Back to Journals » OncoTargets and Therapy » Volume 13
NCAPG Induces Cell Proliferation in Cardia Adenocarcinoma via PI3K/AKT Signaling Pathway
Authors Zhang X , Wang H, Han Y, Zhu M, Song Z, Zhan D, Jia J
Received 14 August 2020
Accepted for publication 18 October 2020
Published 4 November 2020 Volume 2020:13 Pages 11315—11326
DOI https://doi.org/10.2147/OTT.S276868
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Geoffrey Pietersz
Xinxin Zhang,1 Hui Wang,1 Yajuan Han,2 Mengqi Zhu,1 Zaozhi Song,1 Dankai Zhan,1 Jianguang Jia1
1Department of Surgical Oncology, The First Affiliated Hospital of Bengbu Medical College, Bengbu 233000, People’s Republic of China; 2Department of Medical Oncology, The First Affiliated Hospital of Bengbu Medical College, Bengbu 233000, People’s Republic of China
Correspondence: Jianguang Jia Department of Surgical Oncology
The First Affiliated Hospital of Bengbu Medical College, 2600 Donghai Road, Bengbu 233000, People’s Republic of China
Email [email protected]
Purpose: Previous studies have shown that non-SMC condensin I complex subunit G (NCAPG) overexpression is correlated to poor prognosis of multiple cancer types. Herein, we explored the underlying mechanism of NCAPG-mediated cardia adenocarcinoma (CA) proliferation and cell cycle regulation.
Methods: The protein profiling technology was used to analyze the gene expression in 20 CA and adjacent tissue samples. Differential genes were identified by bioinformatic analysis. Western blot and qRT-PCR-based analysis assessed the NCAPG expression levels in multiple CA cell lines. CA cell lines, SGC-7901 and AGS, were transfected with Lip 2000, and stably transfected cell lines were screened for NCAPG overexpression and downregulation. MTT and clone formation assays were employed to detect cell proliferation, and cell cycle phases were analyzed using flow cytometry. Western blot was performed to determine the NCAPG gene expression levels. Finally, we studied the tumorigenic effects of NCAPG in the mouse model and validated the cell experiment results using immunohistochemistry.
Results: A significant overexpression of NCAPG was found in CA tissues and CA cell lines. The outcomes of MTT and clone formation assays showed that NCAPG upregulation promoted cell proliferation. The outcomes of these analyses were further validated using nude mice as an in vivo tumor model. As per the outcome of Western blot and flow cytometry analysis, NCAPG regulated the G1 phase through the cyclins (CDK4, CDK6, and cyclin D1) overexpression and cell cycle inhibitors (P21 and P27) downregulation. Overexpressed NCAPG and silenced NCAPG, both in vitro and in vivo, resulted in abnormal activation of the PI3K/AKT signaling pathway in CA cells. We observed that NCAPG overexpression increased the levels of phosphorylated PI3K, AKT, and GSK3β; however, their total protein levels remained unchanged in CA cells.
Conclusion: As a CA oncogene, NCAPG promoted cell proliferation and regulated cell cycle through PI3K/AKT signaling pathway activation.
Keywords: NCAPG, cardia adenocarcinoma, PI3K/AKT pathway, cell cycle, proliferation
Introduction
Gastric cancer is a highly prevalent malignancy and one of the most common causes of cancer-related death across the world.1,2 China accounts for around 24% of the gastric cancer cases.3 Cardia adenocarcinoma (CA) is a subtype of gastric cancer that occurs 5 cm above and below the junction of the esophagus and the stomach.4 CA can be defined anatomically as the diaphragmatic space, and histologically as the transition of the esophageal squamous epithelium to gastric gland cells. Due to this histological transformation, the mucosa at the junction of the lower esophagus becomes particularly vulnerable to the destruction by gastric acid reflux, which increases the risk of tumors and malignant transformation.5 A sharp rise in the incidence of CA has been observed during the last decade in western countries.6 Clinical symptoms of this malignancy include upper abdominal discomfort, fullness after light eating, indigestion, and front stomach pain. Different therapeutic strategies, i.e., surgery, radiotherapy, chemotherapy, and immunotherapy, have been used to manage CA; even then, the overall five-year survival rate of this malignancy is as low as 24%. Thus, unraveling the internal mechanism of CA can improve the treatment effect in CA patients.
