Back to Journals » Infection and Drug Resistance » Volume 12
Molecular Characterization Of Vancomycin-Resistant Enterococcus faecalis Among Inpatients At Iranian University Hospitals: Clonal Dissemination Of ST6 And ST422
Authors Zalipour M, Esfahani BN, Halaji M , Azimian A , Havaei SA
Received 30 May 2019
Accepted for publication 5 September 2019
Published 25 September 2019 Volume 2019:12 Pages 3039—3047
DOI https://doi.org/10.2147/IDR.S217718
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Joachim Wink
Mehrdad Zalipour,1 Bahram Nasr Esfahani,1 Mehrdad Halaji,1 Amir Azimian,2 Seyed Asghar Havaei1,3
1Department of Microbiology, School of Medicine, Isfahan University of Medical Sciences, Isfahan, Iran; 2Department of Pathobiology and Laboratory Sciences, North Khorasan University of Medical Sciences, Bojnurd, Iran; 3Infectious Diseases and Tropical Medicine Research Center, Isfahan University of Medical Sciences, Isfahan, Iran
Correspondence: Seyed Asghar Havaei
Department of Microbiology, School of Medicine, Isfahan University of Medical Sciences, Hezar-Jerib Avenue, Isfahan 81746 73461, Iran
Tel +983137922478
Email [email protected]
Purpose: Over the past two decades, enterococci have emerged as an important opportunistic pathogen causing life-threatening infections in hospitals. The purpose of the present study was to examine the prevalence of genes encoding virulence factor and molecular characterization of vancomycin-resistant E. faecalis strains isolated from hospitalized patients in Isfahan, the central city of Iran.
Patients and methods: A total of 53 vancomycin-resistant E. faecalis isolates (VRE) obtained from clinical samples of hospitalized patients were characterized by phenotypic and genotypic methods, and 25 selected VRE isolates from internal and ICU wards were typed by multilocus sequence typing.
Results: The efa was the most prevalent virulence gene (100%) among isolates, followed by gelE (92.45%), asa1 (90.56%), ace (86.79%), esp (75.47%), cylA (39.62%), and hyl (18.86%). More than 80% of the isolates were HLGR. Multilocus sequence typing showed eight different sequence types including ST6, ST422, ST28, ST448, ST531, ST328, ST421, and ST495. STs were grouped into two clonal complex (CC) including CCA (ST6, ST422, ST448, ST531) and CCF (ST28, ST421) and two singletons (ST328, ST495).
Conclusion: Our data indicated a high prevalence of virulence genes among STs described in this study. In addition, the molecular analysis demonstrated a relatively high genetic diversity among selected VRE strains from the ICU in comparison with the internal ward. Therefore, in order to prevent the colonization of virulent strains in the hospital environment, infection control procedures should be performed.
