Back to Journals » OncoTargets and Therapy » Volume 12
MicroRNA-769-5p Promotes The Growth Of Glioma Cells By Targeting Lysine Methyltransferase 2A
Authors Chang M, Yan P, Zhang B, Zhang G, Wang J, Ge H, Han N, Du C, Shi W, Tian Y
Received 11 July 2019
Accepted for publication 18 October 2019
Published 5 November 2019 Volume 2019:12 Pages 9177—9187
DOI https://doi.org/10.2147/OTT.S222836
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Dr Sanjeev K. Srivastava
This paper has been retracted.
Mingze Chang, 1, 2 Peng Yan, 3 Bei Zhang, 4 Gejuan Zhang, 1 Juanhong Wang, 5, 6 Hanming Ge, 1 Nannan Han, 1 Chengxue Du, 1 Wenzhen Shi, 1 Ye Tian 2
1Department of Neurology, Xi’an No. 3 Hospital, Xi’an 710021, People’s Republic of China; 2Department of Neurology, The Affiliated Hospital of Northwest University, Xi’an 710021, People’s Republic of China; 3The College of Life Sciences, Northwest University, Xi’an 710069, People’s Republic of China; 4Department of Neurology, The First Affiliated Hospital of Xi’an Medical University, Xi’an 710077, People’s Republic of China; 5Department of Pathology, Xi’an No.3 Hospital, Xi’an 710021, People’s Republic of China; 6Departments of Pathology, Xi’an Central Hospital, Xi’an 71000, People’s Republic of China
Correspondence: Ye Tian
Department of Neurology, the Affiliated Hospital of Northwest University, 10 East Section of Fengcheng 3rd Road, Xi’an 710021, People’s Republic of China
Tel +86-029-87268355
Email [email protected]
Background: Accumulating evidence supports the involvement of microRNAs (miRNAs) in the progression of human cancers including glioma. Recently, miR-769-5p has been reported to play a tumor suppressive role in colorectal cancer and lung cancer, whereas it exerts an oncogenic role in melanoma. However, the role of miR-769-5p and its related mechanism are poorly elucidated.
Methods: The levels of miR-769-5p in glioma tissues and adjacent non-tumor tissues were detected by qRT-PCR. In addition, the effects of miR-769-5p on cell proliferation and apoptosis were evaluated by CCK-8, EdU, colony formation and flow cytometric assays, respectively. Meanwhile, the dual-luciferase reporter assay was used to investigate the interaction of miR-769-5p and lysine methyltransferase 2A (KMT2A) in glioma.
Results: We found that miR-769-5p expression was strongly upregulated in glioma tissues and cell lines. Glioma tissues with high World Health Organization (WHO) grades had obvious higher levels of miR-769-5p compared to samples with low WHO grades. Interestingly, glioma patients highly expressing miR-769-5p showed prominent poorer survivals. Knockdown of miR-769-5p significantly suppressed cell proliferation and resulted in apoptosis in glioma cells. Additionally, miR-769-5p silencing restrained in vivo growth of glioma cells in mice. Interestingly, KMT2A was identified to be a direct target of miR-769-5p in glioma cells. The expression of KMT2A mRNA was downregulated in glioma tissues and inversely correlated with miR-769-5p level. KMT2A overexpression inhibited cell proliferation and induced the apoptosis of A172 cells. Moreover, siRNA-mediated KMT2A silencing could partially abolish miR-769-5p knockdown-induced suppressive effects on A172 cells.
Conclusion: In summary, our findings suggest that targeting miR-769-5p/KMT2A axis may be a promising therapeutic target for glioma treatment.
Keywords: miR-769-5p, glioma, KMT2A, tumor growth, apoptosis
Corrigendum for this paper has been published
Introduction
Glioma is one of the most serious leading causes of cancer-related mortality worldwide.1 Glioblastoma multiforme (GBM), a malignant grade IV tumor, is the most aggressive type of glioma, which accounts for 60–70% of all gliomas.2 Surgical resection remains the main choice of glioma treatment, but the clinical outcome for glioma patients remains poor.3 Hence, there is an urgent need to identify both novel sensitive biomarkers for diagnosis and new therapeutic targets for treatment in glioma.
