Back to Journals » Drug Design, Development and Therapy » Volume 15
Paeoniflorin Sensitizes Breast Cancer Cells to Tamoxifen by Downregulating microRNA-15b via the FOXO1/CCND1/β-Catenin Axis
Authors Wang Y, Wang Q, Li X, Luo G, Shen M, Shi J, Wang X, Tang L
Received 20 August 2020
Accepted for publication 12 December 2020
Published 22 January 2021 Volume 2021:15 Pages 245—257
DOI https://doi.org/10.2147/DDDT.S278002
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Anastasios Lymperopoulos
Yanhong Wang,1,2 Qian Wang,1 Xibei Li,3 Gongwen Luo,2 Mou Shen,2 Jia Shi,4 Xueliang Wang,5 Lu Tang6
1Department of Basic Medicine, Medical College of Yunnan University of Economics and Management, Kunming, Yunnan 650000, People’s Republic of China; 2Second Department of Internal Medicine, Chongming Branch of Yueyang Integrated Hospital of Traditional Chinese and Western Medicine Affiliated to Shanghai University of Traditional Chinese Medicine, Chongming, Shanghai, 202150, People’s Republic of China; 3Department of Stomatology, Jining Medical College, Jining, Shandong 272000, People’s Republic of China; 4Department of Information, The First Affiliated Hospital of Naval Military Medical University (Shanghai Changhai Hospital), Shanghai 200433, People’s Republic of China; 5Department of Nephrology and Rheumatology, Zhaotong Traditional Chinese Medicine Hospital of Yunnan Province, Zhaotong, Yunnan 657000 People’s Republic of China; 6Department of Traditional Chinese Medicine, Kunming Second People’s Hospital, Kunming, Yunnan, 650000 People’s Republic of China
Correspondence: Lu Tang
Department of Traditional Chinese Medicine, Kunming Second People’s Hospital, No. 672 Longquan Road, Panlong District, Kunming, Yunnan 650000, People’s Republic of China
Email [email protected]
Background: Paeoniflorin (Pae) possesses anti-tumor activity in various malignancies. However, it is unclear whether Pae plays a sensitizer role in breast cancer (BC) and the molecular mechanisms involved in this process. Our oligonucleotide microarray revealed that microRNA (miR)-15b is the most significantly downregulated miRNA in MCF-7/4-hydroxytamoxifen (4-OHT) cells treated with Pae. This paper summarized the relevance of Pae in BC cell endocrine resistance to tamoxifen (Tam) and the molecular mechanisms involved miR-15b expression.
Materials and Methods: 4-OHT-resistant BC cell lines were developed and treated with different concentrations of Pae. Flow cytometry, lactose dehydrogenase activity, caspase-3 activity, colony formation, and EdU assays were carried out to assess the impact of Pae on BC cells. Differentially expressed miRNAs in BC cells treated with Pae were analyzed by microarray. Targeting mRNAs of screened miR-15b as well as the binding of forkhead box O1 (FOXO1) to the cyclin D1 (CCND1) promoter sequence were predicted through bioinformatics analysis. Finally, the expression of β-catenin signaling-related genes in cells was detected by Western blotting.
Results: Pae (100 μg/mL) inhibited the clonality and viability of BC cells, while enhancing apoptosis in vitro. Pae also repressed miR-15b expression. Overexpression of miR-15b restored the growth and resistance of BC cells to 4-OHT. Moreover, Pae promoted FOXO1 expression by downregulating miR-15b, thereby transcriptionally inhibiting CCND1 and subsequently blocking β-catenin signaling.
Conclusion: Pae inhibits 4-OHT resistance in BC cells by regulating the miR-15b/FOXO1/CCND1/β-catenin pathway.
Keywords: paeoniflorin, breast cancer, tamoxifen, microRNA-15b, FOXO1
Introduction
Breast cancer (BC) represented the most prevalent cancer diagnosed among the female population (excluding skin cancers) in 2019 and ranked second in cancer-related deaths in women just after lung cancer.1 Estrogen receptor (ER) signaling plays a crucial role in BC progression. While endocrine therapy which blocks ER activity is of great importance, its effectiveness is limited due to frequent intrinsic and acquired resistance.2 Tamoxifen (Tam), a selective ER modulator, has been widely used to treat patients with ERα-positive BC for over 40 years in both the adjuvant and the recurrent settings.3 It significantly lowers disease recurrence and mortality of patients with BC.4 Unfortunately, endocrine resistance interferes with the therapeutic effects of Tam, since approximately 30% of women develop local recurrence or distant metastasis despite the initial beneficial response.5
Paeoniflorin (Pae), a water-soluble monoterpene glycoside derived from Paeonia lactiflora Pall, possesses a wide spectrum of medicinal properties including anti-inflammatory, antioxidant, analgesic, cardioprotective, neuroprotective, antidepressant-like, and immune-regulatory activities.6 Moreover, Pae has been reported to modulate multidrug resistance in gastric cancer cells,7 which allows us to hypothesize whether it exerts a similar function in BC. Pae has been reported to regulate the function and activation of immune cells, interfere with the generation of inflammatory mediators, and restore abnormal intracellular signaling, and is thus involved in a great range of autoimmune diseases.8 More recently, the anti-tumor effects of Pae have been described in nasopharyngeal carcinoma9 and non-small cell lung cancer.10 Moreover, Pae protects BC cells from undergoing epithelial-mesenchymal transition stimulated by hypoxia.11 Nonetheless, its role in BC endocrine resistance, especially resistance to Tam, remains undefined. Interestingly, Pae has been proposed to repress cell proliferation in multiple myeloma and to facilitate apoptosis through its effects on microRNA-29b.12 As a consequence, we postulate that Pae inhibits endocrine resistance of BC cells to 4-hydroxytamoxifen (4-OHT) by interacting with certain miRNAs. Our oligonucleotide microarray findings indicated that miR-15b is the most significantly downregulated miRNAs in an MCF-7/4-OHT cell line treated with Pae. Moreover, miR-15b was upregulated in BC and accelerated BC progression by targeting a tumor suppressor PAQR3.13 In addition, the upregulation of miR-15b is associated with lung adenocarcinoma cell chemoresistance to cisplatin and metastasis both in vitro and in vivo.14 Thus, miR-15b regulation by Pae is believed to be engaged in the endocrine resistance observed in BC cells by interacting with its target gene. Since the first Tam-resistant cell model was developed in 1981, resistance has been stimulated in MCF7 cells, an ER+ BC cell line, by prolonged incubation with 4-OHT.15 In this context, our present study explored the effects of Pae on BC cells in 4-OHT-induced endocrine resistance in vitro, in addition to the underlying mechanisms involving miR-15b.
