Back to Journals » OncoTargets and Therapy » Volume 13
Mechanism of Sanguinarine in Inhibiting Macrophages to Promote Metastasis and Proliferation of Lung Cancer via Modulating the Exosomes in A549 Cells
Authors Yu Y , Luo Y, Fang Z, Teng W, Yu Y, Tian J, Guo P , Xu R , Wu J , Li Y
Received 3 May 2020
Accepted for publication 16 August 2020
Published 10 September 2020 Volume 2020:13 Pages 8989—9003
DOI https://doi.org/10.2147/OTT.S261054
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 3
Editor who approved publication: Dr Leo Jen-Liang Su
Yuanyuan Yu,1,* Yingbin Luo,1,* Zhihong Fang,1 Wenjing Teng,1 Yongchun Yu,2 Jianhui Tian,3 Peng Guo,1 Rongzhong Xu,1 Jianchun Wu,1 Yan Li1
1Department of Oncology, Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai University of Traditional Chinese Medicine, Shanghai 200071, People’s Republic of China; 2Institute for Thoracic Oncology, Shanghai Chest Hospital, Shanghai Jiao Tong University, Shanghai 200030, People’s Republic of China; 3Institute of Traditional Chinese Medicine in Oncology, Department of Oncology, Longhua Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai 200032, People’s Republic of China
*These authors contributed equally to this work
Correspondence: Yan Li; Jianchun Wu
Department of Oncology, Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai University of Traditional Chinese Medicine, No. 274, Zhijiang Road, Jing’an District, Shanghai 200071, People’s Republic of China
Tel/Fax +86-21-63925588
Email [email protected]; [email protected]
Objective: Sanguinarine (SNG) is a benzophenanthridine alkaloid obtained from the roots of Sanguinaria canadensis and has an anticancer effect. The aim of this study was to explore the mechanism of SNG in inhibiting macrophages via regulating the exosomes derived from lung carcinoma cells to reduce metastasis and proliferation of lung carcinoma.
Methods: Human lung cancer cells (A549 cells) were treated with 4μM of SNG. Exosomes of A549 cells were extracted from A549 cells supernatant, and THP-1 cells were cultured with exosomes. Then, the supernatant of THP-1 cells was collected and cultured with A549 cells. Cell proliferation was measured via plate clone formation and CCK-8 assays. Migration was assessed by using Transwell assay and scratch test. Cellular invasion was detected by Transwell assay. Apoptosis was determined using flow cytometry. Moreover, the protein expressions of GAPDH, P65 and P-P65 in THP-1 cells were measured by Western blot. Levels of tumor necrosis factor-α (TNF-α), interleukin-6 (IL-6), and chemotactic cytokines ligand 2 (CCL-2) extracted from THP-1 cells were determined by reverse transcription-polymerase chain reaction (RT-PCR).
Results: Compared to the control group, exosomes could activate THP-1 cells, and the invasion, migration, and proliferation of A549 cells were consequently enhanced. Exosomes could increase the protein expression of p-p65 and the RNA expression levels of TNF-α, IL-6, and CCL-2 in THP-1 cells. Compared with the exosome group, SNG-treated exosomes inhibited THP-1 cells so that the invasion, proliferation, and migration of A549 cells were attenuated and apoptosis was promoted. In THP-1 cells, SNG-treated exosomes inhibited P-P65 expression and the RNA expression levels of TNF-α, IL-6, and CCL-2.
Conclusion: Exosomes treated by SNG inhibited THP-1 cells so that the invasion, proliferation, and migration of A549 cells were inhibited, and the apoptosis was promoted. The mechanism is possibly associated with the inhibition of NF-κB pathway in THP-1 cells.
Keywords: SNG, exosomes, THP-1 cells, NF-κB signaling pathway
Introduction
Exosome, a kind of extracellular vesicles (EV), has a lipid-bilayer spheroid structure without replicating capacity and contains proteins, nucleic acids, lipids, etc. The sizes of the exosomes range from 30 nm to 100nm.1 Exosomes are regarded as key mediators in intercellular communications because their outer membrane structure can stably preserve the contents and deliver them to the receptor cells. It has been shown that exosomes from human lung cancer cells could promote the proliferation and ameliorate the apoptosis of A549 cells, thus enhancing development of malignancy.2 Furthermore, it is confirmed that the exosomes derived from tumor cells could change the microenvironment surrounding the tumors and establish a pro-cancer environment.3 There are various components in the tumor microenvironment, including stromal cells, endothelial cells, and immune cells. The interaction between these cells contributes to inflammatory responses in the microenvironment, which promotes development of the tumors, angiogenesis, and tumor metastasis.4,5 Macrophages respond to different signals in the microenvironment and have functional plasticity.6 In the microenvironment of tumors, macrophages can induce the development, metastasis, development and drug resistance of lung carcinoma.7 Therefore, inhibiting the influence of exosomes in tumor cells can reduce the connection between tumor cells and macrophages, thus inhibiting tumor cell metastasis and proliferation.
