Back to Journals » Infection and Drug Resistance » Volume 13
Optimized Production of the Allylamine Antifungal “Terbinafine” by Lysinibacillus Isolate MK212927 Using Response Surface Methodology
Authors El-Sayed SE, El-Housseiny GS , Abdelaziz NA , El-Ansary MR , Aboshanab KM
Received 20 June 2020
Accepted for publication 15 September 2020
Published 15 October 2020 Volume 2020:13 Pages 3613—3626
DOI https://doi.org/10.2147/IDR.S267590
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Professor Suresh Antony
Sayed E El-Sayed,1 Ghadir S El-Housseiny,2 Neveen A Abdelaziz,1 Mona R El-Ansary,3 Khaled M Aboshanab2
1Department of Microbiology and Immunology, Faculty of Pharmacy, Ahram Canadian University, Giza, Egypt; 2Department of Microbiology and Immunology, Faculty of Pharmacy, Ain Shams University, Cairo 11566, Egypt; 3Department of Biochemistry, Modern University for Technology and Information (MTI), Cairo, Egypt
Correspondence: Khaled M Aboshanab Department of Microbiology and Immunology, Faculty of Pharmacy
Ain Shams University, Organization of African Unity St., Abbassia, Cairo 11566, Egypt
Tel +011201001547800
Fax +01120224051107
Email [email protected]
Purpose: We aimed to optimize the factors affecting the production of the allylamine antifungal, terbinafine, by Lysinibacillus isolate MK212927, a natural producer of this broad-spectrum fungicidal compound.
Methods: We employed a central composite design to optimize the five most important variables influencing the production of terbinafine which were carbon source, nitrogen source, temperature, pH and agitation.
Results: The optimum conditions were found to be starch 5 g/L, ammonium chloride 5 g/L, temperature 32°C, agitation 150 rpm and pH 7. The actual response (inhibition zone diameter) was highly comparable to the value predicted by the model, indicating a valid model. Using the standard calibration curve of terbinafine, the optimized conditions resulted in an increase in the antifungal metabolite production (terbinafine) by about 1.6-fold (1814.662 μg/mL compared to 1165.550 μg/mL under standardized conditions).
Conclusion: This is the first report, to the best of our knowledge, on optimized production of terbinafine by Lysinibacillus species. Hence, these findings may be useful as baseline data for scaling up the production of terbinafine from a natural microbial source.
Keywords: fungicidal, optimization, central composite design, terbinafine
Introduction
Invasive fungal infections (IFIs) have elevated consistently with the increase in at-risk immunocompromised patients, patients suffering from malignancies and those with severe illnesses. Most of these infections are caused by Candida and Aspergillus species.1,2 Twenty percent of the infections reported in intensive care unit (ICU) were attributed to invasive candidiasis.3 Oropharyngeal candidiasis remains the prominent opportunistic infection in HIV-positive patients with 95% of the cases caused by C. albicans.4 Increased resistance to antifungal agents such as azoles both in C. albicans and non-albicans Candida spp. is extensively reported.5 Aspergillus causes infections ranging from invasive aspergillosis (IA) to extreme complications. Huge elevation in drug-resistant isolates of Aspergillus species poses an extra danger to humans.6 The current pharmaceutical practices visualize fungi/yeast as a difficult challenge to common drugs. Not only do these drugs have adverse effects but also present regimes of antifungals are becoming gradually inefficient. Moreover, the capability of Candida sp. to form drug-resistant biofilm is a driving factor for fungal pathogenicity.7 Indeed, there is an increased need for new drugs to treat fungal diseases in humans.
Many bacterial species are recognized to produce antifungal agents. Among the soil bacteria is Lysinibacillus (gram-positive spore-forming rods) which was previously studied and known for its ability to produce antifungal metabolites against plant pathogens.8–10 Recent studies reported that Lysinibacillus fusiformis displayed strong antifungal activity against aflatoxigenic Aspergillus flavus and Aspergillus parasiticus.11 Lysinibacillus isolate MK212927 is a novel bacterial isolate recovered through a screening program and shown to produce a metabolite with promising antifungal activities against human fungal pathogens. Upon characterization and structure elucidation, it was found to be the allylamine antifungal (terbinafine). The present study aimed to optimize the various factors affecting terbinafine production. Since many factors might affect the complexity of experimental results, experimental design may be applied to perform data optimization.12 Response surface methodology (RSM), a superior form of experimental design, can be defined as a hybrid of both mathematical and statistical methods for designing, developing, analysis and optimization of multifaceted processes.13 It simultaneously identifies the result of altering different factors and their influence on the required outcome of a procedure.14,15 This method is more efficient than the traditional optimization method of one-factor-at-a-time since it requires fewer trials, has the ability to investigate the interactive effects among the factors and avoids wasting unnecessary time and materials.16 Response surface methodology is a very beneficial tool to optimize numerous parameters, to find relativeness among the factors, to find the best combination of parameters and for prediction of responses. Several types of designs are available for the optimization of important fermentation parameters, and central composite design (CCD) is one of the most useful ones.17 Therefore, the goal of this study was to optimize the various physical and environmental parameters using RSM for maximum production of the terbinafine metabolite produced by Lysinibacillus isolate MK212927.