Keywords: Enterococcus faecalis, virulence factors, vancomycin resistant, multilocus sequence typing, Iran
Introduction
Nosocomial infections (NIs) are a widespread problem in the current clinical setting and occurs in approximately 4% to 10% of hospitalized patients annually.1,2 Previous studies indicate that the most common bacterial isolates from hospitalized patients are methicillin-resistant Staphylococcus aureus (MRSA), vancomycin-resistant enterococci (VRE), multidrug-resistant gram-negative bacteria, and Clostridium difficile.3 Enterococci, in particular, VRE, is one of the most important nosocomial pathogens, that causes serious infections such as bloodstream infections (BSIs), catheter-associated urinary tract infections (CAUTIs), and intra-abdominal and intra-pelvic abscesses.4,5 According to previous reports, prolonged hospitalization, immunodeficiency, and uncontrolled antibiotic administration as a high risk of conditions are responsible for acquiring enterococcal infections in patients.6 Moreover, their ability to acquire specific genetic traits, such as virulence and antibiotic resistance determinants, plays an important role in the success and survival of enterococci in the hospital environment.7 Virulence factors through the ability to adhere to a range of extracellular matrix proteins and followed by colonization and invasion into the host tissues lead to modulation of the host immunity and production of pathological changes directly through the production of enzymes and toxins or indirectly through induction of inflammation, contributing to pathogenesis and severity of enterococcal infections.8
Therefore, characterization of adhesions and invasion factors such as enterococcal surface protein (Esp), aggregation substance (AS) proteins (Asa1), collagen-binding protein (Ace), gelatinase (GelE), hyaluronidase (Hyl) and cytolysin (CylA) can be useful to improve our understanding and assessment of the pathogenicity of enterococcal infections.9,10,11 Different molecular typing methods have been developed for the epidemiological investigations of enterococci.12 Among the known enterococcal molecular typing methods, pulsed-field gel electrophoresis (PFGE) was approved as a standard and efficient typing method with a high degree of discrimination,13 but PFGE remains difficult to standardize between laboratories due to its unsuitability for long-term epidemiology studies or for phylogenetic or population structure studies; this method is more susceptible to small genetic changes.14,15 However, the most appropriate technique for global and long-term epidemiology studies is multilocus sequence typing (MLST).15,16 In addition, MLST provides an unambiguous nomenclature for genotypes, and clones and data are easily stored in databases that can be exchanged between different laboratories via the Internet.16,17,18 The emergence of antimicrobial resistant virulent enterococci is a serious problem for hospital infection control practitioners and clinicians treating infected patients.19,20 Previous studies have demonstrated that Enteroccus faecalis is an important and virulent pathogen causing various infections.21 Although there are several reports on the endemicity of vancomycin-resistant enterococci (VRE) in Iran, limited information is available in relation to the virulence determinants and molecular relatedness of vancomycin-resistant E. faecalis isolates in Isfahan (Center city of Iran). Therefore, in the present study, we investigated the molecular characterization of vancomycin-resistant E. faecalis isolates among Iranian hospitalized patients as a first study using MLST technique.
Materials And Methods
Study Design And Samples
This cross-sectional study was performed during the 7-month period from April 2017 to October 2017 at four teaching hospitals affiliated to Isfahan University of Medical Science, Isfahan, Iran. The study was approved by the Ethics Committee and was in accordance with the declaration of Helsinki [IR.MUI.REC.1396.3.066].
A total of 53 vancomycin-resistant E. faecalis isolates (VRE) were obtained from true infections including urinary tract infections (UTIs) (n = 35), respiratory tract infections (n = 5), bloodstream infections (n = 4), wound infections (n=4), abdominal infections (n = 3), eye infection (n = 1), and meningitis (n=1).
The samples were cultured on blood agar (Merck, Germany) and then were incubated at 37°C for 24 hrs. Enterococcal isolates were identified according to conventional microbiological tests such as Gram staining (FARA CO.) (Gram positive), catalase reaction (catalase negative), growth on brain heart Infusion agar (Conda, Madrid, Spain) with 6.5% NaCl, and bile-esculin test (positive) (Merck, Germany). Furthermore, E. faecaliswas confirmed by amplification of the ddlE gene using species-specific primers (ddl E. faecalis F- 5ʹ- ATCAAGTACAGTTAGTCT-3ʹ and R-5ʹ-ACGATTCAAAGCTAACTG-3ʹ).22 The PCR protocol consisted of a pre-denaturation step at 95°C for 5 mins, followed by 30 cycles of 60 s at 95°C, 45 s at 48°C and 1 min at 72°C. A final extension step was performed at 72°C for 5 mins. However, high levels of vancomycin resistance and gentamicin resistance were detected by the E-test strips (Liofilchem, Italy) and gentamicin (120 μg) disk (Mast Group Ltd., UK.), respectively, on the Mueller–Hinton agar (Merck, Germany) in accordance with Clinical and Laboratory Standards Institute (CLSI) guidelines.23
DNA Extraction And Detection Of Virulence Genes
Genomic DNA was extracted from fresh colonies as described previously.21 PCR was performed for detecting seven of the genes encoding virulence factors including enterococcal surface protein (esp), gelatinase (gelE), aggregation substance (asa1), collagen-binding protein (ace), Enterococcus faecalis endocarditis antigen (efa), hyaluronidase (hyl), and cytolysin (cylA). Primers were synthesized as previously described and were as follows: esp, hyl, gelE, asa1,24 cylA,25 ace,26 and efa.27 The PCR products were separated by electrophoresis in 1% agarose gels with 1× TAE (Tris/Acetate/EDTA) buffer, stained with safe stain load dye (CinnaGen Co., Tehran, Iran) and visualized under ultraviolet illumination.
