Back to Journals » OncoTargets and Therapy » Volume 12
Oral Microbiota as Promising Diagnostic Biomarkers for Gastrointestinal Cancer: A Systematic Review
Authors Chen Y, Chen X , Yu H, Zhou H , Xu S
Received 8 September 2019
Accepted for publication 21 November 2019
Published 18 December 2019 Volume 2019:12 Pages 11131—11144
DOI https://doi.org/10.2147/OTT.S230262
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Geoffrey Pietersz
Yanwei Chen,1,* Xuechen Chen,2,* Haixin Yu,2 Haibo Zhou,3 Shu Xu4
1Infection Control Department of Shenzhen Hospital of University of Chinese Academy of Sciences, Shenzhen, People’s Republic of China; 2Medical Faculty Heidelberg, University of Heidelberg, Heidelberg, Germany; 3Institute of Pharmaceutical Analysis, College of Pharmacy, Jinan University, Guangzhou, People’s Republic of China; 4Oncology Department of Shenzhen Hospital of University of Chinese Academy of Sciences, Shenzhen, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Shu Xu
Oncology Department of Shenzhen Hospital of University of Chinese Academy of Sciences, No. 4253 Songbai Road, Guangming District, Shenzhen 518106, People’s Republic of China
Tel +86 18126081801
Email [email protected]
Abstract: Emerging evidence has shown the potential of oral microbiota as a noninvasive diagnostic tool in gastrointestinal (GI) cancer. PubMed, Web of Science, and Embase were systematically searched for eligible studies published until May 31, 2019. Of the 17 included studies published between 2011 and 2019, five kinds of GI cancer, including colorectal cancer (n=6), pancreatic cancer (n=5), gastric cancer (n=4), esophageal cancer (n=2) and liver cancer (n=1), were reported. Generally, the diagnostic performance of the multi-bacteria model for GI cancer was strong with the best area under the receiver operator characteristic curve (AUC) exceeding 0.90, but only one study had a validation phase. Pathogens involved in periodontal disease, such as Porphyromonas gingivalis and Tannerella forsythia, were linked to various kinds of GI cancer. Besides, more oral bacteria significantly differed between cases with upper digestive cancer and healthy controls when compared to colorectal cancer (the most common form of lower digestive cancer), probably indicating a different mechanism due to anatomical and physiological differences in the digestive tract. Oral microbiota changes were associated with risk of various kinds of GI cancer, which could be considered as a potential tool for early prediction and prevention of GI cancer, but validation based on a large population, reproducible protocols for oral microbiota research and oral-gut microbiota transmission patterns are required to be resolved in further studies.
Keywords: gastrointestinal cancer, oral microbiota, detection
Introduction
Even with improvement in health care and advanced treatment means, the outcome of gastrointestinal (GI) cancer is still disappointing. Colorectal cancer (CRC) and gastric cancer (GC), the two most common types of GI cancer, accounted for about 2.8 million new cases and 1.6 million deaths worldwide, respectively, in 2018. Cancers of the pancreas and esophagus were less common, but their poor survival landed them on the list of leading causes of cancer-related deaths.1 Population-based screening programs have dramatically decreased disease burden, yet the participation rates are low, even in the high-risk regions.2–4 The reference standards for diagnosis and screening for GI cancer are mostly based on endoscopies. Lack of awareness in people and the invasiveness and cost-effectiveness issues of endoscopies have been recognized as major barriers to screening.5–7 This fact drives the development of more powerful diagnostic tools with higher compliance to help us detect patients in the early stages.
Periodontal disease or tooth loss has been found to be associated with increased risk of systemic disease, including diabetes,8,9 cardiovascular disease,10,11 oral and GI cancers,12–14 and some other diseases and conditions.15,16 It is well established that bacterial infection resulting from dysbiosis of oral microbiota is the main cause of periodontal disease.17 The chronic inflammation and immune dysregulation resulting from oral bacteria or their products may have systemic effects, which could be the latent factors associated with health and disease.18–20 In addition, intestinal colonization by bacteria of oral origin has been correlated with the development of GI cancer. One recent study by Atarashi et al showed strains of salivary bacteria, Klebsiella spp., could induce chronic intestinal inflammation when colonizing the gut.21 A striking overrepresentation of oral microbes in carcinoma samples might also promote the development of GI cancer through eliciting severe gut inflammation,21 increasing cell proliferation and invasive ability,22 or modulating the tumor-immune microenvironment.23
More than 700 bacterial species inhabit the human oral cavity, including at least 11 bacterial phyla and 70 genera.24 High-throughput genetic-based assays and more sophisticated analytical techniques now make it possible to comprehensively survey the human oral microbiome.25,26 Concerning the potential of the oral microbiome as a noninvasive alternative in population screening and diagnosis of GI cancer, we conducted this systematic review to present the performance of bacteria from the oral cavity to discriminate patients with GI cancer from healthy individuals. Due to the existence of oral-gut bacteria transmission, only bacteria obtained from samples in the oral cavity were considered in this review.