As a fatal malignancy, lung cancer owns the highest morbidity and mortality among all cancers across the world. Non-small cell lung cancer (NSCLC) accounts for about 85% among all kinds of lung cancers and can be treated by surgical resection, radiotherapy, chemotherapy, molecular targeted therapy and immunotherapy.8 NF-κB, a pro-inflammatory transcription factor, is a key activator in inflammation and decisively promotes tumor cell survival, proliferation, invasion, angiogenesis and metastasis. NF-κB can regulate expressions of inflammatory genes,9 including TNF-α, IL-6, CCL-2 and other inflammatory factors. Inflammatory factors in tumor microenvironment play an important role in the development, invasion and metastasis of tumors. TNF-α enhances tumor-producing inflammatory pathways such as NF-κB signaling pathway, which in turn stimulates the proliferation of malignant cells.10 IL-6 promotes the progression of colorectal cancer, tumor development and metastasis by regulating local and systemic inflammation and angiogenesis.11 CCL2 promotes tumor invasion and metastasis by recruiting mononuclear macrophages.12 In recent years, great attention has been paid to the natural products derived from plants to develop candidate anticancer drugs. SNG is a benzophenanthridine alkaloid obtained from the roots of Sanguinaria canadensis.13 A number of studies have shown that SNG had a therapeutic potential in a variety of human diseases, including lung cancer,14 cervical cancer,15 gastric cancer,16 hepatocellular carcinoma.17 However, the effect of SNG on exosomes in tumor microenvironment and the mechanism of its regulation on immunity are still unclear. It was found that SNG suppressed invasion, migration and proliferation of A549 cells and promoted cell apoptosis via regulating the exosomes which inhibited NF-κB signaling pathway in THP-1 cells. This study aimed to explore the mechanisms of SNG in ameliorating metastasis and proliferation of lung carcinoma cells through inhibiting macrophages via regulating the exosomes derived from lung carcinoma cells.
Materials and Methods
Reagents and Drugs
SNG (Figure 1) was obtained from Selleck (Shanghai, China). Dimethyl sulfoxide (DMSO) purchased from Sigma-Aldrich (St. Louis, MO, USA), was used to dissolve the agent (stock concentration: 10 mM). Annexin V-FITC/PI and CCK-8 assay kit was purchased from SigmaAldrich (St. Louis, MO, USA). Fetal bovine serum (FBS) was provided by GIBCO/BRL (Grand Island, NY, USA) and RPMI-1640 cell culture medium was purchased from HyClone (Logan, Utah, USA). The flow cytometry antibodies CD206, CD68, CD14 were purchased from Peprotech (New Jersey, NJ, USA). Primary antibodies of human p-NF-κB (1:1000), NF-κB (1:1000), and GAPDH (1:2000) were bought from Cell Signaling Technology (Boston, Massachusetts, USA). Primary antibodies of CD63 (dilution), Alix (dilution), and TSG101 (dilution) were bought from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Primary antibody of Calnexin (1:1000) was bought from Proteintech company (Chicago, Illinois, USA). Total exosome isolation reagent and exosome-depleted FBS media supplement were purchased from SBI (Johnston, Pennsylvania, USA).
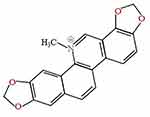 |
Figure 1 Chemical structure of sanguinarine. |
Cell Culture
Human acute monocytic leukemia (THP-1) and human lung cancer (A549) cell lines were purchased from the cell bank of Shanghai institute of life sciences (Shanghai, China). THP-1 and A549 cell lines were cultured by using Roswell Park Memorial Institute 1640 (RPMI-1640) medium added with 10% exosomes-depleted FBS (SBI, Johnston, Pennsylvania, USA).18 All cells were cultured in an incubator (37 °C) with humidified environment and CO2 (5%).