Multilocus Sequence Typing (MLST)
A total of 25 vancomycin-resistant E. faecalis isolates (VRE) with high levels of gentamicin resistance from internal (n=12) and intensive care units (ICU) (n=13) wards were subjected to MLST. Seven of E. faecalis housekeeping genes including gdh (glucose-6-phosphate dehydrogenase), gyd (glyceraldehyde-3-phosphate dehydrogenase), pstS (phosphate ATP-binding cassette transporter), gki (glucokinase, putative), aroE (shikimate 5-dehydrogenase), xpt (xanthine phosphoribosyl transferase), and yqil (acetyl-CoA acetyltransferase) were amplified by PCR method. Information of the E. faecalis MLST scheme, sequences of PCR primers, and the PCR conditions are available at MLST website (https://pubmlst.org/efaecalis/). STs were analyzed to determine clonal complex (CCs), single-locus variants (SLV), double-locus variants (DLV), and singletons using the eBURST algorithm (http://eburst.mlst.net/). Sequence types that were not grouped into a CC were defined as singletons. Also, the genetic relationship among strains was analyzed by constructing a dendrogram using eBURST program.
Statistical Analysis
The analysis was performed by using SPSSTM software, version 21.0 (IBM Corp., USA). The results are presented as descriptive statistics in terms of relative frequency.
Results
During 6 months of the study, a total of 53 VRE isolates were collected randomly with the possibility of true infections. Totally, of 53 confirmed VRE isolates, 56.6% and 43.4% isolates were obtained from female and male subjects, respectively.28 The most frequent isolates 35 (66%) were obtained from UTIs and followed that from respiratory infections and bloodstream infections. Meanwhile, 32% (17/53) and 26.4% (14/53) isolates were obtained from hospitalized patients in ICUs and internal wards, respectively. According to MDR definition, all of 53 E. faecalis strains were MDR (100%) and more than 80% of the isolates were high-level gentamicin resistance (HLGR) (Table 1).28
 |
Table 1 The Phenotypic And Genotypic Characteristics Of The 53 Vancomycin-resistant E. faecalis Isolates From Clinical Samples |
Distribution Of Virulence Genes
According to the results of PCR assay, the efa was most prevalent virulence gene (100%), followed by gelE (92.4%), asa1(90.6%), ace (86.8%), esp (75.5%), cylA (39.6%), and hyl (18.9%). However, none of the isolates contained all the virulent genes. The distribution of genes encoding virulence factors among VRE isolates and different STs is shown in Table 1.
Clonal Lineages Identified By MLST
MLST analysis was performed for 25 selected VRE isolates from internal and ICU wards that were classified into eight different STs: ST6, ST422, ST28, ST448, ST531, ST328, ST421, and ST495. Using the BURST v3 algorithm, STs were grouped into two CCs including CCA (ST6, ST422, ST448, ST531) and CCF (ST28, ST421) and two singletons (ST328, ST495). Two STs were found to be highly prevalent and comprised more than half of the isolates; 40% (10 isolates) of the isolates were ST6 and 24% (six isolates) were ST422, followed by ST28, ST448, and ST531 (two isolates), whereas other three STs were represented by a single isolate (ST328, ST421, and ST495). 91.6% (11/12) of the selected strains from internal wards were ST6 (seven isolates) and ST422 (four isolates), while selected isolates of ICU ward showed a higher diversity.