Materials and Methods
This systematic review was conducted following the procedure recommended by the Cochrane Collaboration27 and was reported according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) checklist.28 Since data extracted in this study were obtained from previous studies, ethical approval and patient informed consent were not necessary.
Literature Search Strategies
PubMed, Embase and Web of Science were searched for eligible papers until May 31, 2019. The search terms used were listed in the Appendix and aimed to cover expressions for oral microbiota, GI cancer and detection abilities. Additionally, reference lists from relevant studies and reviews were scanned to identify articles related to the topic. Duplicates were removed, and then abstracts and titles were browsed according to the inclusion and exclusion criteria mentioned below. The full texts of the remaining papers were scrutinized, and finally, those meeting the pre-defined criteria were included in this review.
Inclusion and Exclusion Criteria
The included studies should meet all of the following criteria: (1) should be relevant to the topics; (2) should be performed in humans; (3) should be published as an original study in a peer-reviewed journal; (4) must include microbiota obtained from samples of the oral cavity; (5) have data from cohort or observational studies or randomized control trials, including cases with GI cancer; and (6) should report results for the differences of oral microbiota between patients with GI cancer and healthy controls or the detection abilities of oral microbiota for GI cancer. Studies were not included if they were published as case reports or conference proceedings or in a language other than English language. Papers without full texts were also excluded.
Data Extraction
Two reviewers (X. C. and Y. C.) independently screened the publications and extracted information for each eligible paper. Extracted variables included first author, year of publication, country, types of GI cancer, number of participants, age and sex of participants, sample types, sample collection time and storage temperature, antibiotic use or treatment prior to the sample collection, database used for taxonomy assignment, microbiome measurement methods, and expected outcomes [bacterial abundance or percentage of carriage, odds ratios, specificity, sensitivity, or area under the receiver operator characteristic curve (AUC) values]. Investigators compared selected data, and discrepancies were resolved by consensus.
Quality Assessment in Eligible Studies
Risk of bias and applicability were assessed according to the Quality Assessment of Diagnostic Accuracy Studies-2 (QUADAS-2).29 QUADAS-2 evaluates the risk level of bias and is composed of four basic components: (1) patient selection, (2) index test, (3) reference standard, and (4) flow and timing. Clinical applicability is also assessed for the first three components. The risk of bias and concerns regarding applicability for each study was then rated as “high”, “low”, or “unclear.”
Results
Literature Search Results
A total of 5230 records were obtained in the initial electronic search, including 544 from PubMed, 3266 from Embase, and 1420 from Web of Science. After removal of duplicates (n=496), the titles and abstracts of 4734 studies were screened for relevance. Studies not relevant to the review topics (n=1914), not original (n=756), not human studies (n=692), not in the English language (n=19), or not including oral microbiota (n=714) and GI cancer (n=585) were excluded. The full texts of 54 studies were further read independently. Of those, 12 full-text articles were assessed for eligibility. Combining with the additional studies30–34 identified from reference, 17 studies30–46 were finally included in this review. The detailed selection process is presented in Figure 1.
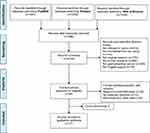 |
Figure 1 PRISMA flow diagram. |
Quality Assessment of Studies
The results for the quality of included studies using the QUADAS-2 tool are presented in Figures S1 and S2. High-risk bias was found in two studies (11.8%), and unclear risk bias was found in five studies (29.4%) in the patient selection domain. One study (5.9%) had high-risk bias in the index test domain, and three studies (17.6%) had unclear risk bias in the flowing and timing domain. For applicability concerns, one study (5.9%) displayed high concerns, and three studies (17.6%) displayed unclear concerns in patient the selection domain. Unclear concern was found in one study (5.9%) in the index test domain.
Study Characteristics
Table 1 describes the characteristics of the included studies published between 2011 and 2019. Of the 17 studies, seven were from the United States of America,30,31,36,39–42 seven were from China,32–34,43–46 and the other three were from France,35 Italy,37 and Ireland,38 respectively. The types of GI cancer included CRC, GC, esophageal adenocarcinoma (EAC), esophageal squamous cell carcinoma (ESCC), liver cancer (LC) and pancreatic cancer (PC). The majority of studies were designed as case–control studies, and only one was a prospective study, in which 1252 postmenopausal females were recruited in the baseline.36 For nested case-control39–41 and prospective cohort designs,36 samples were collected at the time of recruitment and stored for a few years before analysis. The time and temperature for sample storage varied a lot and might have had an impact on the quality of samples and the results of microbiome analysis. Two studies30,33 were designed as screening experiments with GI status confirmed by endoscopies for all participants. The median number (range) of cases and controls was 41 (8, 231) and 54 (10, 461), respectively. The mean age varied greatly between cases and controls in the study (66 vs 52) from Ireland.38
 |
Table 1 Characteristics of Population in the Included Studies |
Oral microbiota composition and variety can be largely affected by the use of antibiotics. Ten studies excluded participants using antibiotics from 1 to 12 weeks prior to the time of sample collection.30,31,33,34,37–39,43,45,46 The other studies did not address antibiotics taken by the participants. Variations in quantity, complexity, and quality of the oral microbiota also occur during cancer treatment. About half of the included studies (n=9) stated that they excluded the participants undergoing cancer therapies before sample collection.30,31,33,34,37,42,43,45,46 Vast variation in most aspects of the studies limited the ability to synthesize the results together or compare the individual study results.
