Back to Journals » International Journal of General Medicine » Volume 14
Overexpression of IRF3 Predicts Poor Prognosis in Clear Cell Renal Cell Carcinoma
Authors Wu J, Leng X, Pan Z, Xu L, Zhang H
Received 8 July 2021
Accepted for publication 19 August 2021
Published 16 September 2021 Volume 2021:14 Pages 5675—5692
DOI https://doi.org/10.2147/IJGM.S328225
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Scott Fraser
Jun Wu,1 Xuefeng Leng,1 Zhengbo Pan,2 Linfei Xu,2 Haitao Zhang2
1Department of Urology, Naval 971 Hospital of Chinese People’s Liberation Army, Qingdao City, Shandong Province, People’s Republic of China; 2Department of Urology, Municipal Hospital Affiliated to Taizhou University, Taizhou City, Zhejiang Province, People’s Republic of China
Correspondence: Haitao Zhang
Department of Urology, Municipal Hospital Affiliated to Taizhou University, Taizhou City, Zhejiang Province, People’s Republic of China
Email [email protected]
Background: Growing findings have demonstrated that interferon regulatory transcription factor (IRF) family members are linked to the progression of various cancers. However, the roles of IRFs in clear cell renal cell carcinoma (ccRCC) remain undefined. Herein, we conducted a comprehensive analysis using the bioinformatics method to evaluate the expression patterns, clinical significance, and regulation of IRFs-related mechanisms in patients with ccRCC.
Methods: Data from the Cancer Genome Atlas (TCGA), International Cancer Genome Consortium (ICGA), and Gene Expression Omnibus (GEO) databases were used for investigation comprehensively. Specifically, we carried out a series of analyses to identify the candidate IRF and to explore its potential action mechanisms using the gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses. What is more, we emphatically investigate the association of candidate IRF with tumor immunity in ccRCC through the CIBERSORT algorithm, TIMER and GEPIA databases.
Results: Herein, IRF3 was identified as candidate IRF, which was highly expressed in ccRCC, and its overexpression was significantly associated with worse clinical outcomes and adverse overall survival. Uni- and multi-variate Cox regression analysis demonstrated that IRF3 overexpression was an independent predictor of worse prognosis. Functional enrichment analysis showed that IRF3 might participate in several cancer-related biological processes and signaling pathways, thereby promoting the progression of ccRCC. Additionally, we found that IRF3 was remarkably associated with tumor-infiltrating immune cells (TIICs) and various immune-related genes.
Conclusion: Herein, we identified IRF3 from the IRF gene family members, which could serve as promising prognostic marker and therapeutic target in ccRCC.
Keywords: kidney renal clear cell carcinoma, bioinformatics, interferon regulatory transcription factor, prognosis
Introduction
Renal cell carcinoma (RCC) is a heterogeneous cancer, making treatment responses difficult to predict. Clear cell renal cell carcinoma (ccRCC) is the most common RCC subtype with the dismal prognosis and few treatment choice.1 Moreover, the 5-year survival rates only 10% to 20% of some patients due to the distant metastases.2 Several targets have been found and widely used in clinical practice to treat advanced ccRCC. However, only a minority of patients benefit from it.3 Therefore, searching for novel targeted agents is one of the most critical things in ccRCC.
Interferon regulatory transcription factors (IRFs) constitute a family of transcription factors, including nine gene members IRF1 to IRF9. The IRFs have been demonstrated that play central roles in immunity and oncogenesis.4 In hepatocellular carcinoma (HCC) cells, interferon regulatory factor 1 (IRF1) and IRF2 were reported to upregulate the PD-L1 expression, affecting the effects of immune checkpoint blockade (ICB).5 Rho et al6 found that IRF1 inhibits tumorigenesis and angiogenesis induced by HPV16 E6 in cervical cancer. Chen et al7 have identified IRF2 as a tumor suppressor, which could suppress the invasion and migration by decreasing the expression of MMP-1 in gastric cancer. In hormone-sensitive prostate cancer cells, IRF3 signaling plays a vital role in TLR3-mediated apoptosis by activating intrinsic and extrinsic apoptotic pathways.8 Additionally, Biswas et al9 found the activation of IRF-3/STAT1 signaling pathway in tumor-associated macrophages (TAMs) of murine fibrosarcoma. IRF4 was identified as a haematopoietic cell-restricted transcription factor essential for haematopoietic development and regulation of the immune response.10 An early clinical study has demonstrated that IRF4 could serve as a tumor suppressor in early B-cell development.11 Furthermore, Ortmann et al12 suggested that IRF-4 promoter methylation might be involved in regulating IRF-4 expression in leukemic cells. In human thyroid cancer cells, ectopic IRF5 promoted both the cells’ proliferation rate and the clonogenic potential.13 Additionally, Guo14 have reported that IRF5 was highly expressed in peripheral blood of patients with non-small cell lung cancer. IRF6 was identified as a tumor-suppressive factor, and Xu et al15 found that IRF6 was decreased in highly aggressive breast cancer cell lines, whereas elevated in poorly invasive ones; upregulated the expression of IRF6 could suppress cell proliferation and tumorigenicity, as well as enhance chemosensitivity. What is more, IRF6 was also reported to decrease in gastric cancer, cervical cancer, and melanoma and associated with worse clinical outcomes.16–18 In breast cancer, increased expression of IRF7 has a significant association with the reduced bone metastases and prolonged survival time.19 Moreover, Li et al20 discovered that IRF7 was remarkably reduced in breast cancer, and upregulated IRF7 expression inhibited tumor cell invasion and proliferation. IRF8 could promote the development of dendritic cells and support anti-tumor adaptive immunity by stimulating T cells; however, breast and pancreatic cancer could interrupt the development of IRF8-dependent dendritic cells to evade immune surveillance;21 in addition, Zhang et al22 have confirmed that IRF8 functions as a tumor suppressor in renal cell carcinoma and its promoter methylation is associated with patient poor prognosis. Tian et al23 have demonstrated that the IRF9 axis played a crucial role in the growth of human acute myeloid leukemia; moreover, Brunn et al24 discovered that IRF9 promoted proliferation and migration in lung cancer via modulating of versican.
