Back to Journals » Cancer Management and Research » Volume 12
miR-302e Suppresses Glioma Progression by Targeting VEGFA
Authors Xie Y, Liu X, Hu T, Wang W
Received 30 June 2020
Accepted for publication 9 September 2020
Published 30 October 2020 Volume 2020:12 Pages 10965—10974
DOI https://doi.org/10.2147/CMAR.S268222
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Antonella D'Anneo
This paper has been retracted.
Yunpeng Xie, 1 Xin Liu, 2 Tiemin Hu, 1 Weixing Wang 1
1Department of Neurosurgery, Chengde Medical College Affiliated Hospital, Chengde, Hebei Province, People’s Republic of China; 2Department of Oncology, Chengde Medical College Affiliated Hospital, Chengde, Hebei Province, People’s Republic of China
Correspondence: Weixing Wang
Department of Neurosurgery, Chengde Medical College Affiliated Hospital, No. 36, Nanyingzi Avenue, Chengde, Hebei Province, People’s Republic of China
Email [email protected]
Background: MiRNA can be involved in regulating tumor genesis and development by regulating the expression of specific genes and regulating corresponding signaling pathways. In this study, we explored the function and mechanisms of miR-302e in glioma progression.
Methods: Experimental methods include the following: real-time quantitative PCR, Western Blot Analysis, CCK8 assay and detection of apoptosis.
Results: MiR-302e was down-regulated in cancer tissues and cell lines, and the expression of miR-302e was negatively correlated with the tumor grade, which indicated poor prognosis in glioma patients. Followed functional analysis showed overexpression of miR-302e inhibited proliferation, migration and invasion but promoted apoptosis of glioma cells, while silencing miR-302e showed the opposite effects. Mechanistic studies have shown that VEGFA was a directed target of miR-302e. Forced expression of VEGFA removed the inhibiting impact of miR-302e on glioma development. In vivo tumorigenesis experiments showed that miR-302e suppressed glioma development by targeting VEGFA.
Conclusion: Present study emphasized miR-302e suppressed glioma development by targeting VEGFA, which might be a valuable target for glioma treatment.
Keywords: glioma, miR-302e, proliferation, migration, VEGFA
Introduction
Glioma is a common primary central nervous system tumor.1 The biological characteristics of glioma, such as abundant blood vessels, invasive growth, and high recurrence rate, make the prognosis of glioma patients poor, which is also a significant problem that affects long-term neurosurgery. The higher the pathological grade of glioma, the stronger its invasion, the more blurred the boundary between tumor tissue and healthy brain tissue, significantly increasing the difficulty of surgical resection and chemoradiotherapy resistance.2 Although in recent years, glioma surgery, chemoradiotherapy and other treatment technology have made significant progress. However, the five-year survival rate for glioma patients is still less than 3%.3 It is found that glioma belongs to a polygenic lesion in nature, and its occurrence and development are regulated by multiple genes.4 Therefore, it has become a novel breakthrough point in glioma treatment to study the relationship between related factors at gene regulation level and to seek new molecular gene targets.
MiRNAs are a class of endogenous non-coding small RNA with a length of 18–24 nucleotides.5 MiRNAs are ubiquitous in viruses and eukaryotes, and their sequences are highly conserved.6 According to statistics, up to 50% of protein-coding genes in all humans are regulated by miRNAs.7 Each miRNA regulates up to hundreds of different RNAs and is associated with tissue specificity and disease conditions. In gliomas, miRNA expression can be altered through a variety of mechanisms, including chromosomal changes, epigenetic defects, and mutations.8 Because of the unique biological properties of miRNAs and the importance of their roles, they can be used as diagnostic biomarkers. In malignant tumors, miRNA s can be used to assess trends in cancer invasion, metastasis, or drug resistance as prognostic indicators.9–11 Therefore, miRNA can be involved in regulating tumor genesis and development by regulating the expression of certain genes and regulating corresponding signaling pathways, which provides a new way for the diagnosis, treatment and prognosis of tumors.
