Back to Journals » Cancer Management and Research » Volume 12
OIP5-AS1/miR-137/ZNF217 Axis Promotes Malignant Behaviors in Epithelial Ovarian Cancer
Authors Guo L , Chen J, Liu D, Liu L
Received 7 November 2019
Accepted for publication 8 June 2020
Published 3 August 2020 Volume 2020:12 Pages 6707—6717
DOI https://doi.org/10.2147/CMAR.S237726
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Antonella D'Anneo
Linlin Guo, Jiabao Chen, Dong Liu, Lili Liu
Department of Obstetrics and Gynecology, First Affiliated Hospital of Jinzhou Medical University, Jinnzhou, Liaoning, People’s Republic of China
Correspondence: Lili Liu
Department of Obstetrics and Gynecology, First Affiliated Hospital of Jinzhou Medical University, No. 2, Section 5, Renming Street, Jinnzhou, Liaoning, People’s Republic of China
Email [email protected]
Background: Long non-coding RNAs (lncRNAs) have been reported to play crucial regulatory roles in cellular activities and are associated with the carcinogenesis of various diseases. OIP5-AS1, as a novel lncRNA, function in epithelial ovarian cancer (EOC) still remains unclear.
Material and Methods: qRT-PCR and Western blot analyses were performed to measure relevant expression, as needed. A series of functional experiments were performed to determine the role of OIP5-AS1 in EOC cells. Luciferase report, RNA pull down and RIP assays were performed to testify the interaction between relevant RNAs.
Results: We found that OIP5-AS1 was significantly overexpressed in EOC. Knockdown of OIP5-AS1 inhibited cell proliferation, migration, invasion and epithelial–mesenchymal transition (EMT) process, yet facilitated apoptosis in vitro. OIP5-AS1 functioned as a competing endogenous RNA (ceRNA) to elevate ZNF217 expression through sponging miR-137. Furthermore, miR-137 inhibition and ZNF217 upregulation can reverse the effects of silencing OIP5-AS1 on the cellular activities of ovarian cancer cells. Also, depleted OIP5-AS1 hindered tumor growth and metastasis in vivo.
Conclusion: OIP5-AS1 regulated ovarian cancer progression via modulating miR-137/ZNF217 signaling, suggesting that targeting OIP5-AS1 could be conducive to EOC clinical treatment.
Keywords: OIP5-AS1, miR-137, ZNF217, epithelial ovarian cancer
Introduction
Epithelial ovarian cancer (EOC) is one of the most common gynecological malignancies in women around forty.1 It is one of the deadliest gynecological malignancies in the United States. According to statistics published in 2018, there are approximately over 22,000 diagnosis cases and 14,000 EOC-related deaths in the United States each year, and less than 50% of patients could survive more than 5 years.2 The high mortality is associated with the inconspicuous symptoms at early phase and frequent spread of metastases at advanced phase.3 Therefore, exploring the molecular mechanism behind EOC carcinogenesis and progression is of vital significance.
The majority transcripts in human genome are non-coding RNAs (ncRNAs), while only approximately 2% of these transcripts are protein-code genes. Among the numerous ncRNAs, lncRNAs, which are over 200 nucleotides (nt) in length, were presumed as “transcription noise”. However, lncRNAs have been discovered as key regulators in biological processes for its diverse regulatory mechanism. Accumulating studies have manifested that lncRNAs are associated with multiple biological activities, including cell proliferation, apoptosis, invasion and migration.4,5 Furthermore, dysregulated lncRNAs have been found to be correlated with tumor progression and poor prognosis outcome.6,7
The competing endogenous theory has revealed that lncRNAs can function as a competing endogenous RNA (ceRNA) to bind with miRNA and reverse the inhibiting effect of the miRNA on target gene. Opa-interacting protein 5 antisense RNA 1 (OIP5-AS1), a novel identified long intergenic noncoding RNA, has been studied in several cancers. It has been found to be overexpressed in some tumor tissues and predict unfavorable prognosis in tumors. For example, OIP5-AS1 is significantly up-regulated in lung cancer tissues and is positively correlated with tumor growth speed and tumor size.8 In addition, mechanically, OIP5-AS1 can target SOX2 via acting as miR-129-5p sponge in breast cancer.9 These findings suggested that OIP5-AS1 could function as an oncogene in the development of cancers. However, the underlying role and mechanism of OIP5-AS1 in EOC has not been learned yet. This study aimed to determine the biological role of OIP5-AS1 in EOC progression.