Non-SMC condensin I complex subunit G (NCAPG), a mitotic associated chromosomal condensing protein, is a polypeptide, which contains 1015 amino acids with a relative molecular weight of 114.1 kDa. In recent years, multiple studies have reported abnormal expression of NCAPG in distinct cancer types, such as gastric cancer,7 lung adenocarcinoma,8 prostate cancer,9 clear cell renal cell carcinoma,10 breast cancer,11 bladder cancer,12 colorectal cancer,13 and hepatocellular carcinoma.14–16 NCAPG participates in multiple biological functions in prostate cancer,17 breast cancer,18 and liver cancer.19–22 Several research studies explored the role of NCAPG and demonstrated its promising treatment prospects. Even then, the functional role of NCAPG in CA remains unclear.
Herein, we screened differential genes in clinical CA tissues. As per the outcome of the analysis, a novel tumor-promoting factor was identified. We found abnormal overexpression of NCAPG in CA tissues and CA cell lines. For the first time, we showed that NCAPG, as an oncogene, plays a crucial role in CA by promoting cell proliferation and regulating cell cycle through PI3K/AKT signaling pathway activation. Thus, this study provides a novel therapeutic strategy for CA.
Patients and Methods
Study Population (Tissue Specimens)
In this study, we analyzed tumor samples from 20 CA confirmed cases from the First Affiliated Hospital of Bengbu Medical College collected during the time period of 2018 to 2019. CA was confirmed in each of these cases, both pre- and post-surgery. None of the patients underwent any type of treatment. Each sample was divided into three equal parts, and each sample was analyzed separately. The protein expression profiles of each sample were detected using mass spectrometry (QE plus), and the differentially expressed proteins were identified using MaxQuant analysis software. The Ethical Review Committee of the First Affiliated Hospital of Bengbu Medical College approved this study.
Cell Culture
The AGS, SGC-7901, and BGC-823 gastric adenocarcinoma cell lines and MKN-45 and MGC-803 GC cell lines were procured from Shanghai Cell Bank, Chinese Academy of Sciences. Normal gastric mucosal cells (GES) were procured from Wuhan Procell Life Science & Technology Co., Ltd (Wuhan, China). These cells were grown in DMEM media (HyClone, USA) with 10% FBS (Gibco, USA) and penicillin/streptomycin at 37°C in a 5% CO2 incubator.
Immunohistochemistry (IHC)
Briefly, the paraffin-embedded CA tissue samples were cut into 4 µm thick sections and placed on glass slides. Subsequently, tissue sections were treated to remove paraffin, later hydrated, and endogenous peroxidase was inactivated using 0.3% H2O2 for 15 min. Antigen retrieval was carried out at 100°C by using 10 mmol/L sodium citrate buffer (pH 6.0). These tissue sections were blocked using goat serum. Later, the tissue sections were washed with PBS thrice, 5–10 min each time, and incubated with primary antibodies (Ki67 polyclonal antibody, 1:200, Abcam, USA; CDK4 polyclonal antibody, 1:200, Abcam) at 4°C and subsequently incubated with biotinylated goat anti-mouse IgG secondary antibody for 30 min at 37°C. The immunohistochemical scoring standard23 coalesced staining intensity, staining grade (0=none, 1=weak, 2=medium, 3=strong), and the percentage of positive tumor cells is (0=none, 1=1–25%, 2=26–50%, 3=51–75%, 4=76–100%). The total score of the immunohistochemical staining was obtained by multiplying the staining intensity and the positive cell score. The total score was divided into high expression group (≥ 4) and low expression group (< 4). Two senior pathologists evaluated the stained tissue sections.
Recombinant Lentivirus Infection
The NCAPG overexpression lentivirus (GeneChem, Shanghai, China) was used for transfecting CA cell lines. For the downregulation of NCAPG, two human shRNA sequences (RNAi1:CCGGGCTATGCAGAAGCATCTTCTTCTCGAGAAGAAGATGCTTCTGCATAGCTTTTTTG; RNAi2:CCGGCGGGCAGTGTTATCATGTATTCTCGAGAATACATGATAACACTGCCCGTTTTTTG) were procured from Invitrogen (CA, USA) and cloned into the pSUPER-puro plasmid (GeneChem, Shanghai, China) to yield RNAi constructs. Lentiviral particle production and transduction were performed, as described previously. Cells exhibiting stable NCAPG overexpression or NCAPG knockdown were selected using puromycin.
