Back to Journals » Infection and Drug Resistance » Volume 13
Molecular Characteristics, Antimicrobial Resistance and Virulence Gene Profiles of Staphylococcus aureus Isolates from Wuhan, Central China
Authors Fu Y, Xiong M, Li X, Zhou J , Xiao X, Fang F, Cheng X, Le Y, Li Y
Received 15 February 2020
Accepted for publication 9 June 2020
Published 30 June 2020 Volume 2020:13 Pages 2063—2072
DOI https://doi.org/10.2147/IDR.S249988
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Professor Suresh Antony
Yu Fu, Mengyuan Xiong, Xuehan Li, Junying Zhou, Xiao Xiao, Fang Fang, Xiaohuan Cheng, Yingbang Le, Yirong Li
Department of Clinical Laboratory, Zhongnan Hospital, Wuhan University, Wuhan, People’s Republic of China
Correspondence: Yirong Li
Department of Clinical Laboratory, Zhongnan Hospital, Wuhan University, Wuhan, People’s Republic of China
Email [email protected]
Purpose: This study aimed to investigate the molecular characteristics, antimicrobial resistance and virulence genes profiles of S. aureus isolates from Wuhan, central China.
Materials and Methods: A total of 302 non-duplicate S. aureus isolates were collected successively during January–December 2018 and subjected to multi-locus sequence typing (MLST), staphylococcal protein A (spa) typing and Panton–Valentine leucocidin (PVL) and staphylococcal enterotoxin A, B, C, D, E, G, H and I (sea, seb, sec, sed, see, seg, seh and sei) detection. All methicillin-resistant S. aureus (MRSA) isolates were additionally subjected to staphylococcal chromosomal cassette mec(SCCmec) typing.
Results: Of the 302 S. aureus isolates, 131 were categorised as MRSA, yielding a rate 1.4 times the average rate in China during 2018 (43.4% vs 30.9%). Thirty-one sequence types (STs) and 82 spa types were identified. The most prevalent clones were ST5-t2460 (10.9%), ST239-t030 (9.3%), ST188-t189 (7.9%) and ST59-t437 (6.3%). Notably, the continued prevalence of ST239-t030 in Wuhan differs from other areas in China. SCCmec types and subtypes I, II, III, IVa and V were present in 0.8%, 36.6%, 26.0%, 20.6% and 8.4% of MRSA isolates. A comprehensive analysis identified ST5-t2460-SCCmec II (25.2%,), ST239-t030-SCCmec III (19.8%) and ST59-t437-SCCmec IVa (7.6%) as the major clones among MRSA isolates. The genes pvl, sea, seb, sec, sed, see, seg, seh and sei were detected at respective frequencies of 11.9%, 42.1%, 49.7%, 45.0%, 20.9%, 33.8%, 60.5%, 25.8% and 66.9%.
Conclusion: ST239-t030 remains one of the most prevalent clones in S. aureus isolates from Wuhan, leading us to conclude that S. aureus isolates from Wuhan possess unique molecular characteristics. The S. aureus isolates also exhibit unique antimicrobial resistance profiles and harbour relatively high numbers of enterotoxin virulence genes, compared with other reports from China.
Keywords: Staphylococcus aureus, molecular characteristics, antimicrobial resistance, virulence
Introduction
Staphylococcus aureus (S. aureus), a common conditionally pathogenetic bacterium that colonises the surface skin and mucosa of humans, is a leading cause of bacteraemia and infective endocarditis, as well as osteoarticular, dermal, soft tissue, pleuropulmonary and device-related infections1 and aerobic vaginitis which is associated with a significantly increased risk of preterm labour and low birth weight and so on.2 S. aureus harbours a diverse range of genotypes and virulence factors, which may influence the severity of infections and the occurrence of associated complications and mortality. For example, infection with S. aureus sequence type 121 (ST121), a strain with a high virulence potential, is often associated with longer durations of hospitalisation and antimicrobial therapy.3 The isolates clonal complex 30 (CC30) and CC5 frequently and easily induce hematogenous complications,4 while bloodstream infection with CC398, a methicillin-susceptible S. aureus (MSSA) strain, is associated with high mortality.5 Therefore, analyses of the molecular characteristics and virulence gene profiles of S. aureus are important for predicting the prognosis of a patient with S. aureus infection.