Exosome Separation and Its Characterization
Exosomes were obtained from the supernatant of A549 cells, which was described previously. A549 cell line was cultured in the medium without exosomes and SNG was added 24 hours later. Then, the supernatant was obtained at 48 h and centrifuged for 15 minutes at 3000g to eliminate cell debris (optimal-90K; BeckmanCoulter, Brea, California, USA). After centrifugation, the solution was filtrated by using filters (0.22-mm pore)(Millipore, St. Louis, MO, USA), and ExoQuick Exosome Precipitation Solution was added for incubation overnight. Afterwards, the solution was centrifuged for 30 minutes at 1500g. Sterile phosphate buffered saline (PBS) was used to dissolve the precipitation and the solution was kept in a −80 °C refrigerator. The concentrations of the proteins from exosomes were measured by BCA protein assay kit (Beyotime Biotechnology, Shanghai, China). Concentrations and sizes of the exosomes were detected by Nanoparticle Tracking Analysis (NTA). The exosome morphology was observed under a transmission electron microscope (Tecnai 12; Philips, Amsterdam, Netherlands).18,19 The marker proteins were detected by Western Blot.
NTA
Exosomes were diluted with PBS to a concentration of 1x107/mL ~1x109/mL and carefully injected into the particle point titration and particle size analyzer without bubbles. Particle matrix ZetaView PMX 110 (Particle Metrix, Meerbusch, Germany) was used to determine the concentration of the exosomes isolated at 405nm. The size and mass were also determined. Meanwhile, the particle trajectory of the exosomes could be analyzed.
Fluorescence Labeling of Exosomes
For staining, 1µM 3, 3ʹ-dioctadecyloxacarbocyanine perchlorate (Dio) was added to the purified exosomes for 10min. Then stained exosomes were centrifuged at 1 x 105g for 40 min at 4 °C for twice. The labeled exosomes in the precipitation were kept and suspended with Dulbecco’s Modified Eagle Medium (DMEM). THP-1 cells were cultured with the labeled exosomes for 24h, and a fluorescence microscope was used to observe exosomes entering the cells. After the culture solution was discarded, the cells were rinsed for 3 times with PBS, fixed for 10min with paraformaldehyde (4%), and then rinsed with PBS for 3 times. The cell nuclei were stained by 4ʹ,6-diamidino-2-phenylindole (DAPI) for 5min in room temperature. After staining, the cells were rinsed with PBS, and confocal microscopy was used to observe whether the Dio labeled exosomes were internalized by THP-1 cells.
THP-1 Cells Treated by Exosomes
THP-1 cells (5×105) were inoculated in 6-well culture plates for 12 hours followed by exosomes treatment. Then, the exosomes (200μg/mL) were added into THP-1 cells. In order to prepare for Western Blot and RT-PCR, THP-1 cells were treated with exosomes for 24h and 12h, respectively.
Preparation of Supernatant
THP-1 cells (5×105) were cultured for 12 h in 6-well culture plates followed by exosome treatment. PBS was used to suspend the exosomes, and the final concentration was 200μg/mL. After 24h treatment, culture medium with the exosomes was replaced. The supernatant was filtered, and then stored in a −80 °C refrigerator. In the NF-κB inhibition group, 10 μM 4.6-Quinazoline diamine (QNZ) (Selleck, Shanghai, China) was added and cultured with the cells for 2 h. In the function experiments, the supernatant was mixed with exosome-free culture medium in equal volume.
Flow Cytometry
THP-1 cells (5×105) were cultured for 12 h in 6-well culture plates followed by exosome treatment. PBS was used to suspend the exosomes, and the final concentration was 200μg/mL. After 24h treatment, differentiated cells were collected and identified by flow cytometry with specific markers (CD68, CD14 for M1, and CD206, CD14 for M2 macrophages). Unstained cells, as negative control, were used to determine the background autofluorescence, which can visualize the cells stained with anti- CD68 (or anti- CD14 or anti- CD206) antibodies.
CCK-8 Assay
After supernatant treatment for 48h, A549 cell line was cultured in 96-well culture plates (5000/200μL) for 24 hours. Then, the supernatant was replaced with the conditioned medium prepared before, and the cells were cultured for 24 hours. Afterwards, the supernatant was replaced with 200µL serum-free medium added with CCK-8 (10%) and incubated for 2 hours. Absorbance was detected at a wavelength of 490 nm by using a microplate reader (Thermo scientific, waltham, Massachusetts, USA). Three wells were set for repeated experiments in every group.
Cell Colony Formation Assay
After supernatant treatment for 48h, A549 cells were cultured in 6-well culture plates (600/well) for 12 days. Paraformaldehyde (4%) was used for cell fixing. After 30 minutes, the cells were stained with crystal violet for 15 minutes and then photographed. Every experiment was repeated for three times.