Cancerous cells are abnormally enhanced in proliferation, invasion, metastasis and angiogenesis, which changes the standard expression regulation mechanism of cells.12,13 The genesis and development of the tumor is a multi-stage complex process, which usually includes unlimited proliferation, decreased apoptosis and angiogenesis. Tumor cells have a strong ability to survive in the body, but the living body itself cannot carry out adequate immune clearance. Early researchers tried to find miRNAs with differential expression in tumor samples and normal samples, so as to find new target indicators for tumor treatment.14,15 The microarray results showed that 30 miRNAs were differentially expressed, some of which were involved in the formation and metastasis of glioma.16
MiR-302e is a broadly conservative miRNA,17 which has been reported to play an essential function in colorectal cancer, ovarian cancer and non-small cell lung cancer.18–20 And it was shown that miR-302e targeted VEGFA in ovarian cancer.19 However, the role of miR-302e in glioma still remains indistinct. In this study, we focused on the functions of miR-302e and the possible mechanisms in glioma, which might contribute to a potential drug target for glioma treatment.
Methods
Clinical Samples
The surgical specimens of 30 glioma patients from our hospital were collected, which were used for follow-up experimental detection. The experiment was permitted by the Ethics Committee of Chengde Medical College Affiliated Hospital, and the patients signed informed consent.
Cell Culture
Cell lines (normal human astrocytes, SHG44, U251, U87 and SHG139) were purchased from CHI Scientific, Inc (Jiangsu, China). The cells were cultured with complete medium including 89% 1640 and 10% FBS, both were purchased from Biological Industries (Beit-Haemek, Israel), and maintained in incubator with 37°C and 5% of CO2 saturated humidity.
Cell Transfection and Treatment
The cells were plated until the cell density reached 80% confluency of dishes to transfect. AgomiR-302e, miR-302e inhibitor, VEGFA plasmid or NC were constructed by Genechem (Shanghai, China). The RNAs or plasmids are transfected with Lipofectamine 2000 (Invitrogen, Carlsbad, CA).
qRT-PCR
RNA extraction was performed using trizol reagent. NanoDrop 8000 (Thermo Scientific, Waltham, MA, USA) was used to detect the concentration and purity of RNA. The single-stranded cDNAs were synthesized from 1 μg of RNA. The expression of mRNAs and miRNAs were quantified by RT-PCR with SYBR Green I (Thermo Fisher Scientific, Inc). Primer list: miR-302e (F: 5′- GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAAGCAT −3′, R: 5′-ATACTCGTAAGTGCTTCCATGCTT −3′), VEGFA (F: 5′- TATTCAGCGGACTCACCAGC −3′, R: 5′- AACCAACCTCCTCAAACCGT −3′), U6 (F: 5′-TCGCCCTTGGCA CAGCA-3′, R: 5′-CGAACCATTCAAGTGTTGCT-3′), GADPH (F: 5′- CTCCTGCACCACCAACTGCT −3′, R: 5′-GGGCCATCCACAGTCTTCTG −3′).
Protein Isolation and Western Blot
After RIPA cleavage, we extracted total protein and measured with BCA method. After quantitative denaturation, protein electrophoresis membrane transfer and blocked. The first incubation and second incubation were carried out according to the operation steps. The expression of the protein was expressed by the gray value. Primary antibodies list: VEGFA (66,828-1-Ig, Proteintech), Bcl2 (ab182858, Abcam), Bax (ab32503, Abcam), Caspase3 (ab13847, Abcam), GADPH (60,004-1-Ig, Proteintech).
CCK8 Assay
Cells were plated in 96-well plates and we used CCK8 assay to detect the cell viability. CCK8 (10 nmol/L; Beyotime Biotechnology, China) was added after curcumin treatment and incubated at 37°C. We measured the absorbance of 450 nm at 24, 48 and 72 h.
Detection of Apoptosis
The Annexin V-FITC/PI apoptosis kit was purchased from Solebao Company (Beijing, China), and an appropriate amount of logarithmic growth phase cells were washed twice with pre-cooled PBS. The MKN-45 cells were suspended with 500 ul of bound buffer, mixed with 5 µl of annexin V-FITC and PI, respectively, and placed at 25°C for 15 min.
Animals
Animal experiments were permitted by the Animal Protection and Ethics Committee of Chengde Medical College Affiliated Hospital. BALB/c nude mice (6–8 weeks) were purchased from Beijing Weitong Lihua Experimental Animal Technology Co., Ltd. (Beijing, China). For the experiment of Xenograft, SHG44 cell cells (5 × 106) were suspended in 200 μL normal saline and subcutaneously injected or through tail vein. Tumor volume (mm3): V (Mm3) = S2 (Mm2) × L (Mm)/2. All animal experiments were performed in compliance with institutional guidelines and had been approved by SJTUSM Institutional Animal Care and Use Committee (Protocol Registry Number: A-2018-014).