Characteristics of Sample Collection and Measurement
An overview of the sample information is shown in Table 2. Samples were all obtained from the oral cavity, most of which were from saliva,30–33,35,37,42 followed by tongue coating,34,43–46 oral washing,39–41 subgingival plaque,33,36 and oral swab (inside of both cheeks).38 Saliva and subgingival plaque were both included in the study by Sun et al33 Participants with saliva samples in a CRC cohort were extracted from the study by Schmidt et al,35 which included several cohorts with different kinds of diseases (rheumatoid arthritis, type 1 diabetes, and CRC) and samples (both stool and saliva). Most of the frozen samples were stored below −80°C or −70°C, except for two studies that stored samples in a temperature of −20°C before analysis.31,32
 |
Table 2 Characteristics of Sample Collection and Measurement in the Included Studies |
16S rRNA gene sequencing was selected to study oral bacterial phylogeny and taxonomy in 15 studies.30–34,37–46 Among these studies, two37,42 also applied quantitative polymerase chain reaction (qPCR) to quantify the abundance of oral bacteria. Indirect immunofluorescence microscopy was used in the study by Mai et al,36 focusing on the presence of bacteria instead of quantity. Reference databases for the assignment of taxonomy varied a lot between studies. Since two of the included studies35,37 did not compare the abundance difference between groups, additional analysis was conducted and presented in the supplementary materials (Tables S1 and S2).
Bacteria Detection
Table 3 presents the significantly higher bacteria in controls or patients with GI cancer at the genus level, which was found in more than two studies. A total of 18 genera were identified across the included studies. They were classified into five phyla Actinobacteria, Bacteroidetes, Firmicutes, Fusobacteria and Proteobacteria. Parvimonas and Leptotrichia were found significantly lower in controls when compared to various kinds of GI cancer. Members of the phylum Proteobacteria, mainly the Neisseria and Haemophilus genera, were the most common bacteria, each of which were reported by six studies, to be significantly more abundant in controls. There were also inconsistent results in genera, such as Streptococcus, which was lower in patients with CRC39 and LC34 but increased in individuals with EAC,32 GC45 and PC,31 when compared to controls. From the distribution of sample types, most of the significant results were from tongue coating (n=17), followed by saliva (n=13) and oral washing (n=5) (Figure 2A). Among the 18 markers presented in Table 3, only four genera differed between cases with CRC and controls (Figure 2B).
 |
Table 3 Genus (Abundance/Carriage) Found Significantly Different Between Cases and Controls in at Least Two Studies |
 |
Figure 2 Distribution of identified genera in Table 3 according to sample types (A) and cancer types (B). Abbreviations: S, saliva; OW, oral washing; TC, tongue coating; CRC, colorectal cancer; EC, esophageal cancer (combine esophageal adenocarcinoma and esophageal squamous cell carcinoma); GC, gastric cancer; LC, liver cancer; PC, pancreatic cancer. |
Association of Oral Microbiota with GI Cancer
Values of the area under the receiver operator characteristic curve (AUC) were reported in four studies.34,38,42,46 Sun et al only calculated sensitivity and specificity for their model.33 The details are presented in Table 4. The model38 combining 16 oral microbiota operational taxonomic units (OTUs) had a high discriminating ability (AUC=0.90) to detect CRC with 53% sensitivity and 96% specificity. The sensitivity increased to 76% when the oral test was combined with the fecal microbiota test. Sun et al33 developed a score based on the combination of 11 genera, with high sensitivity (97%) and specificity (92%) to discriminate patients with GC from controls. Farrell et al42 identified and validated the performance of two single species, Neisseria elongata and Streptococcus mitis, significantly higher in patients with PC with AUC values equal to 0.66 and 0.68, respectively. The performance (AUC=0.90) increased a lot while including these two species in a single model. Models with the combination of Haemophilus, Porphyromonas, Leptotrichia and Fusobacterium also performed well in discriminating PC from healthy controls with AUC equals to 0.80 but without a validation phase.46 Among these five studies in Table 4, the detection model established by Lu et al only included a single genus in the model, Oribacterium or Fusobacterium, but achieved relatively good performance in detecting LC with an AUC value up to 0.81.34
 |
Table 4 Models for Detection of Gastrointestinal Cancer |
Odds ratios (ORs) reported in more than two studies at the genus or species level are presented in Table 5. Five studies32,36,39–41 reported the ORs, and no additional information, such as sensitivity, specificity or other parameters, indicating discriminating abilities were included in these publications. Periodontal disease-associated species, Porphyromonas gingivalis, Tannerella forsythia and Prevotella intermedia, were correlated with a risk of GI cancer regardless of cancer type.36,39–41 However, no statistically significant results for this association were found in this prospective study, with a model adjusted by age and smoking.36 There were also inconsistent results. Both Peptococcus and Lautropia genera were correlated with a higher risk of CRC,39 but largely lowered the risk of ESCC.32
 |
Table 5 Odds Ratios Reported in More Than Two Studies in Genus or Species Level |
Discussion
To our knowledge, this is the first systematic review that focuses exclusively on the effect of microbiota from samples obtained from the oral cavity on the risk of GI cancer. The available studies suggest that a single oral bacterium has limited ability to detect GI cancer, and multi-bacteria models have better performance (the best AUC > 0.90) but need further validation in large populations. Pathogens involved in periodontal disease are linked to various kinds of GI cancer, such as PC, EA and CRC, which is worthy of attention in further studies. More oral bacteria are significantly higher or lower in cases with upper digestive cancer, when compared to CRC (the most common form of cancer in the lower digestive tract), probably indicating a different mechanism due to anatomical and physiological differences. However, the suggested association should be interpreted with caution, considering that they were mostly reported from a single study without further replication or validation.