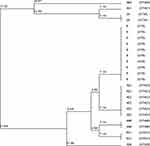
Phylogenetic analysis of the concatenated sequences, including all seven MLST genes, from the 25 VRE isolates examined in this study is demonstrated in Figure 1. The phylogenetic tree showed that there is a close phylogenetic relationship between most of the strains in our study.
Overall, the distribution of virulence genes indicated a high frequency of those among different STs, so that the asa1, efa, gelE, and ace genes were present in all STs and the prevalence of esp, cylA and hyl genes was 84.0%, 44.0% and 28.0%, respectively.
Discussion
Despite the fact that many studies have investigated antibiotic resistance patterns and the virulence factors of Enterococcus spp. isolates, only few studies have systematically investigated vancomycin-resistant E. faecalis isolates originating from clinical specimens in our country. Therefore, we characterized the vancomycin-resistant E. faecalis (VRE) collected from different clinical specimens during a 7-month period in 2017 from various teaching hospitals in Isfahan city, Iran. Similar to other reports from Iran, resistance to multiple classes of antibiotics was common in E. faecalis strains as observed in the current study.21,29,30 In this study, 81% of VRE strains were HLGR. Resistance to high concentrations of gentamicin among E. faecalis isolates has been showed in previous researches.30,31 High-level resistance to gentamicin is caused by the aminoglycoside-modifying enzymes, reducing the effect of aminoglycosides except for streptomycin.32 Antibiotic resistance is not sufficient for the pathogenesis of enterococcal infections and the presence of virulence factors plays an important role in the severity of infection.10 The VRE isolates characterized in this study showed a high prevalence of virulence genes. Our findings indicated that all of the isolates harbored efa gene and similar frequencies of the efaA were reported in various studies.21,26,33 It seems that the efaA gene is always present in clinical E. faecalis strains, whereas the strains isolated from other sources possessed the less frequency of EfaA determinant.26 As reported in other studies, gelE was identified as the second most common virulence factor among the isolates in the current study (92.45%).31,33 GelE seems to mediate virulence through effects such as degradation of host tissues and participates in the activation of autolysin, leading to the release of extracellular DNA and formation of a biofilm.11 The aggregation substance gene (asa1) was present in 90.56% of all isolates. These results are in agreement with those obtained by Choi et al and Nasaj et al who showed a high incidence of asa1 genes in clinical isolates of E. faecalis.31,34 AS proteins encoded by asa1 cause clumping of E. faecalis cells and mediate the high-frequency transfer of plasmid DNA between donor and recipient bacteria.8 These high rates of asa1 gene among all of our isolates can facilitate the exchange of resistance and virulence-associated genes via pheromone-responsive plasmids in a hospital setting. The results of our study indicated that the ace gene was present in 86.79% of isolates. Similarly, a high incidence of this gene in E. faecalis has been reported in previous studies.21,35,36 E. faecalis Ace is a collagen and laminin adhesin, which seems to be effective in endocarditis and UTIs.11 In the current study, the prevalence of the esp gene was also 75.47% in VRE strains. This finding in accordance with other studies indicate a possible role of the Esp as a colonization factor in UTI.10,26,34 The frequency of cylA and hyl genes was 39.62% and 18.86%, respectively. Previous studies by Heidari et al in Tehran and Shiraz showed that the virulence genes cylA and hyl were present at varying levels in E. faecalis isolates.21,33 These findings are in agreement with the results of Wanxiang et al, who showed that cylA and hyl genes were 19.4% and 19.6% in the Enterococcus isolates, respectively.37 During this study, we also identified that hyl-positive strains were HLGR, and this may be due to the co-presence of the hyl and aac(60)-Ie-aph(200)-Ia genes in a common transmissible plasmid.