MicroRNAs (miRNAs) are single-stranded non-coding RNAs that bind to the 3ʹ untranslated region (3ʹUTR) of target mRNAs to induce degradation and interfere in the translation process.4 More and more studies provide evidence to support that miRNAs are aberrantly expressed in human cancers and function as tumor suppressors or drivers.5–11 In gliomas, miRNAs are crucial regulators in the tumorigenesis and tumor progression.12 The expression of miR-708 is down-regulated in glioma tissues and it suppresses the growth and epithelial-to-mesenchymal transition (EMT) of glioma cells by targeting sphingosine kinase 2 (SPHK2)-mediated AKT/β-catenin pathway.13 miR-4500 functions as a tumor suppressor by attenuating insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1) in glioma cells.14 Our previous study reveals that miR-33a promotes cell proliferation and induces the apoptosis of glioma cells by directly suppressing sirtuin 6 (SIRT6).15 Recently, miR-769-5p, a newly emerging cancer-associated miRNA, caught our attention. In oral squamous cell carcinoma (OSCC) tissues, the downregulated expression of miR-769-5p has a hopeful diagnosis potential.16 The increased expression of let-7d-5p combining with miR-769-5p under-expression indicates poor prognosis of non-small cell lung cancer (NSCLC).17 miR-769-5p expression is frequently reduced in colorectal cancer (CRC), and suppresses cell proliferation and invasion by inhibiting cyclin-dependent kinase 1 (CDK1) and hes related family bHLH transcription factor with YRPW motif 1 (HEY1).18,19 Moreover, miR-769-5p suppressed the proliferation, migration and invasion of NSCLC cells by inhibiting transforming growth factor beta receptor 1 (TGFBR1).20 Additionally, long noncoding RNA (lncRNA) LINC00460 functions as a molecular sponge for miR-769-5p to promote epidermal growth factor receptor (EGFR) and accordingly exerts an oncogenic role in NSCLC.21 Conversely, miR-769-5p expression is remarkably increased in melanoma and facilitates cancer cell proliferation by targeting glycogen synthase kinase-3 beta (GSK3β).22 miR-769-5p plays a tumor promoting role in hepatocellular carcinoma (HCC) via targeting RING1 and YY1 binding protein (RYBP).23 However, the role of miR-769-5p in glioma remains unexplored yet.
In the present study, we detected the expression levels of miR-769-5p in glioma tissues and cell lines. Moreover, the biological role of miR-769-5p in the proliferation and apoptosis of glioma cells and its relevant mechanism were studied. Our study suggested that miR-769-5p might act as a novel potential therapeutic target for glioma.
Materials And Methods
Patients And Sample Collection
The study was approved by the Research Ethics Committee of Xi’an No.3 Hospital and with the 1964 Helsinki declaration and its later amendments. Sixty paired glioma tissues and adjacent non-tumor tissues (glioma borders) were collected from patients during surgery in the Xi’an No.3 Hospital and Xi'an Central Hospital. Written informed consent was obtained from all patients, who did not receive treatment before surgery. The clinicopathologic information of glioma patients were previously described.15 Tissue specimens were immediately snap frozen in liquid nitrogen and stored at −80°C until RNA extraction.