Materials and Methods
Microorganisms
Lysinibacillus isolate MK212927 is a novel bacterial isolate proven to be a natural producer of the allylamine antifungal terbinafine with broad-spectrum fungicidal activities. Lysinibacillus isolate MK212927 (NCBI GenBank accession number, MK212927.1) was deposited in the Culture Collection Ain Shams University (CCASU) belonging to the WDCM (strain number, CCASU-MK212927). The target strain used for testing the antifungal activity of the produced metabolite was the standard strain Candida albicans ATCC 10231.
Fermentative Production of the Antifungal Compound
The seed culture was prepared by transferring a loopful from a new culture of Lysinibacillus isolate MK212927 into a flask containing 50 mL trypticase soy broth and incubated at 250 rpm and 30°C for 15–18 hours. One milliliter of the culture obtained was then centrifuged for 10 minutes at 16,000 rpm using a micro centrifuge (Centurion Scientific-K240R) and the obtained cells were washed once with 1 mL sterile normal saline and resuspended in the production media. The basal minimal medium used for the production of the antifungal metabolite was prepared according to Singh et al18 and consisted of: 5g/L glucose, 6 g/L Na2HPO4, 3g/L K2HPO4, 1 g/L NH4Cl, 0.5 g/L NaCl, 0.12 g/L MgSO4, 0.2 g/L CaCl2 and distilled H2O to 1 liter. KOH pellets were used to adjust the pH to 7 before sterilization.
Production was carried out by inoculating 30 mL aliquots of the basal medium (in 250 mL Erlenmeyer flasks) by the seed culture to give a final count of 1x 107cfu/mL and incubating in a shaking incubator at 150 rpm and 30°C.
Estimation of Antifungal Concentration
Antifungal concentrations were quantified from the linear equation of a standard calibration curve of terbinafine (Lamisil®). To obtain a standard calibration curve we plotted known concentrations of terbinafine (Lamisil®) obtained from NOVARTIS Pharmaceutical Company (Cairo, Egypt) versus the corresponding inhibition zone diameters obtained against C. albicans ATCC 10231.
Studying Different Factors Affecting Antifungal Production
Time Course of Antifungal Production
To obtain the time required for maximum antifungal production, seven Erlenmeyer flasks (250 mL) were prepared as mentioned above. One flask was sampled on daily basis for determination of maximum level of the antifungal metabolite produced over an incubation period of 7 days. In brief, one mL of the fermentation broth was centrifuged at 16,000 rpm and the supernatant (100 µL/well) was tested using the agar cup-plate method against Candida albicans ATCC 10231 (0D= 0.1) in sabouraud`s dextrose agar. The diameters of inhibition zones against Candida albicans ATCC 10231 were determined after 24 hours of incubation at 28°C. Each experiment was performed in triplicate and the mean values of the inhibition zone diameters were recorded.
Effect of Different Media Components
Screening of essential media components for enhanced antifungal production was done following “one-factor-at-a-time” approach. The carbon source (glucose) present in basal medium was substituted with 4 other carbon sources (glycerol, starch, lactose or sucrose) all at 5 g/L. Likewise; the nitrogen source (ammonium chloride) present in basal medium was replaced with 4 other nitrogen sources (casein, urea, yeast or peptone) all at 1 g/L, using the best carbon source chosen. For each source, three experiments were carried out and the mean values of corresponding inhibition zones were recorded.18 The carbon and nitrogen sources which proved to yield maximum antifungal production were selected for further optimization experiments.