Statistical Analysis
Significant differences were calculated using two-tailed t-test through Graphpad 7.0 and SPSS 22.0. The results were presented in form of mean ± standard deviation (SD). Comparison between the 2 sets of data was performed using Student’s t-test, and differences between the 3 or more groups of data were compared by one‐way ANOVA. All experiments were carried repeatedly for 3 times. P < 0.05 signified a statistical significance (*p<0.05, **p<0.01).
Results
MiR-302e is Decreased in Glioma Tissues and Associated with Poor Prognosis in Patients
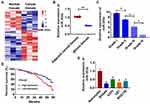
We collected normal and cancer tissues from 30 glioma patients (Table 1), and microarray was used to determine differential expression of miRNAs. And we found miR-302e was a differential expressed miRNA with a lower level in glioma tissues (Figure 1A). Followed qRT-PCR data showed that miR-302e increased in cancer tissues than normal tissues (Figure 1B). We then analyzed the expression of miR-302e in patients with different grades of glioma and found that the miR-302e was negatively correlated with the tumor grade (Figure 1C). According to the median expression level of miR-302e, the level of miR-302e in glioma tissues was divided into low expression and high expression. Kaplan-Meier curves indicated that 5-year survival rate high expression of miR-302e was significantly lower than low expression patients (Figure 1D). Also, miR-302e was decreased in glioma cell lines (SHG44, U251, U87 and SHG139) comparing with human astrocytes (Figure 1E). The results indicated that miR-302e may play a key role in glioma progression.
 |
Table 1 Clinical Characteristics of Glioma Patients |
MiR-302e Suppresses Proliferation and Promotes Apoptosis of Glioma Cells
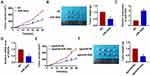
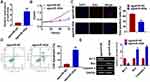
As shown in Figure 1D, SHG44 cells own the lowest expression of miR-302e, and we chose SHG44 to perform the following experiments. We constructed agomiR-302e to force expression of miR-302e in SHG44 cells (Figure 2A). CCK-8 assay manifested that miR-302e inhibited SHG44 cell ability (Figure 2B). Edu staining showed that miR-302e significantly inhibited proliferation of SHG44 cells (Figure 2C). Flow cytometry revealed that miR-302e accelerated glioma cell apoptosis (Figure 2D). In addition, miR-302e remarkably induced Bax and caspase3 expression, but reduced Bcl2 level (Figure 2E).
MiR-302e Inhibits Migration and Invasion of Glioma Cells
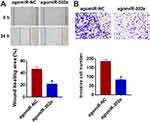
Considering the importance of migration and invasion in cancer progression, we performed wound healing and transwell assay, and found that miR-302e decreased the wound healing area and invasive cell numbers (Figure 3A and B).
Inhibition of miR-302e Promotes Glioma Progression
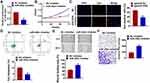
Furthermore, we inhibited miR-302e in U251 cells to evaluate its effect comprehensively. We constructed miR-302e inhibitor and transfected into U251 cells to suppress miR-302e expression (Figure 4A and B). The data of CCK8 showed that inhibition of miR-302e significantly promoted glioma cell growth. And Edu staining also exhibited a remarkable increase of proliferative cells number in miR-302e transfected cells (Figure 4C). Meanwhile, miR-302e inhibitor decreased the apoptotic cells number in flow cytometry assay (Figure 4D). The wound healing analysis showed that loss of miR-302e elevated glioma migratory ability (Figure 4E). Transwell assay suggested showed that deficiency of miR-302e significantly promoted cell invasion in U251 cells (Figure 4F).
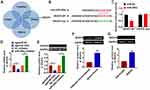
VEGFA is a Direct Target of miR-302e in Glioma Progression
We used 4 databases to identify the target of miR-302e, and their common intersection showed VEGFA was the most potential target (Figure 5A), which showed that the 3ʹUTR of VEGFA possessed directed target site for miR-302e (Figure 5B). And luciferase activity of WT VEGFA, but not mutant VEGFA, was decreased in the miR-302e group compared with NC group (Figure 5C). Furthermore, qRT-PCR analysis showed that agomiR-302e significantly inhibited the expression of VEGFA, while miR-302e inhibitor increased their expression level in SHG44 cells (Figure 5D). In accordance with PCR data, Western blot results exhibited a decrease of VEGFA in miR-302e upregulation cells, and an increase in miR-302e downregulation cells (Figure 5E). These data suggested that VEGFA might be the targets of miR-302e in glioma modification. What’s more, we detected VEGFA expression in clinical glioma tissues and glioma cells. As expected, the protein level of VEGFA was significantly increased in glioma tissue and cells compared with normal tissues and cells (Figure 5F and G).