Materials and Methods
Human Samples
Forty OC tissues and adjacent healthy tissues were gathered from June 2013 to May 2018 from First Affiliated Hospital of Jinzhou Medical University. Before surgery, no patients acquired any therapy. Permission of the guide was provided by Institutional Review Committee of First Affiliated Hospital of Jinzhou Medical University, with the approval numbered JZ-19-25. Every participant signed informed consent prior to surgery. When the surgical resection was over, tissues were frozen in liquid nitrogen and stored at −80°C in requirements. The relationship between the expression of OIP5-AS1 and EOC clinicopathologic characteristics was provided in Supplementary Table 1.
Cell Culture
Human ovarian epithelial cell (IOSE) and ovarian cancer cells (HEY, A2780, SKOV3, OVCAR3) were bought from Chinese Academy of Sciences (Beijing, China) and cultured in DMEM (Invitrogen, Carlsbad, CA, USA) which was mixed with 10% fetal bovine serum (FBS; Invitrogen), 1% penicillin and streptomycin (Sigma-Aldrich, Milan, Italy). Besides, an incubator containing 5% CO2 was employed at 37°C. HEY and OVCAR-3 represent high grade ovarian serous adenocarcinoma; A2780 is representative of ovarian endometrioid adenocarcinoma; SKOV3 is representative of ovarian serous cystadenocarcinoma.
Cell Transfection
Specific shRNAs targeting OIP5-AS1 (shOIP5-AS1#1/#2) and their corresponding NC (shCtrl) were synthesized by GenePharma (Shanghai, China). The shRNAs were designed against sequences shared by all transcript variants of OIP5-AS1. To overexpress ZNF217 expression, the full-length cDNA sequence of ZNF217 was amplified by PCR and then sub-cloned into pcDNA3.1 vector (GenePharma), termed pcDNA3.1/ZNF217. The empty pcDNA3.1 vector was used in control group. The miR-137 mimics, miR-137 inhibitor, NC mimics and NC inhibitor were gained from GenePharma, too. Each plasmid was stably transfected into SKOV3 and OVCAR3 cells using Lipofectamine 2000 (Invitrogen). The stable cells selected using G418 (Clontech, CA, USA) were collected for the following use.
qRT-PCR
The extraction of total RNA was performed utilizing TRIzol reagent (Invitrogen). Total RNA was reverse-transcribed into cDNA via Reverse Transcription Kit. Then, RT-qPCR was progressed using a SYBR Green PCR kit (QIAGEN, Shanghai, China) and carried out in Bio-Rad CFX96 system. Fold-changes were calculated with utilization of 2−∆∆Ct method and GAPDH/U6 was the internal references. Sequences of relative PCR primers are provided in Supplementary Table 2.
Cell Counting Kit-8 (CCK-8) Assay
Briefly, SKOV3 and OVCAR3 cells were loaded in fresh 96-well plates and cultured for 24, 48, 72 and 96 h. 10 µL CCK-8 solution was added for incubating for another 4 h. Absorbance was determined at 450 nm via a microplate reader (Olympus, Tokyo, Japan).
EdU Incorporation Assay
SKOV3 and OVCAR3 cells were loaded into 24-well plates and incubated for 48 h. After incubation with EdU, cells were fixed with 4% paraformaldehyde (Solarbio, Beijing, China) and dyed by Apollo Dye Solution. Nucleic acid was stained by the utilization of DAPI (Invitrogen). Images were captured with an inverted fluorescence microscope (Olympus).
Flow Cytometry Analysis
The FITC Annexin V/Dead Cell Apoptosis Kits (Invitrogen) were used in this experiment. Following cultivation for 24 h, transfected SKOV3 and OVCAR3 cells were collected and washed twice using PBS (Sigma-Aldrich). Next, 5 μL of FITC annexin V plus I μL PI solution were added. Upon 15 min of incubation, flow cytometry was chosen for analyzing the stained cells.
Transwell Assay
In migration assay, the top compartments were added with transfected SKOV3 and OVCAR3 cells, while the lower chambers were full of medium with 10% FBS; 24 h later, cells were immobilized by methanol (Sigma-Aldrich) and dyed in crystal violet (Sigma-Aldrich). Furthermore, under a microscope (Olympus), cells in 5 randomly selected fields were counted. As for invasion assay, the top chambers were pre-coated with Matrigel, and the other procedures were in accordance with the migration assay.