The molecular characteristics of S. aureus differ among countries. ST239 has been the most common clone in China and other Asian countries since the 1970s, where it accounts for approximately 75% of identified cases of hospital-acquired methicillin-resistant S. aureus (HA-MRSA).6 In contrast, ST8 (USA300) is the most frequently identified clone in the United States.7 What’s more, the distinction between the CA-MRSA and HA-MRSA has eroded slowly over time8 and it showed that the staphylococcus aureus strains have changed. Even within China, the molecular characteristics of S. aureus isolates vary among cities. For example, the predominant types are ST188 in Wenzhou,9 ST398 in Hainan,10 ST5 in Dalian and Shenyang11 and ST59 in Chengdu.12 Moreover, the most prevalent staphylococcal protein A (spa) types are t030 in Asia,13,14 t032 in Europe,15 t008 in America, t037 in Africa and t020 in Australia.16 The molecular characteristics of S. aureus also change over time. For example, ST239-t030 has rapidly replaced ST239-t037 since 2000 and has become the major clone among S. aureus isolates,17 but reports from some areas of China indicate that this clone was substituted with ST59-t437 in 2013 and with ST5-t2460 in 2017.18 Therefore, it is preferable to focus on a specific region of interest at a particular time when monitoring the molecular characteristics of S. aureus isolates.
The definition for community-associated methicillin-resistant Staphylococcus aureus(CA-MRSA) is the development of infection by individuals in an outpatient setting or by inpatients discharged within 48 hours of admission without a history of MRSA infection, surgery, dialysis, the presence of an indwelling catheter or devices that penetrate through the skin, or admission to an extended care facility.19 A previous epidemiological study conducted in Wuhan aimed simply to investigate the molecular characteristics and virulence gene profiles of community-acquired MRSA (CA-MRSA) isolates that caused skin and soft tissue infections. That study identified the ST59-SCCmecIVa-PVL-positive clone as predominant among the isolates.20 Another multi-centre study that included Wuhan, Changsha and Zhengzhou identified ST239-t030-SCCmecIII-agr-I and ST188- t189-agr-I as the most common MRSA and MSSA clones, respectively.21 Wuhan is the capital of Hubei province and the largest city in central China, with a population that has increased to 15 million. However, no previous study focused on the molecular characteristics and virulence gene profiles of S. aureus isolates, including MRSA and MSSA. Moreover, the molecular characteristics, antimicrobial resistance and virulence gene profiles of S. aureus isolates vary dramatically over time in the same region. Therefore, successive surveillance is an extremely critical component of S. aureus control and prevention. This study aimed to investigate the molecular characteristics, antimicrobial resistance and virulence gene profiles of S. aureus isolates sampled in Wuhan during 2018.
Materials and Methods
Clinical Isolates and Primers
From January to December 2018, 302 non-duplicate clinical isolates of S. aureus were collected successively from Zhongnan Hospital, Wuhan University. These isolates were derived from wound secretions (n = 111), sputum (n = 95), blood (n = 27) and other samples (n = 69), including tissues, cerebrospinal fluid, pleural effusion and ascites, catheter tips and urine. All isolates were first identified using traditional microbiological approaches, including Gram staining and coagulase and catalase tests, and were further identified using a VITEK 2 Compact system and the VITEK 2 GP Test Kit (bioMerieux, Inc., Durham, NC, USA). All samples were then stored at −80°C for further molecular experiments. All primers used in the present study were synthesised by Tianyihuiyuan Co. Ltd. (Wuhan, China). This study was approved by the Ethics Committee of Zhongnan Hospital, Wuhan University, which waived the requirement for informed consent because the details of patients with S. aureus infection were anonymised.
S. aureus Chromosomal DNA Extraction
All S. aureus isolates were cultured overnight on tryptic soy broth (TSB) agar plates at 35°C. Subsequently, single colonies were transferred into 5 mL of TSB and grown overnight at a rotation speed of 200 rpm. Chromosomal DNA was extracted from S. aureus pellets using TIANamp Bacteria DNA kit (Tiangen, China), as described previously.10,22 The extracted DNAs were stored at −20°C for further studies.
Staphylococcal Protein A (Spa) Typing
The X region, which comprises a variable repeat region of the spa, was amplified using a PCR assay as described previously.10,22 The resultant amplicons were then purified and subjected to Sanger dideoxy DNA sequencing (Tianyihuiyuan, China). All sequences were analysed using the Ridom web server (http://www.ridom.de/spaserver), and each clinical isolate was assigned to a spa type.
Multi-Locus Sequence Typing (MLST)
Seven house-keeping genes (arcC, aroE, glpF, gmk, pta, tpi and yqil) were amplified from S. aureus chromosomal DNA for MLST as described previously.10,22-24 The amplified products were also purified and subjected to Sanger dideoxy DNA sequencing (Tianyihuiyuan, China). The sequences of these genes were then compared with known alleles at each locus, which were stored in the MLST database (http://www.mlst.net). Each clinical isolate was then explained by the sequence type.