Cell Culture And Transfection
These four glioma cell lines (A172, T98, U87 and U251) and a normal human astrocyte (NHA) cell line, which were purchased from the Cell Bank of Shanghai Institute of Cell Biology (Chinese Academy of Medical Science, Shanghai, China), were maintained in our lab and cultured as previously described.15 The miR-769-5p inhibitors (anti-miR-769-5p) and negative control inhibitors (NC) were obtained from RiboBio (Guangzhou, China). Lentiviral vector-mediated miR-769-5p inhibitors were purchased from GeneCopoeia (Guangzhou, China). pcDNA3.1-lysine methyltransferase 2A (KMT2A) and empty vector (EV) were obtained from Genechem (Shanghai, China). KMT2A siRNA (siKMT2A; 5′-GGT GTT GTC GTC GTT GCA AAT-3′) and corresponding scrambled control siRNA (Scr siRNA; 5′-TTC TCC GAA CGT GTC ACG T-3′) were synthesized by RiboBio. These plasmids and oligos were transfected into glioma cells using lipofectamine 2000 (Thermo Fisher Scientific, Waltham, MA, USA).
RNA Extraction And Quantitative Real-Time PCR (qRT-PCR)
Briefly, total RNA was extracted from glioma tissues and cell lines with TRIzol reagent (Invitrogen, Carlsbad CA, USA). RNA was reversely transcribed to cDNA using a Reverse Transcription Kit (Tiangen, China). Real-time PCR assay was performed with SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) in the ABI 7500 Real-time PCR System. The miRNA level was determined using qRT-PCR analysis with TaqMan MicroRNA Assay Kit (Applied Biosystems). The primer sequences were listed as follows: miR-769-5p RT 5´-CTC AAC TGG TGT CGT GGA GTC GGC AAT TCA GTT GAG AAC CCA TG-3´, forward 5´-ACA CTC CAG CTG GGT GAG ACC TCT GGG TTC TG-3´ and reverse 5´-CTC AAC TGG TGT CGT GGA-3´; U6 forward 5ʹ-CTC GCT TCG GCA GCA CA-3ʹ and reverse 5ʹ-AAC GCT TCA CGA ATT TGC GT-3ʹ; KMT2A forward 5ʹ-CCC CTT CGG GTC TCC TCA TT −3ʹ and reverse 5ʹ-ATC CTT CGA GGG CTT TGT CTG-3ʹ; protein kinase AMP-activated catalytic subunit alpha 1 (PRKAA1) forward 5′-TGT CTC TGG AGG AGA GCT ATT TGA-3′ and reverse 5′-GGT GAG CCA CAG CTT GTT CTT-3′; GAPDH forward 5ʹ-TGT TCG TCA TGG GTG TGA AC-3ʹ and reverse 5ʹ-ATG GCA TGG ACT GTG GTC AT-3ʹ.
Cell Proliferation
Cell Counting Kit-8 (CCK-8), colony formation and EdU assays were performed to detect the proliferation of glioma cells. CCK-8 reagent (10μl/well, Dojindo Laboratories, Dojindo, Japan) was added to the plate and cells were cultured for 1.5 hrs. Then, the absorbance of the samples at 450 nm was detected by microplate reader (Bio-Rad, Hercules, CA, USA). For colony formation assay, glioma cells were seeded in six-well plates (200 cells per well) and incubated for 2 weeks until the visible colonies were formed. The colonies with crystal violet staining were counted. The EdU assay was carried out using the Cell-Light™ EdU Apollo®488 In Vitro Imaging Kit (RioBio) following the manufacturer’s recommendations.
Flow Cytometry
The apoptotic glioma cells were detected by using the PE Annexin V Apoptosis Detection Kit I (BD Biosciences, San Jose, CA, USA) as previously described.10
Luciferase Reporter Assay
Glioma cells were con-transfected with miR-769-5p inhibitors or NC inhibitors and luciferase reporter plasmids containing wild-type (wt) or mutant type (mt) 3ʹUTR of KMT2A. After transfection, the luciferase activity was determined by using a dual-luciferase reporter assay system (Promega, Madison, WI, USA).
