Back to Journals » Cancer Management and Research » Volume 13
NR2F1-AS1/miR-140/HK2 Axis Regulates Hypoxia-Induced Glycolysis and Migration in Hepatocellular Carcinoma
Authors Li X, Li Y, Bai S, Zhang J, Liu Z, Yang J
Received 8 June 2020
Accepted for publication 9 September 2020
Published 15 January 2021 Volume 2021:13 Pages 427—437
DOI https://doi.org/10.2147/CMAR.S266797
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Seema Singh
Xiao Li,1 Yize Li,2 Shuang Bai,2 Jing Zhang,2 Zhengcai Liu,1 Jingyue Yang2
1Department of Hepatobiliary Surgery, Xijing Hospital, Fourth Military Medical University, Xian 710032, People’s Republic of China; 2Department of Clinical Oncology, Xijing Hospital, Fourth Military Medical University, Xian 710032, People’s Republic of China
Correspondence: Jingyue Yang
Department of Clinical Oncology, Xijing Hospital, Fourth Military Medical University, 17 Chonglexi Road, Xian, Shaanxi Province 710032, People’s Republic of China
Tel +86-02984775412
Email [email protected]
Background: Hypoxia is an important feature for the progression of hepatocellular carcinoma (HCC). Long noncoding RNA nuclear receptor subfamily 2 group F member 1 antisense RNA 1 (NR2F1-AS1) is dysregulated in HCC. However, the role and mechanism of N2RF1-AS1 in hypoxia-induced glycolysis and migration remain unclear.
Materials and Methods: Tumor tissues and adjacent samples were harvested from 40 HCC patients. HCC cells were treated by hypoxia. The levels of NR2F1-AS1, microRNA (miR)-140, and hexokinase 2 (HK2) were examined via quantitative reverse transcription polymerase chain reaction or Western blot. Glycolysis was analyzed via glucose uptake, lactate production, and adenosine triphosphate (ATP) levels. Cell migration was analyzed via transwell assay. The target association was analyzed via dual-luciferase reporter assay and RNA immunoprecipitation.
Results: NR2F1-AS1 level was enhanced in HCC tissues and cells. High expression of NR2F1-AS1 indicated poor overall survival. Silence of NR2F1-AS1 repressed hypoxia-induced glycolysis and migration in HCC cells. NR2F1-AS1 could regulate HK2 expression by modulating miR-140. miR-140 down-regulation or HK2 up-regulation mitigated the influence of NR2F1-AS1 silence on hypoxia-induced glycolysis and migration in HCC cells.
Conclusion: NR2F1-AS1 knockdown restrained hypoxia-induced glycolysis and migration in HCC cells via increasing miR-140 and decreasing HK2.
Keywords: hepatocellular carcinoma, NR2F1-AS1, miR-140, HK2, glycolysis, migration
Introduction
Hepatocellular carcinoma (HCC) is a dominant liver cancer with leading cause of cancer-related death.1 Through great effort in recent years, significant insight has been gained on the pathogenesis, prevention, and management of HCC.2 Hypoxic environment is a feature of the angiogenesis and development of human cancers, including HCC.3 During hypoxia, glucose metabolism is altered, and many malignancies are triggered in HCC.4 Thus, exploring hypoxia-related mechanism and therapeutic option is helpful for the treatment of HCC.
Long noncoding RNAs (lncRNAs, >200 nucleotides) play key roles in the tumorigenesis of HCC.5 LncRNAs can regulate microRNA (miRNA)/mRNA axes to take part in the tumorigenesis and disease progression in HCC.6 Moreover, lncRNAs levels are changed in response to the stimulus of hypoxia.7 LncRNA nuclear receptor subfamily 2 group F member 1 antisense RNA 1 (NR2F1-AS1) exhibits an oncogenic role in multiple cancers, including endometrial cancer, thyroid cancer, and osteosarcoma.8,9,10 Moreover, NR2F1-AS1 is highly expressed, and NR2F1-AS1 knockdown represses migration, invasion, and oxaliplatin resistance via regulating miR-363/ATP Binding Cassette Subfamily C Member 1 (ABCC1) in HCC.11 Under hypoxic stress, glycolysis and migration are triggered in HCC.12,13 However, it is unknown whether NR2F1-AS1 can regulate glycolysis and migration under hypoxia in HCC.