Molecular typing of 25 strains of VRE isolated in Isfahan using MLST provided the first data for the knowledge on the genetic population structure for this species in Iran, allowing us to compare it to those obtained worldwide, and to know the spread of some E. faecalis clones to Iran. In the current study, eight different STs were found among the 25 VRE isolates, and distributed into two CCs, including CCA (ST6, ST422, ST448, and ST531), CCF (ST28, ST421), and two singletons (ST328, ST495). ST422 and ST531 represented an SLV of ST6 and ST448, respectively. ST531 is a DLV of ST422 belonging to CCA and is also reported in Denmark.38 Furthermore, ST531 represented SLV of ST6 belonging to CCA. ST28 is an SLV of ST421 belonging to CCF. However, our study results showed that most of the strains examined had a common genetic origin. ST6, which is the ancestor of CCA, was the most frequently found ST compared to other STs (40%), and ST422 was the second most detected ST among the isolates (24%). Furthermore, it was found that CCA contained the greatest number of STs identified in this study (20/25; 80%). E. faecalis strains identified as ST6 showed similar characteristics to ST6 clones isolated in Portugal and Poland39,40 and were characterized by high-level resistance to aminoglycosides, and vancomycin, but differed from the strains described in Cuba and Spain where they were vancomycin susceptible.12,41 ST6 has been reported worldwide, and it is probably the major lineage in CCA, which is often associated with invasive infections and resistance to multiple drugs, owing to its adaptability to hospital environment as a result of the acquisition of pathogenicity islands and antimicrobial resistance genes by recombination and horizontal gene transfer.14,42,43,44,45 Notably, 6 of the 25 isolates were found to be ST422 (Table 1). According to a review of studies worldwide, there are few reports of ST422 in E. faecalis, so that Hammerum et al report ST422 as HLGR, agreeing with our results.38 Other STs reported in this study were also found previously from different parts of the world.12,31,38,42,46 A high prevalence of virulence genes was also detected among STs described in this study, and there was no significant difference in the distribution of virulence factors between HLGR and non-HLGR isolates. There were some limitations related to the present study. First, a small number of VRE isolates were typed by MLST. Second, identification of the source of infection, taking preventive measures in the hospital setting, and molecular analysis of environmental specimens were required.
Conclusion
In summary, this research provided the first insight into the population structure of E. faecalis in Isfahan, Iran, and most of the strains examined in this study were related to European strains, and CCA was evidently circulating in Isfahan hospitals, being associated with MDR and virulence traits. In addition, our data indicate that E. faecalis strains isolated from clinical samples possess distinctive patterns of potential virulence factors with a high incidence of genes encoding virulence factors among isolates. It is recommended that some programs be performed to prevent the colonization of such virulent strains in the hospital environment, including better stewardship of antimicrobial agents and better awareness of the source for pathogen transmission in the hospital environment.
Ethics approval
The study protocol was approved by the Ethics Committee of Isfahan University of Medical Sciences (IR.MUI.REC.1396.3.066). However, we did not have human participants. The study used bacteria isolated from clinical samples in the clinical microbiology laboratory.
Acknowledgment
We are thankful to all members of the Department of Microbiology, School of Medicine, at Isfahan University of Medical Sciences. This was an original research paper from a Ph.D thesis.
Funding
This study was funded, in part, by a grant from the Isfahan University of Medical Sciences, [Grant no 396066].
Disclosure
The authors declare that they have no competing interests in this work.
References
1. Sligl WI, Dragan T, Smith SW. Nosocomial gram-negative bacteremia in intensive care: epidemiology, antimicrobial susceptibilities, and outcomes. Int J Infect Dis. 2015;37:129–134. doi:10.1016/j.ijid.2015.06.024
2. Schabrun S, Chipchase L. Healthcare equipment as a source of nosocomial infection: a systematic review. J Hosp Infect. 2006;63(3):239–245. doi:10.1016/j.jhin.2005.10.013
3. Bereket W, Hemalatha K, Getenet B, et al. Update on bacterial nosocomial infections. Eur Rev Med Pharmacol Sci. 2012;16(8):1039–1044.