Materials and Methods
Reagents, Drugs, Antibodies and Plasmids
Pae purchased from MedChemExpress (HY-N0293, Monmouth Junction, NJ, USA) was diluted into a 1 mg/mL solution with DMSO. The lactose dehydrogenase (LDH) assay kit was purchased from Beyotime Biotechnology Co., Ltd. (Shanghai, China, C0016) and the Caspase-3 kit from Abcam Inc. (ab39401, Cambridge, UK). The primary antibodies used in Western blotting analysis included FOXO1 (2880; Cell Signaling Technologies, Beverly, MA, USA), GSK3β (ab32391, Abcam), GSK3β (phosphoY216, ab75745, Abcam), and β-catenin (8480, Cell Signaling Technologies). The secondary antibody was horseradish peroxidase-conjugated goat anti-rabbit antibody against IgG (ab6721, Abcam). miR-15b mimic and mimic control were purchased and synthesized by Sangon (Shanghai, China). A siRNA and a pcDNA3.1 overexpression plasmid targeting FOXO1 were purchased from Shanghai GenePharma Co., Ltd. (Shanghai, China). The β-catenin signaling agonist WAY-262,611 was purchased from MedChemExpress (HY-11,035) and was diluted into 5 μM solution. The GSK3β-specific inhibitor lithium chloride (LiCl) was purchased from Abcam (ab120853). 4-OHT was purchased from Sigma-Aldrich (68,392-35-8) and was dissolved in methanol in concentrations ranging from (0.01–10 μM).
Culture and Treatment of BC Cells
ER-positive T47D/4-OHT and MCF-7/4-OHT resistant cell lines were established as previously reported.16 BC cell lines MCF-7 and T47D were obtained from the American Type Culture Collection (Manassas, VA, USA) and were grown in DMEM (Hyclone, Marlborough, MA, USA) containing 10% fetal bovine serum (FBS, Clark Bioscience, Richmond, VA, USA). 4-OHT-resistant cell lines (MCF-7/4-OHT and T47D/4-OHT) were derived from parental cell lines by continuous and gradual exposure to 4-OHT to reach a final concentration of 1 μM 4-OHT in methanol in 6 months. The established MCF-7/4-OHT and T47D/4-OHT cells were maintained in medium with 1 μM 4-OHT. Changes in cell sensitivity to 4-OHT were detected by the Cell Counting Kit-8 (CCK-8) assay (Sigma, St. Louis, MO, USA) to determine the successful establishment of drug-resistant cell lines. Afterwards, the cells were maintained in RPMI-1640 medium (Hyclone) containing 10% fetal bovine serum (FBS) (Clark Bioscience) and 5 mg/mL insulin in a 5% CO2 atmosphere incubator at 37°C. MCF-7/4-OHT cells were treated with DMSO, Pae (at different concentrations), Pae (100 μg/mL) + Mock (miR-15b control), Pae + miR-15b (50 ng), Pae + scramble (Scr, 100 ng scrambled siRNA), Pae + si-FOXO1, Pae + DMSO or Pae + WAY-262,611 (1 μM). In addition, T47D/4-OHT cells were treated with DMSO, Pae (at different concentrations), Pae + Mock, Pae + miR-15b, Pae + pCDNA3.1 (100 ng pCNDA3.1 empty plasmid), Pae + CCND1, Pae + DMSO or Pae + LiCl (5 μM LiCl).
CCK-8 Assays
MCF-7/4-OHT and T47D/4-OHT cells were plated in a 96-well plate at a concentration of 5 × 103 cells/well. Following cell attachment, Pae at 0, 20, 40, 80 and 100 μg/mL was added. After 24 h, cell viability was assessed by the CCK-8 assay. Cell viability was subsequently evaluated by CCK-8 assay after 24 h of 4-OHT treatment (8 μM). The optical density (OD) value of each well at 450 nm (A450) was read on a spectrophotometer. The median inhibition concentration of each drug (IC50) was estimated by the relative survival curve.
Caspase-3 Activity Assay
A total of 5 × 104 cells were plated in a 10-cm petri dish for a 24-hour growth period. After 24 hours of Pae and 4-OHT treatment, as indicated above, the medium was refreshed with serum-free DMEM or complete DMEM. Cells were harvested after 24 hours. Caspase-3 activity was determined using the Caspase-3 assay kit (ab39401, Abcam, Cambridge, UK).