Wound Healing Assay
After supernatant treatment for 48h, A549 cells were cultured in 6-well culture plates (8×105/well) for 24 h. When the confluence of the cells was 80%, vertical scratches on the cell layer were made by using a 20μL sterile tip. PBS was used to wash the cells once and the picture was taken under the microscope. The cells were cultured for another 24 h and photographed.
Transwell Migration/Invasion Assay
After supernatant treatment for 48h, A549 cells were inoculated in 6-well culture plates (8×105/well) for 24 h. Culture medium without serum was used to suspend the cells and the concentration was adjusted to 5×105/mL. In the invasion experiment, the matrigel basement membrane (provided by BD Biosciences, Franklin Lake, New Jersey, USA) was diluted to 300µL/mL by using the pre-cooled culture medium without serum. Diluted matrigel (100μL) was placed in each chamber 30 min before the incubation (if Transwell migration assay was performed, this step was skipped). Afterwards, 200µL cells were added in the upper compartment and 500µL culture medium supplemented with 10% serum was added in the lower compartment for culture at 37°Cfor 24h. The cells on the bottom surface of the upper compartment were fixed by using paraformaldehyde (4%) for 30 minutes. The chambers were photographed after stained with crystal violet for 15 minutes.
Apoptosis Assay
After supernatant treatment for 48h, A549 cells were cultured with DMEM medium in 6-well plates (2×105 cells/mL) for 24 hours. A549 cells were cultured with the prepared conditioned medium for 36 hours. Afterwards, staining buffer (200 μL) was added into A549 cells (1×106 cells/mL). Annexin V (2 μL) was incubated with the cells at 2~8°C in darkness for 15 minutes. Afterwards, PI solution (4 μL) was added for incubation at 2~8°C in darkness for 5 minutes. A549 cell apoptosis was detected by a flow cytometer (BD, Franklin Lake, New Jersey, USA).
Western Blot Assay
Exosomes or cells were lysed by using RIPA buffer supplemented with protease inhibitor (Beyotime Biotechnology, Shanghai, China). Protein concentrations were detected by using BCA Protein Assay Kit (Beyotime Biotechnology, Shanghai, China). Proteins were subjected to electrophoresis by using 10% SDS-polyacrylamide gels (SDS-PAGE) and transferred onto Polyvinylidene Fluoride (PVDF) membranes. Afterwards, primary antibodies (human p-NF-κB, NF-κB, Calnexin, CD63, Alix, TSG101), GAPDH and secondary antibodies (Anti-rabbit IgG, HRP-linked Antibody) were incubated with the membranes. The protein bands were scanned by using ChemDocTM XRS+ and image LabTM software.
Extraction of RNA and Real-Time PCR
Trizol (Thermo scientific, Waltham, Massachusetts, USA) was used to obtain total RNA. Reverse transcription was conducted by using 1µL RNA and reverse transcriptase (TaKaRa, Japan). SYBR Green I real-time detection kit (TaKaRa, Japan) was used to conduct real-time PCR by using QuantStudio 6 Flex. GAPDH was used for normalization. Specific primers used in this experiment were listed as follows: IL-6 forward, GGTGTTGCCTGCTGCCTTCC, reverse, GCACAGCTCTGGCTTGTTCCTC;TNF-α forward, CCACGCTCTTCTGCCTGCTG, reverse, GACGGCGATGCGGCTGATG;CCL-2 forward, AGTCTCTGCCGCCCTTCTGTG, reverse, GGGTTTGCTTGTCCAGGTGGTC.
Statistics
All results were obtained from three repeated experiments. Experimental results were presented as average value ± standard deviation. One-way ANOVA was used to compare the difference between groups, and P <0.05 indicated significant difference. All statistical analyses were performed using SPSS23.0 or Graphpad 8.0 software.
Results
The Features of the Exosomes in the Lung Cancer Cells
Exosomes were extracted from the cell supernatant by using ExoQuick Exosome Precipitation Solution. The Nanoparticle Tracking Analysis demonstrated that size of the particles varied from 50nm to 200nm. The average particle size and concentration were 138.4nm and 8.3×1010 particles/mL, respectively (Figure 2A). It was proved by transmission electron microscopy that the particle has a typical cup-shaped structure (Figure 2B). Exosome positive marker proteins, CD63, Alix and TSG101 were expressed, while negative marker proteins, calnexin were not (Figure 2C). As a kind of extracellular vesicles, exosome does not contain mitochondrial proteins.