Western Blot
Total protein was extracted from cells which were lysed by RIPA lysis buffer containing protease inhibitors. The total protein were quantified and then separated by 12% SDS-PAGE. After moving proteins to PVDF membranes, 5% fat-free milk was employed to seal the membranes, which were further cultured with the corresponding primary antibodies and secondary antibodies. Antibodies against ZNF217 (ab136678, 1:2000), E-cadherin (ab40772, 1:10,000), N-cadherin (ab202030, 1:2000), Vimentin (ab193555, 1:2000), Slug (ab51772, 1:1000), Twist (ab187008, 1:2000) and GAPDH (ab8245, 1:10,000) were all from Abcam (Cambridge, UK). GAPDH served as internal control. Furthermore, protein bands were detected by chemiluminescence detection system.
Subcellular Fractionation
RNAs from SKOV3 and OVCAR3 cells were isolated using the PARIS kit (Invitrogen) for nuclear and cytoplasmic fraction separation. RNA expression level of OIP5-AS1 in nuclear and cytoplasmic fractions was assayed through qRT-PCR. GAPDH and U6 were seen as the cytoplasmic and nuclear controls, respectively.
Fluorescence in situ Hybridization (FISH) Assay
SKOV3 and OVCAR3 cells were added to 24-well plates before being rinsed with PBS, along with fixation by using 4% formaldehyde, followed by permeabilized in 70% ethanol. Later, cells were cleaned with PBS twice and then hybridization solution with fluorescently labeled OIP5-AS1 probe was added for further incubation overnight. Following washing using saline-sodium citrate (SSC; Sigma-Aldrich), cells were dyed in DAPI and finally photographed via a fluorescence microscope (Olympus).
RNA Pull Down Assay
The wild type or mutated sequences of OIP5-AS1 (OIP5-AS1-WT, OIP5-AS1-MUT) with or without miR-137 binding sites were synthesized for RNA pull down assay. The OIP5-AS1-Wt, OIP5-AS1-Mut and the nonsense RNA sequences (negative control) were labeled with the Biotin into Bio-OIP5-AS1-WT, Bio-OIP5-AS1-MUT and Bio-NC, respectively. Cell lysates of SKOV3 and OVCAR3 cells were incubated with above biotin-labelled RNAs and streptavidin beads (Invitrogen) overnight. qRT-PCR was used to analyze expression levels of various miRNAs.
Luciferase Reporter Assay
The wild-type and mutant binding sites of miR-137 in OIP5-AS1 sequence or ZNF217 3ʹUTR were sub-cloned into pmirGLO dual-luciferase vector to construct OIP5-AS1-WT/MUT or ZNF217-WT/MUT and then co-transfected with miR-137 mimics or NC mimics into SKOV3 and OVCAR3 cells, appropriately. Finally, the luciferase activity was examined by dual luciferase system (Promega, Madison, WI, USA).
RNA Immunoprecipitation (RIP) Assay
The experiment was achieved utilizing the Magna RNA-binding protein immunoprecipitation kit (Invitrogen). Magnetic beads covering anti-IgG or anti-Ago2 were used for culturing lysates in RIP buffer. The normal IgG was negative control. Afterwards, the immunoprecipitated RNAs were detected by qRT-PCR.
Tumor Growth and Metastasis in Nude Mice
Nude mice were purchased from Shi Laike Company (Shanghai, China). The stably transfected SKOV3 cells (a pool of 1×107 cells) were injected subcutaneously into the axilla of nude mice. Tumor growth was observed by recording tumor volume based on the width and length every 4 days, and the calculating formula was: volume = length × width2 × 0.5. Four weeks later, the mice were euthanized and the tumor was weighed and measured. In the liver metastasis study, a pool of 2 × 107 stably transfected SKOV3 cells (with shCtrl or shOIP5-AS1#1) were injected into the tail vein of nude mice. Eight weeks later, mice were killed via cervical decapitation. Livers were surgically excised and fixed in formalin for further analysis. The welfare of animals applied in this study followed the Animals in Research: Reporting In Vivo Experiments (ARRIVE) guideline. The experiment was approved by the Ethics Committee for Animal Research of First Affiliated Hospital of Jinzhou Medical University with the protocol approval number of JZ-19-25.
Hematoxylin-Eosin (HE) Staining
The liver tissues obtained from mice in the in vivo metastasis experiment were fixated by 4% paraformaldehyde and then embedded via paraffin. Thereafter, tissues were cut into 3–5μm slices, and then the slices were processed with hematoxylin- and eosin-staining of nucleus and cytoplasm, respectively. Finally, the images were captured under a microscope.