Optimization of Antifungal Production Using RSM
RSM was employed using a face-centered central composite design to represent the interactions among the various nutritional and physical variables in a cuboidal experimental space.19 The five variables included carbon source (coded as variable A), nitrogen source (coded as variable B), pH (coded as variable C), temperature (coded as variable D) and agitation rate (coded as variable E). The variables were tested at three levels, as shown in Table 1 and a total of 32 runs were carried out (Table 2). Production was carried out as mentioned previously, and after 3 days of incubation, one response value, the inhibition zone diameter was measured accordingly. The design of experiments was performed using Design Expert® v. 7.0 (Design Expert® Software, Stat-Ease Inc., Statistics Made Easy, Minneapolis, MN, USA).
 |
Table 1 Variables and Levels Used in RSM Central Composite Design for Optimization of Antifungal Production |
 |
Table 2 Central Composite Design Runs for the 5 Different Factors Tested Showing the Observed and Predicted Responses |
Confirmation of the RSM Optimization Results
We exploited the numerical optimization function in the software to obtain a set of optimal culture conditions and a new experiment was performed using these optimal conditions. Afterwards, we compared the production of the antifungal metabolite using these conditions versus the standardized ones.
Statistical Analysis
All the values plotted are the means of triplicate experiments. The statistical validation of the experimental data was performed by the analysis of variance (ANOVA), while the significance of the model was evaluated by the lack of fit test.
Results
Estimation of Antifungal Concentration
Concentrations of the antifungal metabolite were estimated using the calibration curve of standard terbinafine (Figure 1). By means of the developed linear equation: Y= 0.0169 X + 5.0022, the concentration of the antifungal compound (Y) expressed as µg/mL was calculated, where (X) is inhibition zone diameter in mm.
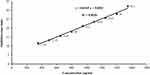 |
Figure 1 Standard calibration curve of terbinafine concentration versus inhibition zones obtained against Candida albicans ATCC 10231. |
Time Course of Antifungal Production by Lysinibacillus Isolate MK212927
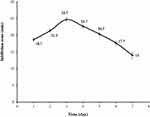
As shown in Figure 2, the maximum antifungal production, indicated by a mean of the inhibition zones of 24.7 mm, was obtained after 3 days of incubation and therefore three days was selected as the optimum incubation time in subsequent experiments.
Effect of Growth Media Composition on the Metabolite Production
We initially followed “one-factor-at-a-time” approach to screen essential medium components required for optimized antifungal production. The results of using different carbon and nitrogen sources on the antifungal activity of Lysinibacillus isolate MK212927 (inhibition zones) against C. albicans are plotted in Figure 3. Enlarged inhibition zones indicated enhancement in the antifungal metabolite production by Lysinibacillus isolate MK212927. As shown in the figure, maximum production was obtained by using starch (5 g/L) and ammonium chloride (1 g/L) as the carbon and nitrogen sources, respectively, and therefore, these were selected for further studies.
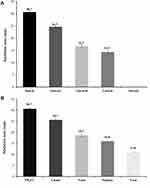 |
Figure 3 Effect of (A) different Carbon sources and (B) Nitrogen sources on the antifungal activity of Lysinibacillus isolate MK212927 against Candida albicans ATCC 10231. |
Optimization of Antifungal Production Using RSM
To optimize the antifungal production conditions by Lysinibacillus isolate MK212927, we undertook a face-centered central composite design. Table 2 shows the actual values of the tested factors, the design, the observed and predicted results. The relationship between the factors was determined and a second-order polynomial equation was fitted by the software using the data obtained from the 32 experiments. The resulting RSM model equation proposed by the software was as follows:
Inhibition zone = −510.17 + 6.08 x Starch + 2.22 x ammonium chloride + 102.59 x pH + 9.92 x Temperature - 0.03 x Agitation + 0.69 x Starch x pH - 0.24 x Starch x Temperature - 0.088 x ammonium chloride x Temperature - 9.17E - 003 x ammonium chloride x Agitation + 0.20 x pH x Temperature + 0.01 x pH x Agitation + 0.68 x ammonium chloride2 - 8.29 x pH2 - 0.15 x Temperature2
To determine the appropriateness of the model we used ANOVA and calculated the P-value which represents the significance of each factor on the antifungal production. ANOVA results are shown in Table 3. The F-value of the model was 34.2, which shows the model is significant. A P-value of less than 0.05 indicates that the model terms are significant. In the current case, the linear terms A, B, C, D and E were significant. The interactive terms AC, AD, BE, CD and the quadratic terms B2, C2and D2 were also significant model terms. Also, the coefficient of variation was 8.46; this low value denotes the good reliability of the experimental values. The fit of the model was expressed with the coefficient of determination R2 that was 0.97, suggesting that 97% of variability in response could be explained by the model. Moreover, a fair agreement was noted between the predicted R-squared (0.833) and the adjusted R-Squared (0.938) values. Lastly, adequate signal was confirmed by the adequate precision ratio which was equal to 25.31. Thus, this model was adequate to navigate the design space.