Based on the above reports, we understood the crucial roles of IRFs in tumor progression and immunity. However, their potential roles in ccRCC remain to be illustrated. Therefore, in the present study, we conducted in-depth and comprehensive analyses to assess the expression patterns, clinical significances, and prognostic values of IRFs in ccRCC. What is more, the candidate IRF was identified, and we focused on its association with tumor immunity.
Materials and Methods
Data Collection
The level 3 data of (HTSeq-FPKM data) and corresponding clinical data of KIRC were retrieved from the TCGA database (https://portal.gdc.cancer.gov/),25 including 530 of the KIRC patients with complete survival data and 72 normal samples. Detailed clinic-pathological information was displayed in Table 1. Data set including 91 cases of KIRC samples and 45 normal samples from the International Cancer Genome Consortium database (ICGC, https://dcc.icgc.org/)26 was used for expression validation. Four gene chips were obtained from the Gene Expression Omnibus database (GEO, https://www.ncbi.nlm.nih.gov/geo/) and were also employed for validation. GSE15641 was from the platform “GPL96[HG-U133A] Affymetrix Human Genome U133A Array”, containing 23 normal renal specimens and 32 ccRCC specimens.27 GSE36895, GSE53757, and GSE66272 datasets were all from the platform GPL570[HG-U133_Plus_2] Affymetrix Human Genome U133 Plus 2.0 Array”, containing 23, 72, and 27 noncancerous tissues, as well as 29, 72 and 27 ccRCC tissues, respectively.28–32
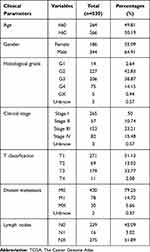 |
Table 1 Characteristics of Clear Cell Renal Cell Carcinoma Patients in the TCGA Database |
Expression Patterns of Interferon Regulatory Transcription Factor (IRF) Family Members in ccRCC
Using the RNA-sequence (RNA-seq) data from the TCGA database, we compared the expression of regulatory transcription factors (IRFs) in ccRCC specimens and corresponding adjacent normal specimens through the Wilcoxon rank-sum test. P-value less than 0.05 was considered statistically significant. Then, the transcriptional data downloaded from the ICGC database was applied for further validation. Heatmaps of all IRFs expressions were drawn using the “heatmap” R package. Correlations between each IRF were estimated using the Spearman coefficient.
Association of IRFs Expression with Overall Survival and Clinicopathological Parameters
According to the median value of each IRF, patients were divided into the high- and low-expression subgroups. Then, the Kaplan–Meier (KM) method and the Log rank test were conducted for overall survival (OS) analysis. Then, the Kaplan–Meier Plotter database (http://www.kmplot.com/)33 was utilized to further estimate the prognostic results. Additionally, the relationship between the expression levels of prognosis-related IRFs and patients’ clinical features was estimated using the Wilcoxon rank-sum test.
Identification of Independent Prognostic Factors
After combining expression data with clinical information, we employed the univariate Cox regression analysis to identify the IRFs and clinicopathologic variables associated with the overall survival of patients with ccRCC. Then, the independent prognostic factors in ccRCC were determined using multivariate Cox regression analysis. The independent prognostic IRF was considered as candidate IRF, which significantly associated with overall survival and clinical outcome of patients with ccRCC.
Expression Was Validated by Various Databases
The transcriptional data of candidate IRF, derived from the GSE15641, GSE36895, GSE53757, and GSE66272, was used to verify the expression. Then, UALCAN database (http://ualcan.path.uab.edu/)34 was employed to assess the protein expression of the candidate IRF. What is more, the representative image was retrieved from the Human Protein Atlas database (HPA, https://www.proteinatlas.org/)35 to further verify the protein expression.