The Internalization of Exosomes in THP-1 Cells
To figure out whether the exosomes of lung cancer cells can be internalized into THP-1 cells, the exosomes were stained by Dio dye and co-cultured with THP-1 cells for 24h. Following co-culture with THP-1 cells for 24h, laser confocal microscope photography showed that the labeled exosomes were absorbed by THP-1 cells (Figure 3), indicating that exosomes can be internalized by THP-1 cells and affect THP-1 cells.
 |
Figure 3 The uptake of Dio-labeled exosomes (green fluorescence) in THP-1 cells was recorded. DAPI (blue fluorescence) was used for nuclei staining. PBS was control group. The bar indicated 50 μm. |
THP-1 Cell Polarization Towards M2 Macrophages Induced by Exosomes
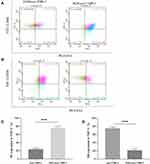
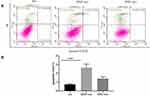
Macrophages can be divided into the classical subtype of activated macrophage (M1) and the alternative subtype of activated macrophages (M2)h.20 M1 macrophages secrete pro-inflammatory cytokines to play an anti-tumor role, while M2 macrophages express immunosuppressive signaling molecules to promote tumor growth, invasion and metastasis.21 The role of M1 and M2 macrophages in the tumor microenvironment is completely different. We examined the morphological changes of THP-1 cells and macrophage markers after treatment of exosomes. No obvious morphological changes were observed. CD14, CD68 and CD206 are the surface markers of human macrophages, M1 macrophages and M2 macrophages, respectively. Therefore, we used CD14 and CD68 double staining for M1-type macrophages, and CD14 and CD206 for M2-type macrophages. Flow cytometry (Figure 4A–D) results showed that most exosomes-treated THP-1 cells can be polarized to M2 macrophages while after SNG-treated exosomes treatment, most THP-1 cells can be polarized to M1 macrophages.
SNG Inhibited the Proliferation of A549 Cells
A549 cells were induced by multiple concentrations of SNG for 24, 48, and 72 hours, respectively. CCK-8 assay was applied to measure the proliferation of A549 cells. In Figure 5, proliferation was remarkably decreased when A549 cells were treated with SNG in a concentration- and time-dependent pattern compared to the control (0 μM SNG).
The Exosomes Induced THP-1 Cells Promote Migration, Development and Invasion of Lung Cancer Cells, Which Could Be Inhibited by SNG
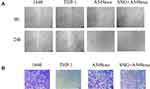
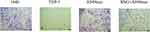
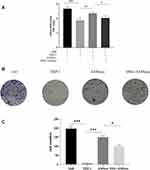
We further investigated the effects of THP-1 cells on lung cancer cells after treatment with exosomes or SNG-treated exosomes. Two groups were set as controls. The conditioned medium was used to stimulate lung cancer cells. After incubation with the supernatant of the THP-1 cells treated by the exosomes, the proliferation (Figure 6A–C), migration (Figure 7A and B) and invasion of A549 cells were remarkably enhanced. After incubation with the supernatant of the THP-1 cells induced by SNG-treated exosomes, the proliferation, migration and invasion (Figure 8) of A549 cells were dramatically reduced. There was no obvious change in the apoptosis after the cells were incubated with SOT induced by the exosomes in lung cancer cells. But the apoptosis of A549 cells was increased after incubated with SOT induced by SNG-treated exosomes in comparison with the exosomes without SNG treatment (Figure 9A and B). In summary, lung cancer cells were activated by SOT so that the development, invasion, and migration of A549 cells were promoted, while SNG inhibited these effects.
Exosomes of Lung Cancer Cells Could Activate NF-κB Pathway and Increase Levels of Pro-Inflammation Cytokines in THP-1 Cells, Which Could Be Reversed by SNG Treatment
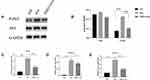
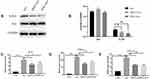
To investigate the possible mechanisms in the inhibitory effects of SNG-treated exosomes on THP-1 cells, the expression level of NF-κB, a common signal transducer related to cancer development and inflammation, was measured. It was demonstrated that there was no obvious change in P65 levels in the THP-1 cells treated by the exosomes in lung carcinoma cells (group A) or by exosomes from lung cancer cells induced by SNG (group B) for 24h. However, the protein level of P-P65 was remarkably elevated in group A and decreased in group B (Figure 10A and B). The mRNA levels of TNF-α, CCL2, and IL-6 in THP-1 cells were detected by RT-PCR. It was demonstrated that the mRNA levels of CCL2, TNF-α, and IL-6 in THP-1 cells were remarkably elevated in group A and decreased in group B (Figure 10C–E). We further investigated whether activated NF-κB contributes to the increase of pro-inflammation cytokines in THP-1 cells after exosomes treatment. After exosome treatment, NF-κB phosphorylation was blocked by QNZ, an inhibitor for NF-κB pathway. In Western blot, the level of p-NF-κB was remarkably decreased by QNZ (Figure 11A and B). In RT-PCR, the levels of TNF-α, CCL2, and IL-6 were also decreased (Figure 11C–E). In summary, the expression levels of pro-inflammation cytokines in THP-1 cells after exosomes treatment relied on the NF-κB activation, and SNG could reverse the changes in the expression.