4. Kafil HS, Asgharzadeh M. Vancomycin-resistant Enterococcus faecium and Enterococcus faecalis isolated from education hospital of iran. Maedica. 2014;9(4):323.
5. Oravcova V, Mihalcin M, Zakova J, Pospisilova L, Masarikova M, Literak I. Vancomycin-resistant enterococci with vanA gene in treated municipal wastewater and their association with human hospital strains. Sci Total Environ. 2017;609:633–643. doi:10.1016/j.scitotenv.2017.07.121
6. O’Driscoll T, Crank CW. Vancomycin-resistant enterococcal infections: epidemiology, clinical manifestations, and optimal management. Infect Drug Resist. 2015;8:217.
7. Sabouni F, Movahedi Z, Mahmoudi S, Pourakbari B, Keshavarz Valian S, Mamishi S. High frequency of vancomycin resistant Enterococcus faecalis in children: an alarming concern. J Prev Med Hyg. 2016;57(4):E201–e204.
8. Vu J, Carvalho J. Enterococcus: review of its physiology, pathogenesis, diseases and the challenges it poses for clinical microbiology. Front Biol. 2011;6(5):357. doi:10.1007/s11515-011-1167-x
9. Upadhyaya PG, Ravikumar K, Umapathy B. Review of virulence factors of enterococcus: an emerging nosocomial pathogen. Indian J Med Microbiol. 2009;27(4):301. doi:10.4103/0255-0857.55443
10. Arbabi L, Boustanshenas M, Rahbar M, et al. Antibiotic susceptibility pattern and virulence genes in Enterococcus spp. isolated from clinical samples of Milad hospital of Tehran, Iran. Arch Clin Infect Dis. 2016;11:3. doi:10.5812/archcid.36260
11. Arias CA, Murray BE. The rise of the Enterococcus: beyond vancomycin resistance. Nat Rev Microbiol. 2012;10(4):266. doi:10.1038/nrmicro2761
12. Quinones D, Kobayashi N, Nagashima S. Molecular epidemiologic analysis of Enterococcus faecalis isolates in Cuba by multilocus sequence typing. Microb Drug Resist. 2009;15(4):287–293. doi:10.1089/mdr.2009.0028
13. Chuang YC, Wang JT, Chen ML, Chen YC. Comparison of an automated repetitive-sequence-based PCR microbial typing system with pulsed-field gel electrophoresis for molecular typing of vancomycin-resistantEnterococcus faecium. J Clin Microbiol. 2010;48(8):2897–2901. doi:10.1128/JCM.00136-10
14. Ruiz-Garbajosa P, Bonten MJ, Robinson DA, et al. Multilocus sequence typing scheme for Enterococcus faecalis reveals hospital-adapted genetic complexes in a background of high rates of recombination. J Clin Microbiol. 2006;44(6):2220–2228. doi:10.1128/JCM.02596-05
15. Tong SY, Xie S, Richardson LJ, et al. High-resolution melting genotyping of Enterococcus faecium based on multilocus sequence typing derived single nucleotide polymorphisms. PloS One. 2011;6(12):e29189. doi:10.1371/journal.pone.0029189
16. Lytsy B, Engstrand L, Gustafsson A, Kaden R. Time to review the gold standard for genotyping vancomycin-resistant enterococci in epidemiology: comparing whole-genome sequencing with PFGE and MLST in three suspected outbreaks in Sweden during 2013–2015. Infect Genet Evol. 2017;54:74–80. doi:10.1016/j.meegid.2017.06.010
17. Werner G. Molecular typing of enterococci/VRE. J Bacteriol Parasitol S5–001. 2013;10:2155–9597.
18. Aanensen DM, Spratt BG. The multilocus sequence typing network: mlst. net. Nucleic Acids Res. 2005;33(suppl_2):W728–W733. doi:10.1093/nar/gki415
19. Miller WR, Munita JM, Arias CA. Mechanisms of antibiotic resistance in enterococci. Expert Rev Anti Infect Ther. 2014;12(10):1221–1236. doi:10.1586/14787210.2014.956092
20. Raza T, Ullah SR, Mehmood K, Andleeb S. Vancomycin resistant Enterococci: A brief review. J Pak Med Assoc. 2018;68(5):768–772.