Cbioportal Database
The cBioPortal for Cancer Genomics (http://cbioportal.org) provides a Web resource for exploring, visualizing, and analyzing multidimensional cancer genomics data. In the study, cbioportal database was utilized to perform analysis of genetic alterations. Specifically, a TCGA-KIRC dataset, contains 537 ccRCC cases, was obtained from the cBioPortal database. The “Cancer Type Summary” module was utilized to analyze the mutation types frequencies. The “comparison/survival” module was used to test differences in overall survival between the altered and unaltered groups.
PPI Network Construction
In order to explore the biological functions of IRFs in ccRCC, we construct a protein–protein interaction (PPI) and identify the potential functional partners using the STRING database (active interaction sources: Experiments; minimum required interaction score greater than 0.400). Additionally, the enriched functional enrichment terms were obtained using the “Analysis” module.
Functional Enrichment Analysis
Based on the median expression values of candidate IRF, we divided the tumor groups into high- and low-expression subgroups and screened differentially expressed genes (DEGs) between two subgroups using the “limma” R package. |logFoldChange (logFC) | more than 1 and false discovery rate (FDR) less than 0.05 were considered statistically significant. Then, Metascape (https://metascape.org/), a platform for functional annotation of genes,36 was used to perform functional enrichment analysis to explore these DEGs’ underlying action mechanisms.
Association of Candidate IRFs with Tumor‐infiltrating Immune Cells (TIICs) and Immune-Related Gene Markers
In this study, we used the TIMER database (https://cistrome.shinyapps.io/timer/) to estimate the association of six types of immunocytes with candidate IRF.37 To further validate the TIMER database’s accuracy, the CIBERSORT algorithm was applied for immune cell quantization.38 Specifically, the normalized gene expression data were uploaded to the CIBERSORT portal, run with LM22 signatures and 1000 permutations to output the fractions of immune cells. To improve the accuracy of the results, only samples with a CIBERSORT P < 0.05 were considered eligible. Then, we utilized the Wilcoxon rank-sum test to analyze the differential abundances of TIICs between low- and high-expression subgroups. Finally, to further understand the association between the candidate IRF expression and various infiltrating immune cell types, we investigated the association between IRF expression and related gene markers for each immune cell type using the TIMER and GEPIA databases. These markers were derived from different immune cells, including CD8+ T cell, T cell (general), B cell, Monocyte, TAM, M1 Macrophage, M2 Macrophage, Neutrophil, Natural killer cell, Dendritic cell, Th1, Th2, Tfh, Th17, Treg, and T cell exhaustion. Correlations were estimated using the Spearman coefficient.
Statistical Analysis
Wilcoxon rank-sum test was employed to compare the expression levels of IRFs in ccRCC samples with that in normal renal samples. The Kaplan–Meier (KM) method and the Log rank test were conducted for overall survival (OS) analysis. Uni- and multivariate Cox regression were applied to identify the independent prognostic factors. The Wilcoxon rank-sum test and Fisher’s exact test or chi-square test were used to explore the association between clinicopathological variables and expression levels of IRFs. Correlations were estimated using the Spearman coefficient in this work. P-value less than 0.05 was considered to be statistically significant. All statistical analyses were carried out using R 3.6.1 software.
Results
IRFs Were Dysregulated in KIRC
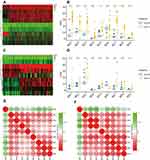
Using the data downloaded from the TCGA database, we plot a heatmap of IRFs expression in ccRCC and noncancerous tissues (Figure 1A). As shown in Figure 1B, all IRFs gene members were dysregulated in KIRC. IRF1/2/3/4/5/7/8/9 were highly expressed in ccRCC, while IRF6 was significantly lower in ccRCC samples than in normal renal samples. Figure 1C reveals the heat map of all IRFs expression in the ICGC database. Additionally, we found the same results using the data from the ICGC database; that is, IRF1/2/3/4/5/7/8/9 was significantly higher in ccRCC specimens by comparison with normal renal tissues, while IRF6 was significantly decreased in ccRCC samples than that in noncancerous samples (Figure 1D). The correlation analysis in the TCGA database showed that IRF1/2/3/4/5/7/8/9 strongly correlates with each other, whereas IRF1/2/3/4/5/7/8/9 were negatively associated with IRF6 (Figure 1E and Supplementary Table 1). The correlation analysis in the ICGC database was presented the same findings (Figure 1F and Supplementary Table 2).