Most of bacteria found in the studies were P. gingivalis, T. forsythia and P. intermedia, which have been established as promising indicators for periodontal disease.47–50 However, not all studies found statistically significant results for the association between these bacteria and GI cancer. The genera Oribacterium and Fusobacterium have also been recognized as pathogens associated with periodontal disease,51,52 which discriminated cases with LC from controls with an AUC value up to 0.81.34 Genera, such Leptotrichia, identified as enriched in cases with GI cancer in this review were also reported to be overabundant in subjects with chronic periodontitis.53 All the evidence suggested that oral health is linked tightly to the risk of GI cancer, regardless of type. Both smoking status and alcohol consumption were adjusted in these three studies,39–41 and Fan et al also included age, sex, race and history of diabetes in the regression model. Besides these factors, it is noteworthy that oral hygiene practices, such as tongue brushing,54 and dietary patterns, such as vegetarian, Western, and hunter-gatherers,55,56 could influence the oral ecosystem, which should be taken into account in future studies.
In terms of multi-bacteria models, very high values of AUC were reported ranging from 0.80 to 0.94,34,38,42,46 among which only one study had a validation phase.42 Their model based on the combination of N. elongata and S. mitis, the predominate genera in the oral cavity,57 achieved great performance in the detection of PC with 96.4% sensitivity and 82% specificity.42 It is well established that models developed from the training samples without validation might face an overfitting problem, which can be hard to generalize to other populations. Such models based on a panel of bacteria may nevertheless provide a clue that a combination of oral microbiota rather than a single species could be useful for cancer screening. Combining the noninvasive methods with endoscopic examination, the gold standard for GI screening, to increase screening compliance and performance has become a major topic as of late. For example, in CRC, stool-based tests, such as fecal immunochemical tests (pooled AUC for different cut-offs: 0.69 [0.64, 0.73])58 and gut microbiome detection in fecal samples (AUC: 0.68–0.95),59 have gained more attention recently for their invasiveness and good performance. The performance of oral multi-bacteria models is comparable to those of noninvasive methods and has the potential to replace these more cumbersome stool-based tests. However, its effectiveness has to be confirmed in further studies.
One of the most important factors that could have affected the outcome might be the variation in methodology. Various kinds of samples, including saliva, oral washing, and tongue coating, were selected in the reviewed studies. Saliva is the preferred sampling site to obtain microbial DNA for further sequencing in oral microbiota research. Bacterial profiles were comparable between salivary samples and oral washing samples in a study that included 10 healthy individuals.60 Besides, salivary microbiota tended to reflect the pathogens from other oral sites and to be associated with the risk of oral disease.61 In terms of temporal stability, temporal shifts were relatively small in saliva (both in the short term and long term).62,63 It seems that saliva might serve as an ideal sample source for oral bacteria related to cancer risk. However, previous studies examining the temporal variability of the oral microbiome using healthy participants has been limited by small numbers of volunteers or by being focused on only one collection site, which could hinder our comprehensive understanding of the clinical value of oral microbiota.
Besides DNA sources, study designs and population characteristics are the other major sources of bias. A majority of included studies were designed as case-control and recruited people in the hospital. Among the 17 included studies, 16 studies were designed as case-control and could probably introduce bias into the results.64 Disease might have occurred before the time of sample collection, which means that there is a large possibility that oral microbiota has been influenced by the treatment or other co-morbidities. As regards controls, while most of the studies included healthy controls, only two studies reported that the controls had undergone endoscopies.30,33 In studies in which participants did not undergo endoscopies, associations could have been underestimated by including patients with precancerous lesions as controls. Furthermore, seven of them used data from studies conducted in America, seven in China, and three in Europe. Cultural aspects, customs, and lifestyles of each geographical area are likely explanations for the differences found across studies.55,65 As a result of the heterogeneity in study designs and population characteristics, a number of suggestions could be made for future research to obtain a more realistic evaluation of the detection abilities of oral microbiota for GI cancer.