We then forced expression of VEGFA in SHG44 cells (Figure 6A and B). Functionally, VEGFA removed the inhibitory role of miR-302e on glioma progression (Figure 6C–F).
MiR-302e Inhibits in vivo Tumor Growth and Metastasis in the Nude Mice
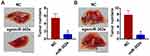
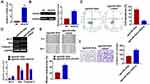
For further explore the function of miR-302e on glioma, we set up xenograft nude mice model. Stable miR-302e transfected SHG44 cell lines were constructed, and were injected into nude mice. And we found miR-302e inhibited the growth of glioma (Figure 7A), miR-302e significantly decreased tumors weight (Figure 7B). The expression of miR-302 was increased in the isolated tumors of injected with SHG44 cell stable transfected miR-302e (Figure 7C). Moreover, miR-302e decreased the mRNA level of VEGFA in tumor tissues (Figure 7D). And then, we set up another xenograft nude mouse model. We injected with SHG44 cell lines firstly. Seven days later, we injected agomiR-302e and its NC into tumors directly. Interestingly, we got the same results. MiR-302e reduced the tumor volume and inhibited tumor growth rate (Figure 7E), miR-302e also significantly decreased tumors weight (Figure 7F).
Next, we examined the effect of miR-302e on glioma metastasis. Stable miR-302e transfected SHG44 cell lines were constructed, and were injected into nude mice through the abdominal cavity (Figure 8A) and the caudal vein (Figure 8B), respectively. And we found that miR-302e significantly inhibited pulmonary metastasis of glioma compared with miR-NC. Together, miR-302e could inhibit glioma growth and metastasis, which may improve the prognosis of glioma patients.
Discussion
Glioma is the deadliest brain tumors.21 The etiology that causes this disease is not clear, with tumor origin, genetic factor, biochemical environment, ionizing radiation, nitroso compound, polluted air, bad life habit, infection and other factors concerned. Even with current treatments that include maximum tumor removal and combined chemoradiotherapy, the prognosis for glioma patients is very poor, with a median survival of only 15 months and only 3% to 5% of patients likely to survive longer.22 Therefore, it is urgent to explore the pathogenesis of glioma and provide targeted therapy strategies from the molecular perspective.
MiRNAs can be completely or incompletely paired with the 3 ‘UTR region of target mRNA,23 and then inhibit the transcriptional translation of target mRNA, participate in the regulation of biological development and cell growth, and are closely related to the occurrence and development of human tumors.24 The discovery of miRNA opens a new window for the study of tumor gene expression regulation. Studies have confirmed the abnormal expression of miRNA in glioma, breast cancer, liver cancer and other malignant tumor stem cells, which affects the proliferation, differentiation, invasion and metastasis of tumor cells by regulating target genes and related signaling pathways.25 MiR-34c was found to reduce the proliferation rate of breast cancer cells.26 MiR-105 expression was significantly down-regulated in stem cell cancer cell lines and clinical liver cancer tissues, while inhibition of miR-105 would promote the proliferation of liver cancer cells and promote tumor formation.27 Abnormal miRNA expression in specific tumors is related to the activation pathway of gene expression, and miRNA is highly conserved. Therefore, miRNAs can be used as high-quality markers to provide theoretical basis for the diagnosis, treatment and prognosis of tumors. MiR-302e is a highly conserved type of RNA.17 However, the role of miR-302e in glioma remains unclear. In present study, we firstly found a disease of miR-302e in glioma clinical tissues and cell lines. Among glioma cell lines, miR-302e was lowest expressed in SHG44 cells, and highest expressed in U251 cells. Thus, we forced miR-302e in SHG44 cells, and miR-302e promoted apoptosis and inhibited the growth, migration and invasion. Then, we silenced miR-302e in U251 cells, miR-302e inhibitor played the opposite function comparing with agomiR-302e. These data suggested that miR-302e was involved in glioma development, and miR-302e might be a diagnose biomarker of glioma.