Identification of IRFs Associated with Overall Survival and Clinical Outcomes of ccRCC
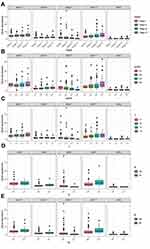
Using the Kaplan–Meier (KM) method, we assessed the association between the expression levels of IRFs with the overall survival of ccRCC. Results showed that high expression of IRF3/5/7/9 (Figure 2C, E, G, I) was associated with a poor prognosis of patients with ccRCC. In contrast, low IRF6 expression indicated a worse prognosis of patients with ccRCC (Figure 2F). However, no statistical significance of IRF1/2/4/8 was found (Figure 2A, B, D, H). The Kaplan–Meier plotter database was used for further validation, and results showed consistent results (Figure 3A–I). Then, we assess the association between prognosis-related IRFs with clinical features of patients with ccRCC. As shown in Figure 4A–E, elevated expression level of IRF5/7 and decreased expression levels of IRF6 indicated the advanced clinical stage, histologic grade, and TNM status, and high expression level of IRF3 was significantly associated with the advanced clinical stage, histologic grade, T, and M status. However, no statistical significance of IRF9 was found. The IRFs associated with overall survival and clinical outcomes of ccRCC, including IRF3/5/6/7, were used for in-depth investigation.
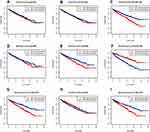 |
Figure 2 Survival Curves of dysregulated IRFs. (A) IRF1; (B) IRF2; (C) IRF3; (D) IRF4; (E) IRF5; (F) IRF6; (G) IRF7; (H) IRF8; (I) IRF9. |
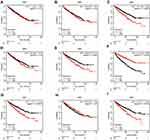 |
Figure 3 Survival Curves of dysregulated IRFs (Kaplan–Meier Plotter). (A) IRF1; (B) IRF2; (C) IRF3; (D) IRF4; (E) IRF5; (F) IRF6; (G) IRF7; (H) IRF8; (I) IRF9. |
Identification of Independent Prognostic Factors
Then, we discovered that IRF3/5/6/7 were significantly associated with overall survival through the univariate Cox regression analysis, including IRF3 (HR = 1.064, p < 0.001), IRF5 (HR = 1.077, p < 0.05), IRF6 (HR = 0.897, p < 0.01) and IRF7 (HR = 1.038, p < 0.05) (Table 2). Additionally, several clinical factors also associated with prognosis, including age (HR = 1.023, p < 0.05), clinical stage (HR = 1.862, p < 0.001), histological grade (HR = 2.242, p < 0.001), T stage (HR = 1.943, p < 0.001), N stage (HR = 2.932, p < 0.01) and M stage (HR = 4.073, p < 0.001). Then, multivariate Cox regression analysis was applied for identification of independent prognostic factors. As presented in Figure 5A, IRF3 (HR = 1.032, p < 0.01), could independently predict overall survival of patient with ccRCC. However, no statistical significance of IRF5/6/7 was found (Figure 5B–D). Thus, IRF3 was defined as a candidate IRF for further exploration.
 |
Table 2 Univariate Cox Proportional Hazards Regression Analyses of IRFs and Clinical Features in ccRCC |
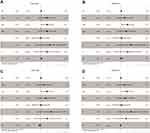 |
Figure 5 Forest plots of the results of multivariate Cox regression analyses of significant prognostic factors: IRF3 (A); IRF5 (B); IRF6 (C); IRF7 (D). Notes: *p < 0.05; **p < 0.01. |
Validation of IRF3 Expression in ccRCC
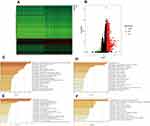
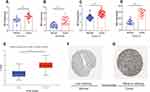
Using the data from the GSE15641, GSE36895, GSE53757, and GSE66272, we further validated the expression levels in ccRCC and normal renal tissues. As illustrated in Figure 6A–D, IRF3 was highly expressed in ccRCC samples compared to in noncancerous samples (both p-value less than 0.001). Protein expression data from the UALCAN database revealed that IRF3 protein was also significantly higher in ccRCC tissues than in adjacent normal tissues (Figure 6E). Representative images revealed that IRF3 showed a “Low” staining in normal renal sample (Figure 6F), whereas displayed a “Medium” staining in ccRCC sample (Figure 6G). Additionally, we estimated the association between IRF3 expression and patients’ clinical features using the Fisher's exact test or chi-square test, and results indicated that IRF3 expression was remarkably correlated with the clinical stage (p < 0.01), histological grade (p < 0.05), and T status (p < 0.01) (Table 3).
 |
Table 3 Association of IRF3 Expression with Clinicopathological Characteristics (Fisher's Exact Test or Chi-Square Test) |
Genetic Mutation Analysis and PPI Network Construction
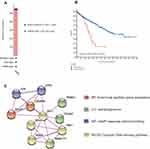
As illustrated in Figure 7A, we found that “mRNA High” was the most frequent genetic alteration overall using the cbioportal database, and the mutation frequency is 5.4% (29 cases); additionally, we observed a case with mutation of “Deep Deletion”. What is more, Kaplan–Meier analysis result show that altered group has a worse overall survival than that of unaltered group (Figure 7B). Next, we construct a PPI network and identify ten functional partners of IRF3 experimentally verified, including JUN, ATF2, EP300, CREBBP, MAVS, TRIM21, IKEKB, TICAM1, TBK1, and TMEM173; further analysis indicates that N-terminal peptidyl-lysine acetylation, autophagosome, cAMP response element binding, and Cytosolic DNA-sensing pathway are the most significant functional enrichment terms of IRF3 and its functional partners (Figure 7C).