Statistical Analysis
Data were expressed as means ± SD. We performed statistical analysis by use of Prism 5 software (Graph-Pad Software, La Jolla, CA, USA). The significance of differences between groups was estimated by Student’s t-test or one-way/two-way ANOVA. Correlations between molecules were estimated by Pearson’s correlation analysis. P < 0.05 was defined to have statistical significance. Experiments were run thrice.
Results
OIP5-AS1 is Highly Expressed in EOC and Knockdown of OIP5-AS1 Can Inhibit the Malignant Behaviors of EOC Cells
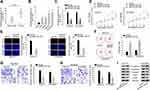
To investigate the expression profile of OIP5-AS1 in EOC, 40 pairs of EOC tissue and adjacent non-tumor tissue specimens were collected. We found that OIP5-AS1 was notably up-regulated in EOC tissues than in non-cancerous tissues (Figure 1A). qRT-PCR also showed an aberrant up-regulation of OIP5-AS1 in EOC cell lines (HEY, A2780, SKOV3 and OVCAR3) compared with normal IOSE cells (Figure 1B). Loss-of-function experiments were done after ensuring the knockdown effects of shOIP5-AS1#1/2 plasmids (Figure 1C). CCK8 and EdU assays showed that knockdown of OIP5-AS1 significantly impaired EOC cell viability (Figure 1D). After calculating the EdU positive rate, we confirmed that OIP5-AS1 deficiency suppressed cell proliferation (Figure 1E). Next, flow cytometry was performed to study the influence of OIP5-AS1 inhibition on cell apoptosis. We found a significant increase in apoptotic cells by OIP5-AS1 depletion (Figure 1F). Likewise, restraining OIP5-AS1 decreased the number of invaded and migrated cells (Figure 1G and H). Epithelial-mesenchymal transition (EMT) is a crucial phase that enables tumor cell migration and metastasis in EOC.10,11 Western blot revealed that silencing OIP5-AS1 notably increased the level of E-cadherin, while decreased the level of N-cadherin, Vimentin, Slug and Twist (Figure 1I). These functional experiments demonstrated that OIP5-AS1 was an oncogene in EOC.
OIP5-AS1 Acts as miR-137 Sponge in EOC
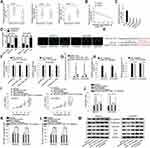
Then, we wondered if OIP5-AS1 could act as a miRNA sponge in EOC. We narrowed the scope of the combinable miRNAs for OIP5-AS1 on starBase database (strict stringency ≥5 in Clip data, low stringency in degradome data ≥1, and 8 cancer types in Pan-cancer). Three miRNAs (miR-141-3p, miR-2682-5p and miR-137) were revealed to bind to OIP5-AS1. qRT-PCR was used to detect their expression in EOC tissues and matched non-tumor tissues. As a result, MiR-137 exhibited a significant down-regulation in EOC tissues compared with normal tissues (Figure 2Aa). No significant expression difference of miR-2682-5p between tumor tissues and normal tissues was identified (Figure 2Ab). MiR-141-3p possessed a significant overexpression in tumor tissues relative to matched control tissues (Figure 2Ac), which was in line with the expression tendency of OIP5-AS1. Besides, Pearson’s correlation analysis manifested a negative correlation between the expression levels of miR-137 and OIP5-AS1 in EOC tissues (Figure 2B). We also observed a significant down-regulated expression of miR-137 in EOC cell lines (Figure 2C). Therefore, we selected miR-137 as the miRNA downstream of OIP5-AS1 in EOC. Subcellular fractionation and FISH assays demonstrated that OIP5-AS1 was principally situated in cytoplasm (Figure 2D). Next, we obtained potential binding sites between miR-137 and OIP5-AS1 by using starBase (Figure 2E). Dual luciferase reporter assays were conducted to verify the combination between OIP5-AS1 and miR-137. We found that miR-137 mimics sharply decreased the luciferase activity of OIP5-AS1-WT, instead of OIP5-AS1-MUT (Figure 2F). RNA pull down assay showed enriched miR-137 in biotinylated OIP5-AS1-WT group (Figure 2G). Collectively, we concluded that miR-137 was down-regulated in EOC tissues and cells and could bind to OIP5-AS1.