Flow Cytometry
The percentage of apoptotic cells was detected using a fluorescein isothiocyanate (FITC)-Annexin V cell apoptosis detection kit (BD Biosciences, Franklin Lakes, NJ, USA). The cells were suspended in 1× binding buffer at a concentration of 1 × 106 cells/mL. Five microliters FITC-Annexin V and 5 μL propidium iodide were supplemented to 100 μL resuspension and incubated at room temperature for 15 minutes. After that, 400 μL 1× binding buffer was added before loading onto the FACSCalibur (BD Biosciences).
Cell Cytotoxicity Assay
The cells were seeded into a 96-well cell culture plate and treated with Pae for 24 hours. After a 48-h treatment with 4-OHT, the LDH release reagent was added at a volume of 10% of the original medium. Cells were suspended by trituration and the cells were incubated for another 30 minutes. The culture plate was then centrifuged using a centrifuge at 400 ×g for 5 minutes. The supernatant (120 μL) was transferred to a new 96-well plate, followed by the sample determination. The OD values of the samples were measured at 490 nm using a microplate reader (1681130a, Bio-Rad Laboratories, Hercules, CA, USA).
5-Ethynyl-2ʹ-Deoxyuridine (EdU) Incorporation Assays
The treated T47D and MCF-7 cells (as described above) were plated into a 96-well plate at 8 × 103 cells/well, followed by incubation with 50 μM EdU reagent (Guangzhou RiboBio Co., Ltd., Guangzhou, Guangdong, China) for 2 h. The cells were then treated with Apollo (RiboBio) and 4ʹ,6-diamidino-2-phenylindole (Sigma-Aldrich). Images were acquired through a fluorescence microscope (Nikon, Tokyo, Japan).
Colony Formation Assay
The cells were transfected and treated with Pae for 24 h, followed by 4-OHT treatment. MCF7 and T47D cells were seeded into 6-cm plates at 103 cells/well. After 2 weeks of culture at 37°C with 5% CO2, the colonies formed were fixed with methanol for 20 minutes and stained with 0.1% crystal violet (Sigma) at room temperature. The number of colonies formed was calculated under a microscope.
Microarray Analysis
Total RNA was isolated from MCF-7/4-OHT cells treated with DMSO or Pae using Trizol reagent (Invitrogen Inc., Carlsbad, CA, USA). Small RNAs (<200 nt) were isolated from total RNA using a mirVana miRNA isolation kit (Ambion, Austin, TX, USA) and subjected to a miRCURYTM LNA Array (Exiqon, Denmark). The miRNA microarray analysis was carried out based on the standard procedure using the MiniSeq System (SY-420-1001, Illumina platform).
Real-Time Quantitative Polymerase Chain Reaction Analysis (RT-qPCR)
Total RNA was isolated using Trizol reagent (Invitrogen). RT-qPCR experiments were performed using the ABI7500 (Applied Biosystems, Foster City, CA, USA) after reverse transcription into cDNA using Evoscript Universal cDNA Master (Roche Diagnostics, Co., Ltd., Rotkreuz, Switzerland) (for mRNA) and a Taqman MicroRNA reverse transcription kit (4,366,596, Applied Biosystems). A SmartChip™ Real-Time PCR System (Takara, 640,022, for mRNA) and a TaqMan™ miRNA ABC Purification Kit (Applied Biosystems, 4,473,088, for miRNA) were used for analysis. The primers used are listed in Table 1. The relative gene expression was standardized to U6 or glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and the Ct value was used for quantification using the 2−ΔΔCt method.
 |
Table 1 List of Oligonucleotides Used in the Study |
Western Blot Analysis
Radioimmunoprecipitation assay buffer (Beyotime, Shanghai, China) was added to extract total protein, and the protein concentration was measured using a bicinchoninic acid kit (Sigma-Aldrich). Protein samples were separated by dodecyl sulfate sodium salt-polyacrylamide gel electrophoresis and then transferred to polyvinylidene difluoride membranes. The membranes were then blocked with 5% skim milk and incubated overnight with the primary antibodies at 4°C and with secondary antibodies for 1 hour at room temperature. Finally, the Enhanced Chemiluminescence Reagent Kit (GE Healthcare, Little Chalfont, Buckinghamshire, UK) was used to detect the signals.
Luciferase Reporter Assay
FOXO1 was predicted to be a target mRNA of miR-15b according to TargetScan (http://www.targetscan.org/). The wild-type (WT) of FOXO1 3ʹuntranslated region (3ʹUTR) was amplified (FOXO1 WT). Site-directed mutagenesis of the miR-15b binding sites was generated (Sangon, Shanghai, China). The FOXO1 WT or mutant (FOXO1 MT) sequences were inserted into pmirGLO vectors for luciferase reporter assays (Promega Corporation, Madison, WI, USA). Then, the pmirGLO recombinant vector was co-transfected with miR-15b mimic or mimic control into HEK293T cells (CRL-1573, American Type Culture Collection), and the luciferase activity was determined 48 h after co-transfection.
Chromatin Immunoprecipitation (ChIP)
We first used the JASPAR website (http://jaspar.genereg.net/) to analyze the binding sites of FOXO1 and CCND1. A total of 1.5 × 108 T47D and MCF-7 cells were subjected to the ChIP-IT Express enzyme (#53009, Active Motif., Carlsbad, CA, USA). FOXO1 antibody or non-specific IgG (I-1000, Vector Laboratories, Inc., Burlingame, CA, USA) was used for pull-down assays. In accordance with the manufacturer’s protocol, DNA was purified using a ChIP DNA purification kit (#5802, Active Motif). The CCND1 promoter region of 560–745 was detected using the following primers, containing multiple common FOXO1 motifs: forward: 5ʹGTGTACGTCAGTCCCTTACATC-3ʹ, reverse: 5ʹ-AGACTCTGACTCTGGGAAGT-3ʹ. The results were calculated as a percentage of Input.