qRT-PCR
RNA was isolated from tissue samples using Trizol (Invitrogen, USA), as per the manufacturer’s instruction. To gauge RNA quality, the A260/A280 ratio was determined using the NanoVue™ Plus (Thermo Fisher Scientific, MA, USA) instrument. Furthermore, cDNA was synthesized from 1 μg of each RNA sample using the PrimeScript™ 1st Strand cDNA Synthesis kit (Thermo Fisher Scientific, USA), and cDNA was stored at −80°C. The qRT-PCR reactions were carried out by using SYBR Premix Ex Taq™ (TaKaRa) and ABI StepOne™ Real-Time PCR System (Applied Biosystems, CA, USA). GAPDH was used as the normalization control. The primer sequences for qRT-PCR (Shanghai GeneChem) were as follows: NCAPG forward: 5ʹ-AAGTTAGACGGGCAGTGTTATC-3ʹ, NCAPG reverse: 5ʹ- CAGCTTTCTGACAGCCTCTT-3ʹ. GAPDH forward: 5ʹ-ACCACAGTCCATGCCATCAC-3ʹ; GAPDH reverse: 5ʹ-TCCACCACCCTG TTGCTGTA-3ʹ.
MTT Assay
5 × 103 cells/well were seeded into 96-well plates (Corning, NY, USA). These cells were transfected with Lip 2000 to overexpress NCAPG or silence NCAPG and incubated for 1, 2, 3, 4, and 5 days. Later, 20 μL MTT was added to each well and incubated for 4 h at 37ºC. Subsequently, this solution was replaced with 150 μL DMSO (Sigma, USA) for each well. Absorbance was recorded using a microplate reader at 490 nm.
Colony Formation Assay
Cells were transfected to either overexpress NCAPG or silence NCAPG, and later these transfected cells were seeded in 6-well plates at a density of 4 ×103 cells/well. These cells were incubated at 37°C and humidified with 5% CO2 for 10 days. These cells were washed with phosphate-buffered saline (PBS), fixed with 4% paraformaldehyde for 20 min, and stained with 5% crystal violet for 20 min. Later, the number of colonies was calculated. Cell colonies > 50 per well were microscopically enumerated.
Western Blotting
Cells were lysed on ice for 30 min using RIPA cell lysate (Solarbio, Beijing, China). The cell lysate was centrifuged at 14,000 g for 15 min at 4°C, and the supernatant was separated. The protein concentration in the supernatant was evaluated by employing the BCA protein detection kit (Beyotime Institute of Biotechnology, China). Protein extracts were resolved on SDS-PAGE gel electrophoresis and later transferred to the PVDF membrane. 5% BSA blocking fluid was applied to PVDF membrane for 45 min, and later PVDF membrane was incubated with the primary antibodies: rabbit anti-NCAPG, -α-Tubulin, -p-GSK3β, -GSK3β, -CDK4, -p-PI3K, -PI3K, -p-AKT and -AKT (Abcam, Cambridge, UK), and rabbit anti-P21, -P27, -CDK6, -cyclinD1, and -Ki67 (Proteintech, Wuhan, China), at 4°C overnight followed by incubation with the rabbit secondary antibody (1:1000, 2 h, RT). The strips were developed using ECL.
In vivo Tumor Formation Experiment in Nude Mice
4-weeks old male BALB/C nude mice were subcutaneously injected into the flanks with 5×106 cells (Cavens Laboratory Animal Center, Changzhou, China). Mice were euthanized after 4–5 weeks under anesthesia (3% pentobarbital sodium), and later tumors were extracted. The volume and weight of the tumor samples were evaluated. These experiments met the requirements of the ethics committee of the institute and approved by the ethics committee of the First Affiliated Hospital of Bengbu Medical College (2020.069).
Flow Cytometry Analysis
Firstly, the harvested cells were thoroughly washed with ice-cold PBS, and each cell was fixed and soaked using 70% ice-cold ethanol. Subsequently, cells were centrifuged at 1000 g for 5 min. The cell pellet was resuspended in ice-cold PBS supplemented with 0.1 mg/mL RNAase and 5 mg/mL propidium iodide (Sigma, USA). Based on the DNA concentration, the cells were flow-cytometrically analyzed to examine the cell cycle phases (BD Accuri C6-plus, USA), and the flow cytometry data were analyzed using FLOWJO software (Tree Star, Inc, Ashland, OR).