The dysregulated miRNAs are also involved in HCC progression, which have important roles in the detection and treatment of HCC.14 Moreover, miRNA expression can be altered via hypoxia treatment.15 Previous studies indicate that miR-140 has an anti-cancer role in HCC by suppressing cancer progression and angiogenesis.16,17 Furthermore, hexokinase 2 (HK2) is a key enzyme associated with glucose metabolism, which is required for glycolysis, migration, and invasion in HCC.18,19 However, whether miR-140 and HK2 are involved in the NR2F1-AS1-mediated mechanism in HCC is unclear.
In this study, we predicted the potential target association among NR2F1-AS1, miR-140, and HK2. Therefore, we hypothesized that NR2F1-AS1 might regulate glycolysis and migration under hypoxia by modulating miR-140 and HK2. Here we first measured the level of NR2F1-AS1 in HCC, and explored the influence of NR2F1-AS1 on glycolysis and migration under hypoxia. Moreover, we explored the regulatory network of NR2F1-AS1/miR-140/HK2 in HCC cells.
Materials and Methods
Patient and Tissues Collection
A total of 40 HCC patients (25 males and 15 females, age range: 45–63 years old) were recruited from Xijing Hospital of Fourth Military Medical University. Tumor tissues and adjacent normal tissues were harvested via surgery. The tissues were confirmed via two histopathologists. The samples were stored in RNA later solution (Thermo Fisher, Wilmington, DE, USA), and snap-frozen in liquid nitrogen until RNA assessment. The RNA and cDNA were stored at −80°C. The patients did not receive preoperative therapy before surgery and they all provided the written informed consents. After the resection, no other therapy was performed before relapse. The patients with relapse were received local or systemic treatment as appropriate, which were excluded in this study. Overall survival of patients was analyzed after a 5-year follow-up after admission. This research was permitted via the Ethics Committee of Fourth Military Medical University.
Cell Culture and Hypoxic Treatment
HCC cell lines (Hep3B, MHCC97-H, Huh7, and SNU-398) and normal liver epithelial cell line THLE-2 were provided via BeNa Culture Collection (Beijing, China). Cells were grown in DMEM (Thermo Fisher) with 10% fetal bovine serum (Biosun, Shanghai, China) and 1% antibiotics (Sigma, St. Louis, MO, USA) at 37°C in 5% CO2. To induce hypoxic condition, Hep3B and Huh7 cells were cultured in a hypoxic chamber with 3% O2 for 24 h.20 Cells maintained in normal condition (21% O2) were used as controls.
Cell Transfection
NR2F1-AS1 and HK2 overexpression vectors were generated via inserting corresponding sequence into pcDNA3.1 (Thermo Fisher). The pcDNA3.1 vector was exploited as negative control (vector). siRNA for NR2F1-AS1 (si-NR2F1-AS1#1, 5ʹ-UAAUAGAAAUAUUGAGAACAU-3ʹ; si-NR2F1-AS1#2, 5ʹ-AUUAAGUUGAAAAGCAAUGCA-3ʹ;), siRNA negative control (si-con, 5ʹ-AAGACAUUGUGUGUCCGCCTT-3ʹ), miR-140 mimic (5ʹ-CAGUGGUUUUACCCUAUGGUAG-3ʹ), mimic negative control (miR-con, 5ʹ-UUCUCCGAACGUGUCACGUTT-3ʹ), miR-140 inhibitor (in-miR-140, 5ʹ-CUACCAUAGGGUAAAACCACUG-3ʹ), inhibitor negative control (in-miR-con, 5ʹ-UGAGCUGCAUAGAGUAGUGAUUA-3ʹ) were synthesized from GenePharma (Shanghai, China). These constructed vectors or oligonucleotides were transfected into Hep3B and Huh7 cells using Lipofectamine 3000 reagent (Thermo Fisher) for 24 h.
Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
RNA was extracted using TRIzol (Thermo Fisher), and the isolated RNA with OD260/280 (1.8–2.0) was reversely transcribed to cDNA through specific reverse transcription kit (Thermo Fisher). The mixture of cDNA, SYBR Green (Thermo Fisher), and specific primers (Genscript, Nanjing, China) was used for qRT-PCR. The primers were listed as follows: NR2F1-AS1 (sense, 5ʹ-CATGCCGTGATGTAAGCTGC-3ʹ; antisense, 5ʹ-GCGACTGTTTCACCTCTCCA-3ʹ), HK2 (sense, 5ʹ-GTGAATCGGAGAGGTCCCAC-3ʹ; antisense, 5ʹ-CAAGCAGATGCGAGGCAATC-3ʹ), miR-140 (sense, 5ʹ-CAGTGGTTTTACCCTATGGTAG-3ʹ; antisense, 5ʹ-ACCATAGGGTAAAACCACTGTT-3ʹ), U6 (sense, 5ʹ-CTCGCTTCGGCAGCACA-3ʹ; antisense, AACGCTTCACGAATTTGCGT), β-actin (sense, 5ʹ-CTCGCCTTTGCCGATCC-3ʹ; antisense, 5ʹ-TCTCCATGTCGTCCCAGTTG-3ʹ). β-actin (for NR2F1-AS1 or HK2) or U6 (for miR-140) was used as an internal reference. The relative RNA expression was calculated according to delta-delta cycle threshold method.21
Glucose Uptake, Lactate Production and Adenosine Triphosphate (ATP) Level
Hep3B and Huh7 cells (5 × 104 cells/well) were added into 24-well plates, and stimulated by hypoxia for 24 h. Next, medium was collected for the detection of glucose uptake and lactate production using Glucose Uptake Assay Kit (Abcam, Cambridge, MA, USA) or Lactate Assay Kit (Abcam) according to the instructions of manufacturer. The concentration of glucose or lactate was normalized to total protein concentration. Moreover, cell lysates were used for detection of ATP level using ATP Assay Kit (Abcam) following the manufacturer’s protocol. The relative ATP level was normalized to normal group.
Western Blot
Cell protein was isolated using RIPA buffer (Beyotime, Shanghai, China) and quantified using BCA assay kit (Sigma). Protein samples (20 μg) were separated via sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred onto nitrocellulose membranes (Solarbio, Beijing, China). Next, the membranes were blocked in 5% skim milk, and then interacted with primary and secondary antibodies, followed via interacting with BeyoECL Plus kit (Beyotime). The antibodies were shown as: anti-HK2 (ab209847, 1:500 dilution, Abcam), and anti-β-actin (ab1801, 1:1000 dilution, Abcam), as well as horseradish peroxidase-conjugated IgG (ab205718, 1:20,000 dilution, Abcam). β-actin was used as a loading reference. HK2 protein level was normalized to the control group.
Transwell Migration Assay
The migrated ability was detected via transwell migration assay using 24-well transwell chambers (Corning, Corning, NY, USA). Hep3B and Huh7 cells (1 × 104 cells/well) were resuspended in serum-free medium and added into upper chambers. The low chambers were added with medium containing 10% serum. Following culture under hypoxic or normoxic condition for 24 h, the migrated cells were dyed with 0.5% crystal violet (Solarbio) and counted under a microscope (Nikon, Tokyo, Japan).
Dual-Luciferase Reporter Analysis and RNA Immunoprecipitation (RIP)
The complementary sequence between miR-140 and NR2F1-AS1 or HK2 was predicted via starBase (http://starbase.sysu.edu.cn/) or DIANA tool (http://diana.imis.athena-innovation.gr/DianaTools/). The wild-type luciferase reporter vectors NR2F1-AS1-WT and HK2-WT were constructed by inserting a sequence of NR2F1-AS1 or HK2 containing miR-140 binding sites into psiCHECK-2 (Promega, Madison, WI, USA). The mutant-type luciferase reporter vectors NR2F1-AS1-MUT and HK2-MUT were generated via inserting the sequence containing mutant binding sites of miR-140. Hep3B and Huh7 cells were co-transfected with wild-type or mutant-type luciferase reporter vectors and miR-140 mimic, miR-con, si-NR2F1-AS1, or si-con for 24 h. Cells in mock group were transfected with luciferase reporter vectors alone. Next, luciferase activity was detected through a dual-luciferase assay kit (Promega).
Magna RIP kit (Sigma) was used for RIP assay. 1 × 107 Hep3B and Huh7 cells were lysed and incubated with magnetic beads coated with anti-Ago2 or anti-IgG antibody for 6 h. Input was used as positive control. RNA in the complex was extracted and abundances of NR2F1-AS1, miR-140, and HK2 were detected via qRT-PCR.
Statistical Analysis
GraphPad Prism 7 (GraphPad, La Jolla, CA, USA) was employed for statistical analysis. Three independent experiments were performed. The data with normal distribution and homogeneity of variance were presented as mean ± SD. Overall survival was analyzed via Kaplan-Meier curve and Log rank test. The linear correlation among NR2F-AS1, miR-140, and HK2 levels was determined via Pearson correlation analysis. The difference was compared via Student’s t-test or ANOVA followed via Tukey post hoc test as appropriate. P<0.05 was considered statistically significant.