21. Heidari H, Emaneini M, Dabiri H, Jabalameli F. Virulence factors, antimicrobial resistance pattern and molecular analysis of Enterococcal strains isolated from burn patients. Microb Pathog. 2016;90:93–97. doi:10.1016/j.micpath.2015.11.017
22. Dutka-Malen S, Evers S, Courvalin P. Detection of glycopeptide resistance genotypes and identification to the species level of clinically relevant enterococci by PCR. J Clin Microbiol. 1995;33(1):24–27.
23. Wayne P. CLSI. Performance Standards For Antimicrobial Susceptibility Testing; 26th Informational Supplement. CLSI Document M100. Clinical and Laboratory Standards Institute (CLSI); 2016.
24. Vankerckhoven V, Van Autgaerden T, Vael C, et al. Development of a multiplex PCR for the detection of asa1, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among European hospital isolates of Enterococcus faecium. J Clin Microbiol. 2004;42(10):4473–4479. doi:10.1128/JCM.42.10.4473-4479.2004
25. Coque TM, Patterson JE, Steckelberg JM, Murray BE. Incidence of hemolysin, gelatinase, and aggregation substance among enterococci isolated from patients with endocarditis and other infections and from feces of hospitalized and community-based persons. J Infect Dis. 1995;171(5):1223–1229. doi:10.1093/infdis/171.5.1223
26. Creti R, Imperi M, Bertuccini L, et al. Survey for virulence determinants among Enterococcus faecalis isolated from different sources. J Clin Microbiol. 2004;53(1):13–20.
27. Dupre I, Zanetti S, Schito AM, Fadda G, Sechi LA. Incidence of virulence determinants in clinical Enterococcus faecium and Enterococcus faecalis isolates collected in Sardinia (Italy). J Med Microbiol. 2003;52(Pt 6):491–498. doi:10.1099/jmm.0.05038-0
28. Zalipour M, Esfahani BN, Havaei SA. Phenotypic and genotypic characterization of glycopeptide, aminoglycoside and macrolide resistance among clinical isolates of Enterococcus faecalis: a multicenter based study. BMC Res Notes. 2019;12(1):292. doi:10.1186/s13104-019-4339-4
29. Asadollahi P, Razavi S, Asadollahi K, Pourshafie M, Talebi M. Rise of antibiotic resistance in clinical enterococcal isolates during 2001–2016 in Iran: a review. New Microbes New Infect. 2018;26:92–99. doi:10.1016/j.nmni.2018.08.018
30. Mohammadi F, Ghafourian S, Mohebi R, et al. Enterococcus faecalis as multidrug resistance strains in clinical isolates in Imam Reza Hospital in Kermanshah, Iran. Br J Biomed Sci. 2015;72(4):182–184.
31. Choi J-M, Woo G-J. Molecular characterization of high-level gentamicin-resistant Enterococcus faecalis from chicken meat in Korea. Int J Food Microbiol. 2013;165(1):1–6. doi:10.1016/j.ijfoodmicro.2013.02.016
32. Emaneini M, Aligholi M, Aminshahi M. Characterization of glycopeptides, aminoglycosides and macrolide resistance among Enterococcus faecalis and Enterococcus faecium isolates from hospitals in Tehran. Pol J Microbiol. 2008;57(2):173–178.