There were also some limited but interesting results, which might lead to more detailed and in-depth research in this field. Most reported bacteria significantly differed between patients with upper digestive cancer and healthy controls in this review. Studies have shown that microbial communities in the esophagus and stomach are more similar in composition to the oral cavity than cancer in the lower digestive tract, indicating that the upper GI tract is seeded, in part, by oral communities.66,67 Transfer of oral bacteria to the gut is common. Oral microbes reach the stomach through swallowed saliva, nutrients, and drinks. Accounting for the microbial composition similarities and common translocation patterns between the oral cavity and upper digestive tract, changes in oral microbes might be more informative when applied for the detection of upper digestive cancer.
In the study by Farrell et al,42 the combination of oral and gut microbiota increased the sensitivity about 23%, which indicated that the combination of analysis of microbiota both from the oral cavity and feces may assist in better early diagnosis. Several studies68–70 have shown that gut microbiota is a promising and noninvasive screening tool for colorectal precancerous lesions and cancer, while reproducible protocols for studying the human gut microbiome have not been developed. Emerging evidence has suggested the existence of oral-gut bacterial translocation. A recent study by Komiya et al71 explored the microbial association between the gut and oral cavity and found identical strains of Fusobacterium nucleatum in cancer tissues and in the oral cavity of patients with CRC. One of the included studies in this review found evidence for extensive oral-gut bacterial transmission, even in healthy people, and cancer-associated strains of several species enriched in the intestines were from the patients’ oral cavities and not from the environment.35 Finding the transmission and interaction patterns between the gut and oral cavity might provide novel clues for cancer screening or diagnosis through approaches such oral-gut bacterial transmission intervention or control of specific bacteria in the source, the oral cavity.
The key limitation on the interpretability and generalization of these results is the heterogeneity between studies selected for inclusion in our review. Due to the heterogeneity across the reviewed studies, we did not conduct a meta-analysis synthesizing the results of all independent studies. Varying sample selection and handling (storage time, collection methods, storage medium, and storage conditions), DNA extraction methods, targeted hypervariable regions, taxonomical assignments, and statistical analyses are all potential sources for bias and heterogeneity.72–75 In addition, these studies might be underpowered because of the limited sample size. Furthermore, studies from different ethnic and geographical regions limit the generalizability to other populations.
Conclusion
In summary, based on the current evidence, there is considerable interest in the use of oral microbiota to assess the likelihood of developing GI cancer, which needs further validation in a large population. The variation in methodology and relatively few studies for each kind of GI cancer among the studies limited the analyses of this review. Therefore, it is strongly recommended that sample types representing oral microbiota profiles should be determined, and the standard collection protocols should be developed so that the results can be more comparable and conclusions can be drawn on a large basis. Integrating both oral and gut markers may represent a promising approach for risk evaluation of GI cancer. Thus, more work needs to be done to unravel the transmission patterns from the oral cavity to gut and the interaction between them. A better knowledge of mechanisms of how the microbiota communities run and how they are involved in the development of disease can help identify novel preventive approaches for GI cancer by modulating the oral microbiota through oral-gut bacterial transmission intervention or the control of specific bacteria in the source, the oral cavity.
Acknowledgments
This research was funded by National Natural Science Foundation of China, grant number 81773684.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.v68.6
2. White A, Thom TD, White MC, et al. Cancer screening test use – United States, 2015. Morbid Mortal Wkly Rep. 2017;66(8):201–206. doi:10.15585/mmwr.mm6608a1
3. Kabat GC, Matthews CE, Kamensky V, Hollenbeck AR, Rohan TE. Adherence to cancer prevention guidelines and cancer incidence, cancer mortality, and total mortality: a prospective cohort study. Am J Clin Nutr. 2015;101(3):558–569. doi:10.3945/ajcn.114.094854