Statistical Analysis
The Statistical Program for Social Sciences (SPSS) version 22.0 software (IBM Corp., Armonk, NY, USA) was used for all statistical analyses of the collected data. Measurement data are expressed as mean ± standard deviation (SD), and were compared by pair or unpaired t-test and one-way or two-way ANOVA followed by Tukey’s multiple comparison. Statistically significant differences were considered at a p-value <0.05.
Results
4-OHT-Resistant BC Cell Lines Were Successfully Established
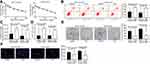
First, the cytotoxic effects of 4-OHT in MCF-7 parental cells (MCF-7/Par), 4-OHT-resistant MCF-7 cells (MCF-7/4-OHT), T47D parent cells (T47D/Par), and 4-OHT-resistant T47D cells (T47D/4-OHT) were evaluated using the CCK-8 assay. We found that the IC50 of MCF-7/4-OHT was 16.43 μM, which was higher than that for MCF-7/Par (3.84 μM). Similar results were observed in T47D cells, where the IC50 of T47D/4-OHT and T47D/Par cells was 6.47 μM and 23.62 μM (Figure 1A). Furthermore, we used flow cytometry to detect changes in apoptosis levels in MCF-7 and T47D cells treated with 4 μM and 8 μM 4-OHT, respectively. We found a significant decline in MCF-7/4-OHT (26.32%) cell apoptosis compared to MCF-7/Par cells (58.16%), and similar results in T47D-7/4-OHT cells (19.63%) and T47D/Par cells (67.71%) (Figure 1B). LDH and Caspase-3 activities presented the same experimental results (Figure 1C and D). Moreover, the data obtained from colony formation and EdU assays indicated that the suppression of 4-OHT (4 μM and 8 μM) on the proliferation of MCF-7 and T47D cells was absent in MCF-7/4-OHT and T47D/4-OHT cells (Figure 1E and F). The above results indicated that MCF-7- and T47D-resistant cell lines were successfully constructed.
Pae Significantly Inhibited 4-OHT Resistance in BC Cells
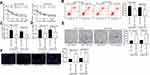
First, to investigate whether Pae could inhibit the proliferative activity of BC cells, we treated parent cells and resistant cells with an increasing concentrations of Pae. The proliferation of MCF-7/Par, MCF-7/4-OHT, T47D/Par, and T47D/4-OHT cells was suppressed in a Pae concentration-dependent manner (Supplementary Figure S1A). Moreover, we found that the proliferation of MCF-7/Par, MCF-7/4-OHT, T47D/Par and T47D/4-OHT cells also decreased significantly after 100 μg/mL Pae treatment (Supplementary Figure S1B, C). Subsequently, we used different concentrations of Pae to treat MCF-7 and T47D-resistant cells to detect the IC50 value of 4-OHT on MCF-7/4-OHT and T47D/4-OHT cells. With increasing Pae concentrations (maximum: 100 μg/mL and minimum: 0 μg/mL), the IC50 of 4-OHT on T47D and MCF-7 cells also progressively decreased (Figure 2A). Consequently, we selected the 100 μg/mL Pae concentration to treat MCF-7/4-OHT and T47D/4-OHT cells for 24 hours, and then used 4 μM and 8 μM 4-OHT to detect apoptosis by flow cytometry, LDH and caspase-3 activities. After Pae treatment, the sensitivity of MCF-7/4-OHT and T47D/4-OHT cells to 4-OHT was significantly improved (Figure 2B–D). In addition, EdU and the colony formation assay were performed to assess the S-phase cell ratio and cell proliferation. Pae treatment also significantly increased 4-OHT inhibition in drug-resistant cells (Figure 2E and F). Furthermore, after Pae treatment in parental MCF-7 and T47D cells, MCF-7 and T47D cells also showed improved sensitivity to 4-OHT (Supplementary Figure S2A-E). Moreover, to further test the safety of Pae, we treated normal human breast epithelial cells MCF10A with an increasing concentrations of Pae, and found that the activity of MCF10A cells had an insignificant decrease, and the number of apoptotic cells did not change significantly after Pae treatment, indicating that Pae had no significant effect on the activity of normal cells (Supplementary Figure S3A-C).
Pae Downregulates miR-15b Expression in 4-OHT-Resistant BC Cells
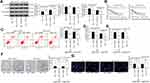
To determine the effects of Pae on drug resistance in BC cells, we used miRNA microarray to analyze the expression of differentially expressed miRNAs in MCF-7/4-OHT cells after Pae treatment. In MCF-7/4-OHT cells, we detected a total of 125 differentially expressed miRNAs, of which 41 were significantly upregulated (log Fold Change >2) and 84 were significantly downregulated (Figure 3A). Subsequently, the expression of the top 10 differentially expressed miRNAs was confirmed by RT-qPCR, which verified the accuracy of microarray data (Figure 3B). To further determine the role of these miRNAs in 4-OHT resistance, we used RT-qPCR to detect the expression of these 10 differentially expressed miRNAs in parental and resistant MCF-7 and T47D cells, which suggested that miR-15b was the most significantly upregulated in MCF-7/4-OHT and T47D/4-OHT cells relative to MCF-7/Par and T47D/Par cells (Supplementary Figure S4). Next, miR-15b expression in parental cells and drug-resistant cells was tested, and the expression of miR-15b in drug-resistant cells was higher than that in parental cells. In contrast, the expression of miR-15b showed a partial decline after further treatment with Pae (Figure 3C). Hence, we hypothesized that Pae promoted sensitivity of drug-resistant cells to 4-OHT by downregulating the expression of miR-15b in cells.