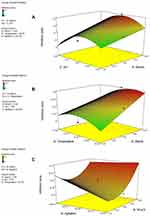
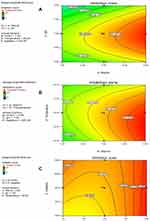
The three-dimensional response surface plots (3D plots) (Figure 4) and contour plots (Figure 5) demonstrate the effect of interactive experimental conditions. By using the numerical optimization function together with these surface plots, the optimum conditions that enhanced terbinafine production were found to be starch and ammonium chloride both at 5 g/L, agitation rate of 150 rpm, a pH of 7 and a temperature of 32°C.
We constructed the following model diagnostic plots to endorse our models and validate our findings:
The normal probability plot of residuals is an important diagnostic method for the model (Figure 6A). It revealed no signs of problems in the data and the normality in the error term could be proved by linear patterns.
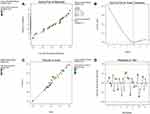 |
Figure 6 Model diagnostic plots. (A) The normal probability plot of residuals, (B) Box Cox plot, (C) Predicted versus actual values plot and (D) Residuals versus Run number plot. |
The Box Cox plot is a valuable tool for the determination of the most suitable power transformation to be applied to response data. Here, the generated plot showed no need for additional transformation and the model was sufficient where the present lambda (λ =1) is within the 95% confidence range of the optimum lambda value (Figure 6B).
The predicted versus actual values plot showed a satisfactory agreement between the outputs from the models and the actual data. This plot also confirmed the proximity of the actual values to the predicted values as the points showed a distribution that was near to the straight line (Figure 6C).
The residuals versus Run number plot showed that the points were randomly scattered around zero (Figure 6D) which indicated that the model fits the data.
Confirmatory Experiment Using Optimal Conditions
An inhibition zone of 35.67 mm was obtained by using the optimum levels of the conditions proposed by the model; temperature 32°C, agitation 150 rpm, Starch 5 g/L, ammonium chloride 5 g/L and pH 7. This resultant inhibition zone is highly comparable to the value predicted by the model, which was 38 mm, indicating a valid model. Using the calibration curve of terbinafine, we concluded that this inhibition zone corresponded to an antifungal concentration of 1814.662 µg/mL. That is to say, the optimal conditions used resulted in a 1.6-fold increase in antifungal production by the Lysinibacillus isolate MK212927 as compared to that produced by using the standardized conditions (1165.550 µg/mL) as shown in Figure 7.
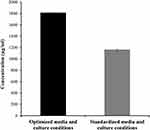 |
Figure 7 Comparison of antifungal production by Lysinibacillus isolate MK212927 using optimized and standardized conditions. |
Discussion
A crucial cause of morbidity and mortality are the invasive, life-threatening fungal infections, especially for immunocompromised patients. The therapeutic options available for the treatment of these infections is rather restricted when compared with those accessible to treat bacterial infections.20 The currently available synthetic antifungal agents have many side effects which minimize their use and safety profile. For example, Itraconazole is a broad-spectrum synthetic triazole whose most common reported adverse effects are gastrointestinal disturbances, hepatic toxicity, and cardiac effects. Thus, it is prohibited to use with patients receiving certain medications.21 Antifungal agents from natural sources can overcome the side effects and the risks associated with synthetic ones. The use of natural antifungals has proven to be an alternate and effective way of treating oral candidiasis in previous reports.22 Therefore, the present investigation aimed to achieve optimal conditions for improved production of the fungicidal metabolite, terbinafine, produced by Lysinibacillus sp MK212927.