Statistical Analysis
Groups were compared using SPSS version 22.0 (SPSS, IL, USA). Data were compared using t-tests and ANOVAs with Tukey’s HSD test as appropriate, with P < 0.05 as the significance threshold. Significance was represented as follows: N.S., nonsignificant; *P < 0.05; **P < 0.01; ***P < 0.001.
Result
NCAPG Overexpression in Cardia Adenocarcinoma
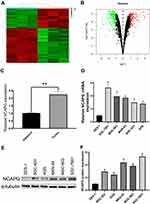
To determine protein profile, 20 CA tissue samples and adjacent tissue samples that were procured from the First Affiliated Hospital of Bengbu Medical College were analyzed, and differentially expressed genes (logFC > 1, P < 0.05) were screened using bioinformatics analysis (Figure 1A and B). We detected NCAPG upregulation in CA tissues as compared to adjacent tissues (Figure 1C). Besides, Western blot and qRT-PCR-based analysis revealed NCAPG overexpression in CA cell lines as compared to normal gastric mucosal cell lines (Figure 1D–F).
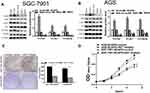
NCAPG Promoted Cardia Adenocarcinoma Proliferation in vitro
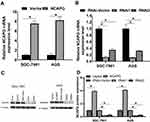
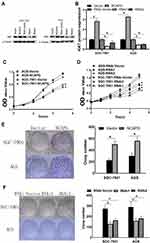
To unravel the role of NCAPG in the CA cell proliferation, based on the NCAPG expression, CA cell lines (SGC-7901 and AGS) were transfected for overexpression or silencing of the NCAPG gene. The transfected CA cell lines were categorized into overexpressed control group (vector), overexpressed experimental group (NCAPG), silenced control group (vector-RNAi), silenced interference sequence 1 (RNAi1), and silenced interference sequence 2 (RNAi2). NCAPG expression levels in the transfected CA cell lines were detected through Western blot and qRT-PCR analysis. NCAPG group showed increased expression levels of NCAPG as compared to the vector group, and the RNAi group showed decreased expression levels of NCAPG as compared to the vector RNAi group. (Figure 2). Ki67 expression level was detected through Western blot analysis to explore the relationship between NCAPG and cell proliferation. The outcome of these analyses showed NCAPG, Ki67 overexpression induced CA cell proliferation; on the contrary, NCAPG, Ki67 gene silencing mitigated CA cell proliferation (Figure 3A and B). Furthermore, the MTT assay revealed the effect of NCAPG on the CA cell proliferation. It showed that the NCAPG overexpression significantly increased the proliferation of CA cells in contrast to the vector group. Also, cell growth in the RNAi group was substantially mitigated than the vector-RNAi group (Figure 3C and D). The outcome of clone formation experiments revealed that NCAPG overexpression promoted CA cell proliferation while the NCAPG knockdown significantly inhibited CA cell proliferation (Figure 3E and F). Concisely, these findings demonstrated NCAPG induced proliferation of cells in CA cell lines.
NCAPG Regulates the Tumorigenesis of Cardia Adenocarcinoma
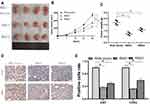
The cell lines with stable NCAPG expression were screened to evaluate the biological role of NCAPG in CA development. To validate the findings of in vitro experiments, SGC-7901 cells were grafted in BALB/C nude mice to establish a xenograft tumor model to assess the NCAPG effect in vivo. Four nude mice were present in each group. As per the tumor weight, volume, and growth curve, cells that were transfected with NCAPG RNAi, when grafted in nude mice, showed significantly reduced tumor formation ability as compared to cells transfected with plain vector (Figure 4A–C). Tumors formed by cells transfected with NCAPG RNAi showed reduced expression levels of Ki67 and CDK4 proteins (Figure 4D and E). Thus, the outcomes of this in vivo experiments indicated that NCAPG gene silencing inhibited the tumorigenicity of CA cells.