Functional Enrichment Analyses Between High and Low IRF3 Expression Subgroups
To explore the regulation of IRF3-related mechanisms. We identified differentially expressed genes (DEGs) between the high and low gene expression subgroups. Heatmaps of DEGs are presented in Figure 8A. Volcano plots of DEGs are shown in Figure 8B. Through the Metascape database, we functionally explored the DEGs using gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analyses. In terms of BP, IRF3 was mainly involved in cellular process, involved in reproduction in multicellular organism, detection of abiotic stimulus, and cilium movement (Figure 8C). In the CC group, IRF3 was significantly involved in cilium, voltage-gated calcium channel complex, and axonemal dynein complex (Figure 8D). As for MF, IRF3 was mainly enriched in channel activity, receptor-ligand activity, and serine-type endopeptidase activity (Figure 8E). KEGG pathway analysis showed that IRF3 was significantly associated with neuroactive ligand–receptor interaction, calcium signaling pathway, and cytokine–cytokine receptor interaction (Figure 8F).
The Relationship of IRF3 with TIICs
Using the TIMER database, we found that the expression levels of IRF3 were positively associated with the immune infiltration levels CD4+ T cells (cor = 0.371, p < 0.001) and Neutrophils (cor = 0.155, p < 0.001) (Figure 9A). Then, we performed CIBERSORT algorithm to further analyze the relationship between IRF3 and TIICs. As illustrated in Figure 9B, the high IRF3 expression group showed higher levels of immune infiltration of Plasma cells (p < 0.01), CD8 + T cells (p < 0.001), T cells follicular helper (p < 0.001), regulatory T cells (Tregs) (p < 0.001), and activated NK cells (p < 0.01), while the low IRF3 expression group exhibited higher levels of immune infiltration of naive B cells (p < 0.001), CD4 memory resting T cells (p < 0.05), Monocytes (p < 0.05), M0 Macrophages (p < 0.05), M2 Macrophages (p < 0.001), resting Dendritic cells (p < 0.05) activated Dendritic cells (p < 0.05), Eosinophils (p < 0.05), and Neutrophils (p < 0.001). These results implied that IRF3 may play a critical role in regulation of the immune microenvironment for ccRCC.
The Association of IRF3 with Immune-Related Genes
To further understand the association between IRF3 expression and various infiltrating immune cell types, we investigated the association between IRF3 expression and relevant gene markers for each immune cell type using the TIMER and GEPIA databases. These markers were derived from different immune cells, including CD8+ T cell, T cell (general), B cell, Monocyte, TAM, M1 Macrophage, M2 Macrophage, Neutrophil, Natural killer cell, Dendritic cell, Th1, Th2, Tfh, Th17, Treg, and T cell exhaustion. Using the TIMER database, we obtained the correlation results after adjustment for tumor purity. As revealed in Table 4, we noticed that IRF3 expression levels were significantly correlated with 37 of the 56 immune cell markers in KIRC. Notably, we found that the expression levels of Treg and T cell exhaustion gene markers showed a strong correlation with IRF3 expression in KIRC. We thus further explored these gene markers with IRF3 expression in KIRC via the GEPIA database, and the results similar to those from the TIMER database (Table 5). The findings investigate that IRF3 might contribute to the immune escape of cancer cells in KIRC.
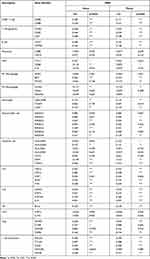 |
Table 4 Correlation Analysis Between IRF3 and Relate Genes and Markers of Immune Cells in TIMER Database |
 |
Table 5 Correlation Analysis Between IRF3 and Relate Genes and Markers of Treg Cells and Exhausted T Cells in GEPIA Database |
Discussion
Clear cell renal cell carcinoma is a highly aggressive malignant tumor with a poor prognosis and increased mortality, and it is an urgent need to find the more promising biomarkers and therapeutic targets for ccRCC patients. Interferon regulatory transcription factor (IRF) family was composed of nine gene members, IRF1 to IRF9. Abundant reports have confirmed their roles in various malignancies. Their clinical implications and mechanisms in ccRCC, however, remain to be elucidated.
Herein, we discovered that all IRFs were abnormally expressed in ccRCC. Among these dysregulated IRFs, IRF1/2/3/4/5/7/8/9 were significantly higher in ccRCC tissues than in noncancerous tissues, whereas IRF6 significantly decreased in ccRCC tissues by comparison with in normal renal tissues. Survival analysis revealed that high expression levels of IRF3/5/7/9 were significantly associated with worse overall survival of patients with ccRCC, while low IRF6 expression indicated an adverse prognosis. Then, the correlation between IRF3/5/6/7/9 expression with clinical features showed that elevated IRF3/5/7 expression and decreased IRF6 expression were remarkably correlated with patient’s clinical outcomes. Additionally, we identified IRF3 as a candidate IRF in ccRCC, which could independently predict ccRCC patients’ OS. Subsequently, Further analyses confirmed that IRF3 was highly expressed in ccRCC and associated with poor clinical outcomes.