Western Blotting
The detailed protocols for immunoblotting analysis were previously mentioned.15 Primary antibodies against KMT2A (#14197), PCNA (#13110), Bcl-2 (#4223), CDK4 (#12790), CDK6 (#3136), β-actin (#3700) and GAPDH (#5174), and a horseradish peroxidase (HRP)-conjugated secondary antibodies (#7074 and #7076) were purchased from Cell Signaling Technology, Inc (Beverly, MA, USA). Bound antibodies were visualized by super ECL detection reagent (Beyotime, Shanghai, China).
Animal Experimental
The subcutaneous xenograft transplantation experiments were performed using co-injection of U251 cells with Matrigel as previously described.15 Immunoblotting analysis of CDK4, CDK6, PCNA and Bcl-2 was carried out in the xenograft tumor tissues. The animal studies were approved by the Institutional Animal Care and Use Committee of The Affiliated Hospital of Northwest University and the protocols complied with the guidelines for the welfare and use of animals in cancer research.
Statistical Analysis
GraphPad Prism 8.0 (GraphPad Inc., San Diego, CA, USA) was used to analyze the data. Data were expressed as the mean ± standard deviation (SD) of three independent experiments and analyzed using Student’s t-test or ANOVA followed by a Tukey’s post-hoc test. Kaplan–Meier method and log-rank test were carried out for survival analysis. The relationship between miR-769-5p and KMT2A mRNA expression was assessed by Spearman correlation test in glioma tissues. P<0.05 was determined to indicate a statistically significant difference.
Results
miR-769-5p Expression Is Upregulated In Gliomas
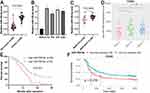
Firstly, qRT-PCR analysis was conducted to determine the expression difference of miR-769-5p between 60 paired glioma tissues and adjacent nontumor tissues. As shown in Figure 1A, the expression of miR-769-5p was significantly higher in glioma tissues compared to adjacent nontumor tissues (P<0.0001). Next, we measured the expressions of miR-769-5p in glioma cell lines including A172, T98, U87 and U251. Consistently, the levels of miR-769-5p in glioma cell lines were markedly higher than that in NHA cell line (P<0.05, Figure 1B). Thus, miR-769-5p expression was frequently increased in gliomas.
The Upregulated Expression Of miR-769-5p Predicts Poor Prognosis Of Glioma
Next, the expressions of miR-769-5p were compared among glioma tissues with different World Health Organization (WHO) grades. Glioma tissues with high WHO (III+IV) grades had obvious higher levels of miR-769-5p compared to samples with low WHO grades (I+II) (P=0.0002, Figure 1C). Interestingly, the data form Chinese Glioma Genome Altas (CGGA) consistently demonstrated that miR-769-5p expression was positively correlated with tumor grades of glioma (P<0.05, Figure 1D). Notably, glioma patients highly expressing miR-769-5p had a prominent poorer overall survival compared to cases with low miR-769-5p level (P=0.007, Figure 1E). Additionally, CGGA data revealed that elevated expression of miR-769-5p predicted poor survival of glioma patients (P=0.016, Figure 1F). These data suggested that miR-769-5p might be a promising prognostic biomarker for glioma.
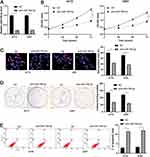
Inhibition Of miR-769-5p Represses The Proliferation Of Glioma Cells
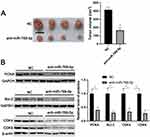
To further investigate the role of miR-769-5p in glioma cells, synthetic miR-769-5p inhibitors were transfected into A172 and U251 cells and miR-769-5p knockdown was confirmed by qRT-PCR (P<0.05, Figure 2A). Then, CCK-8 and EdU assays indicated that miR-769-5p knockdown significantly suppressed the proliferation of A172 and U251 cells (P<0.05, Figure 2B and C). Furthermore, the colonies formed by glioma cells with miR-769-5p knockdown were obviously less than those formed by control cells (P<0.05, Figure 2D). Subsequently, we investigated the effect of miR-769-5p silencing on glioma cell apoptosis. As shown in Figure 2E, miR-769-5p inhibition markedly induced the apoptosis of A172 and U251 cells (P<0.05). Additionally, silencing of miR-769-5p obviously reduced the growth of U251 cells in vivo (P<0.05, Figure 3A). The levels of CDK4, CDK6, PCNA and Bcl-2 in xenograft tumor tissues arising from miR-769-5p knockdown group were prominently lower than those in control group (P<0.05, Figure 3B). These results showed that miR-769-5p knockdown suppressed cell proliferation and induced apoptosis in glioma cells.