Results
The Level of NR2F1-AS1 is Up-Regulated in HCC
To explore the abundance of NR2F1-AS1 in HCC, 40 paired tumor and adjacent samples were collected from HCC patients. As shown in Figure 1A, NR2F1-AS1 level was higher in tumor tissues than that in adjacent samples. Next, patients were divided into high (n=23) or low NR2F1-AS1 expression (n=17) group according to the mean value of NR2F1-AS1 in tumor tissues. High level of NR2F1-AS1 was correlated with lower overall survival of patients (Figure 1B). In addition, the level of NR2F1-AS1 was measured in four HCC cell lines. Results showed that NR2F1-AS1 expression was markedly elevated in HCC cell lines in comparison to that in THLE-2 cells (Figure 1C). These data suggested that high expression of NR2F1-AS1 was associated with HCC development. Hep3B and Huh7 cells with relative highest NR2F1-AS1 level were chosen for further experiments.
NR2F1-AS1 Knockdown Inhibits Hypoxia-Induced Glycolysis and Migration in HCC Cells
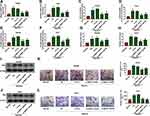
To explore the role of NR2F1-AS1 in HCC development under hypoxic condition, Hep3B and Huh7 cells were transfected with si-NR2F1-AS1#1, si-NR2F1-AS1#2, or si-con before treatment of hypoxia. As exhibited in Figure 2A and B, NR2F1-AS1 expression was evidently enhanced under hypoxic condition compared with that in normal group, and it was effectively decreased by transfection of si-NR2F1-AS1#1 or si-NR2F1-AS1#2. Moreover, glucose uptake, lactate production, and ATP level were markedly increased in Hep3B and Huh7 cells after stimulation of hypoxia, which were weakened via knockdown of NR2F1-AS1 (Figure 2C–H). In addition, glycolysis-associated marker HK2 was measured, and results showed that HK2 protein abundance was remarkably elevated by stimulation of hypoxia in the two cell lines, which was attenuated via NR2F1-AS1 inhibition (Figure 2I and J). Besides, transwell migration assay displayed the migrated abilities of Hep3B and Huh7 cells were enhanced under hypoxic condition, and these events were mitigated via silence of NR2F1-AS1 (Figure 2K and L). These results indicated that NR2F1-AS1 knockdown suppressed HCC development under hypoxic condition. si-NR2F1-AS1#1 with higher efficacy of NR2F1-AS1 knockdown was used for further experiments and named as si-NR2F1-AS1.
NR2F1-AS1 is a Sponge of miR-140
To explore the mechanism addressed via NR2F1-AS1, the targets of miR-140 were predicted via starBase. miR-140 was a predicted target, and the binding sequence of NR2F1-AS1 and miR-140 is shown in Figure 3A. To identify the relationship between NR2F1-AS1 and miR-140, NR2F1-AS1-WT, and NR2F1-AS1-MUT were constructed and used for dual-luciferase reporter assay. miR-140 overexpression obviously decreased the luciferase activity in NR2F1-AS1-WT group, but the effect was abolished in NR2F1-AS1-MUT group (Figure 3B and C). Moreover, RIP assay displayed that there were higher enrichment levels of NR2F1-AS1 and miR-140 in the same complex of Ago2 RIP (Figure 3D and E). Additionally, miR-140 level was detected in HCC tissues and cell lines. Results displayed miR-140 level was notably declined in HCC tissues and cells in comparison to that in normal samples or THLE-2 cells (Figure 3F and G). Furthermore, miR-140 expression was evidently reduced by treatment of hypoxia in Hep3B and Huh7 cells (Figure 3H). These findings indicated that miR-140 was a target of NR2F1-AS1 in HCC cells.
HK2 is Targeted by miR-140
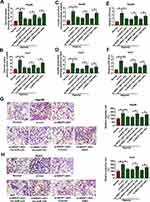
To further study the mechanism mediated by NR2F1-AS1/miR-140 axis, the target of miR-140 was analyzed via DIANA tool. HK2 as an important biomarker for glycolysis was predicted to be a candidate target of miR-140, and the complementary sequence between miR-140 and HK2 was exhibited in Figure 4A. Moreover, we constructed HK2-WT and HK2-MUT, and then transfected them into Hep3B and Huh7 cells. The data of dual-luciferase reporter assay showed that miR-140 overexpression or NR2F1-AS1 silence evidently decreased luciferase activity in HK2-WT group, whereas it did not alter the activity in HK2-MUT group (Figure 4B–E). In addition, RIP assay indicated that miR-140 and HK2 could be enriched in the same complex by Ago2 RIP (Figure 4F and G). Furthermore, HK2 level was increased in HCC tissues and cells compared with that in adjacent samples or THLE-2 cells (Figure 4H and I). Besides, HK2 protein level in Hep3B and Huh7 cells was significantly increased via treatment of hypoxia (Figure 4J). These results displayed that HK2 was directly targeted via miR-140.