Staphylococcus Chromosomal Cassette mec(SCCmec) Typing
All MRSA isolates were subjected to SCCmec typing as described previously.10,25 In brief, amultiple PCR assay containing nine pairs of primers specific for S CCmec types and subtypes, including I, II, III, IVa, IVb, IVc, IVd and V and mecA was used for SCCmec typing. Subsequently, the classification of SCCmec types and subtypes I, II, III, IVa, IVb, IVc, IVd and V was performed based on the molecular weights of the amplified products. MRSA isolates without any specific band were classified as non-typable (NT).
Detection of pvl and Enterotoxin Genes
The S. aureus isolates were screened for the presence of the Panton–Valentine leucocidin (PVL) and enterotoxin genes, including staphylococcal enterotoxins A, B, C, D, E, G, H and I (sea, seb, sec, sed, see, seg, seh and sei, respectively), using nine independent PCR assays with specific primers and reaction conditions as described previously.26 In brief, all PCR reaction mixtures contained 1 μL of DNA template, 2 μL of a primer pair (10 μM), 12.5 μL of 2× Taq Master Mix and 9.5 μL of double-distilled water. All PCR products were detected by electrophoresis on 1% agarose gels.
Antimicrobial Susceptibility Test (AST)
AST was performed using a VITEK 2 Compact system and the VITEK 2 AST-GP67 Test Kit (bioMerieux, Inc., Durham, NC, USA). The VITEK 2 AST-GP67 Test Kit contains 16 antibiotics: levofloxacin (LVX), gentamicin (GEN), trimethoprim-sulfamethoxazole (TST), penicillin (PEN), ciprofloxacin (CIP), vancomycin (VAN), erythromycin (ERY), clindamycin (CLI), linezolid (LZD), tetracycline (TET), rifampin (RIF), oxacillin (OXA), cefoxitin (FOX), nitrofurantoin (NIT), quinupristin/dalfopristin (QUD) and moxifloxacin (MOF). S. aureus strains ATCC 25923 and 29213 were used as the quality control strains for AST. The results were analysed according to the Clinical and Laboratory Standards Institute (CLSI) M100-S27 standard.
Statistical Analysis
The statistical analysis was performed using SPSS Statistics 24.0 for Windows (IBM Corp., Armonk, NY, USA). The chi-square test was used to compare the resistance rates, prevalence of MDR and distributions of PVL and enterotoxin genes between the MSSA and MRSA isolates. A p value <0.05 was considered to indicate a statistically significant difference.
Results
The results of AST
The detailed results from the AST of 302 S. aureus isolates are presented in Table 1. Notably, all tested S. aureus isolates were susceptible to VAN, LZD and QUD, and most were also susceptible to NIT (99.3%, 300/302), TST (92.1%, 278/302) and RIF (87.1%, 263/302). However, most isolates were resistant to PEN (93.0%, 281/302). The rates of susceptibility to the remaining antibiotics ranged from 37.4% to 76.5%. Of the 302 S. aureus isolates, 131 (43.4%) were categorised as MRSA based on the OXA susceptibility test and FOX screening test, including 116 HA-MRSA and 15 CA-MRSA isolates as determined from patient medical records. The frequency of MRSA isolates was 1.4 times the average level of 30.9% reported in China during 2018 (www.carss.cn).
 |
Table 1 The Distribution of Antimicrobial Resistance of MRSA, MSSA and S. aureus Isolates and Its Statistical Significance |
The resistance rates of the MRSA isolates to CLI, ERY, PEN, RIF, LVX, TET, CIP, GEN and MOF were 78.6% (103/131), 80.2% (105/131), 100.0% (131/131), 28.2% (37/131), 60.3% (79/131), 58.0% (76/131), 60.3% (79/131), 42.0% (55/131) and 58.8% (77/131). These rates were significantly higher than those observed for MSSA (Table 1). The multidrug resistance (MDR) rates among all S. aureus, MRSA and MSSA isolates were 66.6%, 89.3% and 49.1%, respectively. The prevalence of MDR was significantly higher among MRSA isolates than among MSSA isolates (χ2 = 53.827, P < 0.001; chi-square test).
MLST, Spaand SCCmec Typing
All 302 S. aureus isolates were successfully typed for MLST and assigned to 31 sequence types. The detailed results of MLST typing are listed in Table 2. The most prevalent ST type was ST5 (20.5%, 62/302), followed by ST239 (11.6%, 35/302), ST59 (10.3%, 31/302) and ST188 (8.9%, 27/302). Notably, 67.7% (42/62) of ST5, 77.4% (24/31) of ST59 and 97.1% (34/35) of ST239 isolates were MRSA, whereas 100% of ST188 isolates were MSSA. Among the 131 MRSA isolates, ST5 (32.1%, 42/131) was the most prevalent, followed by ST239 (26.0%, 34/131) and ST59 (18.3%, 24/131). Among MSSA isolates, ST188 (15.8%, 27/171) was the most common clone, followed by ST7 (14.0%, 24/171) and ST5 (11.7%, 20/171). In addition to ST1281, 30 other STs were identified as belonging to 16 clonal complexes (CCs) by eBURST. As shown in Fig. 1 and Table 2, CC5 (22.5%, 68/302) was the most prevalent clone complex, followed by CC8 (15.9%, 48/302), CC59 (10.6%, 32/302) and CC188 (8.9%, 27/302).