VEGFA, a vital gene in angiogenesis, can induce endothelial cell proliferation, promote cell migration.28,29 Some studies have found that VEGFA expression is increased in tumors, which is closely related to tumor staging and progression.30,31 Furthermore, VEGFA has been reported to be a target of miR-302e in ovarian cancer, and miR-302e inhibited VEGFA expression.19 In our study, we surprisedly found that there were paired bases between miR-302e and VEGFA. And further experiments revealed that miR-302e directly targeted VEGFA. Functional experiments showed that VEGFA removed the inhibitory effect of miR-302e on glioma progression. Furthermore, we constructed two in vivo tumor formation models to examine the role of miR-302e on glioma tumorigenesis. And we got the same conclusion, which was that miR-302e could significantly inhibit the glioma growth, which was conducive to clinical targeted therapy. In addition, because metastasis is an important part of tumor progression, we performed animal experiments to assess pulmonary metastasis of glioma. We found that miR-302e could inhibit glioma metastasis, which may improve the prognosis of glioma patients.
Through understanding relevant studies on the involvement of miRNA in the regulation mechanism of glioma, the pathological processes such as proliferation, apoptosis, invasion and metastasis of glioma cells were deepened. At present, studies on the molecular level of glioma are still insufficient, so how to make miRNA play a better clinical application value in the diagnosis of glioma, evaluation of the efficacy of chemotherapy drugs, anti-angiogenesis, treatment and prognosis judgment, etc. Whether it can provide a new way for the development of cancer treatment drugs, these will be a long way to go. It is believed that with the further study of miRNA in the function and mechanism of action of glioma, these problems will be gradually solved, which will bring new opportunities for clinical diagnosis and treatment.
In conclusion, present study revealed that miR-302e contributed to the progression of glioma, and miR-302e acted as an anti-tumor gene by targeting VEGFA.
Disclosure
The authors declare no conflicts of interest for this work.
References
1. Haider AS, van den Bent M, Wen PY, et al. Towards a standard pathological and molecular characterization of recurrent glioma in adults: a RANO effort. Neuro Oncol. 2019.
2. Arnold A, Yuan M, Price A, Harris L, Eberhart CG, Raabe EH. Synergistic activity of mTORC1/2 kinase and MEK inhibitors suppresses pediatric low-grade glioma tumorigenicity and vascularity. Neuro Oncol. 2020;22(4):563–574.
3. Yekula A, Minciacchi VR, Morello M, et al. Large and small extracellular vesicles released by glioma cells in vitro and in vivo. J Extracell Vesicles. 2020;9(1):1689784. doi:10.1080/20013078.2019.1689784
4. Han F, Zhang L, Chen C, et al. GLTSCR1 negatively regulates BRD4-dependent transcription elongation and inhibits CRC metastasis. Adv Sci (Weinh). 2019;6(23):1901114. doi:10.1002/advs.201901114
5. Han M, Wang S, Fritah S, et al. Interfering with long non-coding RNA MIR22HG processing inhibits glioblastoma progression through suppression of Wnt/beta-catenin signalling. Brain. 2020;143(2):512–30.
6. Xiong Y, Chen L, Yan C, et al. Circulating exosomal miR-20b-5p inhibition restores wnt9b signaling and reverses diabetes-associated impaired wound healing. Small. 2019;e1904044.
7. Park K, Kim KB. miRTar hunter: a prediction system for identifying human microRNA target sites. Mol Cells. 2013;35(3):195–201. doi:10.1007/s10059-013-2165-4
8. Deng Q, Hu H, Yu X, et al. Tissue-specific microRNA expression alters cancer susceptibility conferred by a TP53 noncoding variant. Nat Commun. 2019;10(1):5061.
9. Huang H, Tang J, Zhang L, Bu Y, Zhang X. miR-874 regulates multiple-drug resistance in gastric cancer by targeting ATG16L1. Int J Oncol. 2018;53(6):2769–2779.
10. Sun W, Ping W, Tian Y, Zou W, Liu J, Zu Y. miR-202 enhances the anti-tumor effect of cisplatin on non-small cell lung cancer by targeting the Ras/MAPK pathway. Cell Physiol Biochem. 2018;51(5):2160–2171. doi:10.1159/000495835
11. Li B, Wang W, Li Z, et al. MicroRNA-148a-3p enhances cisplatin cytotoxicity in gastric cancer through mitochondrial fission induction and cyto-protective autophagy suppression. Cancer Lett. 2017;410:212–227. doi:10.1016/j.canlet.2017.09.035
12. Xie C, Ji N, Tang Z, Li J, Chen Q. The role of extracellular vesicles from different origin in the microenvironment of head and neck cancers. Mol Cancer. 2019;18(1):83. doi:10.1186/s12943-019-0985-3
13. Xin Z, Jiang S, Jiang P, et al. Melatonin as a treatment for gastrointestinal cancer: a review. J Pineal Res. 2015;58(4):375–387.