miR-15b Confers BC Cell Resistance to 4-OHT
First, we transfected MCF-7/Par, MCF-7/4-OHT, T47D/Par, and T47D/4-OHT with miR-15b mimic and verified transfection success by RT-qPCR (Supplementary Figure S5A). We then examined the IC50 value of MCF-7/4-OHT and T47D/4-OHT cells treated with Pae and transfected with miR-15b mimic using the CCK-8 assay. The miR-15b mimic significantly inhibited the effects of Pae exerted favoring cell drug sensitivity (Figure 4A). Moreover, when the miR-15b was overexpressed, the cells were treated with Pae for 24 h and then exposed to 4-OHT for 48 h. Flow cytometry, LDH release and caspase-3 activity assessment revealed that the miR-15b mimic reduced the number of apoptotic cells induced by Pae treatment (Figure 4B–D) and restored the cell proliferation repressed by 4-OHT (Figure 4E and F). Similarly, overexpression of miR-15b significantly weakened the promoting effects of Pae on MCF-7 and T47D cell sensitivity to 4-OHT (Supplementary Figure S5B-F).
miR-15b Induced β-Catenin Signaling by Targeting FOXO1 and Transcriptionally Activating CCND1
To further investigate the downstream molecular mechanism of miR-15b underlying BC cell resistance, we analyzed the targeting gene of miR-15b through the Star base. miR-15b targeted the mRNA 3ʹUTR sequence of FOXO1 (Figure 5A). The results of dual-luciferase assays showed that miR-15b mimic could significantly inhibit the luciferase activity of FOXO1 WT (Figure 5B). Furthermore, FOXO1 is significantly downregulated in BC tissues according to the GEPIA website (http://gepia.cancer-pku.cn/index.html) (Figure 5C). RT-qPCR and Western blot results indicated that the FOXO1 expression decreased significantly in BC resistant cells, while it was restored after Pae treatment. However, further transfection of miR-15b mimic inhibited the stimulative role of Pae in FOXO1 expression (Figure 5D and E).
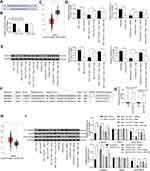 |
Figure 5 miR-15b potentiates the β-catenin signaling by targeting FOXO1 and transcriptionally activating CCND1. (A) Mapping of the binding sites between miR-15b and FOXO1 predicted by StarBase bioinformatics website; (B) the luciferase activity of FOXO1 in cells transfected with miR-15b mimic or mimic control; (C) FOXO1 expression in BC patients revealed by GEPIA website (http://gepia.cancer-pku.cn/index.html); (D) FOXO1 mRNA expression in T47D and MCF-7 cells under different treatments assessed by RT-qPCR; (E) FOXO1 protein expression in T47D and MCF-7 cells under different treatments assessed by Western blot analysis; (F) the promoter sequence binding sites of FOXO1 to CCND1 analyzed by JASPAR bioinformatics website (http://jaspar.genereg.net/); (G) FOXO1 binding sites to CCND1 promoter sequence confirmed by ChIP-qPCR; (H) CCND1 expression in BC patients revealed by GEPIA website (http://gepia.cancer-pku.cn/index.html); (I) the β-catenin and GSK3β expression as well as the extent of GSK3β phosphorylation in MCF-7 and T47D parents and resistant cells. Data are displayed in the form of mean ± SD. All experiments were repeated at least three times. In panel (B) two-way ANOVA along with Tukey’s multiple comparison was applied, while in panel (D, E and I) one-way ANOVA along with Tukey’s multiple comparison was used. *p < 0.05, **p < 0.01. Abbreviations: miR, microRNA; FOXO1, forkhead box O1; CCND1, cyclin D1; SD, standard deviation; ANOVA, analysis of variance. |
We predicted that FOXO1 could bind to the promoter sequence of CCND1 through the JASPAR website (http://jaspar.genereg.net/) (Figure 5F). Thus, we verified the binding relationship between FOXO1 and CCND1 promoter sequence by ChIP-qPCR assays, which supported our prediction (Figure 5G). The GEPIA website indicated that CCND1 expression was significantly higher in BC patients than in normal subjects (Figure 5H). Next, we evaluated the expression of β-catenin signaling-related proteins in MCF-7 and T47D parental and drug-resistant cells by Western blotting analysis. The extent of GSK3β phosphorylation was reduced in drug-resistant cells, whereas β-catenin expression was elevated. After the addition of 100 μg/mL Pae, the phosphorylation of GSK3β was increased, while β-catenin expression was decreased. However, further transfection of miR-15b reversed the inhibition of Pae on β-catenin signaling (Figure 5I).
Silencing of FOXO1 or Overexpression of CCND1 Suppressed the Effects of Pae on 4-OHT Resistance in BC Cells
In order to verify our proposed roles of FOXO1 and CCND1 on BC cell endocrine resistance, we transfected three siRNAs targeting FOXO1 into Pae-treated MCF-7/4-OHT and MCF-7/Par cells and pcDNA3.1-CCND1 overexpression plasmids into T47D/4-OHT and T47D/Par cells. The transfection efficiency verified by RT-qPCR revealed that FOXO1-siRNA-2# had the highest interference efficiency (Figure 6A, Supplementary Figure S6A). We noted that si-FOXO1 or overexpression of CCND1 partially reversed the effects of Pae on 4-OHT sensitivity in both parental and resistant cells (Figure 6B–G, Supplementary Figure S6B-F).