Of the 302 isolates, 294 were successfully typed and assigned to 82 spa types, whereas eight isolates were NT (Table 2). The most prevalent spa types were t2460 (11.2%, 33/302), t030 (9.5%, 28/302), t189 (8.2%, 24/302) and t437 (6.8%, 20/302). The MRSA isolates were assigned to 33 spa types, of which t2460 was the most prevalent. The MSSA isolates yielded 63 spa types, among which t189 was dominant (Table 2). Fourteen spa types—t002, t030, t062, t084, t127, t701, t309, t3527, t437, t441, t2091, t13828, t034 and t4549—were identified in both MRSA and MSSA isolates. In a combined analysis of the ST and spa types, strong associations were observed between the main ST and spa types. Specifically, ST188 and ST239 were primarily associated with t189 (88.9%, 24/27) and t030 (80.0%, 28/35), respectively, whereas ST5 and ST59 were mainly associated with t2460 (53.2%, 33/62) and t437 (61.3%, 19/31), respectively. Therefore, the major epidemic clones were ST5-t2460 (10.9%, 33/302), ST239-t030 (9.3%, 28/302), ST188-t189 (7.9%, 24/302) and ST59-t437 (6.3%, 19/302).
SCCmectyping was performed successfully for 121 out of the 131 MRSA isolates(92.4%). The SCCmec types and subtypes I, II, III, IVa and V were identified in 1 (0.8%), 48 (36.6%), 34 (26.0%), 27 (20.6%) and 11 (8.4%) MRSA isolates, respectively (Table 2). SCCmec type II was the most prevalent followed by III and IVa. The majority(85.4%, 41/48) of the SCCmec type IIMRSA isolates were ST5, while most type III MRSA isolates (94.1%, 32/34) were ST239 and IVa was primarily associated with ST59 (74.1%, 20/27). A comprehensive analysis of the results from MLST, spa and SCCmec typing identified ST5-t2460-SCCmec II (25.2%, 33/131), ST239-t030-SCCmec III (19.8%, 26/131) and ST59-t437-SCCmec IVa (7.6%, 10/131) as the major clones of MRSA isolates.
 |
Table 2 The Results of Different Typing of All the 302 Non-Duplicate S. aureus Isolates |
PVL and Enterotoxin Gene Profiles
The frequencies of the PVL and enterotoxin genes in the 302 S. aureus isolates are listed in Table 3. Thirty-six of the 302 isolates (11.9%) were positive for PVL, and 14 of these were identified as MRSA (including nine CA-MRSA isolates). The frequency of PVL positivity differed significantly between the 15 CA-MRSA and 116 HA-MRSA isolates (64.3% vs 4.3%, χ2 = 37.520, p < 0.001). The most frequently identified enterotoxin gene was sei (66.9%), followed by seg (60.5%). The frequencies of the other enterotoxin genes ranged from 20.9% to 49.7%. Compared with the MSSA isolates, the MRSA isolates had higher frequencies of sec and see positivity and a lower rate of seb and seh positivity (Table 3). On average, each isolate harboured 3.5 enterotoxin genes, although 156 isolates harboured four or more enterotoxin virulence genes. Specifically, 5, 13, 55 and 83 isolates harboured seven, six, five or four enterotoxin virulence genes, respectively. The frequency of clones harbouring four or more enterotoxin genes did not differ significantly between the MRSA and MSSA isolates (χ2 = 2.438, p > 0.05). The most common enterotoxin virulence genotype was seb-sec-seg-sei (8.9%, 27/302), followed by seb-seg-sei (3.0%, 9/302).