Next, we silenced miR-137 expression by using miR-137 inhibitor, and identified that miR-137 inhibition had no influence on OIP5-AS1 expression (Figure 2H). OIP5-AS1 knockdown inhibited cell viability while miR-137 knockdown elevated cell viability in SKOV3 and OVCAR3 cells, and the suppressive effects of OIP5-AS1 knockdown on cell viability were restored by further inhibition of miR-137 (Figure 2I). Flow cytometry result demonstrated that apoptotic cells increased after silencing OIP5-AS1 but decreased after silencing miR-137, while co-transfecting with miR-137 inhibitor recovered OIP5-AS1 depletion-accelerated apoptosis (Figure 2J). Apart from this, miR-137 inhibition promoted cell migration and invasion, as well as reversed the anti-migration and anti-invasion ability of shOIP5-AS1#1 (Figure 2K and L). Moreover, miR-137 inhibitor accelerated cell EMT process and could offset the suppressive effects of OIP5-AS1 knockdown on cell EMT process as well (Figure 2M). Based on these findings, miR-137 inhibitor played an oncogenic role and rescued the anti-cancer ability of silenced OIP5-AS1. In other words, OIP5-AS1 regulated EOC progression via sponging miR-137.
OIP5-AS1 Positively Regulates ZNF217 via Competitively Binding to miR-137
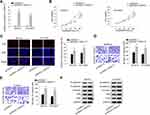
We have proved that OIP5-AS1 could bind with miR-137, whether OIP5-AS1 impacted downstream target gene of miR-137 still needed to be investigated. We narrowed the range of the combinable mRNAs for miR-137 from starBase (medium stringency ≥2 on clip data, low stringency on degradome data, 6 cancer types in Pan-cancer and selecting miRmap, PITA, miRanda, PicTar and TargetScan). Thirteen mRNAs were identified. Then, we used miR-137 mimics to study its effects on expression of the thirteen mRNAs. The overexpression effect of miR-137 mimics was verified by qRT-PCR (Figure 3A). We noticed that over-expression of miR-137 caused significant down-regulation of ZNF217 whereas had no or limited impacts on the expression of other candidates (Figure 3B). Thus, we selected ZNF217 as the focus of subsequent research. We performed qRT-PCR to investigate the relative expression of ZNF217 in EOC tissues and cell lines. Results exhibited an evident up-regulation of ZNF217 in EOC tissues and cell lines compared with corresponding controls (Figure 3C and D). Then, we obtained putative binding site of miR-137 within 3ʹUTR of ZNF217 from starBase (Figure 3E). From luciferase report assay, we observed that miR-137 mimics could significantly attenuate the luciferase activity of ZNF217-WT (Figure 3F). RIP assay further certified the abundant enrichments of miR-137, ZNF217and OIP5-AS1 in Ago2-precipitated RISCs (RNA induced silence complexes) (Figure 3G). More significantly, ZNF217 expression was hampered by OIP5-AS1 deficiency as well as by miR-137 suppression (Figure 3H). Further, the lessened ZNF217 level caused by suppressed OIP5-AS1 was reversed after inhibiting miR-137 (Figure 3I). Altogether, OIP5-AS1 served as a ceRNA against miR-137 to up-regulate ZNF217.
ZNF217 Exerts a Pro-Tumor Function in EOC
Further, we sought to examine the function of ZNF217 in malignant behaviors of EOC cells. Prior to that, the overexpression efficiency of ZNF217 was verified in SKOV3 and OVCAR3 cells after transfected with plasmids containing pcDNA3.1/ZNF217 (Figure 4A). The following CCK-8 and EdU assays revealed that cell viability and proliferation were enhanced by up-regulated ZNF217 (Figure 4B and C). Besides, overexpression of ZNF217 facilitated cell migration and invasion capacities (Figure 4D and E). Consistently, E-cadherin expression was inhibited while the levels of N-cadherin, Vimentin, Slug and Twist were elevated by overexpression of OIP5-AS1 (Figure 4F). To sum up, ZNF217 targeted by miR-137 served a carcinogenic part in EOC.