4. Inadomi JM, Vijan S, Janz NK, et al. Adherence to colorectal cancer screening: a randomized clinical trial of competing strategies. Arch Intern Med. 2012;172(7):575–582. doi:10.1001/archinternmed.2012.332
5. Jones RM, Devers KJ, Kuzel AJ, Woolf SH. Patient-reported barriers to colorectal cancer screening: a mixed-methods analysis. Am J Prevent Med. 2010;38(5):508–516. doi:10.1016/j.amepre.2010.01.021
6. Saumoy M, Schneider Y, Shen N, Kahaleh M, Sharaiha RZ, Shah SC. Cost effectiveness of gastric cancer screening according to race and ethnicity. Gastroenterology. 2018;155(3):648–660. doi:10.1053/j.gastro.2018.05.026
7. Park YM, Cho E, Kang HY, Kim JM. The effectiveness and safety of endoscopic submucosal dissection compared with endoscopic mucosal resection for early gastric cancer: a systematic review and metaanalysis. Surg Endosc. 2011;25(8):2666–2677. doi:10.1007/s00464-011-1627-z
8. Lalla E, Papapanou PN. Diabetes mellitus and periodontitis: a tale of two common interrelated diseases. Nat Rev Endocrinol. 2011;7(12):738–748. doi:10.1038/nrendo.2011.106
9. Chapple ILC, Genco R; for working group 2 of the joint EFP/AAP workshop. Diabetes and periodontal diseases: consensus report of the Joint EFP/AAP Workshop on Periodontitis and Systemic Diseases. J Periodontol. 2013;84(4 Suppl):S106–S112. doi:10.1902/jop.2013.1340011
10. Rydén L, Buhlin K, Ekstrand E, et al. Periodontitis increases the risk of a first myocardial infarction: a report from the PAROKRANK study. Circulation. 2016;133(6):576–583. doi:10.1161/CIRCULATIONAHA.115.020324
11. Tonetti MS, Van Dyke TE; for working group 1 of the joint EFP/AAP workshop. Periodontitis and atherosclerotic cardiovascular disease: consensus report of the Joint EFP/AAP Workshop on Periodontitis and Systemic Diseases. J Periodontol. 2013;84(4 Suppl):S24–S29. doi:10.1902/jop.2013.1340019
12. Michaud DS, Lu JY, Peacock-Villada AY, et al. Periodontal disease assessed using clinical dental measurements and cancer risk in the ARIC study. J Natl Cancer Inst. 2018;110(8):843–854. doi:10.1093/jnci/djx278
13. Momen-Heravi F, Babic A, Tworoger SS, et al. Periodontal disease, tooth loss and colorectal cancer risk: results from the Nurses’ Health Study. Int J Cancer. 2017;140(3):646–652. doi:10.1002/ijc.v140.3
14. Maisonneuve P, Amar S, Lowenfels AB. Periodontal disease, edentulism, and pancreatic cancer: a meta-analysis. Ann Oncol. 2017;28(5):985–995. doi:10.1093/annonc/mdx019
15. Winning L, Linden GJ. Periodontitis and systemic disease. BDJ Team. 2015;2:15163. doi:10.1038/bdjteam.2015.163
16. Loos BG. Periodontal medicine: work in progress! J Clin Periodontol. 2016;43(6):470–471. doi:10.1111/jcpe.12550
17. Hajishengallis G, Darveau RP, Curtis MA. The keystone-pathogen hypothesis. Nat Rev Microbiol. 2012;10(10):717–725. doi:10.1038/nrmicro2873
18. Ahn J, Chen CY, Hayes RB. Oral microbiome and oral and gastrointestinal cancer risk. Cancer Causes Control. 2012;23(3):399–404. doi:10.1007/s10552-011-9892-7
19. Karpinski TM. Role of oral microbiota in cancer development. Microorganisms. 2019;7(1):20. doi:10.3390/microorganisms7010020
20. Mascitti M, Togni L, Troiano G, et al. Beyond head and neck cancer: the relationship between oral microbiota and tumour development in distant organs. Front Cell Infect Microbiol. 2019;9:232. doi:10.3389/fcimb.2019.00232
21. Atarashi K, Suda W, Luo C, et al. Ectopic colonization of oral bacteria in the intestine drives TH1 cell induction and inflammation. Science. 2017;358(6361):359–365. doi:10.1126/science.aan4526
22. Yang Y, Weng W, Peng J, et al. Fusobacterium nucleatum increases proliferation of colorectal cancer cells and tumor development in mice by activating TLR4 signaling to NFκB, upregulating expression of microRNA-21. Gastroenterology. 2017;152(4):851–866.e824. doi:10.1053/j.gastro.2016.11.018
23. Kostic AD, Chun E, Robertson L, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. 2013;14(2):207–215. doi:10.1016/j.chom.2013.07.007
24. Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43(11):5721–5732. doi:10.1128/JCM.43.11.5721-5732.2005
25. Pozhitkov AE, Beikler T, Flemmig T, Noble PA. High-throughput methods for analysis of the human oral microbiome. Periodontol 2000. 2011;55(1):70–86. doi:10.1111/prd.2010.55.issue-1
26. Goodwin S, McPherson JD, McCombie WR. Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet. 2016;17(6):333–351. doi:10.1038/nrg.2016.49
27. Higgins JP, Green S, editors. Cochrane Handbook for Systematic Reviews of Interventions. Hoboken, NJ: The Cochrane Collaboration/Wiley-Blackwell; 2011.