33. Heidari H, Hasanpour S, Ebrahim-Saraie HS, Motamedifar M. High incidence of virulence factors among clinical Enterococcus faecalis isolates in Southwestern Iran. J Infect Chemother. 2017;49(1):51–56. doi:10.3947/ic.2017.49.1.51
34. Nasaj M, Mousavi SM, Hosseini SM, Arabestani MR. Prevalence of virulence factors and vancomycin-resistant genes among Enterococcus faecalis and Enterococcus faecium isolated from clinical specimens. Iran J Public Health. 2016;45(6):806–813.
35. Iweriebor BC, Obi LC, Okoh AI. Virulence and antimicrobial resistance factors of Enterococcus spp. isolated from fecal samples from piggery farms in Eastern Cape, South Africa. BMC Microbiol. 2015;15(1):136. doi:10.1186/s12866-015-0468-7
36. Strateva T, Atanasova D, Savov E, Petrova G, Mitov I. Incidence of virulence determinants in clinical Enterococcus faecalis and Enterococcus faecium isolates collected in Bulgaria. Braz J Infect Dis. 2016;20(2):127–133. doi:10.1016/j.bjid.2015.11.011
37. Li W, Li J, Wei Q, et al. Characterization of aminoglycoside resistance and virulence genes among Enterococcus spp. isolated from a hospital in China. Int J Environ Res Public Health. 2015;12(3):3014–3025. doi:10.3390/ijerph120303014
38. Hammerum AM, Lester CH, Olsen SS, Holm A, Hansen DS, Schønheyder HC. Molecular characterisation of high-level gentamicin-resistant enterococci from bloodstream infections in Denmark: first description of clonal spread of aph (2′′)-Ib. Int J Antimicrob Agents. 2012;39(3):266–268. doi:10.1016/j.ijantimicag.2011.11.007
39. Freitas AR, Novais C, Ruiz-Garbajosa P, Coque TM, Peixe L. Clonal expansion within clonal complex 2 and spread of vancomycin-resistant plasmids among different genetic lineages of Enterococcus faecalis from Portugal. J Antimicrob Chemother. 2009;63(6):1104–1111. doi:10.1093/jac/dkp103
40. Kawalec M, Pietras Z, Daniłowicz E, et al. Clonal structure of Enterococcus faecalis isolated from Polish hospitals: characterization of epidemic clones. J Clin Microbiol. 2007;45(1):147–153. doi:10.1128/JCM.01704-06
41. Muruzábal‐Lecumberri I, Girbau C, Canut A, Alonso R, Fernández‐Astorga A. Spread of an Enterococcus faecalis sequence type 6 (CC 2) clone in patients undergoing selective decontamination of the digestive tract. Apmis. 2015;123(3):245–251. doi:10.1111/apm.12336
42. Bourafa N, Abat C, Loucif L, et al. Identification of vancomycin-susceptible major clones of clinical Enterococcus from Algeria. J Glob Antimicrob Resist. 2016;6:78–83. doi:10.1016/j.jgar.2016.03.009
43. Weng PL, Ramli R, Shamsudin MN, Cheah Y-K, Hamat RA. High genetic diversity of Enterococcus faecium and Enterococcus faecalis clinical isolates by pulsed-field gel electrophoresis and multilocus sequence typing from a hospital in Malaysia. Biomed Res Int. 2013;2013.
44. Santos BA, Oliveira JS, Cardoso NT, et al. Major globally disseminated clonal complexes of antimicrobial resistant enterococci associated with infections in cancer patients in Brazil. Infect Genet Evol. 2017;55:56–62. doi:10.1016/j.meegid.2017.08.027
45. Zheng JX, Wu Y, Lin ZW, et al. Characteristics of and virulence factors associated with biofilm formation in clinical Enterococcus faecalis isolates in China. Front Microbiol. 2017;8:2338. doi:10.3389/fmicb.2017.02338
46. Said LB, Klibi N, Lozano C, et al. Diversity of enterococcal species and characterization of high-level aminoglycoside resistant enterococci of samples of wastewater and surface water in Tunisia. Sci Total Environ. 2015;530:11–17. doi:10.1016/j.scitotenv.2015.05.091
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.