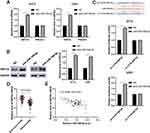
miR-769-5p Directly Targets KMT2A In Glioma Cells
To investigate the molecular mechanism underlying the role of miR-769-5p in glioma cells, we predicted the target genes of miR-769-5p based on starBase V3.0 online platform.24,25 After analysis, 22 genes were predicted by four databases cross-validation. Among these, PRKAA1 and KMT2A are previously reported to be tumor suppressors in glioma.26,27 Next, we found that miR-769-5p knockdown prominently increased the expression of KMT2A mRNA in A172 and U251 cells (P<0.05, Figure 4A). However, miR-769-5p inhibition did not impact the level of PRKAA1 mRNA in glioma cells (Figure 4A). Furthermore, immunoblotting analysis showed that the expression of KMT2A protein was remarkedly upregulated by miR-769-5p knockdown in A172 and U251 cells (P<0.05, Figure 4B). To validate whether KMT2A is a direct target gene of miR-769-5p, we inserted the 3′UTR of KMT2A containing binding sites for miR-769-5p into the luciferase system. As shown in Figure 4C, miR-769-5p knockdown significantly enhanced the luciferase activities of vectors carrying wt 3′UTR of KMT2A (P<0.05), but not those of vectors containing mutant binding sites in 3′UTR of KMT2A. Notably, qRT-PCR data showed that the expression of KMT2A mRNA was significantly downregulated in glioma tissues when compared with adjacent nontumor tissues (P<0.0001, Figure 4D). Further analysis indicated that the level of KMT2A mRNA was negatively correlated with miR-769-5p expression in glioma tissues (r=−0.5636, P<0.0001, Figure 4E). Therefore, our data demonstrated that miR-769-5p directly targeted KMT2A and inhibited its abundance in glioma cells.
KMT2A Partially Mediates The Role Of miR-769-5p In Glioma Cells
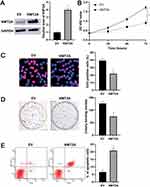
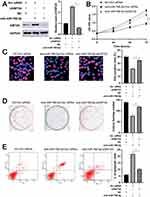
As mentioned above, KMT2A was a direct target of miR-769-5p in glioma cells. Thus, we further disclosed whether KMT2A mediated the role of miR-769-5p in glioma cells. The expression of KMT2A was dramatically up-regulated by plasmid transfection in A172 cells (P<0.05, Figure 5A). Functional assays indicated that KMT2A overexpression significantly inhibited cell proliferation and induced apoptosis in A172 cells (P<0.05, Figure 5B–E). Next, KMT2A was knocked down by a synthetic siRNA in A172 cells with miR-769-5p silencing (P<0.05, Figure 6A). Interestingly, KTM2A knockdown partially abolished miR-769-5p silencing-induced proliferation inhibition and apoptosis in A172 cells (P<0.05, Figure 6B–E). Collectively, these results revealed that miR-769-5p facilitated glioma cell growth and reduced apoptosis by targeting KMT2A.
Discussion
In this study, miR-769-5p was found to be upregulated in glioma tissues and cell lines. The upregulated expression of miR-769-5p was positively correlated with tumor grades and predicted poor prognosis of glioma. Furthermore, knockdown of miR-769-5p could suppress cell proliferation and induced apoptosis in glioma cells. KMT2A was identified as a direct target of miR-769-5p. KMT2A overexpression exerted similar effects with miR-769-5p silencing on glioma cells. Importantly, knockdown of KMT2A partially reversed the suppressive effects of miR-769-5p silencing on glioma cells.