IRF3 is a well-characterized signaling transcription factor that plays a crucial role in the innate antiviral response.39 In human host cells, viral DNA and RNA can activate TLRs and promote signal transduction, which subsequently phosphorylates and activates IRF3 through a series of reactions.40 Activated IRF3 forms a dimer and enters the nucleus to regulate type I interferon and interferon-stimulated genes.41 Interestingly, danger signals of self-origin such as DNA damage also promote anti-tumor immunity by activating the IRF3-related signal transduction pathways in cancer.42 This finding suggests that IRF3 might play a crucial role in cancer progression and immunity. In this work, we found that IRF3 was abnormally upregulated in ccRCC and resulted in a worse prognosis and advanced clinical status, indicating that IRF3 might serve as an oncogene in ccRCC. Furthermore, genetic analysis revealed that genetic mutations may play a crucial role in the regulation of IRF3 expression in ccRCC. Then, JUN, ATF2, EP300, CREBBP, MAVS, TRIM21, IKEKB, TICAM1, TBK1, and TMEM173 were found to be functional partners of IRF3. Functional analysis showed that these genes are mainly involved in N-terminal peptidyl-lysine acetylation, autophagosome, cAMP response element binding, and cytosolic DNA-sensing pathway.
Then, we conducted GO and KEGG analyses to further investigate the regulation of IRF3-related mechanisms. Interestingly, we found that IRF3 was significantly enriched in receptor-ligand activity, cytokine–cytokine receptor interaction, channel activity, and calcium signaling pathway. Some ligand/receptor signaling pathways played critical roles in regulating the immune system and cancer progression;43 cytokine–cytokine receptor interaction was a crucial immune signaling pathway and also significantly associated with cancer progression.44 Changes of channel activity and dysregulation of the calcium signaling pathway were essential characteristics of solid tumors associated with invasiveness and uncontrolled malignant behaviors.45 These signaling pathways were related to cancer progression and also were the key points to learn about the pathophysiological mechanism of IRF3 in ccRCC.
Then, we focused on the relationship between IRF3 and the TIICs in ccRCC, and we discovered the differences in infiltrating immune cells between the high and low IRF3 expression subgroups. The differences might be one reason for the different clinical outcomes of the different groups of patients. Another interesting finding of this study is that most of the immune-related gene markers were significantly associated with the IRF3 expression, especially for those gene markers for regulatory T (Treg) cells and exhausted T cells, such as FOXP3, TGFB1, PDCD1, CTLA4, LAG3, and GZMB. Treg cells were involved in tumor development by suppressing anti-tumor immunity; FOXP3 was highly expressed on highly immunosuppressive Treg cells, and the increased levels of FOXP3 within the tumor microenvironment were remarkably correlated with poor prognosis in various cancer patients.46 TGF-β1 bond to the membrane protein GARP and subsequently was activated on the surface of Treg cells, which exerted contact-dependent inhibition on immune cells.47 PDCD1, CTLA4, LAG3, and GZMB were classical immune checkpoints that have the tumor-promoting phenotype during tumor progression and assist the immune escape of tumor cells.48 The results revealed that IRF3 might play critical role in regulating the immunosuppressive phenotypes in the TME of ccRCC.
Herein, we identified IRF3 as a potential prognostic biomarker and therapeutic target for ccRCC through comprehensive bioinformatics analysis. Nevertheless, some limitations should not be disregarded. Larger sample size and sufficient clinical information were required to confirm our findings because of the limited clinical and pathological information. Additionally, the precise mechanism of IRF3 in ccRCC progression and tumor immunity has not been fully illustrated. Therefore, more substantial evidence should be performed to validate our findings.
Ethical Approval and Consent to Participate
The public database mentioned in this study is publicly available for re-analyzing, and no ethical approval was required by the local ethics committees, so that this study does not require the ethics approval.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
There is no funding to report.
Disclosure
The authors declare that they have no competing interests.
References
1. Grignon DJ, Che M. Clear cell renal cell carcinoma. Clin Lab Med. 2005;25(2):305–316. doi:10.1016/j.cll.2005.01.012
2. Hsieh JJ, Purdue MP, Signoretti S, et al. Renal cell carcinoma. Nat Rev Dis Primers. 2017;3:17009.
3. Barata PC, Rini BI. Treatment of renal cell carcinoma: current status and future directions. CA Cancer J Clin. 2017;67(6):507–524. doi:10.3322/caac.21411
4. Tamura T, Yanai H, Savitsky D, et al. The IRF family transcription factors in immunity and oncogenesis. Annu Rev Immunol. 2008;26:535–584. doi:10.1146/annurev.immunol.26.021607.090400
5. Yan Y, Zheng L, Du Q, et al. Interferon regulatory factor 1 (IRF-1) and IRF-2 regulate PD-L1 expression in hepatocellular carcinoma (HCC) cells. Cancer Immunol, Immunother. 2020;69(9):1891–1903. doi:10.1007/s00262-020-02586-9
6. Rho SB, Lee SH, Byun HJ, et al. IRF-1 inhibits angiogenic activity of HPV16 E6 oncoprotein in cervical cancer. Int J Mol Sci. 2020;21(20):7622.