NR2F1-AS1 Regulates HK2 Expression by Sponging miR-140
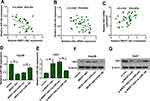
To explore whether and how NR2F1-AS1 regulated HK2 expression, the association among NR2F1-AS1, miR-140, and HK2 was analyzed. In HCC tissues, miR-140 expression was negatively correlated with NR2F1-AS1 or HK2 level (Figure 5A and B), while HK2 expression was positively associated with NR2F1-AS1 abundance (Figure 5C). Moreover, Hep3B and Huh7 cells were transfected with vector, NR2F1-AS1 overexpression vector, NR2F1-AS1 overexpression vector + miR-con, NR2F1-AS1 overexpression vector + miR-140 mimic, si-con, si-NR2F1-AS1, si-NR2F1-AS1 + in-miR-con or si-NR2F1-AS1 + in-miR-140. As shown in Figure 5D and E, miR-140 expression was decreased or increased via NR2F1-AS1 overexpression or knockdown, which was weakened via transfection of miR-140 mimic or in-miR-140, respectively. Meanwhile, HK2 protein expression was positively regulated via NR2F1-AS1, which was attenuated by miR-140 up-regulation or down-regulation (Figure 5F and G). These findings indicated that NR2F1-AS1 positively regulated HK2 expression by competitively modulating miR-140 in HCC cells.
NR2F1-AS1 Silence Suppresses Hypoxia-Induced Glycolysis and Migration via Regulating miR-140 and HK2 in HCC Cells
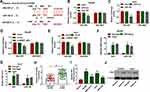
To explore whether miR-140 and HK2 were required for NR2F1-AS1-mediated glycolysis and migration of HCC under hypoxic condition, Hep3B and Huh7 cells were transfected with si-con, si-NR2F1-AS1, si-NR2F1-AS1 + in-miR-con, si-NR2F1-AS1 + in-miR-140, si-NR2F1-AS1 + vector or si-NR2F1-AS1 + HK2 overexpression vector. As shown in Figure 6A–F, miR-140 knockdown or HK2 overexpression reversed silence of NR2F1-AS1-mediated inhibition of glycolysis in Hep3B and Huh7 cells under hypoxic condition, revealed via restoration of glucose uptake, lactate production, and ATP levels. Furthermore, miR-140 knockdown or HK2 overexpression attenuated the interference of NR2F1-AS1-induced migration inhibition in hypoxia-treated cells (Figure 6G and H). These data suggested that NR2F1-AS1 silence repressed HCC development under hypoxic condition by modulating miR-140 and HK2.
Discussion
HCC is a main type of liver cancer with high mortality worldwide.22 Hypoxia is implicated in the pathogenesis of liver disorders, including HCC.23 LncRNAs play important roles in the development and prognosis of HCC.24 Moreover, lncRNAs take part in hypoxia-driven cancer progression.25 A previous study reported that NR2F1-AS1 knockdown could decrease cell migration, invasion, and drug resistance in HCC.11 However, the role of NR2F1-AS1 in glycolysis and migration under hypoxia and the related mechanism in HCC are unknown. In this research, we found that NR2F1-AS1 silence suppressed hypoxia-induced glycolysis and migration in HCC cells through regulating miR-140 and HK2. The novelty of our study was that we first explored the role of NR2F1-AS1 under hypoxia and the regulatory network of NR2F1-AS1/miR-140/HK2.