Specific Activation of the β-Catenin Signaling Blocked the Effects of Pae on 4-OHT Resistance in BC Cells
We applied Western blotting analysis to determine the expression of β-catenin signaling-associated proteins in MCF-7/4-OHT cells with FOXO1 knockdown and in T47D/4-OHT cells overexpressing CCND1. Either FOXO1 knockdown or CCND1 overexpression inhibited the phosphorylation level of GSK3β, while either treatment significantly improved the expression of β-catenin (Figure 7A). We subsequently added the β-catenin signaling agonist WAY-262,611 to MCF-7/4-OHT cells and the GSK3β-specific inhibitor LiCl to T47D/4-OHT cells. We determined that activation of β-catenin or inhibition of GSK3β activity interfered with the effects of Pae on 4-OHT-resistant cells (Figure 7B–G)with identical trends in the activities of Caspase-3 and LDH as well as cell proliferation and apoptosis as Figure 6B–G.
Discussion
Tam lowers the 10-year risk of recurrence by almost a half and the risk of death for BC patients to around one-third.17 Even though Tam stands out as a classic example of a targeted drug, a considerable amount of ERα-positive BC patients do not respond adequately to treatment.18 Although the causes of acquiring Tam resistance differ, the principal mechanisms attributed to the resistance remain basically unclear, therefore exploring novel therapeutic targets for Tam-resistant BC is urgently needed.19 In this study, Pae was shown to inhibit endocrine resistance of BC cells to 4-OHT by blocking proliferation and facilitating apoptosis. Furthermore, Pae downregulated miR-15b expression to restore FOXO1 expression, which reduced CCND1 transcription and β-catenin activation.
Pae boasts capable anticancer functions on diverse tumors, including BC.20 In BC, Pae has been validated to disrupt cell proliferation and invasion via the impairment of the Notch-1 signaling pathway.21,22 Moreover, Pae at 20 μmol/L could effectively promote caspase-3 activity and apoptosis in gastric carcinoma cells, yet inhibited cell proliferation through the regulation of miR-124.23 Our oligonucleotide microarray findings demonstrated that miR-15b was the most significantly upregulated miRNA in BC resistant cells and the most downregulated miRNAs by Pae in the same cell line. The overexpression of miR-15b has been confirmed in ovarian cancer24 and gastric cancer.25 Moreover, upregulation of miR-15b has been established to promote the resistance of renal cell carcinoma cells to sunitinib.26 While in tissues collected from patients with highly aggressive BC, the overexpression of miR-15b was also observed.27 Griñán-Lisón et al showed that miR-15b is tightly correlated with radio-resistance, stemness properties, and metastasis in BC, and could be applied as an attractive biomarker in the clinical arena, predominantly in radiotherapy so as to predict and monitor the tumor response to radiation.28 However, whether miR-15b contributes to acquired endocrine resistance in BC and the potential mechanisms involved, remain unknown.
FOXO1, a transcription factor and member of the FOXO subtype of the forkhead/winged-helix family,29 was identified as a target of miR-15b in BC using an online bioinformatics tool and by luciferase reporter assays in our study. FOXO1 has been indicated to augment the efficacy of chemotherapy in digestive malignancies.30 In addition, FOXO1 was negatively regulated by miR-3188 and involved in nasopharyngeal carcinoma cell resistance to 5-FU.31 Furthermore, FOXO1 enhanced the transcriptional expression of Sry-related high mobility box 2 in BC cells overexpressing tribbles homolog 3 (TRB3).32 While the tumor inhibitory role of LINC01355 in BC was achieved through the transcriptional suppression of CCND1 in a FOXO3-dependent manner.33 Consistently, we presented evidence illustrating that miR-15b targeted FOXO1 to transcriptionally activate CCND1 in BC cells. Our rescue experiments evidenced that silencing of FOXO1 or overexpression of CCND1 reversed the suppressive role of Pae on BC cell endocrine resistance to 4-OHT. Similarly, CCND inhibition has been reported to be beneficial for facilitating human glioblastoma cell sensitivity to temozolomide.34
Varma et al suggested that Wnt signaling is activated in BC tissues, and contributed to abnormal expression of β-catenin and subsequent CCND1 upregulation, resulting in the occurrence and development of BC.35 ICG-001, an inhibitor of β-catenin diminished the proliferation of MDA-MB-231 cells by synergizing with cisplatin and doxorubicin.36 β-catenin, the essential factor of the canonical Wnt signaling, was increased in Tam-resistant BC cell lines, while the inhibition of β-catenin by XAV939 partially reversed Tam resistance by its impact on the cell cycle by lowering the expression of β-catenin and β-catenin-modulated downstream targets such as CCND1.37 Moreover, CCND1 has been proposed to accelerate binding between β-catenin and Nanog in the nucleus, thus facilitating cervical carcinoma cell colony formation.38 However, the specific regulatory mechanism of CCND1 on β-catenin signaling was not evaluated herein. Nevertheless, we used WAY-262,611 (a β-catenin signaling agonist) and LiCl (a GSK3β-specific inhibitor) to validate the participation of the β-catenin signaling pathway in Pae-modulated 4-OHT resistance. Both WAY-262,611 and LiCl exerted pro-resistance roles in BC cells in the presence of Pae. Thus, we may conclude that miR-15b interacts with FOXO1 to upregulate the expression of CCND1 in BC cells, leading to the β-catenin signaling pathway overactivation.