ZNF217 Up-Regulation Restores the Suppressive Function of Inhibited OIP5-AS1 on Cell Malignant Behaviors
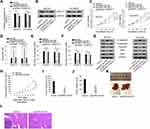
To understand whether ZNF217 was responsible for OIP5-AS1-mediated biological functions in EOC, a series of rescue functional experiments were performed. Firstly, qRT-PCR results indicated that the expression of ZNF217 was inhibited after knockdown of OIP5-AS1, but increased again when co-transfected with pcDNA3.1/ZNF217 (Figure 5A). Western blot showed the same trend in ZNF217 protein expression (Figure 5B). CCK8 assay showed that silencing OIP5-AS1-dampened cell viability was revived under ZNF217 overexpression (Figure 5C). In contrast, cell apoptosis facilitated by OIP5-AS1 knockdown was normalized in the presence of pcDNA3.1/ZNF217 (Figure 5D). Cell migration and invasion ability as well as EMT process were assessed by Transwell and Western blot assays. All the results manifested that ZNF217 overexpression reversed the suppressive effects of OIP5-AS1 deficiency on cell migration, invasion and EMT process (Figure 5E–G). Then, we further validated the function of OIP5-AS1 in in vivo tumor growth and metastasis. As anticipated, we found that tumor grew at a much slower speed in shOIP5-AS1#1 group (Figure 5H). The tumor volume was apparently smaller and tumor weight was much lessened in shOIP5-AS1#1 group than negative control group (Figure 5I and J). As for in vivo metastasis, the result showed significant less metastasized liver particles after knockdown of OIP5-AS1 than the livers from control group (Figure 5K). Moreover, HE staining presented that the metastatic nodules in livers from mice inoculated with OIP5-AS1-silenced cells were evidently reduced microscopically (Figure 5L). These experiments validated that OIP5-AS1 promoted EOC tumor growth and metastasis in vivo. More importantly, OIP5-AS1 exerted its effects on the malignant behaviors of EOC cells by up-regulating ZNF217 through sponging miR-137.
Discussion
LncRNAs have been found to be aberrantly expressed in various human malignancies, and the abnormal expressed lncRNAs can be used as clinical biomarkers at early diagnosis period.12–14 Recently, OIP5-AS1, as a novel lncRNA, was found to be aberrantly expressed in multiple tumors and could be viewed as a clinical biomarker in bladder cancer and osteosarcoma.15,16 However, its expression pattern and biological effects on EOC have not been studied yet. In the present study, we identified an obvious overexpression of OIP5-AS1 in EOC tissues and cell lines. Loss-of-function and gain-of-function experiments suggested that OIP5-AS1 played an oncogenic role in EOC. Knockdown of OIP5-AS1 significantly impaired EOC malignant behaviors, including cell proliferation, invasion, migration, and EMT process, and facilitated tumor cell apoptosis. The functional role of OIP5-AS1 in cancers is also supported in other literatures. OIP5-AS1 is up-regulated in undifferentiated oral tumors and predicts enhanced cancer stemness.17 Perturbations targeting OIP5-AS1, TUG1 and WT1-AS altered proliferation of breast and gynecologic cancer cells.18
Mounting studies have found that lncRNAs could act as a ceRNA and influence the function of downstream genes.19–22 Functioning as a ceRNA, the biological role of some lncRNAs has been analyzed in EOC.23,24 After determining the cytoplasmic location of OIP5-AS1, we found that OIP5-AS1 sponged miR-137. MiR-137 exhibited a remarkable down-regulation in EOC tissues and cells. Besides, miR-137 has been reported to be a tumor suppressor in ovarian cancer.25–27 Additionally, although the interaction between OIP5-AS1 and miR-137 has been testified in valve interstitial cells,28 such relationship was first identified in cancer cells until this study.
MiRNAs can bind to 3ʹUTR region of downstream target gene with micro response elements (MREs). We selected ZNF217 as candidate target gene of miR-137, as it exhibited the most significant down-regulation among the thirteen predicted target genes after overexpressing miR-137. Additionally, ZNF217 has been reported to play an oncogenic role and predict poor prognosis in ovarian cancer. For instance, overexpressing ZNF217 contributes to ovarian cancer invasion and metastasis progression.29 In the present study, we observed that ZNF217 was up-regulated in EOC tissues and cell lines. We also found that overexpression of ZNF217 abolished the tumor-inhibiting ability of silenced OIP5-AS1. In other words, ZNF217 up-regulation can rescue the oncogenic function of OIP5-AS1. Together, our finding initially suggested that lncRNA OIP5-AS1 overexpression aggravated EOC malignant activities via up-regulating ZNF217 by sponging miR-137.
Conclusions
Our study found the OIP5-AS1/miR-137/ZNF217 axis in EOC for the first time. We identified OIP5-AS1 as an oncogene in EOC progression. OIP5-AS1 could hopefully become a meaningful biomarker for EOC.