In this work, high expression of NR2F1-AS1 was measured in HCC tissues and cells, which was also consistent with a former report.11 In addition, a previous work indicated that lncRNA could be modulated via hypoxia in cancers.26 Here we found that NR2F1-AS1 expression was changed in response to the stimulation of hypoxia, indicating that NR2F1-AS1 might play an important role in hypoxia-driven HCC progression. The glucose metabolism is reprogrammed in HCC under hypoxia.27 Through measuring glucose uptake, lactate production, ATP generation, and HK2 expression, which acted as important factors of glycolysis,28,29 we found that NR2F1-AS1 knockdown suppressed hypoxia-driven glycolysis in HCC. Furthermore, hypoxia could promote cancer metastasis and migration in HCC.13,30 Here we found that NR2F1-AS1 silence inhibited HCC cell migration, which was also consistent with previous studies in other cancers.8–10 Collectively, NR2F1-AS1 knockdown could repress hypoxia-induced glycolysis and migration in HCC, indicating that NR2F1-AS1 might serve as a promising target for the treatment of HCC.
The network of lncRNA-miRNA-mRNA is an important mechanism underlying the role of lncRNA. Previous studies indicated multiple regulatory networks of NR2F1-AS1, such as NR2F1-AS1/miR-363/SRY-related high-mobility group box 4 (SOX4), NR2F1-AS1/miR-338-3p/cyclin D1 (CCND1), NR2F1-AS1/miR-423-5p/SRY-box 12 (SOX12), NR2F1-AS1/miR-483-3p/forkhead box A1 (FOXA1), and NR2F1-AS1/miR-363/ABCC1.8–11,31 There would be multiple networks because of the presence of numerous binding sites. To explore a novel mechanism addressed via NR2F1-AS1, here we were the first to confirm NR2F1-AS1 could target miR-140. Low expression of miR-140 was measured in HCC, which was also in agreement with former works.16,32 Moreover, miR-140 expression was reduced in HCC after stimulation of hypoxia. A previous study suggested that miR-140 could inhibit HCC cell migration.33 In addition, miR-140 was reported to repress glycolysis in sepsis and breast cancer.34,35 Similarly, we also found the anti-migration and anti-glycolysis roles of miR-140 in HCC under hypoxia, revealed via which miR-140 knockdown reversed the anti-cancer role of NR2F1-AS1 in HCC.
HK2 is a key enzyme associated with glycolysis, which contributes to multiple cell processes, like cell proliferation, invasion, migration, angiogenesis, and drug resistance in HCC.36 Moreover, Huang et al showed that the up-regulation of HK2 increased glycolysis to promote cell migration and invasion in HCC.37 Furthermore, multiple evidences suggested that HK2 could promote glycolysis and was responsible for migration and invasion in HCC.18,19,38,39,40 These all indicated the oncogenic role of HK2 in HCC via increasing glycolysis and migration. Consistent with these reports, our study also confirmed the carcinogenic function of HK2, and NR2F1-AS1 could control hypoxia-driven HCC progression by regulating HK2. In addition, we identified HK2 was targeted by miR-140, and NR2F1-AS1 could up-regulate the HK2 level via modulating miR-140 in HCC cells. In this way, NR2F1-AS1 could promote hypoxia-driven HCC progression via regulating miR-140/HK2 axis.
In conclusion, NR2F1-AS1 expression was enhanced in HCC under hypoxia. Moreover, NR2F1-AS1 knockdown suppressed hypoxia-induced glycolysis and migration in HCC, possibly via modulating the miR-140/HK2 axis. This research indicated a novel mechanism for understanding the hypoxia-driven HCC progression.
Highlights
1. NR2F1-AS1 expression is increased in HCC tissues and cells.
2. NR2F1-AS1 knockdown inhibits hypoxia-induced glycolysis and migration in HCC.
3. NR2F1-AS1 regulates HK2 by sponging miR-140.
Funding
No financial support was received.
Disclosure
The authors declare that there are no competing interests associated with the manuscript.
References
1. Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391:1301–1314. doi:10.1016/S0140-6736(18)30010-2
2. Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol. 2019;16:589–604. doi:10.1038/s41575-019-0186-y
3. Choi SH, Park JY. Regulation of the hypoxic tumor environment in hepatocellular carcinoma using RNA interference. Cancer Cell Int. 2017;17:3. doi:10.1186/s12935-016-0374-6
4. Xiong XX, Qiu XY, Hu DX, Chen XQ. Advances in hypoxia-mediated mechanisms in hepatocellular carcinoma. Mol Pharmacol. 2017;92:246–255. doi:10.1124/mol.116.107706
5. Lim LJ, Wong SYS, Huang F, et al. Roles and regulation of long noncoding RNAs in hepatocellular carcinoma. Cancer Res. 2019;79:5131–5139. doi:10.1158/0008-5472.CAN-19-0255
6. Huo X, Han S, Wu G, et al. Dysregulated long noncoding RNAs (lncRNAs) in hepatocellular carcinoma: implications for tumorigenesis, disease progression, and liver cancer stem cells. Mol Cancer. 2017;16:165. doi:10.1186/s12943-017-0734-4
7. Dong J, Xu J, Wang X, Jin B. Influence of the interaction between long noncoding RNAs and hypoxia on tumorigenesis. Tumour Biol. 2016;37:1379–1385. doi:10.1007/s13277-015-4457-0
8. Wang L, Zhao S, Mingxin YU. LncRNA NR2F1-AS1 is involved in the progression of endometrial cancer by sponging miR-363 to target SOX4. Pharmazie. 2019;74:295–300.