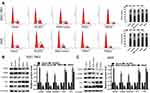
NCAPG Regulates the Cell Cycle by Arresting Cell Cycle at G1 Phase
Flow cytometry-based analysis was employed to investigate the underlying mechanism of NCAPG-mediated CA cell proliferation. A higher cell count in the G1 phase and a lower cell count in the S and G2/M phase were observed in the NCAPG gene silenced SGC-7901 and AGS cells than the vector-RNAi group of cells as per the flow cytometry-based analysis. However, cells overexpressing NCAPG showed opposite outcomes (Figure 5A). It indicated that inhibition of NCAPG expression in CA cells arrested the G1 phase and thus inhibited cell proliferation. To validate these outcomes, expression levels of cell-cycle related proteins were detected by employing Western blot analysis. The outcomes revealed that the cell cycle inhibitors (p21 and p27) levels were reduced in CA cells overexpressing NCAPG, but their levels were increased in NCAPG knockdown CA cells. The cell cycle molecules, i.e., cyclinD1, CDK4, and CDK6, were up-regulated in the CA cells overexpressing NCAPG and down-regulated in the NCAPG gene silenced group of CA cells (Figure 5B and C). In summary, these data indicated that NCAPG might regulate the cell cycle in CA cell lines.
AKT/PI3K Signaling Pathway is Regulated by NCAPG
AKT/PI3K pathway plays a crucial role in tumor progression of distinct tumor types. To evaluate the influence of NCAPG expression on the AKT/PI3K signaling pathway, key molecular proteins of the AKT/PI3K signaling pathway were analyzed using Western blot. The outcomes demonstrated escalated levels of phosphorylated AKT and PI3K in CA cells with NCAPG overexpression. Conversely, NCAPG gene silenced CA cells showed reduced levels of phosphorylated AKT and PI3K. Concomitantly, the activation of the AKT’s downstream target GSK3β corresponded with AKT and PI3K phosphorylation levels. It indicates that NCAPG regulated the AKT and PI3K activity in CA cells (Figure 6A and B). To delineate the crucial role of AKT and PI3K in NCAPG-regulated proliferation with tumorigenesis, CA cells overexpressing NCAPG were treated with AKT inhibitor (Perofosine). The clone formation and the MTT assay outcomes showed that AKT inhibition reduced cell proliferation (Figure 6C and D). In summary, as per the outcome of this study, NCAPG regulated CA through the AKT/PI3K pathway (Figure 7).
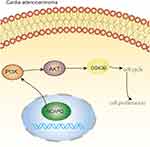 |
Figure 7 NCAPG regulates the cell cycle and promotes cell proliferation through PI3K/AKT signaling pathway in cardio adenocarcinoma. Pictorial representation of the outcome of our study. |
Discussion
The clinical manifestation of early-stage CA mimics with peptic ulcers. Early-stage CA is responsive to symptomatic treatment. Thus, most of the patients afflicted with early-stage CA fail to undergo further clinical examination until the disease is highly advanced. In addition, it is difficult to detect a small cavity in the abdomen due to the complex anatomy of the cardia region with an irregular cavity shape.24,25 It delays the diagnosis of CA, making the targeted treatment of CA difficult. Thus, the targeted treatment of CA demands urgent investigation.
NCAPG is a subunit of the condensin complex that encodes a cohesive complex and is responsible for chromosomal cohesion and stability during the process of mitosis and meiosis. As per previous research investigations, NCAPG is also involved in distinct tumor types. In the current study, NCAPG overexpression was found in 20 age-sex matched CA tissues and corresponding adjacent normal tissues using protein profiling technology. Concurrently, we also found that as compared to normal gastric mucosal cells, the NCAPG expression levels in CA cell lines were abnormally increased. As per the outcome of this analysis, we speculated that NCAPG might promote tumor formation in CA. Through the MTT assay and clone formation experiments, we confirmed that NCAPG mediated the CA cell proliferation. In addition, our experimental data demonstrated that NCAPG affected cell-cycle in CA cells in vitro. Flow cytometry and Western blot were employed to analyze the cell cycle in CA cells. The analysis results showed that NCAPG promoted cell proliferation by regulating cell cycle molecules, i.e., CDK4 and CDK6. NCAPG gene silencing altered the xenograft tumor size in nude mice, which validated the oncogenic role of NCAPG in CA. Through the immunohistochemical analysis of the tumor cells, we further validated the outcomes of the cell experiments, which showed that NCAPG promoted CA cell proliferation and regulated the G1 phase of the cell cycle by regulating CDK4. The findings of our study were in line with the previous research reports, which stated abnormal overexpression of NCAPG in gastric cancer.7 However, until now, the underlying mechanism (direct or indirect) of NCAPG in cellular proliferation and cell cycle regulation in CA remains ambiguous.
