et al. Pathogens in COPD exacerbations identified by comprehensive real-time PCR plus older methods. Int J Chron Obstruct Pulmon Dis. 2015;10:2009–2016. | ||
Leung JM, Tiew PY, Mac Aogáin M, et al. The role of acute and chronic respiratory colonization and infections in the pathogenesis of COPD. Respirology. 2017;22(4):634–650. | ||
Rangelov K, Sethi S. Role of infections. Clin Chest Med. 2014;35(1):87–100. | ||
Mohan A, Chandra S, Agarwal D, et al. Prevalence of viral infection detected by PCR and RT-PCR in patients with acute exacerbation of COPD: a systematic review. Respirology. 2010;15(3):536–542. | ||
Rutten FH, Cramer MJ, Lammers JW, Grobbee DE, Hoes AW. Heart failure and chronic obstructive pulmonary disease: an ignored combination? Eur J Heart Fail. 2006;8(7):706–711. | ||
Buch P, Friberg J, Scharling H, Lange P, Prescott E. Reduced lung function and risk of atrial fibrillation in the Copenhagen City Heart study. Eur Respir J. 2003;21(6):1012–1016. | ||
Hurst JR, Vestbo J, Anzueto A, et al. Susceptibility to exacerbation in chronic obstructive pulmonary disease. N Engl J Med. 2010;363(12):1128–1138. | ||
Aleva FE, Voets L, Simons SO, de Mast Q, van der Ven A, Heijdra YF. Prevalence and localization of pulmonary embolism in unexplained acute exacerbations of COPD: a systematic review and meta-analysis. Chest. 2017;151(3):544–554. | ||
Buchan A, Bennett R, Coad A, Barnes S, Russell R, Manuel AR. The role of cardiac biomarkers for predicting left ventricular dysfunction and cardiovascular mortality in acute exacerbations of COPD. Open Heart. 2015;2(1):e000052. | ||
Hawkins NM, Khosla A, Virani SA, McMurray JJ, FitzGerald JM. B-type natriuretic peptides in chronic obstructive pulmonary disease: a systematic review. BMC Pulm Med. 2017;17(1):11. | ||
Chang CH, Tsao KC, Hu HC, et al. Procalcitonin and C-reactive protein cannot differentiate bacterial or viral infection in COPD exacerbation requiring emergency department visits. Int J Chron Obstruct Pulmon Dis. 2015;10:767–774. | ||
Kwak HJ, Park DW, Kim JE, et al. Prevalence and risk factors of respiratory viral infections in exacerbations of chronic obstructive pulmonary disease. Tohoku J Exp Med. 2016;240(2):131–139. | ||
Fiore AE, Fry A, Shay D, et al. Antiviral agents for the treatment and chemoprophylaxis of influenza – recommendations of the Advisory Committee on Immunization Practices (ACIP). MMWR Recomm Rep. 2011;60(1):1–24. | ||
Almagro P, Cabrera FJ, Diez J, et al. Comorbidities and short-term prognosis in patients hospitalized for acute exacerbation of COPD: the EpoC en Servicios de medicina interna (ESMI) study. Chest. 2012;142(5):1126–1133. | ||
Hansell AL, Walk JA, Soriano JB. What do chronic obstructive pulmonary disease patients die from? A multiple cause coding analysis. Eur Respir J. 2003;22(5):809–814. | ||
Chang CL, Robinson SC, Mills GD, et al. Biochemical markers of cardiac dysfunction predict mortality in acute exacerbations of COPD. Thorax. 2011;66(9):764–768. | ||
Marin A, Garcia-Aymerich J, Sauleda J, et al. Effect of bronchial colonisation on airway and systemic inflammation in stable COPD. COPD. 2012;9(2):121–130. | ||
Goddard AF, Staudinger BJ, Dowd SE, et al. Direct sampling of cystic fibrosis lungs indicates that DNA-based analyses of upper-airway specimens can misrepresent lung microbiota. Proc Natl Acad Sci U S A. 2012;109(34):13769–13774. | ||
Gutierrez JP, Grimwood K, Armstrong DS, et al. Interlobar differences in bronchoalveolar lavage fluid from children with cystic fibrosis. Eur Respir J. 2001;17(2):281–286. | ||
Dollow JM, Green JA. Significant roadblocks exist in developing sputum sample libraries for clinical validation of novel in vitro diagnostics. Drug Des Devel Ther. 2014;8:175–182. | ||
Singh S, Singh A, Prajapati S, et al. Xpert MTB/RIF assay can be used on archived gastric aspirate and induced sputum samples for sensitive diagnosis of paediatric tuberculosis. BMC Microbiol. 2015;15:191. | ||
Willner D, Daly J, Whiley D, Grimwood K, Wainwright CE, Hugenholtz P. Comparison of DNA extraction methods for microbial community profiling with an application to pediatric bronchoalveolar lavage samples. PLoS One. 2012;7(4):e34605. | ||
Zemanick ET, Wagner BD, Robertson CE, et al. Assessment of airway microbiota and inflammation in cystic fibrosis using multiple sampling methods. Ann Am Thorac Soc. 2015;12(2):221–229. | ||
Wu GD, Lewis JD, Hoffmann C, et al. Sampling and pyrosequencing methods for characterizing bacterial communities in the human gut using 16S sequence tags. BMC Microbiol. 2010;10:206. | ||
Ó Cuív P, Aguirre de Cárcer D, Jones M, et al. The effects from DNA extraction methods on the evaluation of microbial diversity associated with human colonic tissue. Microb Ecol. 2011;61(2):353–362. | ||
Bag S, Saha B, Mehta O, et al. An improved method for high quality Metagenomics DNA extraction from human and environmental samples. Sci Rep. 2016;6:26775. | ||
von Wintzingerode F, Göbel UB, Stackebrandt E. Determination of microbial diversity in environmental samples: pitfalls of PCR-based rRNA analysis. FEMS Microbiol Rev. 1997;21(3):213–229. | ||
Kajiura LN, Stewart SD, Dresios J, Uyehara CFT. Simultaneous extraction of viral and bacterial nucleic acids for molecular diagnostic applications. J Biomol Tech. 2015;26(4):118–124. | ||
Frickmann H, Masanta WO, Zautner AE. Emerging rapid resistance testing methods for clinical microbiology laboratories and their potential impact on patient management. Biomed Res Int. 2014;2014(6):1–19. |
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.