We combined the traditional one-factor-at-a-time approach and experimental design to optimize the conditions for antifungal production. The one-factor-at-a-time experiments were used to select the best medium components for production, while the response surface methodology, which included the central composite design was used to build a model to select the optimum levels of the chosen medium components, select optimum environmental conditions and study the interactions across the several factors with antifungal metabolite production as the response. Carbon and nitrogen sources are the key nutritional components in the medium for bacterial cultivation; bacteria need both of these sources for their metabolite production.15 It was deemed necessary to examine the effect of varying the nitrogen and carbon sources on the production of the antifungal metabolite. Previous studies stated that various bacteria showed an increase in secondary metabolite production utilizing a complex carbon source like starch.23 On the other hand, simple sources of carbon caused catabolic repression leading to reduced levels of metabolites.24 In agreement with these reports, maximum production of the antifungal metabolite was obtained by using starch (5 g/L) as the carbon source. Ammonium chloride (1 g/L) was proven to be the suitable nitrogen source; this is in accordance with reports from earlier studies.24
The classical methods used for optimizing fermentation conditions, in particular physical and nutritional parameters, are slow, tedious and expensive; also, they neglect the combined interactions among different variables.25 The statistical methodologies are favorable because they are fast, reliable, help understand the effect of the nutrients at varying concentrations and result in a substantial reduction in total number of experiments therefore saving time, glassware, chemicals and manpower.26 These statistical methods based on experimental design, such as the RSM, allowed the evaluation of the simultaneous effects of different variables including environmental conditions and culture medium components on the antifungal metabolite production.18 It was important to use such methods for a better perception of the likely interactions between evaluated variables.27,28 RSM has been intensely used in previous studies for the optimization of the fermentation conditions and the medium composition to maximize the production of antifungal metabolites by different soil bacteria.29 For example, the central composite design of RSM was used to improve the antifungal activity of a metabolite produced by Streptomyces diastatochromogenes KX852460 by optimizing the nutrient components in submerged fermentation in a previous study.17 Moreover, in a recent study, the co-culture medium components and fermentation conditions of Trichoderma atroviride SG3403 and Bacillus subtilis 22 were optimized using RSM for the production of antifungal metabolites.30
Using central composite design, a total of 32 runs were conducted to study the interactions between the selected 5 variables (starch, ammonium chloride, pH, temperature and agitation) and to determine how they influenced the metabolite yield. To investigate the significance of the design, we used ANOVA which provides a better understanding about the sources of variation.31 The Fisher’s test compares between the mean square values of these two sources of variance; the model and residual error.32 The attained F-value, 34.2 proved that the model developed in this study was significant (P-value <0.0001) and could be used to explain the variation in the response under investigation (antifungal production). Alternatively, the lack of fitness should be non-significant for the model to fit well with the experimental design. In our results, the non-significant lack of fit (P-value >0.05) indicated that the model was appropriate for the current study. Normally, an R2 value exceeding 0.9 reflects great correlation for any regression model.33 Hence, the obtained R2 value (0.97) reveals a very good fit between the predicted and the observed responses. The ability of the model to precisely predict a response value can be expressed as the predicted R2 which should be in good agreement with the adjusted R2; difference between both should not exceed 0.2.31 Accordingly, our results displayed a fair agreement between the adjusted and predicted R2 values (0.833 and 0.938, respectively). The ratio of adequate precision, a measure of signal-to-noise ratio, we obtained was 25.309. This ratio demonstrated a satisfactory model discrimination since it was greater than 4.34 In addition, the low value of coefficient of variation (CV=8.46) indicated that these experiments were precise and reliable. Usually, the CV value and the reliability of the experiment are inversely proportional.35
Moreover, the significance of each factor was reported by P-value. The smaller the P-value and larger the sum of squares, the more significant the corresponding factor is.36,37 As shown in Table 3, starch (A) was the most significant factor affecting antifungal production, followed by ammonium chloride (B), pH (C), temperature (D) and finally agitation (E). Temperature and pH are vital physiological parameters influencing the metabolic pathways, hence the generation of various metabolites.38 Metabolite production from our Lysinibacillus species was strongly dependent upon pH and temperature and showed maximum production at 32°C and pH 7. Previous studies observed optimum yield of metabolites near or slightly lower than optimum growth temperature and around the neutral pH (pH 6.8–7.4).39,40
Moreover, the interaction between starch and temperature (AD) was the most significant, followed by the interaction between starch and pH (AC) and ammonium chloride and agitation (BE), then interaction between pH and temperature (CD). The other interactions were not significant. A significant interaction between 2 factors means the effect of one factor depends on the level of the other.41 As shown in the equation representing the model, all factors and interactions had positive synergistic effects on antifungal yield except agitation (E) and the interactions between starch and temperature (AD), ammonium chloride and temperature (BD) and ammonium chloride and agitation (BE), which had negative antagonistic effects on antifungal production.
