Back to Journals » OncoTargets and Therapy » Volume 12
MAGI1 Inhibits the Proliferation, Migration and Invasion of Glioma Cells
Authors Li ZY, Li XH, Tian GW, Zhang DY , Gao H, Wang ZY
Received 7 September 2019
Accepted for publication 5 December 2019
Published 19 December 2019 Volume 2019:12 Pages 11281—11290
DOI https://doi.org/10.2147/OTT.S230236
Checked for plagiarism Yes
Review by Single anonymous peer review
Peer reviewer comments 2
Editor who approved publication: Prof. Dr. Takuya Aoki
Zhong-Yan Li,1,2 Xue-Hua Li,2 Guang-Wei Tian,3 Dong-Yong Zhang,4 Hai Gao,1 Zhen-Yu Wang1
1Department of Anatomy, College of Basic Medical Sciences, China Medical University, Shenyang, People’s Republic of China; 2Department of Neurosurgery, Fuxin Central Hospital, Fuxin, People’s Republic of China; 3Department of Radiation Oncology, The First Affiliated Hospital of China Medical University, Shenyang, People’s Republic of China; 4Department of Neurosurgery, First Affiliated Hospital of China Medical University, Shenyang, People’s Republic of China
Correspondence: Zhen-Yu Wang
Department of Anatomy, College of Basic Medical Sciences, China Medical University, Shenyang, People’s Republic of China
Email [email protected]
Background: Membrane-associated guanylate kinase inverted repeat member 1 (MAGI1) acts as a tumor suppressor in a variety of tumors; however, its expression and biological function in glioma are still unknown.
Methods: MAGI1 expression in glioma was examined by immunohistochemistry. In addition, overexpression of MAGI1 in U87 and U373 cells, colony formation and MTT assays were used to evaluate cell proliferation, Transwell assays to determine cell migration and invasion, and a xenograft model established using U87 cells to evaluate the effect of MAGI1 overexpression in vivo. Western blot assays were used to analyze the Akt, MMP2, MMP9 and E-cadherin/N-cadherin/vimentin pathway changes after overexpression of MAGI1.
Results: We demonstrated that MAGI1 was expressed at low levels in glioma. Low MAGI1 expression was positively correlated with the malignant progression of glioma and indicated a poor prognosis. Moreover, we found that overexpressed MAGI1 inhibited the proliferation, migration and invasion of glioma cells by regulating cell growth and EMT through Akt, MMP2, MMP9 and the E-cadherin/N-cadherin/vimentin pathway.
Conclusion: These findings demonstrate a novel function of MAGI1 in glioma progression and suggest that MAGI1 might be a target for the diagnosis and treatment of glioma.
Keywords: membrane-associated guanylate kinase inverted repeat member 1, glioma, proliferation, migration, invasion
Introduction
Glioma is one of the most common tumors of the human central nervous system. Although there are treatments including microsurgery, chemotherapy and radiotherapy, the prognosis of glioma patients is still poor.1–4 The World Health Organization (WHO) classifies glioma tumors into four grades (I–IV) according to their prognostic histopathological characteristics, of which grade IV glioblastoma (GBM) is the most lethal. Current studies show that the 5-year survival rate of patients is only 10%.2,5 Therefore, a more in-depth study of the pathogenesis of glioma and the discovery of markers closely related to prognosis may provide a theoretical basis for the early diagnosis of glioma, as well as new therapeutic targets for its treatment.
Membrane-associated guanylate kinase inverted repeat member 1 (MAGI1) is an important protein that communicates the extracellular environment and intracellular signaling pathways as well as the cytoskeleton in synapses and tight junctions.6–8 MAGI1 consists of PDZ domains, a guanylate kinase domain and two WW domains.7,8 MAGI1 is widely expressed in multiple human tissues of almost all epithelial cell types. Dysregulated expression of MAGI1 has been reported, and MAGI1 acts as a tumor suppressor in a variety of tumors, such as, colorectal cancer, hepatocellular carcinoma, T-cell leukemia, cervical cancer and gastric cancer.9–18 MAGI1 has since been shown to participate in a wide range of biological functions, such as cell junctions, adhesion, proliferation, apoptosis, migration and invasion.9,19–21 However, its expression and roles in glioma, as well as its specific molecular mechanisms are unclear.
Our research aimed to confirm the expression of MAGI1 in gliomas and its relationship to clinicopathological features. In addition, we explored the biological functions and mechanisms of MAGI1 in gliomas. Our findings provide theoretical and experimental evidence for the diagnosis and treatment of glioma patients.
Materials and Methods
Patients and Tissue Specimens
Eighty-six glioma tissue samples were collected from the First Affiliated Hospital of China Medical University from 2009 to 2013. Patients that had received radiotherapy or chemotherapy before the surgery were excluded. Tumor samples removed by surgery were fixed in 4% paraformaldehyde or liquid nitrogen for morphological or molecular biological experiments respectively. This study was approved by the Ethical Committee of the First Affiliated Hospital of China Medical University at Shenyang. Written informed consent was obtained from all participants.
Cell Culture and Transfection
The cell lines astrocyte, U251, U87, U373, A172 and U118 were cultured in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum and 100 units/mL penicillin/streptomycin at 37°C in a 5% CO2 incubator. U87 and U373 cells were purchased from the Cell Bank of Shanghai Chinese Academy of Sciences. U251, A172 and U118 cells were purchased from Shanghai HonSun Biological Technology Co., Ltd. Astrocyte cells were purchased from Procell Life Science and Technology Co., Ltd. MAGI1 plasmids were purchased from Shanghai GeneChem Company. The U87 and U373 cell lines were transfected with MAGI1 plasmids and screened by puromycin.