Data Sharing Statement
All related data and materials have been enclosed in the manuscript and additional files.
Ethics Approval and Informed Consent
Approval on human sample collection was provided by Institutional Review Committee of First Affiliated Hospital of Jinzhou Medical University. The animal experiment was approved by the Ethics Committee for Animal Research of First Affiliated Hospital of Jinzhou Medical University. The approval supported this study was numbered JZ-19-25.
Acknowledgments
We are very grateful to all individuals and groups involved in this study.
Author Contributions
Linlin Guo contributed to study design and conducted the mechanistic experiments while Jiabao Chen performed all the functional assays. All authors contributed to data analysis, drafting or revising the article, gave final approval of the version to be published, and agree to be accountable for all aspects of the work.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Webb PM, Jordan SJ. Epidemiology of epithelial ovarian cancer. Best Pract Res Clin Obstet Gynaecol. 2017;41:3–14. doi:10.1016/j.bpobgyn.2016.08.006
2. Moufarrij S, Dandapani M, Arthofer E, et al. Epigenetic therapy for ovarian cancer: promise and progress. Clin Epigenetics. 2019;11(1):7. doi:10.1186/s13148-018-0602-0
3. Pradeep S, Kim SW, Wu SY, et al. Hematogenous metastasis of ovarian cancer: rethinking mode of spread. Cancer Cell. 2014;26(1):77–91. doi:10.1016/j.ccr.2014.05.002
4. Yang MH, Hu ZY, Xu C, et al. MALAT1 promotes colorectal cancer cell proliferation/migration/invasion via PRKA kinase anchor protein 9. Biochim Biophys Acta. 2015;1852(1):166–174. doi:10.1016/j.bbadis.2014.11.013
5. Liu Q, Liu N, Shangguan Q, et al. LncRNA SAMD12-AS1 promotes cell proliferation and inhibits apoptosis by interacting with NPM1. Sci Rep. 2019;9(1):11593. doi:10.1038/s41598-019-48116-1
6. Wang AH, Tan P, Zhuang Y, Zhang XT, Yu ZB, Li LN. Down-regulation of long non-coding RNA HOTAIR inhibits invasion and migration of oesophageal cancer cells via up-regulation of microRNA-204. J Cell Mol Med. 2019;23(10):6595–6610.
7. Li M, Chen Y, Zhu J, Gao Z, Wang T, Zhou P. Long noncoding RNA CASC15 predicts unfavorable prognosis and exerts oncogenic functions in non-small cell lung cancer. Am J Transl Res. 2019;11(7):4303–4314.
8. Wang M, Sun X, Yang Y, Jiao W. Long non-coding RNA OIP5-AS1 promotes proliferation of lung cancer cells and leads to poor prognosis by targeting miR-378a-3p. Thorac Cancer. 2018;9(8):939–949. doi:10.1111/1759-7714.12767
9. Zeng H, Wang J, Chen T, et al. Downregulation of long non-coding RNA Opa interacting protein 5-antisense RNA 1 inhibits breast cancer progression by targeting sex-determining region Y-box 2 by microRNA-129-5p upregulation. Cancer Sci. 2019;110(1):289–302. doi:10.1111/cas.13879
10. Loret N, Denys H, Tummers P, Berx G. The role of epithelial-to-mesenchymal plasticity in ovarian cancer progression and therapy resistance. Cancers. 2019;11(6):838. doi:10.3390/cancers11060838
11. Vergara D, Merlot B, Lucot JP, et al. Epithelial-mesenchymal transition in ovarian cancer. Cancer Lett. 2010;291(1):59–66. doi:10.1016/j.canlet.2009.09.017
12. Luo W, Wang M, Liu J, Cui X, Wang H. Identification of a six lncRNAs signature as novel diagnostic biomarkers for cervical cancer. J Cell Physiol. 2020;235(2):993–1000.
13. Liu Y, Feng W, Liu W, et al. Circulating lncRNA ABHD11-AS1 serves as a biomarker for early pancreatic cancer diagnosis. J Cancer. 2019;10(16):3746–3756. doi:10.7150/jca.32052
14. Zhang Z, Li J, He T, et al. The competitive endogenous RNA regulatory network reveals potential prognostic biomarkers for overall survival in hepatocellular carcinoma. Cancer Sci. 2019;110(9):2905–2923. doi:10.1111/cas.14138
15. Wang Y, Shi F, Xia Y, Zhao H. LncRNA OIP5-AS1 predicts poor prognosis and regulates cell proliferation and apoptosis in bladder cancer. J Cell Biochem. 2019;120(5):7499–7505.