For a better understanding of the variables’ effects on the antifungal metabolite production and their interactions, the predicted model was presented as 3D-plots and contour plots. The 3D plots can directly reflect the effect of different levels of the factors on the response and therefore pinpoint their optimum levels. The 2D contour plots, on the other hand, can reflect the significance of the interaction between any two factors: an elliptical or saddle contour suggests that the interaction between the two factors is significant, whereas a circular contour suggests that the interaction is weak.36,37,42 From the 3D figures and numerical optimization function, the recommended optimum conditions for maximum antifungal metabolite yield were determined and attempted experimentally. In addition, the contour plots obtained for AD, AC and BE were elliptical in shape indicating significant interaction between these variables.
In summary, the model has significant F-value, high R2 value, a low coefficient of variance and an insignificant lack-of-fit. These results point towards the accuracy of the model in predicting the antifungal metabolite yield. Moreover, the model diagnostic plots generated also verified the soundness of the model constructed in our study.
A successful validation of the model was performed as the actual results obtained experimentally were in reasonable agreement with the values predicted by the model for the maximum metabolite yield. The optimal antifungal metabolite concentration produced by Lysinibacillus isolate MK212927 was 1814.662 µg/mL which is equivalent to about 1.6-fold increase in production as compared to that produced by using the standardized conditions (1165.550µg/mL).
Conclusion
In the present work, we demonstrated the impact of multiple factors on the production of terbinafine antifungal metabolite produced by Lysinibacillus isolate MK212927 and defined the optimum levels required for maximum production. Central composite factorial design in RSM was found to be very effective in determining the interaction between these significant variables and resulted in about 1.6-fold increase in productivity. Hence, these results may serve as baseline data for scaling up the production of terbinafine antifungal metabolite by the respective isolate as a natural source for its production.
Author Contributions
All authors made substantial contributions to conception and design, acquisition of data, or analysis and interpretation of data; took part in drafting the article or revising it critically for important intellectual content; agreed to submit to the current journal; gave final approval of the version to be published; and agree to be accountable for all aspects of the work.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Disclosure
The authors declare that they have no conflicts of interest.
References
1. Pagano L, Mayor S. Invasive fungal infections in high-risk patients: report from TIMM-8 2017. Future Sci OA. 2018;4(6):FSO307–FSO307. doi:10.4155/fsoa-2018-0019
2. Stohs E, Zimmer A. An approach to suspected invasive fungal infection in patients with hematologic malignancy and HCT recipients with persistent neutropenic fever despite mold-active prophylaxis. Curr Fungal Infect Rep. 2020;14(1):89–98. doi:10.1007/s12281-020-00375-6
3. Lamoth F, Lockhart SR, Berkow EL, Calandra T. Changes in the epidemiological landscape of invasive candidiasis. J Antimicrob Chemother. 2018;73(suppl_1):i4–i13. doi:10.1093/jac/dkx444
4. Quindós G, Gil-Alonso S, Marcos-Arias C, et al. Therapeutic tools for oral candidiasis: current and new antifungal drugs. Med Oral Patol Oral Cir Bucal. 2019;24(2):e172–e180. doi:10.4317/medoral.22978
5. Barbosa F, Pinto E, Kijjoa A, Pinto M, Sousa E. Targeting antimicrobial drug resistance with marine natural products. Int J Antimicrob Agents. 2020;106005. doi:10.1016/j.ijantimicag.2020.106005
6. Andes DR, Safdar N, Baddley JW, et al. Impact of treatment strategy on outcomes in patients with candidemia and other forms of invasive candidiasis: a patient-level quantitative review of randomized trials. Clin Infect Dis. 2012;54(8):1110–1122. doi:10.1093/cid/cis021
7. Alves R, Barata-Antunes C, Casal M, Brown AJP, Van Dijck P, Paiva S. Adapting to survive: how Candida overcomes host-imposed constraints during human colonization. PLoS Pathog. 2020;16(5):e1008478. doi:10.1371/journal.ppat.1008478
8. Nam Y-D, Seo M-J, Lim S-I, Lee S-Y. Genome sequence of Lysinibacillus boronitolerans F1182, isolated from a traditional Korean fermented soybean product. J Bacteriol. 2012;194(21):5988. doi:10.1128/JB.01485-12.