28. Moher D, Liberati A, Tetzlaff J, Altman DG; for the PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. BMJ. 2009;339:b2535. doi:10.1136/bmj.b2535
29. Whiting PF, Rutjes AW, Westwood ME, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155(8):529–536. doi:10.7326/0003-4819-155-8-201110180-00009
30. Torres PJ, Fletcher EM, Gibbons SM, Bouvet M, Doran KS, Kelley ST. Characterization of the salivary microbiome in patients with pancreatic cancer. Peer J. 2015;3:e1373. doi:10.7717/peerj.1373
31. Olson SH, Satagopan J, Xu Y, et al. The oral microbiota in patients with pancreatic cancer, patients with IPMNs, and controls: a pilot study. Cancer Causes Control. 2017;28(9):959–969. doi:10.1007/s10552-017-0933-8
32. Chen X, Winckler B, Lu M, et al. Oral microbiota and risk for esophageal squamous cell carcinoma in a high-risk area of China. PLoS One. 2015;10(12):e0143603. doi:10.1371/journal.pone.0143603
33. Sun JH, Li XL, Yin J, Li YH, Hou BX, Zhang Z. A screening method for gastric cancer by oral microbiome detection. Oncol Rep. 2018;39(5):2217–2224. doi:10.3892/or.2018.6286
34. Lu HF, Ren ZG, Li A, et al. Deep sequencing reveals microbiota dysbiosis of tongue coat in patients with liver carcinoma. Sci Rep. 2016;6:33142. doi:10.1038/srep33142
35. Schmidt TSB, Hayward MR, Coelho LP, et al. Extensive transmission of microbes along the gastrointestinal tract. Elife. 2019;8:e42693. doi:10.7554/eLife.42693
36. Mai X, Genco RJ, LaMonte MJ, et al. Periodontal pathogens and risk of incident cancer in postmenopausal females: the buffalo osteoperio study. J Periondontol. 2016;87(3):257–267. doi:10.1902/jop.2015.150433
37. Russo E, Bacci G, Chiellini C, et al. Preliminary comparison of oral and intestinal human microbiota in patients with colorectal cancer: a pilot study. Front Microbiol. 2018;8:2699. doi:10.3389/fmicb.2017.02699
38. Flemer B, Warren RD, Barrett MP, et al. The oral microbiota in colorectal cancer is distinctive and predictive. Gut. 2018;67(8):1454–1463. doi:10.1136/gutjnl-2017-314814
39. Yang YH, Cai QY, Shu XO, et al. Prospective study of oral microbiome and colorectal cancer risk in low-income and African American populations. Int J Cancer. 2019;144(10):2381–2389. doi:10.1002/ijc.31941
40. Peters BA, Wu J, Pei ZH, et al. Oral microbiome composition reflects prospective risk for esophageal cancers. Cancer Res. 2017;77(23):6777–6787. doi:10.1158/0008-5472.CAN-17-1296
41. Fan XZ, Alekseyenko AV, Wu J, et al. Human oral microbiome and prospective risk for pancreatic cancer: a population-based nested case-control study. Gut. 2018;67(1):120–127. doi:10.1136/gutjnl-2016-312580
42. Farrell JJ, Zhang L, Zhou H, et al. Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut. 2012;61(4):582–588. doi:10.1136/gutjnl-2011-300784
43. Hu J, Han S, Chen Y, Ji Z. Variations of tongue coating microbiota in patients with gastric cancer. Biomed Res Int. 2015;2015:173729. doi:10.1155/2015/173729
44. Han S, Yang X, Qi Q, et al. Potential screening and early diagnosis method for cancer: tongue diagnosis. Int J Oncol. 2016;48(6):2257–2264. doi:10.3892/ijo.2016.3466
45. Wu J, Xu S, Xiang C, et al. Tongue coating microbiota community and risk effect on gastric cancer. J Cancer. 2018;9(21):4039–4048. doi:10.7150/jca.25280
46. Lu HF, Ren ZG, Li A, et al. Tongue coating microbiome data distinguish patients with pancreatic head cancer from healthy controls. J Oral Microbiol. 2019;11(1):1563409. doi:10.1080/20002297.2018.1563409
47. Slots J, Ting M. Actinobacillus actinomycetemcomitans and Porphyromonas gingivalis in human periodontal disease: occurrence and treatment. Periodontol 2000. 1999;20:82–121. doi:10.1111/j.1600-0757.1999.tb00159.x
48. Socransky SS, Smith C, Haffajee AD. Subgingival microbial profiles in refractory periodontal disease. J Clin Periodontol. 2002;29(3):260–268. doi:10.1034/j.1600-051x.2002.290313.x
49. Amaliya A, Laine ML, Delanghe JR, Loos BG, Van Wijk AJ, Van der Velden U. Java project on periodontal diseases: periodontal bone loss in relation to environmental and systemic conditions. J Clin Periodontol. 2015;42(4):325–332. doi:10.1111/jcpe.2015.42.issue-4
50. Socransky SS, Haffajee AD, Cugini MA, Smith C, Kent RL
51. Kinney JS, Morelli T, Braun T, et al. Saliva/pathogen biomarker signatures and periodontal disease progression. J Dent Res. 2011;90(6):752–758. doi:10.1177/0022034511399908
52. Huang S, Li R, Zeng XW, et al. Predictive modeling of gingivitis severity and susceptibility via oral microbiota. ISME J. 2014;8(9):1768–1780. doi:10.1038/ismej.2014.32
53. Chen C, Hemme C, Beleno J, et al. Oral microbiota of periodontal health and disease and their changes after nonsurgical periodontal therapy. ISME J. 2018;12:1210–1224. doi:10.1038/s41396-017-0037-1
54. Matsui M, Chosa N, Shimoyama Y, Minami K, Kimura S, Kishi M. Effects of tongue cleaning on bacterial flora in tongue coating and dental plaque: a crossover study. BMC Oral Health. 2014;14:4. doi:10.1186/1472-6831-14-4
55. Lassalle F, Spagnoletti M, Fumagalli M, et al. Oral microbiomes from hunter-gatherers and traditional farmers reveal shifts in commensal balance and pathogen load linked to diet. Mol Ecol. 2018;27(1):182–195. doi:10.1111/mec.2018.27.issue-1
56. David R. Diet and oral microbiota go hand in hand. Nat Rev Microbio. 2013;11:223.doi:10.1038/nrmicro3000
57. Mason MR, Chambers S, Dabdoub SM, Thikkurissy S, Kumar PS. Characterizing oral microbial communities across dentition states and colonization niches. Microbiome. 2018;6(1):67. doi:10.1186/s40168-018-0443-2
58. Aniwan S, Ek T R, Pongprasobchai S, et al. The optimal cut-off level of the fecal immunochemical test for colorectal cancer screening in a country with limited colonoscopy resources: a multi-center study from Thailand. Asian Pac J Cancer Prev. 2017;18(2):405–412.