Inhibited NF-κB Could Reverse the Effects of Exosomes on Development, Migration, Invasion, and Apoptosis of Lung Carcinoma Cells
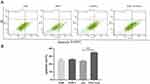
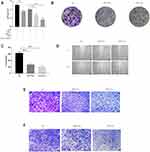
The role of NF-κB inhibitors in the effects of exosomes on development, invasion, migration, and apoptosis of A549 cells was investigated. THP-1 cells were pretreated with NF-κB inhibitor before exosome treatment. CCK-8 and cell colony formation assays demonstrated that proliferation (Figure 12A–C), migration (Figure 12D and E) and invasion (Figure 12F) of A549 cells were remarkably enhanced and apoptosis (Figure 13A and B) reduced after incubation with the supernatant of the THP-1 cells treated by the exosomes in lung cancer cells, which was remarkably ameliorated by using NF-κB inhibitor. In summary, it was indicated that exosomes in cancer cells could activate THP-1 cells, which could subsequently promote tumor growth, invasion and migration of lung carcinoma cells and inhibit the apoptosis via NF-κB pathway.
Discussion
Cancer progression is related to microenvironment.22 Exosomes are the key substance of intercellular communication.19 Exosomes derived from cancer cells consist of mRNAs, noncoding RNAs, proteins, and DNA. There were specific mechanisms in inserting chosen molecules into the exosomes.23 In this study, THP-1 cells simulated the human mononuclear macrophage system. The basic activities of cancer cells include development, differentiation, division, migration, invasion and death. A series of complex signal transmission and biological activities can increase the expression levels of the associated genes, regulate the biological activities of cells and determine the progress of cancer. In the present research, it was demonstrated that there were communications between macrophages and exosomes, and macrophages treated by exosomes affected tumor development. It has been discovered that exosomes in lung carcinoma cells were effectively absorbed into macrophages. It was also illustrated that macrophages activated by lung cancer exosomes could promote the development, invasion, and migration, and SNG could reverse these effects. In addition, the apoptosis was enhanced after the cells were incubated with the supernatant of the THP-1 cells induced by SNG-treated exosomes compared with the supernatant of the THP-1 cells induced by exosomes.
Macrophages are important drivers of inflammation in tumors.24 The functions of macrophage are diverse and the phenotypes are different in various tumors or different parts of the same tumor.25 Macrophages can polarize to two types under different stimulation: M1 type (pro-inflammation) or M2 (anti-inflammation).26 Advances have been achieved in identifying molecular pathways in macrophages related to tumors, including NF-kB signaling pathway.24
The study showed that NF-κB pathway could regulate the expression levels of inflammation-related genes.27 NF-κB plays a critical part in modulating the immediate reactions to pathogens, cellular proliferation, and survival.28 The study showed that NF-κB could regulate levels of inflammation-related genes.9 NF-κB p65, as the most common activated form in NF-κB family, is an important component in NF-κB pathway.29 In our research, we investigated the mechanisms of NF-κB pathway in the activated macrophages induced by exosomes. It was indicated that exosomes in lung carcinoma cells could facilitate NF-κB pathway activation in macrophages. Furthermore, the exosomes could not deliver the activated molecules via NF-κB pathway to the macrophages after NF-κB pathway was sufficiently suppressed in the macrophages pretreated by NF-κB inhibitor. SNG can inhibit the exosomes in lung carcinoma cells so as to inhibit NF-κB pathway activation in macrophages. SNG is a benzophenanthridine alkaloid obtained from the roots of Sanguinaria canadensis and has anticancer effect.13,30 It was reported that SNG could act on the Shh-Gli-Nanog pathway and inhibit the proliferation of pancreatic cancer stem cells.30 It was reported that SNG inhibited proliferation of non-small cell lung cancer cells by upregulating Fas-associated factor 1.14 Therefore, the discovery of new anticancer drugs from Chinese herbal medicine resources is promising. Our study suggested that SNG could suppress macrophages, and hence inhibit metastasis and proliferation of lung cancer through modulating the exosomes from the lung cancer cells. However, changes in exosomes were not explored in this study. Therefore, the morphology, quantity and substance of exosomes under the treatment of SNG and how do exosomes affect THP −1 cells remain to be further explored. This research will help to seek effective anticancer drugs from traditional Chinese medicine and find more therapeutic targets.