PI3Ks are encoded by multiple heterodimeric lipid kinase genes. AKT is a downstream effector of PI3K, which regulates a myriad of biological processes, such as proliferation,26 apoptosis,27 and growth,28 and acts as a crucial mediator of the cell cycle.29 Thr308 and Ser473 phosphorylation activate AKT. The PI3K/AKT signaling pathway is regulated abnormally in multiple tumor types, such as breast cancer,30 colon cancer,31 liver cancer,32 ovarian cancer,33 and pancreatic cancer.34 Previous studies have also stated an abnormal activation of the PI3K/AKT signaling pathway in gastric cancer.35,36 Interestingly, our results also demonstrated that NCAPG overexpression activated the PI3K/AKT signaling pathway in the CA cells. Western blot analysis showed that NCAPG up-regulated the expression levels of phosphorylated AKT, PI3K, and GSK3β, and activated the PI3K/AKT signaling pathway. Correlation between NCAPG and PI3K/AKT pathways in the pathogenesis of CA has not been established so far. Thus, in this study, we revealed that NCAPG promoted cell proliferation through PI3K/AKT signaling pathway in CA cells. By adding perifosine, an AKT inhibitor, to CA cells overexpressing NCAPG, we found that cell proliferation was inhibited in these cells, which indicated that the oncogenic function of NCAPG in CA depends on PI3K/AKT signaling.
Conclusion
In the current research, we found that NCAPG is expressed in CA tissues and CA cell lines. Functional studies validated the carcinogenic effects of NCAPG in CA. NCAPAG promoted the CA cell proliferation and regulated the G1 phase of the cell cycle by regulating CDK4 and CDK6 molecules. It is worth noting that, for the first time, we clarified that NCAPG has a tumor-promoting role in the pathogenesis of CA through the PI3K/AKT signaling pathway (Figure 7). This indicates that NCAPG may serve as a prognostic treatment target for CA.
Data Sharing Statement
All data generated or analyzed during this study are included in this published article.
Ethics Approval and Consent to Participate
The present study was granted ethical approval by the Institutional Review Board of Bengbu Medical College. Written informed consent was obtained from all participants involved in the study. This research was conducted in accordance with the Declaration of Helsinki. Animal experiments were as per the “Regulations on the Administration of Laboratory Animals” of the State Council of the People’s Republic of China and the “Detailed Rules for the Administration of Medical Laboratory Animals” of the Ministry of Health. The certificate number BYYFY-2017KY21 and 2020.069 were used for the collection of clinical samples and animal experimentations, respectively.
Acknowledgments
We thank Mr. Liu Hao’s research group from Anhui Biochemical Pharmaceutical Engineering Technology Research Center of Bengbu Medical College for their help in the experiment.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Funding
This study was supported by the Key Project of the Education Department of Anhui Province (KJ2017A240) and 2019 Bengbu Science and Technology Innovation Guidance Project and the 2019 College Student Innovation Training Program of Bengbu Medical College (Byycx1919).