9. Che J, Liu B, Liu G, Chen Q, Lan J. Volatile organic compounds produced by Lysinibacillus sp. FJAT-4748 possess antifungal activity against Colletotrichum acutatum. Biocontrol Sci Technol. 2017;27:1–14. doi:10.1080/09583157.2017.1397600
10. Naureen Z, Rehman NU, Hussain H, et al. Exploring the potentials of Lysinibacillus sphaericus ZA9 for plant growth promotion and biocontrol activities against phytopathogenic fungi. Front Microbiol. 2017;8(1477). doi:10.3389/fmicb.2017.01477
11. Khadka S, Adhikari S, Thapa A, et al. Screening and optimization of newly isolated thermotolerant Lysinibacillus fusiformis strain SK for protease and antifungal activity. Curr Microbiol. 2020;77(8):1558–1568. doi:10.1007/s00284-020-01976-7
12. Ziegel ER. Statistics and chemometrics for analytical chemistry. Technometrics. 2004;46(4):498–499. doi:10.1198/tech.2004.s248
13. Elibol M. Optimization of medium composition for actinorhodin production by Streptomyces coelicolor A3 (2) with response surface methodology. Process Bioch. 2004;39(9):1057–1062. doi:10.1016/S0032-9592(03)00232-2
14. Ganesan G, Velayudhan SS, David JSR. Statistical optimization of medium constituents and conditions for improved antimicrobial compound production by marine Streptomyces sp. JRG-04. Arch Biol Sci. 2017;69(4):723–731. doi:10.2298/ABS170224019G
15. Sa-Uth C, Rattanasena P, Chandrapatya A, Bussaman P. Modification of medium composition for enhancing the production of antifungal activity from Xenorhabdus stockiae PB09 by using response surface methodology. Int J Microbiol. 2018;2018:3965851. doi:10.1155/2018/3965851
16. Bezerra MA, Santelli RE, Oliveira EP, Villar LS, Escaleira LA. Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta. 2008;76(5):965–977. doi:10.1016/j.talanta.2008.05.019
17. Ahsan T, Chen J, Wu Y, Irfan M. Application of response surface methodology for optimization of medium components for the production of secondary metabolites by Streptomyces diastatochromogenes KX852460. AMB Express. 2017;7(1):96. doi:10.1186/s13568-017-0388-z
18. Singh RK, Kumar DP, Solanki MK, et al. Optimization of media components for chitinase production by chickpea rhizosphere associated Lysinibacillus fusiformis B‐CM18. J Basic Microbiol. 2013;53(5):451–460. doi:10.1002/jobm.201100590
19. Zhang Y, Zhang J. Optimization of headspace solid-phase microextraction for analysis of ethyl carbamate in alcoholic beverages using a face-centered cube central composite design. Anal Chim Acta. 2008;627(2):212–218. doi:10.1016/j.aca.2008.08.014
20. Roemer T, Krysan DJ. Antifungal drug development: challenges, unmet clinical needs, and new approaches. Cold Spring Harb Perspect Med. 2014;4(5):a019703. doi:10.1101/cshperspect.a019703.
21. Girois SB, Chapuis F, Decullier E, Revol BGP. Adverse effects of antifungal therapies in invasive fungal infections: review and meta-analysis. Eur J Clin Microbiol Infect Dis. 2006;25(2):138. doi:10.1007/s10096-005-0080-0
22. Gowhar O, Singh N, Sultan S, Ain T, Mewara A, Shah I. Natural herbs as alternative to synthetic antifungal drugs – the future challenging therapy. Br Biomed Bull. 2015;3:440–452.
23. Zhao Y, Liang Y, Liu L, Cheng J, Yuan Y. Medium optimization for antifungal active substance production from Streptomyces lydicus using response surface methodology. Trans Tianjin Univ. 2017;23(1):78–86. doi:10.1007/s12209-016-0023-0
24. Fukuda T, Matsumoto A, Takahashi Y, Tomoda H, Ōmura S. Phenatic acids A and B, new potentiators of antifungal miconazole activity produced by Streptomyces sp. K03-0132. J Antibiot (Tokyo). 2005;58(4):252. doi:10.1038/ja.2005.29
25. Li M, Penner G, Hernandez‐Sanabria E, Oba M, Guan L. Effects of sampling location and time, and host animal on assessment of bacterial diversity and fermentation parameters in the bovine rumen. J Appl Microbiol. 2009;107(6):1924–1934. doi:10.1111/j.1365-2672.2009.04376.x
26. Vaidya R, Macmil S, Vyas P, Chhatpar H. The novel method for isolating chitinolytic bacteria and its application in screening for hyperchitinase producing mutant of Alcaligenes xylosoxydans. Lett Appl Microbiol. 2003;36(3):129–134. doi:10.1046/j.1472-765x.2003.01274.x
27. Nawani N, Kapadnis B. Optimization of chitinase production using statistics based experimental designs. Process Bioch. 2005;40(2):651–660. doi:10.1016/j.procbio.2004.01.048
28. Mechri S, Kriaa M, Berrouina MBE, et al. Optimized production and characterization of a detergent-stable protease from Lysinibacillus fusiformis C250R. Int J Biol Macromol. 2017;101:383–397. doi:10.1016/j.ijbiomac.2017.03.051
29. Souagui Y, Tritsch D, Grosdemange-Billiard C, Kecha M. Optimization of antifungal production by an alkaliphilic and halotolerant actinomycete, Streptomyces sp. SY-BS5, using response surface methodology. J Mycol Med. 2015;25(2):108–115. doi:10.1016/j.mycmed.2014.12.004
30. Li T, Tang J, Karuppiah V, Li Y, Xu N, Chen J. Co-culture of Trichoderma atroviride SG3403 and Bacillus subtilis 22 improves the production of antifungal secondary metabolites. Biol Control. 2020;140:104122. doi:10.1016/j.biocontrol.2019.104122.