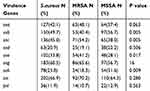 |
Table 3 The Frequencies of PVL and Enterotoxin Genes of the 302 Non-Duplicate S. aureus Strains Both in MRSA and MSSA |
Discussion
In this study, 302 S. aureus isolates were subjected to AST, and 131 (43.4%) were classified as MRSA according to OXA and FOX susceptibility. All 131 isolates were mecA-positive, which further supported methicillin resistance. Notably, the prevalence of MRSA in this study was 1.4 times the average level of 30.9% in China during 2018 (maximum, 46.8%; minimum, 15.5%) (www.carss.cn). In 2017, the respective frequencies of MRSA in Russia, Japan and Australia were 16.0%, 41.0% and 19.0% (resistancemap.cddep.org/AntibioticResistance.php). In other words, the prevalene of MRSA varies among countries and cities. Most MRSA isolates evaluated in this study were classified as SCCmec II, although III and IVa were also abundant. In contrast, types I and Vwere less represented. Our finding differ from the recent studies that the most common SCCmec types were III in Guangdong and Xinjiang27,28 and IV in Shanghai29 and Trento.30 In this study, AST also revealed that all or most of the isolates were susceptible to VAN, LZD, QUD, TST and NIT but were resistant to PEN, consistent with the results of most previous studies conducted in China and worldwide.31–33 Our findings support previous evidence suggesting that QUD and LZD are valuable substitutes for VAN in the treatment of MRSA infections.34 In addition, the rates of resistance to five antibiotics including CIP, CLI, GEN, LVF and RIF among the MRSA isolates in this study differed from the average levels reported in China in 2018 (www.chinets.com). Therefore, we conclude that S. aureus isolates in Wuhan has an unique antibiotic susceptibility, which might be associated with differences in health care programmes and drug usage patterns.
Thirty-one different sequence types were identified among the 302 S. aureus isolates in this study. The most prevalent sequence types in both the total S. aureus and MRSA isolates were ST5 and ST239, and this finding differs from other reports.27 Specifically, the predominant MRSA isolates were ST5 and ST398 in Shanghai in 2017,18 ST239 and ST338 in Guangdong during 2010–2016,28 ST239 and ST15 in Iran during 2015–1635 and ST5 and ST8 in the United States during 2010–2011.36 The predominant MSSA genotypes were ST188 and ST7, which was in accordance with the majority of previous studies.37 Therefore, the common STs may change over time and are frequently associated with specific geographic areas.
In this study, 294 S.aureus isolates were assigned to 82 spa types, indicating a high level of diversity and complexity with respect to the genetic backgrounds. We identified t2460, t030 and t189 as the predominant spa types among the S. aureus isolates. Of these three types, t2460 was predominant among MRSA isolates, whereas t189 was most common among MSSA isolates. We reviewed 253 articles that describe studies in Europe, Asia, America, Africa and Australia and were published online before 17 December 2017. According to these reports, the most prevalent spa types were t030, t037 and t002 in Asia; t032, t008 and t002 in Europe; t008, t002 and t242 in America; t037, t84 and t064 in Africa; and t202, t037 and t437 in Australia.16 Our study and other studies conducted in China during 2018–2019 identified t2460, rather than t030, as the most prevalent spa type among S. aureus and MRSA isolates.18,22,38,39 Therefore, we conclude that both the common spa type and common sequence type also change over time and are frequently associated with specific geographic areas. These observations suggest that the polymorphic X region may be associated with the ability of the organism to adapt to conditions such as diverse geographic areas, temperatures and host populations.
In this study, ST188, ST239, ST5 and ST59 were associated primarily with t189, t030, t2460 and t437, respectively, consistent with the findings of most previous studies.10,18,40 We then combined the sequence and spa types and identified ST5-t2460 as the most prevalent pandemic clone, followed by ST239-t030, ST188-t189 and ST59-t437. A multi-centre study of S. aureus bacteraemia at eight hospitals from four cities in China during 2017 also identified ST5-t2460 as the most common genotype, followed by ST188-t189 and ST59-t437.40 Another study of S. aureus in Shanghai during 2017 also identified ST5-t2460 as the predominant genotype, followed by ST5-t002, ST398-t034 and ST7-t091.18 Our results further demonstrate that ST239-t030, a truly pandemic MRSA isolate that has circulated in China and worldwide since the 1970s, remains one of the most prevalent clones in Wuhan, despite its exclusion from the top rank. This observation differs from those reported in other areas of China,18,40 leading us to conclude that S. aureus isolates from Wuhan possess unique molecular characteristics.