16. Dai J, Xu L, Hu X, et al. Long noncoding RNA OIP5-AS1 accelerates CDK14 expression to promote osteosarcoma tumorigenesis via targeting miR-223. Biomed Pharmacother. 2018;106:1441–1447. doi:10.1016/j.biopha.2018.07.109
17. Arunkumar G, Anand S, Raksha P, et al. LncRNA OIP5-AS1 is overexpressed in undifferentiated oral tumors and integrated analysis identifies as a downstream effector of stemness-associated transcription factors. Sci Rep. 2018;8(1):7018. doi:10.1038/s41598-018-25451-3
18. Chiu HS, Somvanshi S, Patel E, et al. Pan-cancer analysis of lncRNA regulation supports their targeting of cancer genes in each tumor context. Cell Rep. 2018;23(1):297–312.e212. doi:10.1016/j.celrep.2018.03.064
19. Zhuang X, Tong H, Ding Y, et al. Long noncoding RNA ABHD11-AS1 functions as a competing endogenous RNA to regulate papillary thyroid cancer progression by miR-199a-5p/SLC1A5 axis. Cell Death Dis. 2019;10(8):620. doi:10.1038/s41419-019-1850-4
20. Zhou RS, Zhang EX, Sun QF, et al. Integrated analysis of lncRNA-miRNA-mRNA ceRNA network in squamous cell carcinoma of tongue. BMC Cancer. 2019;19(1):779. doi:10.1186/s12885-019-5983-8
21. Zhou JM, Liang R, Zhu SY, et al. LncRNA WWC2-AS1 functions as a novel competing endogenous RNA in the regulation of FGF2 expression by sponging miR-16 in radiation-induced intestinal fibrosis. BMC Cancer. 2019;19(1):647. doi:10.1186/s12885-019-5754-6
22. Guo J, Li Y, Duan H, Yuan L. LncRNA TUBA4B functions as a competitive endogenous RNA to inhibit gastric cancer progression by elevating PTEN via sponging miR-214 and miR-216a/b. Cancer Cell Int. 2019;19(1):156. doi:10.1186/s12935-019-0879-x
23. Fan MJ, Zou YH, He PJ, Zhang S, Sun XM, Li CZ. Long non-coding RNA SPRY4-IT1 promotes epithelial-mesenchymal transition of cervical cancer by regulating the miR-101-3p/ZEB1 axis. Biosci Rep. 2019;39(6). doi:10.1042/BSR20181339
24. Zhu L, Zhang Q, Li S, Jiang S, Cui J, Dang G. Interference of the long noncoding RNA CDKN2B-AS1 upregulates miR-181a-5p/TGFbetaI axis to restrain the metastasis and promote apoptosis and senescence of cervical cancer cells. Cancer Med. 2019;8(4):1721–1730. doi:10.1002/cam4.2040
25. Sun J, Cai X, Yung MM, et al. miR-137 mediates the functional link between c-Myc and EZH2 that regulates cisplatin resistance in ovarian cancer. Oncogene. 2019;38(4):564–580. doi:10.1038/s41388-018-0459-x
26. Dong P, Xiong Y, Watari H, et al. MiR-137 and miR-34a directly target snail and inhibit EMT, invasion and sphere-forming ability of ovarian cancer cells. J Exp Clin Cancer Res. 2016;35(1):132. doi:10.1186/s13046-016-0415-y
27. Zhang L, Li Z, Gai F, Wang Y. MicroRNA-137 suppresses tumor growth in epithelial ovarian cancer in vitro and in vivo. Mol Med Rep. 2015;12(2):3107–3114. doi:10.3892/mmr.2015.3756
28. Zheng D, Wang B, Zhu X, et al. LncRNA OIP5-AS1 inhibits osteoblast differentiation of valve interstitial cells via miR-137/TWIST11 axis. Biochem Biophys Res Commun. 2019;511(4):826–832. doi:10.1016/j.bbrc.2019.02.109
29. Li J, Song L, Qiu Y, Yin A, Zhong M. ZNF217 is associated with poor prognosis and enhances proliferation and metastasis in ovarian cancer. Int J Clin Exp Pathol. 2014;7(6):3038–3047.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.