7. Chen YJ, Liang L, Li J, et al. IRF-2 inhibits gastric cancer invasion and migration by down-regulating MMP-1. Dig Dis Sci. 2020;65(1):168–177. doi:10.1007/s10620-019-05739-8
8. Gambara G, Desideri M, Stoppacciaro A, et al. TLR3 engagement induces IRF-3-dependent apoptosis in androgen-sensitive prostate cancer cells and inhibits tumour growth in vivo. J Cell Mol Med. 2015;19(2):327–339. doi:10.1111/jcmm.12379
9. Biswas SK, Gangi L, Paul S, et al. A distinct and unique transcriptional program expressed by tumor-associated macrophages (defective NF-kappaB and enhanced IRF-3/STAT1 activation). Blood. 2006;107(5):2112–2122. doi:10.1182/blood-2005-01-0428
10. Huber M, Lohoff M. IRF4 at the crossroads of effector T-cell fate decision. Eur J Immunol. 2014;44(7):1886–1895. doi:10.1002/eji.201344279
11. Acquaviva J, Chen X, Ren R. IRF-4 functions as a tumor suppressor in early B-cell development. Blood. 2008;112(9):3798–3806. doi:10.1182/blood-2007-10-117838
12. Ortmann CA, Burchert A, Lzle HK, et al. Down-regulation of interferon regulatory factor 4 gene expression in leukemic cells due to hypermethylation of CpG motifs in the promoter region. Nucleic Acids Res. 2005;33(21):6895–6905. doi:10.1093/nar/gki1001
13. Massimino M, Vigneri P, Fallica M, et al. IRF5 promotes the proliferation of human thyroid cancer cells. Mol Cancer. 2012;11(21):1–8. doi:10.1186/1476-4598-11-21.
14. Guo J, Wang X, Wang Y, et al. A promising role of interferon regulatory factor 5 as an early warning biomarker for the development of human non-small cell lung cancer. Lung Cancer. 2019;135:47–55. doi:10.1016/j.lungcan.2019.07.008
15. Xu HF, Huang TJ, Yang Q, et al. Candidate tumor suppressor gene IRF6 is involved in human breast cancer pathogenesis via modulating PI3K-regulatory subunit PIK3R2 expression. Cancer Manag Res. 2019;11:5557–5572. doi:10.2147/CMAR.S203060
16. Li D, Cheng P, Wang J, et al. IRF6 is directly regulated by ZEB1 and ELF3, and predicts a favorable prognosis in gastric cancer. Front Oncol. 2019;9:220. doi:10.3389/fonc.2019.00220
17. Ren Y, Dong J, He P. miR-587 promotes cervical cancer by repressing interferon regulatory factor 6. J Gene Med. 2020;22(11):e3257.
18. Nobeyama Y, Nakagawa H. Silencing of interferon regulatory factor gene 6 in melanoma. PLoS One. 2017;12(9):e0184444. doi:10.1371/journal.pone.0184444
19. Bidwell BN, Slaney CY, Withana NP, et al. Silencing of Irf7 pathways in breast cancer cells promotes bone metastasis through immune escape. Nat Med. 2012;18(8):1224–1231. doi:10.1038/nm.2830
20. Li Y, Huang R, Wang L, et al. microRNA-762 promotes breast cancer cell proliferation and invasion by targeting IRF7 expression. Cell Prolif. 2015;48(6):643–649. doi:10.1111/cpr.12223
21. Meyer MA, Baer JM, Knolhoff BL, et al. Breast and pancreatic cancer interrupt IRF8-dependent dendritic cell development to overcome immune surveillance. Nat Commun. 2018;9(1):1250. doi:10.1038/s41467-018-03600-6
22. Zhang Q, Zhang L, Li L, et al. Interferon regulatory factor 8 functions as a tumor suppressor in renal cell carcinoma and its promoter methylation is associated with patient poor prognosis. Cancer Lett. 2014;354(2):227–234. doi:10.1016/j.canlet.2014.07.040
23. Tian WL, Guo R, Wang F, et al. The IRF9-SIRT1-P53 axis is involved in the growth of human acute myeloid leukemia. Exp Cell Res. 2018;365(2):185–193. doi:10.1016/j.yexcr.2018.02.036
24. Brunn D, Turkowski K. Interferon regulatory factor 9 promotes lung cancer progression via regulation of versican. Cancers. 2021;13(2):208.