9. Guo F, Fu Q, Wang Y, Sui G. Long non-coding RNA NR2F1-AS1 promoted proliferation and migration yet suppressed apoptosis of thyroid cancer cells through regulating miRNA-338-3p/CCND1 axis. J Cell Mol Med. 2019;23:5907–5919. doi:10.1111/jcmm.14386
10. Li S, Zheng K, Pei Y, Wang W, Zhang X. Long noncoding RNA NR2F1-AS1 enhances the malignant properties of osteosarcoma by increasing forkhead box A1 expression via sponging of microRNA-483-3p. Aging (Albany NY). 2019;11:11609–11623. doi:10.18632/aging.102563
11. Huang H, Chen J, Ding CM, Jin X, Jia ZM, Peng J. LncRNA NR2F1-AS1 regulates hepatocellular carcinoma oxaliplatin resistance by targeting ABCC1 via miR-363. J Cell Mol Med. 2018;22:3238–3245. doi:10.1111/jcmm.13605
12. Li Q, Pan X, Zhu D, Deng Z, Jiang R, Wang X. Circular RNA MAT2B promotes glycolysis and malignancy of hepatocellular carcinoma through the miR-338-3p/PKM2 axis under hypoxic stress. Hepatology. 2019;70:1298–1316. doi:10.1002/hep.30671
13. Chen Y, Huang F, Deng L, et al. HIF-1-miR-219-SMC4 regulatory pathway promoting proliferation and migration of HCC under hypoxic condition. Biomed Res Int. 2019;2019:8983704. doi:10.1155/2019/8983704
14. Yang N, Ekanem NR, Sakyi CA, Ray SD. Hepatocellular carcinoma and microRNA: new perspectives on therapeutics and diagnostics. Adv Drug Deliv Rev. 2015;81:62–74. doi:10.1016/j.addr.2014.10.029
15. Truettner JS, Katyshev V, Esen-Bilgin N, Dietrich WD, Dore-Duffy P. Hypoxia alters MicroRNA expression in rat cortical pericytes. Microrna. 2013;2:32–44.
16. Li Y, Liu G, Li X, Dong H, Xiao W, Lu S. Long non-coding RNA SBF2-AS1 promotes hepatocellular carcinoma progression through regulation of miR-140-5p-TGFBR1 pathway. Biochem Biophys Res Commun. 2018;503:2826–2832. doi:10.1016/j.bbrc.2018.08.047
17. Hou ZH, Xu XW, Fu XY, Zhou LD, Liu SP, Tan DM. Long non-coding RNA MALAT1 promotes angiogenesis and immunosuppressive properties of HCC cells by sponging miR-140. Am J Physiol Cell Physiol. 2020;318:649–663. doi:10.1152/ajpcell.00510.2018
18. DeWaal D, Nogueira V, Terry AR, et al. Hexokinase-2 depletion inhibits glycolysis and induces oxidative phosphorylation in hepatocellular carcinoma and sensitizes to metformin. Nat Commun. 2018;9:446. doi:10.1038/s41467-017-02733-4
19. Chai F, Li Y, Liu K, Li Q, Sun H. Caveolin enhances hepatocellular carcinoma cell metabolism, migration, and invasion in vitro via a hexokinase 2-dependent mechanism. J Cell Physiol. 2019;234:1937–1946. doi:10.1002/jcp.27074
20. Zhang CY, Jiang ZM, Ma XF, et al. Saikosaponin-d inhibits the hepatoma cells and enhances chemosensitivity through SENP5-dependent inhibition of Gli1 SUMOylation under hypoxia. Front Pharmacol. 2019;10:1039. doi:10.3389/fphar.2019.01039
21. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi:10.1006/meth.2001.1262
22. Villanueva A, Longo DL. Hepatocellular carcinoma. N Engl J Med. 2019;380:1450–1462. doi:10.1056/NEJMra1713263
23. Nath B, Szabo G. Hypoxia and hypoxia inducible factors: diverse roles in liver diseases. Hepatology. 2012;55:622–633. doi:10.1002/hep.25497
24. Abbastabar M, Sarfi M, Golestani A, Khalili E. lncRNA involvement in hepatocellular carcinoma metastasis and prognosis. EXCLI J. 2018;17:900–913.