Disclosure
The authors report no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.21492
2. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30. doi:10.3322/caac.21442
3. Feng RM, Zong YN, Cao SM, Xu RH. Current cancer situation in China: good or bad news from the 2018 global cancer statistics. Cancer Commun. 2019;39(1):22. doi:10.1186/s40880-019-0368-6
4. Jia J, Zhang X, Zhan D, et al. LncRNA H19 interacted with miR-130a-3p and miR-17-5p to modify radio-resistance and chemo-sensitivity of cardiac carcinoma cells. Cancer Med. 2019;8(4):1604–1618. doi:10.1002/cam4.1860
5. Chevallay M, Bollschweiler E, Chandramohan SM, et al. Cancer of the gastroesophageal junction: a diagnosis, classification, and management review. Ann N Y Acad Sci. 2018;1434(1):132–138. doi:10.1111/nyas.13954
6. Matsuno K, Ishihara R, Ohmori M, et al. Time trends in the incidence of esophageal adenocarcinoma, gastric adenocarcinoma, and superficial esophagogastric junction adenocarcinoma. J Gastroenterol. 2019;54(9):784–791. doi:10.1007/s00535-019-01577-7
7. Song B, Du J, Song DF, Ren JC, Feng Y. Dysregulation of NCAPG, KNL1, miR-148a-3p, miR-193b-3p, and miR-1179 may contribute to the progression of gastric cancer. Biol Res. 2018;51(1):44. doi:10.1186/s40659-018-0192-5
8. Li S, Xuan Y, Gao B, et al. Identification of an eight-gene prognostic signature for lung adenocarcinoma. Cancer Manag Res. 2018;10:3383–3392. doi:10.2147/CMAR.S173941
9. Yamamoto S, Takayama KI, Obinata D, et al. Identification of new octamer transcription factor 1-target genes upregulated in castration-resistant prostate cancer. Cancer Sci. 2019;110(11):3476–3485. doi:10.1111/cas.14183
10. Wei W, Lv Y, Gan Z, Zhang Y, Han X, Xu Z. Identification of key genes involved in the metastasis of clear cell renal cell carcinoma. Oncol Lett. 2019;17(5):4321–4328. doi:10.3892/ol.2019.10130
11. Chen J, Qian X, He Y, Han X, Pan Y. Novel key genes in triple-negative breast cancer identified by weighted gene co-expression network analysis. J Cell Biochem. 2019;120(10):16900–16912. doi:10.1002/jcb.28948
12. Pan S, Zhan Y, Chen X, Wu B, Liu B. Identification of biomarkers for controlling cancer stem cell characteristics in bladder cancer by network analysis of transcriptome data stemness indices. Front Oncol. 2019;9:613. doi:10.3389/fonc.2019.00613
13. Ding X, Duan H, Luo H. Identification of core gene expression signature and key pathways in colorectal cancer. Front Genet. 2020;11:45. doi:10.3389/fgene.2020.00045
14. Liu ZK, Zhang RY, Yong YL, et al. Identification of crucial genes based on expression profiles of hepatocellular carcinomas by bioinformatics analysis. Peer J. 2019;7:e7436. doi:10.7717/peerj.7436
15. Wan Z, Zhang X, Luo Y, Zhao B. Identification of hepatocellular carcinoma-related potential genes and pathways through bioinformatic-based analyses. Genet Test Mol Biomarkers. 2019;23(11):766–777. doi:10.1089/gtmb.2019.0063
16. Zhou L, Du Y, Kong L, Zhang X, Chen Q. Identification of molecular target genes and key pathways in hepatocellular carcinoma by bioinformatics analysis. Onco Targets Ther. 2018;11:1861–1869. doi:10.2147/OTT.S156737
17. Arai T, Okato A, Yamada Y, et al. Regulation of NCAPG by miR-99a-3p (passenger strand) inhibits cancer cell aggressiveness and is involved in CRPC. Cancer Med. 2018;7(5):1988–2002. doi:10.1002/cam4.1455
18. Jiang L, Ren L, Chen H, et al. NCAPG confers trastuzumab resistance via activating SRC/STAT3 signaling pathway in HER2-positive breast cancer. Cell Death Dis. 2020;11(7):547. doi:10.1038/s41419-020-02753-x
19. Ai J, Gong C, Wu J, et al. MicroRNA‑181c suppresses growth and metastasis of hepatocellular carcinoma by modulating NCAPG. Cancer Manag Res. 2019;11:3455–3467. doi:10.2147/CMAR.S197716
20. Gong C, Ai J, Fan Y, et al. NCAPG promotes the proliferation of hepatocellular carcinoma through PI3K/AKT signaling. Onco Targets Ther. 2019;12:8537–8552. doi:10.2147/OTT.S217916