31. Mourabet M, El Rhilassi A, El Boujaady H, Bennani-Ziatni M, Taitai A. Use of response surface methodology for optimization of fluoride adsorption in an aqueous solution by brushite. Arab J Chem. 2017;10:S3292–S3302. doi:10.1016/j.arabjc.2013.12.028
32. Kasiri MB, Modirshahla N, Mansouri H. Decolorization of organic dye solution by ozonation; optimization with response surface methodology. Int J Ind Chem. 2013;4(1):3. doi:10.1186/2228-5547-4-3
33. Chen X-C, Bai J-X, Cao J-M, et al. Medium optimization for the production of cyclic adenosine 3′, 5′-monophosphate by Microbacterium sp. no. 205 using response surface methodology. Bioresour Technol. 2009;100(2):919–924. doi:10.1016/j.biortech.2008.07.062
34. Abdel-Hafez SM, Hathout RM, Sammour OA. Towards better modeling of chitosan nanoparticles production: screening different factors and comparing two experimental designs. Int J Biol Macromol. 2014;64:334–340. doi:10.1016/j.ijbiomac.2013.11.041
35. El-Housseiny GS, Aboulwafa MM, Aboshanab KA, Hassouna NAH. Optimization of rhamnolipid production by P. aeruginosa Isolate P6. J Surfactants Deterg. 2016;19(5):943–955. doi:10.1007/s11743-016-1845-4
36. Wang Z-W, Liu X-L. Medium optimization for antifungal active substances production from a newly isolated Paenibacillus sp. using response surface methodology. Bioresour Technol. 2008;99(17):8245–8251. doi:10.1016/j.biortech.2008.03.039
37. Zhou Q, Ding L, Zhu Y, Zhong M, Yang C. Process parameters optimization of gallic acid removal from water by MIEX resin based on response surface methodology. Processes. 2020;8(3):273. doi:10.3390/pr8030273
38. Yun TY, Feng RJ, Zhou DB, et al. Optimization of fermentation conditions through response surface methodology for enhanced antibacterial metabolite production by Streptomyces sp. 1–14 from cassava rhizosphere. PLoS One. 2018;13(11):e0206497. doi:10.1371/journal.pone.0206497
39. Aasen I, Møretrø T, Katla T, Axelsson L, Storrø I. Influence of complex nutrients, temperature and pH on bacteriocin production by Lactobacillus sakei CCUG 42687. Appl Microbiol Biotechnol. 2000;53(2):159–166. doi:10.1007/s002530050003
40. Iyapparaj P, Maruthiah T, Ramasubburayan R, et al. Optimization of bacteriocin production by Lactobacillus sp. MSU3IR against shrimp bacterial pathogens. Aquat Biosyst. 2013;9(1):1–10. doi:10.1186/2046-9063-9-12
41. Rajendran D, Venkatachalam P, Ramakrishnan J. Response surface methodology: optimisation of antifungal bioemulsifier from novel Bacillus thuringiensis. Sci World J. 2014;2014:423289. doi:10.1155/2014/423289
42. Wang L, Zhang B, Han J, Zheng Y, Li J, Shan A. Optimization of hydrolysis condition of blood meal by Bacillus subtilis with response surface methodology. Int Biodeterior Biodegradation. 2015;104:112–117. doi:10.1016/j.ibiod.2015.05.018
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.