Recently, miR-769-5p has been reported to be a tumor suppressor or oncogene in human cancers.18–23 For instance, miR-769-5p suppresses CRC cell proliferation and invasion via attenuating CDK1 and HEY1.18,19 Furthermore, miR-769-5p inhibited NSCLC cell proliferation, migration and invasion by targeting TGFBR1 and EGFR.20,21 However, miR-769-5p plays a tumor promoting role during the tumor progression of melanoma22 and HCC.23 There results indicate that miR-769-5p has different functions in different tumor types. Currently, the expression of miR-769-5p and its role in glioma remain unclear yet. Here, we found the upregulated expression of miR-769-5p in glioma tissues and cell lines. Both our data and CGGA data consistently indicated that high miR-769-5p level conferred to high tumor grades and poor clinical outcomes of glioma. Moreover, knockdown of miR-769-5p suppressed cell proliferation and induced apoptosis in glioma cells. These data suggested that miR-769-5p might play an essential role in the progression of glioma.
The biological role of KMT2A has been widely investigated in human cancers.27–32 In melanoma, KMT2A contributes to tumor growth via inducing the activation of hTERT signaling28 KMT2A fusions and rearrangements are frequently detected in hematological malignancies.29–31 Moreover, KMT2A is highly expressed in CRC tissues and promotes cancer cell migration and invasion via epigenetically regulating cathepsin Z (CTSZ)32 Notably, KMT2A knockdown facilitates glioma cell proliferation and promotes tumor growth in vivo by epigenetically regulating notch receptor 1 (NOTCH1) and NOTCH3, suggesting a tumor suppressive role of KMT2A in glioma.27 However, whether KMT2A is a functional target of miR-769-5p in glioma cells remains to be elucidated. In this study, miR-769-5p knockdown increased the abundance of KMT2A in glioma cells. The expression of KMT2A mRNA was downregulated in glioma tissues and negatively correlated with miR-769-5p level. Herein, KMT2A was validated as a target gene of miR-769-5p in glioma cells. Moreover, ectopic expression of KMT2A exerted the anti-proliferative and pro-apoptotic roles in glioma cells. KMT2A knockdown partially abrogated miR-769-5p silencing-induced glioma cell growth arrest and apoptosis. These data suggested that miR-769-5p might promote the growth of glioma cells possibly by targeting KMT2A.
In conclusion, we found that miR-769-5p level was increased in glioma tissues and cell lines. The upregulated expression of miR-769-5p correlated with poor prognosis of glioma. More importantly, knockdown of miR-769-5p repressed cell proliferation and induced apoptosis possibly by increasing KMT2A expression in glioma cells. These results suggest that miR-769-5p/KMT2A axis may represent potential therapeutic targets for glioma.
Conclusion
To conclude, we demonstrate that the expression of miR-769-5p is dramatically upregulated in glioma, and closely correlates with poor prognosis of patients. In vitro and in vivo assays reveal that miR-769-5p knockdown suppresses cell proliferation and induces apoptosis of glioma cells. Interestingly, KMT2A is directly regulated by miR-769-5p, and partially mediates the tumor promoting role of miR-769-5p in glioma cells. These data provide novel insights into the complex regulatory network of glioma pathogenesis.
Acknowledgment
This study was supported by grants from the Key Research and Development Program of Shaanxi Province (2018SF-035; 2018SF-087) and Xi’an Science and Technology Plan Project [2017118SF/YX012(1)].