Conclusion
This study focused on the role of Pae in the Tam sensitivity of BC and demonstrated that Pae downregulated the expression of miR-15b. Furthermore, miR-15b targeted FOXO1 to promote the transcription of CCND1 in BC through β-catenin signaling. Accordingly, our findings suggest that Pae plays an essential role in the sensitivity of BC cells to Tam and should be further studied to better understand its role in resistance to endocrine therapies in BC. However, the molecular mechanism of the pro-resistance effect of CCND1/β-catenin axis requires further elucidation.
Disclosure
All authors declare that they have no conflicts of interest in this work.
References
1. DeSantis CE, Ma J, Gaudet MM, et al. Breast cancer statistics, 2019. CA Cancer J Clin. 2019;69(6):438–451. doi:10.3322/caac.21583
2. Osborne CK, Schiff R. Mechanisms of endocrine resistance in breast cancer. Annu Rev Med. 2011;62(1):233–247. doi:10.1146/annurev-med-070909-182917
3. Jordan VC. Tamoxifen as the first targeted long-term adjuvant therapy for breast cancer. Endocr Relat Cancer. 2014;21(3):R235–246. doi:10.1530/ERC-14-0092
4. Jager NG, Linn SC, Schellens JH, Beijnen JH. Tailored tamoxifen treatment for breast cancer patients: a perspective. Clin Breast Cancer. 2015;15(4):241–244. doi:10.1016/j.clbc.2015.04.005
5. Xing J, Li J, Fu L, Gai J, Guan J, Li Q. SIRT4 enhances the sensitivity of ER-positive breast cancer to tamoxifen by inhibiting the IL-6/STAT3 signal pathway. Cancer Med. 2019;8(16):7086–7097. doi:10.1002/cam4.2557
6. Zhou YX, Gong XH, Zhang H, Peng C. A review on the pharmacokinetics of paeoniflorin and its anti-inflammatory and immunomodulatory effects. Biomed Pharmacother. 2020;130:110505. doi:10.1016/j.biopha.2020.110505
7. Fang S, Zhu W, Zhang Y, Shu Y, Liu P. Paeoniflorin modulates multidrug resistance of a human gastric cancer cell line via the inhibition of NF-kappaB activation. Mol Med Rep. 2012;5(2):351–356. doi:10.3892/mmr.2011.652
8. Zhang L, Wei W. Anti-inflammatory and immunoregulatory effects of paeoniflorin and total glucosides of paeony. Pharmacol Ther. 2020;207:107452. doi:10.1016/j.pharmthera.2019.107452
9. Chen Y, Zhang R, Zhao W, et al. Paeoniflorin exhibits antitumor effects in nasopharyngeal carcinoma cells through downregulation of NEDD4. Am J Transl Res. 2019;11(12):7579–7590.
10. Gao J, Song L, Xia H, Peng L, Wen Z. 6ʹ-O-galloylpaeoniflorin regulates proliferation and metastasis of non-small cell lung cancer through AMPK/miR-299-5p/ATF2 axis. Respir Res. 2020;21(1):39. doi:10.1186/s12931-020-1277-6
11. Zhou Z, Wang S, Song C, Hu Z. Paeoniflorin prevents hypoxia-induced epithelial-mesenchymal transition in human breast cancer cells. Onco Targets Ther. 2016;9:2511–2518. doi:10.2147/OTT.S102422
12. Wang S, Liu W. Paeoniflorin inhibits proliferation and promotes apoptosis of multiple myeloma cells via its effects on microRNA29b and matrix metalloproteinase2. Mol Med Rep. 2016;14(3):2143–2149. doi:10.3892/mmr.2016.5498
13. Qi LQ, Sun B, Yang BB, Lu S. MiR-15b facilitates breast cancer progression via repressing tumor suppressor PAQR3. Eur Rev Med Pharmacol Sci. 2020;24(2):740–748. doi:10.26355/eurrev_202001_20054
14. Zhao Z, Zhang L, Yao Q, Tao Z. miR-15b regulates cisplatin resistance and metastasis by targeting PEBP4 in human lung adenocarcinoma cells. Cancer Gene Ther. 2015;22(3):108–114. doi:10.1038/cgt.2014.73
15. Guney Eskiler G, Cecener G, Tunca B, Egeli U. An in vitro model for the development of acquired tamoxifen resistance. Cell Biol Toxicol. 2016;32(6):563–581. doi:10.1007/s10565-016-9355-8
16. Wang L, Zhang X, Wang ZY. The Wilms’ tumor suppressor WT1 regulates expression of members of the epidermal growth factor receptor (EGFR) and estrogen receptor in acquired tamoxifen resistance. Anticancer Res. 2010;30(9):3637–3642.