59. Amitay EL, Krilaviciute A, Brenner H. Systematic review: gut microbiota in fecal samples and detection of colorectal neoplasms. Gut Microbes. 2018;9(4):293–307. doi:10.1080/19490976.2018.1445957
60. Fan XZ, Peters BA, Min D, Ahn J, Hayes RB. Comparison of the oral microbiome in mouthwash and whole saliva samples. PLoS One. 2018;13(4):e0194729. doi:10.1371/journal.pone.0194729
61. Slots J, Slots H. Bacterial and viral pathogens in saliva: disease relationship and infectious risk. Periodontol 2000. 2011;55(1):48–69. doi:10.1111/prd.2010.55.issue-1
62. Hall MW, Singh N, Ng KF, et al. Inter-personal diversity and temporal dynamics of dental, tongue, and salivary microbiota in the healthy oral cavity. NPJ Biofilms Microbiomes. 2017;3:2. doi:10.1038/s41522-016-0011-0
63. Belstrøm D, Holmstrup P, Bardow A, Kokaras A, Fiehn NE, Pasteur BJ. Temporal stability of the salivary microbiota in oral health. PLoS One. 2016;11(1):e0147472. doi:10.1371/journal.pone.0147472
64. Moss SM. Case-control studies of screening. Int J Epidemiol. 1991;20(1):1–6. doi:10.1093/ije/20.1.1
65. Adler CJ, Dobney K, Weyrich LS, et al. Sequencing ancient calcified dental plaque shows changes in oral microbiota with dietary shifts of the Neolithic and Industrial revolutions. Nat Genet. 2013;45:450–455. doi:10.1038/ng.2536
66. Tsuda A, Suda W, Morita H, et al. Influence of proton-pump inhibitors on the luminal microbiota in the gastrointestinal tract. Clin Transl Gastroenterol. 2015;6:e89. doi:10.1038/ctg.2015.20
67. Gall A, Fero J, Mccoy C, et al. Bacterial composition of the human upper gastrointestinal tract microbiome is dynamic and associated with genomic instability in a Barrett’s esophagus cohort. PLoS One. 2015;10(6):e0129055. doi:10.1371/journal.pone.0129055
68. Baxter NT, Ruffin MT
69. Liang QY, Chiu J, Chen YX, et al. Fecal bacteria act as novel biomarkers for noninvasive diagnosis of colorectal cancer. Clin Cancer Res. 2017;23(8):2061–2070. doi:10.1158/1078-0432.CCR-16-1599
70. Zackular JP, Rogers MA, Ruffin MT
71. Komiya Y, Shimomura Y, Higurashi T, et al. Patients with colorectal cancer have identical strains of Fusobacterium nucleatum in their colorectal cancer and oral cavity. Gut. 2019;68(7):1335–1337. doi:10.1136/gutjnl-2018-316661
72. Goodrich JK, Di Rienzi SC, Poole AC, et al. Conducting a microbiome study. Cell. 2014;158(2):250–262. doi:10.1016/j.cell.2014.06.037
73. Luo T, Srinivasan U, Ramadugu K, et al. Effects of specimen collection methodologies and storage conditions on the short-term stability of oral microbiome taxonomy. Appl Environ Microbiol. 2016;82(18):5519–5529. doi:10.1128/AEM.01132-16
74. Teng F, Nair SS, Zhu PF, et al. Impact of DNA extraction method and targeted 16S-rRNA hypervariable region on oral microbiota profiling. Sci Rep. 2018;8:16321. doi:10.1038/s41598-018-34294-x
75. Koren O, Knights D, Gonzalez A, et al. A guide to enterotypes across the human body: meta-analysis of microbial community structures in human microbiome datasets. PLoS Comput Biol. 2013;9(1):e1002863. doi:10.1371/journal.pcbi.1002863
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