25. Shih JW, Kung HJ. Long non-coding RNA and tumor hypoxia: new players ushered toward an old arena. J Biomed Sci. 2017;24:53. doi:10.1186/s12929-017-0358-4
26. Chang YN, Zhang K, Hu ZM, et al. Hypoxia-regulated lncRNAs in cancer. Gene. 2016;575:1–8. doi:10.1016/j.gene.2015.08.049
27. Shang RZ, Qu SB, Wang DS. Reprogramming of glucose metabolism in hepatocellular carcinoma: progress and prospects. World J Gastroenterol. 2016;22:9933–9943. doi:10.3748/wjg.v22.i45.9933
28. Jin F, Wang Y, Zhu Y, et al. The miR-125a/HK2 axis regulates cancer cell energy metabolism reprogramming in hepatocellular carcinoma. Sci Rep. 2017;7:3089. doi:10.1038/s41598-017-03407-3
29. Ren L, Yao Y, Wang Y, Wang S. MiR-505 suppressed the growth of hepatocellular carcinoma cells via targeting IGF-1R. Biosci Rep. 2019;39:BSR20182442. doi:10.1042/BSR20182442
30. Wong CC, Kai AK, Ng IO. The impact of hypoxia in hepatocellular carcinoma metastasis. Front Med. 2014;8:33–41. doi:10.1007/s11684-013-0301-3
31. Yang C, Liu Z, Chang X, et al. NR2F1-AS1 regulated miR-423-5p/SOX12 to promote proliferation and invasion of papillary thyroid carcinoma. J Cell Biochem. 2020;121:2009–2018. doi:10.1002/jcb.29435
32. Wang ZY, Zhu Z, Wang HF, et al. Downregulation of circDYNC1H1 exhibits inhibitor effect on cell proliferation and migration in hepatocellular carcinoma through miR-140-5p. J Cell Physiol. 2019;234:17775–17785. doi:10.1002/jcp.28403
33. Yan X, Zhu Z, Xu S, et al. MicroRNA-140-5p inhibits hepatocellular carcinoma by directly targeting the unique isomerase Pin1 to block multiple cancer-driving pathways. Sci Rep. 2017;7:45915. doi:10.1038/srep45915
34. Liu L, Li TM, Liu XR, et al. MicroRNA-140 inhibits skeletal muscle glycolysis and atrophy in endotoxin-induced sepsis in mice via the WNT signaling pathway. Am J Physiol Cell Physiol. 2019;317:189–199. doi:10.1152/ajpcell.00419.2018
35. He Y, Deng F, Zhao S, et al. Analysis of miRNA-mRNA network reveals miR-140-5p as a suppressor of breast cancer glycolysis via targeting GLUT1. Epigenomics. 2019;11:1021–1036. doi:10.2217/epi-2019-0072
36. Feng J, Li J, Wu L, et al. Emerging roles and the regulation of aerobic glycolysis in hepatocellular carcinoma. J Exp Clin Cancer Res. 2020;39:126.
37. Huang M, Xiong H, Luo D, Xu B, Liu H. CSN5 upregulates glycolysis to promote hepatocellular carcinoma metastasis via stabilizing the HK2 protein. Exp Cell Res. 2020;388:111876. doi:10.1016/j.yexcr.2020.111876
38. Ding Z, Guo L, Deng Z, Li P. Circ-PRMT5 enhances the proliferation, migration and glycolysis of hepatoma cells by targeting miR-188-5p/HK2 axis. Ann Hepatol. 2020;19:269–279. doi:10.1016/j.aohep.2020.01.002
39. Wang J, Chen J, Sun F, et al. miR-202 functions as a tumor suppressor in hepatocellular carcinoma by targeting HK2. Oncol Lett. 2020;19:2265–2271.
40. Ye J, Xiao X, Han Y, Fan D, Zhu Y, Yang L. MiR-3662 suppresses cell growth, invasion and glucose metabolism by targeting HK2 in hepatocellular carcinoma cells. Neoplasma. 2020;67:773–781. doi:10.4149/neo_2020_190730N689
 © 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2021 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.