S. aureus isolates that harbour pvl can potentially secrete the cytotoxin PVL, which causes tissue necrosis and leukocyte destruction. This gene is generally considered a stable marker of CA-MRSA in some European countries and the United States.41–43 In this study, pvl was detected in 36 (11.9%) S. aureus isolates, of which 14 were MRSA and 5 were HA-MRSA. Our statistical analysis revealed a higher rate of pvl positivity among CA-MRSA relative to HA-MRSA isolates, suggesting that this gene could be used to identify CA-MRSA. Previous studies revealed low rates of pvl positivity among MRSA isolates in Korea (1.7%)44 and various European countries, including Germany and Spain (0–4.4%).45 In contrast, MRSA was isolated from 48% of patients with skin and skin structural infections at an emergency department in the United States, and 92.8% of these MRSA isolates harboured pvl.46 Therefore, the prevalence of pvl positivity differs among countries. In this study, we tested eight enterotoxin virulence genes and identified sei as the most prevalent, followed by seg. Of the tested enterotoxin virulence genes, six were present in >25.8% of S. aureus isolates, whereas in a previous study conducted in Shanghai, 13 enterotoxin genes (including those tested in our study) were present in 5.1–23.1% of S. aureus isolates.29 Moreover, we observed an average rate of 3.6 enterotoxin genes per isolate, which was about two times the rate reported previously in Shanghai.29 Another study performed in Guangzhou reported that the prevalence of sea, seb, sec and sed in MRSA isolates ranged from 0.97% to 21.4%,29 whereas the corresponding rates in this study were 42.1%, 49.7%, 45.0% and 20.9%, respectively. Therefore, we conclude that S. aureus isolates from Wuhan more frequently harbour enterotoxin genes than isolates from Shanghai and Guangdong.
In conclusion, this study is the first to simultaneously reveal the antibiotic resistance, molecular characteristics and virulence genes profiles of S. aureus isolates from Wuhan, China. ST239-t030 remains one of the most prevalent clones in S. aureus isolates from Wuhan. The S. aureus isolates from this city exhibit unique antimicrobial resistance profiles and harbour relatively high numbers of enterotoxin virulence genes, compared with other reports from China.
Disclosure
The authors report no conflicts of interest in this work.
References
1. Tong SYC, Davis JS, Emily E, Holland TL, Fowler VG. Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev. 2015;28(3):603–661. doi:10.1128/CMR.00134-14
2. Gajdacs M, Urban E. Epidemiology and resistance trends of Staphylococcus aureus isolated from vaginal samples: a 10-year retrospective study in Hungary. Acta Dermatovenerologica Alpina Pannonica Et Adriatica. 2019;28(4):143–147. doi:10.15570/actaapa.2019.35
3. Rao Q, Shang W, Hu X, Rao X. Staphylococcus aureus ST121: a globally disseminated hypervirulent clone. J Med Microbiol. 2015;64(12):1462. doi:10.1099/jmm.0.000185
4. Chen H, Liu Y, Jiang X, Chen M, Wang H. Rapid change of methicillin-resistant staphylococcus aureus clones in a Chinese Tertiary Care Hospital over a 15-year period. Antimicrob Agents Chemother. 2010;54(5):1842–1847.
5. Bouiller K, Gbaguidi-Haore H, Hocquet D, Cholley P, Bertrand X, Chirouze C. Clonal complex 398 methicillin-susceptible Staphylococcus aureus bloodstream infections are associated with high mortality. Clin Microbiol Infect. 2016;22(5):451–455. doi:10.1016/j.cmi.2016.01.018
6. Wang Z, Zhou H, Wang H, et al. Comparative genomics of methicillin-resistant Staphylococcus aureus ST239: distinct geographical variants in Beijing and Hong Kong. BMC Genomics. 2014;15(1):1–10. doi:10.1186/1471-2164-15-529
7. O’Hara FP, Amrinemadsen H, Mera RM, et al. Molecular characterization of Staphylococcus aureus in the United States 2004–2008 reveals the rapid expansion of USA300 among inpatients and outpatients. Microb Drug Resist. 2012;18(6):555. doi:10.1089/mdr.2012.0056