Conclusions
To conclude, our research implied that exosomes could activate THP-1 cells to promote the proliferation, invasion and migration and inhibit the apoptosis of A549 cells. SNG-treated exosomes could inhibit THP-1 cells so as to ameliorate cancer progression. The mechanism may have relationships with suppressed NF-κB pathway in macrophages.
Acknowledgments
The authors appreciated the Shanghai Municipal Hospital of Traditional Chinese Medicine, Shanghai University of Traditional Chinese Medicine. Yuanyuan Yu and Yingbin Luo are co-first authors. Both Yan Li and Jianchun Wu are corresponding authors. This study was supported by the National Nature Science Foundation of China (81973795 and 81673947), the Shanghai Natural Science Fund (17ZR1428000 and 19ZR1452200), Shanghai Municipal Commission of Health and Family Planning (20154Y0060).
Disclosure
The authors declare no conflicts of interest.
References
1. Li K, Chen Y, Li A, Tan C, Liu X. Exosomes play roles in sequential processes of tumor metastasis. Int J Cancer. 2019;144(7):1486–1495. doi:10.1002/ijc.31774
2. Huang J, Ding Z, Luo Q, Xu W. Cancer cell-derived exosomes promote cell proliferation and inhibit cell apoptosis of both normal lung fibroblasts and non-small cell lung cancer cell through delivering alpha-smooth muscle actin. Am J Transl Res. 2019;11(3):1711–1723.
3. Li C, Liu DR, Li GG, et al. CD97 promotes gastric cancer cell proliferation and invasion through exosome-mediated MAPK signaling pathway. World J Gastroenterol. 2015;21(20):6215–6228. doi:10.3748/wjg.v21.i20.6215
4. Yang TT, Zhang X, Wang M, et al. Activation of mesenchymal stem cells by macrophages prompts human gastric cancer growth through NF-κB pathway. PLoS One. 2014;9(5):e97569. doi:10.1371/Journal.pone.0097569
5. Luga V, Zhang L, Viloria-petit AM, et al. Exosomes mediate stromal mobilization of autocrine Wnt-PCP signaling in breast cancer cell migration. Cell. 2012;151(7):1542–1556. doi:10.1016/j.cell.2012.11.024
6. Li YS, Zhao LM, Shi BH, et al. Functions of miR-146a and miR-222 in tumor-associated macrophages in breast cancer. Sci Rep. 2015;5:18648–18663. doi:10.1038/srep18648
7. Cho HJ, Jung JI, Lim DY, et al. Bone marrow-derived, alternatively activated macrophages enhance solid tumor growth and lung metastasis of mammary carcinoma cells in a Balb/C mouse orthotopic model. Breast Cancer Res. 2012;14(3):R81. doi:10.1186/bcr3195
8. Hirsch FR, Scagliotti GV, Mulshine JL, et al. Lung cancer: current therapies and new targeted treatments. Lancet. 2017;389:299–311. doi:10.1016/S0140-6736(16)30958-8
9. Scirpo R, Fiorotto R, Villani A, Amenduni M, Spirli C, Strazzabosco M. Stimulation of nuclear receptor peroxisome proliferator-activated receptor-γ limits NF-κB-dependent inflammation in mouse cystic fibrosis biliary epithelium. Hepatology. 2015;62(5):1551–1562. doi:10.1002/hep.28000
10. Murata M, Thanan R, Ma N, et al. Role of nitrative and oxidative DNA damage in inflammation-related carcinogenesis. J Biomed Biotechnol. 2012;10:623019. doi:10.1155/2012/623019
11. Guthrie GJK, Roxburgh CSD, Richards CH, et al. Circulating IL-6 concentrations link tumour necrosis and systemic and local inflammatory responses in patients undergoing resection for colorectal cancer. Br J Cancer. 2013;109(1):131–137. doi:10.1038/bjc.2013.291
12. Roca H, Varsos ZS, Sud S, et al. CCL2 and interleukin-6 promote survival of human CD11b + peripheral blood mononuclear cells and induce M2-type macrophage polarization. J Biol Chem. 2009;284(49):34342–34354. doi:10.1074/jbc.M109.042671
13. Akhtar S, Achkar IW, Siveen KS, et al. Sanguinarine induces apoptosis pathway in multiple myeloma cell lines via inhibition of the JaK2/STAT3 signaling. Front Oncol. 2019;9:285. doi:10.3389/fonc.2019.00285