21. Wang Y, Gao B, Tan PY, et al. Genome-wide CRISPR knockout screens identify NCAPG as an essential oncogene for hepatocellular carcinoma tumor growth. FASEB J. 2019;33(8):8759–8770. doi:10.1096/fj.201802213RR
22. Zhang Q, Su R, Shan C, Gao C, Wu P. Non-SMC condensin I complex, subunit G (NCAPG) is a novel mitotic gene required for hepatocellular cancer cell proliferation and migration. Oncol Res. 2018;26(2):269–276. doi:10.3727/096504017X15075967560980
23. Song S, Peng P, Tang Z, et al. Decreased expression of STING predicts poor prognosis in patients with gastric cancer. Sci Rep. 2017;7:39858. doi:10.1016/j.repc.2014.02.022
24. Pontillo D, Zampi G, Pergolini A, Polizzi V, Tinti D, Spadaccia P. Cardiac metastasis from colorectal cancer: to be or not to be …. Rev Port Cardiol. 2014;33(9):569–570. doi:10.1016/j.repc.2014.02.022
25. Yu L, Gu T, Shi E, et al. Primary malignant cardiac tumors. J Cancer Res Clin Oncol. 2014;140(6):1047–1055. doi:10.1007/s00432-014-1651-1
26. Si X, Xu F, Xu F, Wei M, Ge Y, Chenge S. CADM1 inhibits ovarian cancer cell proliferation and migration by potentially regulating the PI3K/Akt/mTOR pathway. Biomed Pharmacother. 2020;123:109717. doi:10.1016/j.biopha.2019.109717
27. Rong L, Li Z, Leng X, et al. Salidroside induces apoptosis and protective autophagy in human gastric cancer AGS cells through the PI3K/Akt/mTOR pathway. Biomed Pharmacother. 2020;122:109726. doi:10.1016/j.biopha.2019.109726
28. Fang S, Zhang D, Weng W, et al. CPSF7 regulates liver cancer growth and metastasis by facilitating WWP2-FL and targeting the WWP2/PTEN/AKT signaling pathway. Biochim Biophys Acta Mol Cell Res. 2020;1867(2):118624. doi:10.1016/j.bbamcr.2019.118624
29. Zhang F, Zhang YY, Sun YS, et al. Asparanin A from Asparagus officinalis L. induces G0/G1 cell cycle arrest and apoptosis in human endometrial carcinoma ishikawa cells via mitochondrial and PI3K/AKT signaling pathways. J Agric Food Chem. 2020;68(1):213–224. doi:10.1021/acs.jafc.9b07103
30. Wang Y, Liu C, Xie Z, Lu H. Knockdown of TRIM47 inhibits breast cancer tumorigenesis and progression through the inactivation of PI3K/Akt pathway. Chem Biol Interact. 2020;317:108960. doi:10.1016/j.cbi.2020.108960
31. Xiao W, Liu Y, Dai M, et al. Rotenone restrains colon cancer cell viability, motility and epithelial‑mesenchymal transition and tumorigenesis in nude mice via the PI3K/AKT pathway. Int J Mol Med. 2020;46(2):700–708. doi:10.3892/ijmm.2020.4637
32. Kattan SW, Nafie MS, Elmgeed GA, Alelwani W, Badar M, Tantawy MA. Molecular docking, anti-proliferative activity and induction of apoptosis in human liver cancer cells treated with androstane derivatives: implication of PI3K/AKT/mTOR pathway. J Steroid Biochem Mol Biol. 2020;198:105604. doi:10.1016/j.jsbmb.2020.105604
33. Zhou J, Jiang YY, Chen H, Wu YC, Zhang L. Tanshinone I attenuates the malignant biological properties of ovarian cancer by inducing apoptosis and autophagy via the inactivation of PI3K/AKT/mTOR pathway. Cell Prolif. 2020;53(2):e12739. doi:10.1111/cpr.12739
34. Jiang J, Bai J, Qin T, Wang Z, Han L. NGF from pancreatic stellate cells induces pancreatic cancer proliferation and invasion by PI3K/AKT/GSK signal pathway. J Cell Mol Med. 2020;24(10):5901–5910. doi:10.1111/jcmm.15265
35. Sun J, Piao J, Li N, Yang Y, Kim KY, Lin Z. Valproic acid targets HDAC1/2 and HDAC1/PTEN/Akt signalling to inhibit cell proliferation via the induction of autophagy in gastric cancer. FEBS J. 2020;287(10):2118–2133. doi:10.1111/febs.15122
36. Sun DP, Lin CC, Hung ST, et al. Aberrant expression of NCAPG is associated with prognosis and progression of gastric cancer. Cancer Manag Res. 2020;12:7837–7846. doi:10.2147/CMAR.S248318
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The
full terms of this license are available at https://www.dovepress.com/terms.php
and incorporate the Creative Commons Attribution
- Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted
without any further permission from Dove Medical Press Limited, provided the work is properly
attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.