8. Gajdacs M. The continuing threat of methicillin-resistant Staphylococcus aureus. Antibiotics-Basel. 2019;8(2):27.
9. Fangyou YU, Tingjian LI, Huang X, et al. Virulence gene profiling and molecular characterization of hospital-acquired Staphylococcus aureus isolates associated with bloodstream infection. Diagn Microbiol Infect Dis. 2012;74(4):363–368. doi:10.1016/j.diagmicrobio.2012.08.015
10. Li X, Huang T, Xu K, Li C, Li Y. Molecular characteristics and virulence gene profiles of Staphylococcus aureus isolates in Hainan, China. BMC Infect Dis. 2019;19(1):873. doi:10.1186/s12879-019-4547-5
11. Yudong L, Hui W, Na D, et al. Molecular evidence for spread of two major methicillin-resistant Staphylococcus aureus clones with a unique geographic distribution in Chinese hospitals. Antimicrob Agents Chemother. 2009;53(2):512–518. doi:10.1128/AAC.00804-08
12. Tan S, Wan C, Wang H, Zhou W, Shu M. Relationship between nasal Carrier isolates and clinical isolates in children with Staphylococcus aureus infections. Microb Pathog. 2019;127:233–238. doi:10.1016/j.micpath.2018.11.032
13. Chen Y, Liu Z, Duo L, et al. Characterization of Staphylococcus aureus from distinct geographic locations in China: an increasing prevalence of spa-t030 and SCCmec type III. PLoS One. 2014;9(4):e96255. doi:10.1371/journal.pone.0096255
14. Ghasemzadeh-Moghaddam H, Ghaznavi-Rad E, Sekawi Z, et al. Methicillin-susceptible Staphylococcus aureus from clinical and community sources are genetically diverse. Int J Med Microbiol. 2011;301(4):347–353. doi:10.1016/j.ijmm.2010.10.004
15. Mehraj J, Akmatov MK, Strompl J, et al. Methicillin-sensitive and methicillin-resistant Staphylococcus aureus nasal carriage in a random sample of non-hospitalized adult population in northern Germany. PLoS One. 2014;9(9):e107937. doi:10.1371/journal.pone.0107937
16. Asadollahi P, Farahani NN, Mirzaii M, et al. Distribution of the most prevalent spa types among clinical isolates of methicillin-resistant and -susceptible staphylococcus aureus around the world: a review. Front Microbiol. 2018;9:163. doi:10.3389/fmicb.2018.00163
17. Hongbin C, Yudong L, Xiuhong J, Minjun C, Hui W. Rapid change of methicillin-resistant Staphylococcus aureus clones in a Chinese tertiary care hospital over a 15-year period. Antimicrob Agents Chemother. 2010;54(5):1842.
18. Dai Y, Liu J, Guo W, et al. Decreasing methicillin-resistant Staphylococcus aureus (MRSA) infections is attributable to the disappearance of predominant MRSA ST239 clones, Shanghai, 2008-2017. Emer Microbes Infect. 2019;8(1):471–478. doi:10.1080/22221751.2019.1595161
19. Tidwell J, Kirk L, Luttrell T, Pike CA. CA-MRSA decolonization strategies do they reduce recurrence rate? J Wound Ostomy Cont Nurs. 2016;43(6):577–582. doi:10.1097/WON.0000000000000277
20. Liu X, Liang J, Jiang Y, et al. Molecular characteristics of community-acquired methicillin-resistant Staphylococcus aureus strains isolated from outpatients with skin and soft tissue infections in Wuhan, China. Pathog Dis. 2016;74:4. doi:10.1093/femspd/ftw026
21. Liu C, Chen ZJ, Sun Z, et al. Molecular characteristics and virulence factors in methicillin-susceptible, resistant, and heterogeneous vancomycin-intermediate Staphylococcus aureus from central-southern China. J Microbiol Immunol Infect. 2015;48(5):490–496. doi:10.1016/j.jmii.2014.03.003
22. Li X, Fang F, Zhao J, et al. Molecular characteristics and virulence gene profiles of Staphylococcus aureus causing bloodstream infection. Braz J Infect Dis. 2018;22.
23. Enright MC, Day NP, Davies CE, Peacock SJ, Spratt BG. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J Clin Microbiol. 2000;38(3):1008–1015. doi:10.1128/JCM.38.3.1008-1015.2000
24. Robinson DA, Enright MC. Multilocus sequence typing and the evolution of methicillin-resistant Staphylococcus aureus. Clin Microbiol Infect. 2004;10(2):92–97. doi:10.1111/j.1469-0691.2004.00768.x
25. Ito T, Kuwahara-Arai K, Katayama Y, Uehara Y, Hiramatsu K. Staphylococcal Cassette Chromosome mec (SCCmec) analysis of MRSA. Methods Mol Biol. 2007;391:131–148.
26. Campbell SJ, Deshmukh HS, Nelson CL, et al. Genotypic characteristics of staphylococcus aureus isolates from a multinational trial of complicated skin and skin structure infections. J Clin Microbiol. 2008;46(2):678. doi:10.1128/JCM.01822-07
27. Yuan W, Liu J, Zhan Y, et al. Molecular typing revealed the emergence of pvl-positive sequence type 22 methicillin-susceptible Staphylococcus aureus in Urumqi, Northwestern China. Infect Drug Resist. 2019;12:1719–1728. doi:10.2147/IDR.S202906
28. Liang Y, Tu C, Tan C, et al. Antimicrobial resistance, virulence genes profiling and molecular relatedness of methicillin-resistant Staphylococcus aureus strains isolated from hospitalized patients in Guangdong Province, China. Infect Drug Resist. 2019;12:447–459. doi:10.2147/IDR.S192611
29. Wang X, Li X, Liu W, Huang W, Fu Q, Li M. Molecular characteristic and virulence gene profiles of community-associated methicillin-resistant Staphylococcus aureus isolates from pediatric patients in Shanghai, China. Front Microbiol. 2016;7:1818. doi:10.3389/fmicb.2016.01818
30. Manara S, Pasolli E, Dolce D, et al. Whole-genome epidemiology, characterisation, and phylogenetic reconstruction of Staphylococcus aureus strains in a paediatric hospital. Genome Med. 2018;10(1):82. doi:10.1186/s13073-018-0593-7
31. McGuinness WA, Malachowa N, DeLeo FR. Vancomycin resistance in Staphylococcus aureus. Yale J Biol Med. 2017;90(2):269–281.