Disclosure
The authors declare no conflicts of interest in this work.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. doi:10.3322/caac.v68.6
2. Wen PY, Kesari S. Malignant gliomas in adults. N Engl J Med. 2008;359(5):492–507. doi:10.1056/NEJMra0708126
3. Chen J, Li Y, Yu TS, et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature. 2012;488(7412):522–526. doi:10.1038/nature11287
4. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–297. doi:10.1016/S0092-8674(04)00045-5
5. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6(11):857–866. doi:10.1038/nrc1997
6. Liu Z, Wang Y, Dou C, et al. MicroRNA-1468 promotes tumor progression by activating PPAR-gamma-mediated AKT signaling in human hepatocellular carcinoma. J Exp Clin Cancer Res. 2018;37(1):49. doi:10.1186/s13046-018-0717-3
7. Xu Q, Zhu Q, Zhou Z, et al. MicroRNA-876-5p inhibits epithelial-mesenchymal transition and metastasis of hepatocellular carcinoma by targeting BCL6 corepressor like 1. Biomed Pharmacother. 2018;103:645–652. doi:10.1016/j.biopha.2018.04.037
8. Xu Q, Liu X, Liu Z, et al. MicroRNA-1296 inhibits metastasis and epithelial-mesenchymal transition of hepatocellular carcinoma by targeting SRPK1-mediated PI3K/AKT pathway. Mol Cancer. 2017;16(1):103. doi:10.1186/s12943-017-0675-y
9. Dou C, Liu Z, Xu M, et al. miR-187-3p inhibits the metastasis and epithelial-mesenchymal transition of hepatocellular carcinoma by targeting S100A4. Cancer Lett. 2016;381(2):380–390. doi:10.1016/j.canlet.2016.08.011
10. Wang Y, Yang L, Chen T, et al. A novel lncRNA MCM3AP-AS1 promotes the growth of hepatocellular carcinoma by targeting miR-194-5p/FOXA1 axis. Mol Cancer. 2019;18(1):28. doi:10.1186/s12943-019-0957-7
11. Zhang L, Wang L, Wang Y, et al. LncRNA KTN1-AS1 promotes tumor growth of hepatocellular carcinoma by targeting miR-23c/ERBB2IP axis. Biomed Pharmacother. 2019;109:1140–1147. doi:10.1016/j.biopha.2018.10.105
12. Anthiya S, Griveau A, Loussouarn C, et al. MicroRNA-based drugs for brain tumors. Trends Cancer. 2018;4(3):222–238. doi:10.1016/j.trecan.2017.12.008
13. Chen Y, Deng X, Chen W, et al. Silencing of microRNA-708 promotes cell growth and epithelial-to-mesenchymal transition by activating the SPHK2/AKT/beta-catenin pathway in glioma. Cell Death Dis. 2019;10(6):448. doi:10.1038/s41419-019-1671-5
14. Li ZW, Xue M, Zhu BX, Yue CL, Chen M, Qin HH. microRNA-4500 inhibits human glioma cell progression by targeting IGF2BP1. Biochem Biophys Res Commun. 2019;513(4):800–806. doi:10.1016/j.bbrc.2019.04.058
15. Chang M, Qiao L, Li B, et al. Suppression of SIRT6 by miR-33a facilitates tumor growth of glioma through apoptosis and oxidative stress resistance. Oncol Rep. 2017;38(2):1251–1258. doi:10.3892/or.2017.5780
16. Pedersen NJ, Jensen DH, Lelkaitis G, et al. MicroRNA-based classifiers for diagnosis of oral cavity squamous cell carcinoma in tissue and plasma. Oral Oncol. 2018;83:46–52. doi:10.1016/j.oraloncology.2018.05.020
17. Gasparini P, Cascione L, Landi L, et al. microRNA classifiers are powerful diagnostic/prognostic tools in ALK-, EGFR-, and KRAS-driven lung cancers. Proc Natl Acad Sci USA. 2015;112(48):14924–14929. doi:10.1073/pnas.1520329112