17. Clarke R, Tyson JJ, Dixon JM. Endocrine resistance in breast cancer – an overview and update. Mol Cell Endocrinol. 2015;418(Pt 3):220–234. doi:10.1016/j.mce.2015.09.035
18. Droog M, Beelen K, Linn S, Zwart W. Tamoxifen resistance: from bench to bedside. Eur J Pharmacol. 2013;717(1–3):47–57. doi:10.1016/j.ejphar.2012.11.071
19. Li G, Zhang J, Xu Z, Li Z. ERalpha36 as a potential therapeutic target for tamoxifen-resistant breast cancer cell line through EGFR/ERK signaling pathway. Cancer Manag Res. 2020;12:265–275. doi:10.2147/CMAR.S226410
20. Xiang Y, Zhang Q, Wei S, Huang C, Li Z, Gao Y. Paeoniflorin: a monoterpene glycoside from plants of Paeoniaceae family with diverse anticancer activities. J Pharm Pharmacol. 2020;72(4):483–495. doi:10.1111/jphp.13204
21. Zhang J, Yu K, Han X, et al. Paeoniflorin influences breast cancer cell proliferation and invasion via inhibition of the notch1 signaling pathway. Mol Med Rep. 2018;17(1):1321–1325. doi:10.3892/mmr.2017.8002
22. Zhang Q, Yuan Y, Cui J, Xiao T, Jiang D. Paeoniflorin inhibits proliferation and invasion of breast cancer cells through suppressing notch-1 signaling pathway. Biomed Pharmacother. 2016;78:197–203. doi:10.1016/j.biopha.2016.01.019
23. Zheng YB, Xiao GC, Tong SL, et al. Paeoniflorin inhibits human gastric carcinoma cell proliferation through up-regulation of microRNA-124 and suppression of PI3K/Akt and STAT3 signaling. World J Gastroenterol. 2015;21(23):7197–7207. doi:10.3748/wjg.v21.i23.7197
24. Miao S, Wang J, Xuan L, Liu X. LncRNA TTN-AS1 acts as sponge for miR-15b-5p to regulate FBXW7 expression in ovarian cancer. Biofactors. 2020;46(4):600–607. doi:10.1002/biof.1622
25. Wei S, Peng L, Yang J, et al. Exosomal transfer of miR-15b-3p enhances tumorigenesis and malignant transformation through the DYNLT1/Caspase-3/Caspase-9 signaling pathway in gastric cancer. J Exp Clin Cancer Res. 2020;39(1):32. doi:10.1186/s13046-019-1511-6
26. Lu L, Li Y, Wen H, Feng C. Overexpression of miR-15b promotes resistance to sunitinib in renal cell carcinoma. J Cancer. 2019;10(15):3389–3396. doi:10.7150/jca.31676
27. Kedmi M, Ben-Chetrit N, Korner C, et al. EGF induces microRNAs that target suppressors of cell migration: miR-15b targets MTSS1 in breast cancer. Sci Signal. 2015;8(368):ra29. doi:10.1126/scisignal.2005866
28. Grinan-Lison C, Olivares-Urbano MA, Jimenez G, et al. miRNAs as radio-response biomarkers for breast cancer stem cells. Mol Oncol. 2020;14(3):556–570. doi:10.1002/1878-0261.12635
29. Zou Y, Lin X, Bu J, et al. Timeless-stimulated miR-5188-FOXO1/beta-catenin-c-jun feedback loop promotes stemness via ubiquitination of beta-catenin in breast cancer. Mol Ther. 2020;28(1):313–327. doi:10.1016/j.ymthe.2019.08.015
30. Shi F, Li T, Liu Z, et al. FOXO1: another avenue for treating digestive malignancy? Semin Cancer Biol. 2018;50:124–131. doi:10.1016/j.semcancer.2017.09.009
31. Zhao M, Luo R, Liu Y, et al. miR-3188 regulates nasopharyngeal carcinoma proliferation and chemosensitivity through a FOXO1-modulated positive feedback loop with mTOR-p-PI3K/AKT-c-JUN. Nat Commun. 2016;7(1):11309. doi:10.1038/ncomms11309
32. Yu JM, Sun W, Wang ZH, et al. TRIB3 supports breast cancer stemness by suppressing FOXO1 degradation and enhancing SOX2 transcription. Nat Commun. 2019;10(1):5720. doi:10.1038/s41467-019-13700-6
33. Ai B, Kong X, Wang X, et al. LINC01355 suppresses breast cancer growth through FOXO3-mediated transcriptional repression of CCND1. Cell Death Dis. 2019;10(7):502. doi:10.1038/s41419-019-1741-8
34. Zhang D, Dai D, Zhou M, et al. Inhibition of cyclin D1 expression in human glioblastoma cells is associated with increased temozolomide chemosensitivity. Cell Physiol Biochem. 2018;51(6):2496–2508. doi:10.1159/000495920
35. Varma K, Chauhan A, Bhargava M, Misra V, Srivastava S. Association of different patterns of expression of beta-catenin and cyclin D1 with pathogenesis of breast carcinoma. Indian J Pathol Microbiol. 2020;63(1):13–18. doi:10.4103/IJPM.IJPM_419_19
36. Fatima I, El-Ayachi I, Playa HC, et al. Simultaneous multi-organ metastases from chemo-resistant triple-negative breast cancer are prevented by interfering with WNT-signaling. Cancers (Basel. 2019;11(12):12. doi:10.3390/cancers11122039
37. Fu Y, Wang Z, Luo C, et al. Downregulation of CXXC finger protein 4 leads to a tamoxifen-resistant phenotype in breast cancer cells through activation of the Wnt/beta-catenin pathway. Transl Oncol. 2020;13(2):423–440. doi:10.1016/j.tranon.2019.12.005
38. Xia X, Xia J, Yang H, et al. Baicalein blocked cervical carcinoma cell proliferation by targeting CCND1 via Wnt/beta-catenin signaling pathway. Artif Cells Nanomed Biotechnol. 2019;47(1):2729–2736. doi:10.1080/21691401.2019.1636055
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.