32. Wu D, Wang Q, Yang Y, et al. Epidemiology and molecular characteristics of community-associated methicillin-resistant and methicillin-susceptible Staphylococcus aureus from skin/soft tissue infections in a children’s hospital in Beijing, China. Diagn Microbiol Infect Dis. 2010;67(1):1–8. doi:10.1016/j.diagmicrobio.2009.12.006
33. Yu Y, Yao Y, Weng Q, et al. Dissemination and molecular characterization of Staphylococcus aureus at a Tertiary Referral Hospital in Xiamen City, China. Biomed Res Int. 2017;2017:1367179. doi:10.1155/2017/1367179
34. Culos KA, Cannon JP, Grim SA. Alternative agents to vancomycin for the treatment of methicillin-resistant Staphylococcus aureus infections. Am J Ther. 2013;20(2):200–212. doi:10.1097/MJT.0b013e31821109ec
35. Goudarzi M, Bahramian M, Satarzadeh Tabrizi M, et al. Genetic diversity of methicillin resistant Staphylococcus aureus strains isolated from burn patients in Iran: ST239-SCCmec III/t037 emerges as the major clone. Microb Pathog. 2017;105:1–7. doi:10.1016/j.micpath.2017.02.004
36. Velasco V, Buyukcangaz E, Sherwood JS, Stepan RM, Koslofsky RJ, Logue CM. Characterization of Staphylococcus aureus from humans and a comparison with isolates of animal origin, in North Dakota, United States. PLoS One. 2015;10(10):e0140497. doi:10.1371/journal.pone.0140497
37. Li S, Sun S, Yang C, et al. The changing pattern of population structure of Staphylococcus aureus from Bacteremia in China from 2013 to 2016: ST239-030-MRSA replaced by ST59-t437. Front Microbiol. 2018;9:332. doi:10.3389/fmicb.2018.00332
38. Li Y, Zhao R, Zhang X, et al. Prevalence of enterotoxin genes and spa genotypes of methicillin-resistant Staphylococcus aureus from a Tertiary Care Hospital in China. J Clin Diagn Res. 2015;9(5):Dc11–4.
39. Yu F, Liu Y, Lu C, et al. Dissemination of fusidic acid resistance among Staphylococcus aureus clinical isolates. BMC Microbiol. 2015;15:210. doi:10.1186/s12866-015-0552-z
40. Li X, Fang F, Zhao J, et al. Molecular characteristics and virulence gene profiles of Staphylococcus aureus causing bloodstream infection. Braz J Infect Dis. 2018;22(6):487–494. doi:10.1016/j.bjid.2018.12.001
41. Strandén AM, Frei R, Adler H, Flückiger U, Widmer AF. Emergence of SCCmec type IV as the most common type of methicillin-resistant Staphylococcus aureus in a university hospital. Infection. 2009;37(1):44–48. doi:10.1007/s15010-008-7430-7
42. Rossney AS, Shore AC, Morgan PM, Fitzgibbon MM, Brian OC, Coleman DC. The emergence and importation of diverse genotypes of methicillin-resistant Staphylococcus aureus (MRSA) harboring the Panton-Valentine leukocidin gene (pvl) reveal that pvl is a poor marker for community-acquired MRSA strains in Ireland. J Clin Microbiol. 2007;45(8):2554. doi:10.1128/JCM.00245-07
43. Voyich JM, Michael O, Barun M, et al. Is Panton-Valentine leukocidin the major virulence determinant in community-associated methicillin-resistant Staphylococcus aureus disease? J Infect Dis. 2006;194(12):1761–1770. doi:10.1086/509506
44. Song KH, Kim ES, Park KH, et al. Clinical and molecular characterization of panton-valentine leukocidin-positive invasive staphylococcus aureus infections in Korea. Microb Drug Resist. 2019;25(3):450–456. doi:10.1089/mdr.2018.0238
45. Gijon M, Bellusci M, Petraitiene B, et al. Factors associated with severity in invasive community-acquired Staphylococcus aureus infections in children: a prospective European multicentre study. Clin Microbiol Infect. 2016;22(7):
46. Immergluck LC, Jain S, Ray SM, et al. Risk of skin and soft tissue infections among children found to be Staphylococcus aureus MRSA USA300 carriers. West J Emerg Med. 2017;18(2):201–212. doi:10.5811/westjem.2016.10.30483
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.