25. Hutter C, Zenklusen JC. The cancer genome atlas: creating lasting value beyond its data. Cell. 2018;173(2):283–285. doi:10.1016/j.cell.2018.03.042
26. Zhang J, Bajari R, Andric D. The international cancer genome consortium data portal. Nature. 2019;37(4):367–369.
27. Jones J, Otu H, Spentzos D, et al. Gene signatures of progression and metastasis in renal cell cancer. Clin Cancer Res. 2005;11(16):5730–5739. doi:10.1158/1078-0432.CCR-04-2225
28. A-llopis PE, Liao A, BAP1 loss defines a new class of renal cell carcinoma. Nat Genet. 2012;44(7):751–759. doi:10.1038/ng.2323
29. A-llopis PE, Brugarolas J. Simultaneous isolation of high-quality DNA, RNA, miRNA and proteins from tissues for genomic applications. Nat Protoc. 2013;8(11):2240–2255. doi:10.1038/nprot.2013.141
30. Von Roemeling CA, Radisky DC, Marlow LA, et al. Neuronal pentraxin 2 supports clear cell renal cell carcinoma by activating the AMPA-selective glutamate receptor-4. Cancer Res. 2014;74(17):4796–4810. doi:10.1158/0008-5472.CAN-14-0210
31. Wotschofsky Z, Gummlich L, Liep J, et al. Integrated microRNA and mRNA signature associated with the transition from the locally confined to the metastasized clear cell renal cell carcinoma exemplified by miR-146-5p. PLoS One. 2016;11(2):e0148746. doi:10.1371/journal.pone.0148746
32. Liep J, Kilic E, Meyer HA, et al. Cooperative effect of miR-141-3p and miR-145-5p in the regulation of targets in clear cell renal cell carcinoma. PLoS One. 2016;11(6):e0157801. doi:10.1371/journal.pone.0157801
33. Nagy Á, Munkácsy G, Győrffy B. Pancancer survival analysis of cancer hallmark genes. Sci Rep. 2021;11(1):6047. doi:10.1038/s41598-021-84787-5
34. Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia (New York, NY). 2017;19(8):649–658. doi:10.1016/j.neo.2017.05.002
35. Colwill K, GR SLUND S. A roadmap to generate renewable protein binders to the human proteome. Nat Methods. 2011;8(7):551–558. doi:10.1038/nmeth.1607
36. Zhou Y, Zhou B, Pache L, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nature. 2019;10(1):1523.
37. Li T, Fan J, Wang B, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–e10. doi:10.1158/0008-5472.CAN-17-0307
38. Newman AM, Liu CL, Green MR. Robust enumeration of cell subsets from tissue expression profiles. Nature. 2015;12(5):453–457.
39. O’neill LA, Bowie AG. Sensing and signaling in antiviral innate immunity. Current Biol. 2010;20(7):R328–R333. doi:10.1016/j.cub.2010.01.044
40. Sharma S, Tenoever BR, Grandvaux N, et al. Triggering the interferon antiviral response through an IKK-related pathway. Science. 2003;300(5622):1148–1151. doi:10.1126/science.1081315
41. Shinobu N, Iwamura T, Yoneyama M, et al. Involvement of TIRAP/MAL in signaling for the activation of interferon regulatory factor 3 by lipopolysaccharide. FEBS Lett. 2002;517(1–3):251–256. doi:10.1016/S0014-5793(02)02636-4
42. Liu H, Zhang H, Wu X, et al. Nuclear cGAS suppresses DNA repair and promotes tumorigenesis. Nature. 2018;563(7729):131–136. doi:10.1038/s41586-018-0629-6
43. Kim JW, Cochran JR. Targeting ligand-receptor interactions for development of cancer therapeutics. Curr Opin Chem Biol. 2017;38:62–69. doi:10.1016/j.cbpa.2017.03.010
44. Spangler JB, Moraga I, Mendoza JL, et al. Insights into cytokine-receptor interactions from cytokine engineering. Annu Rev Immunol. 2015;33:139–167. doi:10.1146/annurev-immunol-032713-120211
45. Cui C, Merritt R, Fu L, et al. Targeting calcium signaling in cancer therapy. Acta Pharmaceutica Sinica B. 2017;7(1):3–17. doi:10.1016/j.apsb.2016.11.001
46. Saleh R, Elkord E. FoxP3(+) T regulatory cells in cancer: prognostic biomarkers and therapeutic targets. Cancer Lett. 2020;490:174–185. doi:10.1016/j.canlet.2020.07.022
47. Cuende J, Li Nart S, Dedobbeleer O, et al. Monoclonal antibodies against GARP/TGF-β1 complexes inhibit the immunosuppressive activity of human regulatory T cells in vivo. Sci Transl Med. 2015;7(284):284ra56. doi:10.1126/scitranslmed.aaa1983
48. Abril-rodriguez G, Ribas A. SnapShot: immune checkpoint inhibitors. Cancer Cell. 2017;31(6):848–e1. doi:10.1016/j.ccell.2017.05.010
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.