14. Selimoglu-Buet D, Riviere J, Ghamlouch H, et al. A miR-150/TET3 pathway regulates the generation of mouse and human non-classical monocyte subset. Nat Commun. 2018;9(1):5455. doi:10.1038/s41467-018-07801-x
15. Ahn JH, Lee HS, Lee JS, et al. Author correction: nc886 is induced by TGF-beta and suppresses the microRNA pathway in ovarian cancer. Nat Commun. 2018;9(1):5458. doi:10.1038/s41467-018-07818-2
16. Wu DM, Wang S, Wen X, et al. MircoRNA-1275 promotes proliferation, invasion and migration of glioma cells via SERPINE1. J Cell Mol Med. 2018;22(10):4963–4974. doi:10.1111/jcmm.13760
17. Chen X, Xu Y, Liao X, et al. Plasma miRNAs in predicting radiosensitivity in non-small cell lung cancer. Tumour Biol. 2016;37(9):11927–11936. doi:10.1007/s13277-016-5052-8
18. Cui D, Qian R, Li Y. Circular RNA circ-CMPK1 contributes to cell proliferation of non-small cell lung cancer by elevating cyclin D1 via sponging miR-302e. Mol Genet Genomic Med. 2020;8(2):e999.
19. Wang LL, Zong ZH, Liu Y, Guan X, Chen S, Zhao Y. CircRhoC promotes tumorigenicity and progression in ovarian cancer by functioning as a miR-302e sponge to positively regulate VEGFA. J Cell Mol Med. 2019;23(12):8472–8481. doi:10.1111/jcmm.14736
20. Li D, Jiang X, Zhang X, Cao G, Wang D, Chen Z. Long noncoding RNA FGD5-AS1 promotes colorectal cancer cell proliferation, migration, and invasion through upregulating CDCA7 via sponging miR-302e. In Vitro Cell Dev Biol Anim. 2019;55(8):577–585. doi:10.1007/s11626-019-00376-x
21. Gritsenko PG, Atlasy N, Dieteren CEJ, et al. p120-catenin-dependent collective brain infiltration by glioma cell networks. Nat Cell Biol. 2020;22(1):97–107. doi:10.1038/s41556-019-0443-x
22. Jaraiz-Rodriguez M, Talaveron R, Garcia-Vicente L, et al. Connexin43 peptide, TAT-Cx43266-283, selectively targets glioma cells, impairs malignant growth and enhances survival in mouse models in vivo. Neuro Oncol. 2020;22(4):493–504.
23. Li J, Zhang S, Wan Y, et al. MISIM v2.0: a web server for inferring microRNA functional similarity based on microRNA-disease associations. Nucleic Acids Res. 2019;47(W1):W536–W541. doi:10.1093/nar/gkz328
24. Huang T, Wan X, Alvarez AA, et al. MIR93 (microRNA −93) regulates tumorigenicity and therapy response of glioblastoma by targeting autophagy. Autophagy. 2019;15(6):1100–1111. doi:10.1080/15548627.2019.1569947
25. Wang R, Zhang S, Chen X, et al. EIF4A3-induced circular RNA MMP9 (circMMP9) acts as a sponge of miR-124 and promotes glioblastoma multiforme cell tumorigenesis. Mol Cancer. 2018;17(1):166.
26. Ilisso CP, Delle Cave D, Mosca L, et al. S-Adenosylmethionine regulates apoptosis and autophagy in MCF-7 breast cancer cells through the modulation of specific microRNAs. Cancer Cell Int. 2018;18:197. doi:10.1186/s12935-018-0697-6
27. Shen G, Rong X, Zhao J, et al. MicroRNA-105 suppresses cell proliferation and inhibits PI3K/AKT signaling in human hepatocellular carcinoma. Carcinogenesis. 2014;35(12):2748–2755.
28. Thompson EM, Keir ST, Venkatraman T, et al. The role of angiogenesis in group 3 medulloblastoma pathogenesis and survival. Neuro Oncol. 2017;19(9):1217–1227. doi:10.1093/neuonc/nox033
29. Liu L, Bi N, Wu L, et al. MicroRNA-29c functions as a tumor suppressor by targeting VEGFA in lung adenocarcinoma. Mol Cancer. 2017;16(1):50.
30. Wang H, Deng Q, Lv Z, et al. N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1. Mol Cancer. 2019;18(1):181.
31. Teufel M, Seidel H, Kochert K, et al. Biomarkers associated with response to regorafenib in patients with hepatocellular carcinoma. Gastroenterology. 2019;156(6):1731–1741.
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.