18. Han C, Song Y, Lian C. MiR-769 inhibits colorectal cancer cell proliferation and invasion by targeting HEY1. Med Sci Monit. 2018;24:9232–9239. doi:10.12659/MSM.911663
19. Wang L, Xu M, Lu P, Zhou F. microRNA-769 is downregulated in colorectal cancer and inhibits cancer progression by directly targeting cyclin-dependent kinase 1. Onco Targets Ther. 2018;11:9013–9025. doi:10.2147/OTT
20. Yang Z, He J, Gao P, et al. miR-769-5p suppressed cell proliferation, migration and invasion by targeting TGFBR1 in non-small cell lung carcinoma. Oncotarget. 2017;8(69):113558–113570. doi:10.18632/oncotarget.v8i69
21. Ma G, Zhu J, Liu F, Yang Y. Long noncoding RNA LINC00460 promotes the gefitinib resistance of nonsmall cell lung cancer through epidermal growth factor receptor by sponging miR-769-5p. DNA Cell Biol. 2019;38(2):176–183. doi:10.1089/dna.2018.4462
22. Qiu HJ, Lu XH, Yang SS, Weng CY, Zhang EK, Chen FC. MiR-769 promoted cell proliferation in human melanoma by suppressing GSK3B expression. Biomed Pharmacother. 2016;82:117–123. doi:10.1016/j.biopha.2016.04.052
23. Xian Y, Wang L, Yao B, et al. MicroRNA-769-5p contributes to the proliferation, migration and invasion of hepatocellular carcinoma cells by attenuating RYBP. Biomed Pharmacother. 2019;118:109343. doi:10.1016/j.biopha.2019.109343
24. Yang JH, Li JH, Shao P, Zhou H, Chen YQ, Qu LH. starBase: a database for exploring microRNA-mRNA interaction maps from argonaute CLIP-seq and degradome-seq data. Nucleic Acids Res. 2011;39(Database issue):D202–209. doi:10.1093/nar/gkq1056
25. Li JH, Liu S, Zhou H, Qu LH, Yang JH. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-seq data. Nucleic Acids Res. 2014;42(Database issue):D92–97. doi:10.1093/nar/gkt1248
26. Pan SJ, Ren J, Jiang H, et al. MAGEA6 promotes human glioma cell survival via targeting AMPKalpha1. Cancer Lett. 2018;412:21–29. doi:10.1016/j.canlet.2017.09.051
27. Huang YC, Lin SJ, Shih HY, et al. Epigenetic regulation of NOTCH1 and NOTCH3 by KMT2A inhibits glioma proliferation. Oncotarget. 2017;8(38):63110–63120. doi:10.18632/oncotarget.18668
28. Zhang C, Song C, Liu T, et al. KMT2A promotes melanoma cell growth by targeting hTERT signaling pathway. Cell Death Dis. 2017;8(7):e2940. doi:10.1038/cddis.2017.285
29. Hyrenius-Wittsten A, Pilheden M, Sturesson H, et al. De novo activating mutations drive clonal evolution and enhance clonal fitness in KMT2A-rearranged leukemia. Nat Commun. 2018;9(1):1770. doi:10.1038/s41467-018-04180-1
30. Kanayama T, Imamura T, Kawabe Y, et al. KMT2A-rearranged infantile acute myeloid leukemia masquerading as juvenile myelomonocytic leukemia. Int J Hematol. 2018;108(6):665–669. doi:10.1007/s12185-018-2522-3
31. Peterson JF, Baughn LB, Pearce KE, et al. KMT2A (MLL) rearrangements observed in pediatric/young adult T-lymphoblastic leukemia/lymphoma: a 10-year review from a single cytogenetic laboratory. Genes Chromosomes Cancer. 2018;57(11):541–546. doi:10.1002/gcc.v57.11
32. Fang Y, Zhang D, Hu T, et al. KMT2A histone methyltransferase contributes to colorectal cancer development by promoting cathepsin Z transcriptional activation. Cancer Med. 2019;8:3544–3552. doi:10.1002/cam4.2019.8.issue-7
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.