14. Wei G, Xu Y, Peng T, Yan J, Wang Z, Sun Z. Sanguinarine exhibits antitumor activity via up-regulation of Fas-associated factor 1 in non-small cell lung cancer. J Biochem Mol Toxicol. 2017;31:e21914. doi:10.1002/jbt.21914
15. Zhang H, Zhang J, Venkat PS, Gu C, Meng Y. Sanguinarine exhibits potent efficacy against cervical cancer cells through inhibiting the STAT3 pathway in vitro and in vivo. Cancer Manag Res. 2019;11:7557–7566. doi:10.2147/CMAR.S212744
16. Fan HN, Chen W, Peng SQ, et al. Sanguinarine inhibits the tumorigenesis of gastric cancer by regulating the TOX/DNA-PKCS/KU70/80 pathway. Pathol Res Pract. 2019;215:152677. doi:10.1016/j.prp.2019.152677
17. Zhang B, Wang X, Deng J, et al. P53-dependent upregulation of miR-16-2 by sanguinarine induces cell cycle arrest and apoptosis in hepatocellular carcinoma. Cancer Lett. 2019;459:50–58. doi:10.1016/j.canlet.2019.05.042
18. Wu L, Zhang X, Zhang B, et al. Exosomes derived from gastric cancer cells activate NF-κB pathway in macrophages to promote cancer progression. Tumor Biol. 2016;37:12169–12180. doi:10.1007/s13277-016-5071-5
19. Thery C, Zitvogel L, Amigorena S. Exosomes: composition, biogenesis and function. Nat Rev Immunol. 2002;2(8):569–579. doi:10.1038/nri855
20. Mills CD. M1 and M2 macrophages: oracles of health and disease. Crit Rev Immunol. 2012;32(6):463–488. doi:10.1615/CritRevImmunol.v32.i6.10
21. Ruffell B, Affara NI, Coussens LM. Differential macrophage programming in the tumor microenvironment. Trends Immunol. 2012;33(3):119–126. doi:10.1016/j.it.2011.12.001
22. Challagundla KB, Wise PM, Neviani P, et al. Exosome-mediated transfer of microRNAs within the tumor microenvironment and neuroblastoma resistance to chemotherapy. J Natl Cancer Inst. 2015;107(7):djv135. doi:10.1093/jnci/djv135
23. Paggetti J, Haderk F, Seiffert M, et al. Exosomes released by chronic lymphocytic leukemia cells induce the transition of stromal cells into cancer-associated fibroblasts. Blood. 2015;126(9):1106–1117. doi:10.1182/blood-2014-12-618025
24. Mantovani A, Allavena P. The interaction of anticancer therapies with tumor-associated macrophages. J Exp Med. 2015;212(4):435–445. doi:10.1084/jem.20150295
25. Ma S, Liu M, Xu Z, et al. A double feedback loop mediated by microRNA-23a/27a/24-2 regulates M1 versus M2 macrophage polarization and thus regulates cancer progression. Oncotarget. 2016;7(12):13502–13519. doi:10.18632/oncotarget.6284
26. Zhou D, Huang C, Lin Z, et al. Macrophage polarization and function with emphasis on the evolving roles of coordinated regulation of cellular signaling pathways. Cell Signal. 2014;26:192–197. doi:10.1016/j.cellsig.2013.11.004
27. Yang Z, Xian H, Hu J, et al. USP18 negatively regulates NF-κB signaling by targeting TAK1 and NEMO for deubiquitination through distinct mechanisms. Sci Rep. 2015;5:12738. doi:10.1038/srep12738
28. Ghosh S, Hayden MS. New regulators of NF-κB in inflammation. Nat Rev Immunol. 2008;8(11):837–848. doi:10.1038/nri2423
29. Liu P, Yuan HB, Zhao S, et al. Activation of GABAB receptor suppresses diabetic neuropathic pain through toll-like receptor 4 signaling pathway in the spinal dorsal horn. Mediators Inflamm. 2018;2018:6016272. doi:10.1155/2018/6016272
30. Ma Y, Yu W, Shrivastava A, et al. Sanguinarine inhibits pancreatic cancer stem cell characteristics by inducing oxidative stress and suppressing sonic hedgehog-Gli-Nanog pathway. Carcinogenesis. 2017;38(10):1047–1056. doi:10.1093/carcin/bgx070
 © 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2020 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.