Immunohistochemical Staining
After paraffin tissue sections were dehydrated by hydration and subjected to antigen retrieval, an UltraSensitive SP (Mouse/Rabbit) IHC Kit (Maxin Bio-Technology Co. Ltd., Beijing, China) was used for immunohistochemistry. The anti-MAGI1 antibody was used at dilution of 1:100 (Abcam, Cambridge, UK), developed with DAB, and counterstained with hematoxylin. The intensity of the staining was scored as follows: 0 (negative), 1 (weakly positive), 2 (moderately positive), and 3 (strongly positive). The percentage of positive cells was scored as follows: 0 (0% cells stained), 1 (1–25% cells stained), 2 (26–50% cells stained), and 3 (50–100% cells stained). The intensity and percentage scores were multiplied and classified into two groups: low expression (0–4.5) and high expression (4.5–9).
Western Blotting
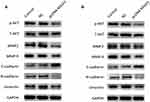
Human glioma tissues and cells were lysed by radioimmunoprecipitation assay (RIPA) lysis buffer (Beyotime, Jiangsu, China) containing protease and phosphatase inhibitors. Equal amounts of protein were separated on 10–12% SDS-PAGE gels and transferred onto polyvinylidene difluoride (PVDF) membranes (Millipore, Billerica, MA, USA). After blocking with 5% nonfat milk at room temperature for 1 h, the membranes were incubated with anti-MAGI1 (1:1000 Abcam, Cambridge, UK), anti-AKT (1:1500, Cell Signaling Technology, Beverly, MA), anti-p-AKT (Ser473, 1:1500, Cell Signaling Technology, Beverly, MA), anti-MMP2 (1:1000 Abcam, Cambridge, UK), anti-MMP9 (1:1000 Abcam, Cambridge, UK), anti-N-cadherin (1:1000 Abcam, Cambridge, UK) anti-vimentin (1:1000 Abcam, Cambridge, UK), anyi-E-cadherin (1:1000 Abcam, Cambridge, UK), GAPDH (1:5000 Abcam, Cambridge, UK) overnight at 4 °C. After washing the membranes with TBST, the protein signals were developed by using an ECL reagent, and the densities were analyzed by Gel-Pro Analyzer software. Protein signals were visualized by Bio-Rad imaging system (Bio-Rad Laboratories, Inc., Hercules, CA, USA).
Colony Formation Assays
For the colony formation assays, 500 cells were seeded in 6-well plates. The cells were cultured in medium with 10% FBS and at 37°C for 14 days for 2 weeks. The colonies were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet for 30 min. The colony forming efficiency was counted manually. The experiment was performed in triplicate.
Migration and Invasion Assays
For the Transwell assay, cells with at a density of 3–5 x 104 suspended in 200 µL of serum-free medium were seeded in the upper chamber of each Transwell, and the lower chamber was filled with 10% FBS medium. The upper chamber was coated with Matrigel (BD Biosciences, San Jose, CA) for invasion assays, and no Matrigel was used for the migration experiment. After culturing for 24 h, the chamber membrane were fixed in 4% paraformaldehyde for 15 min and stained with a 0.1% crystal violet. Each experiment was repeated three times.
Xenograft Model in Nude Mice
U87 cells (1 × 107 in 0.1mL PBS) were injected into nude mouse. Tumor length (L) and width (W) were measured every 5 days, and the volume was calculated by using the following formula: volume = L×W2×0.5. All mice were sacrificed on day 35 and tumor volume and weight were measured. The animal experiments conformed to the guidelines for the care and use of laboratory animals, and were approved by the Ethics Committee of China Medical University.
Statistical Analysis
Differences between the groups were evaluated by one-way ANOVA with Tukey’s post hoc test. The Kaplan–Meier method was used for the survival analysis, and univariate and multivariate Cox regressions were used to explore the association between the biomarkers and the outcomes of patient characteristics. All statistical analyses were performed using SPSS 19.0 software (IBM Corporation, Armonk, NY, USA). The results were considered significant if P <0.05, and the data are presented as the mean ± SD from at least three independent experiments.
Results
MAGI1 Expression Is Downregulated in Glioma Patients and Correlated with Poor Prognosis
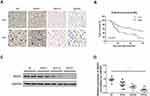
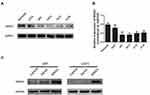
To determine the role of MAGI1 in glioma, the protein expression of MAGI1 was examined in 86 glioma tissues. The clinicopathological characteristics of the patients are listed in Table 1. The expression of MAGI1 in these glioma tissues and 7 normal brain tissues was detected by immunohistochemistry. The results showed that MAGI1 was strongly expressed in normal brain tissues and expression was significantly higher in the low-grade glioma tissues (WHO II) than in the high-grade tissues (WHO III and WHO IV, Figure 1A). We examined 7 normal brain tissues and 29 glioma tissues by Western blot (Figure 1C and D, *P <0.05), and the results were consistent with the immunohistochemistry findings. As presented in Table 2, MAGI1 expression scores correlated significantly with tumor stage (p=0.008) and tumor size (p=0.039). Other clinicopathological features, including patient age, gender, and Karnofsky Performance Scale (KPS) score showed no statistically significant correlation. Importantly, Kaplan-Meier survival analysis revealed that glioma patients with lower MAGI1 expression were significantly correlated with worse prognosis compared with patients with higher MAGI1 expression (P <0.05, Figure 1B). We also measured MAGI1 protein expression in 5 glioma cell lines (U251, U87, U118, U373 and A172) and an astrocyte cell line, and the Western blot result showed that MAGI1 expression in the five glioma cell lines was clearly lower than that in the astrocyte cell line (Figure 2A and B).
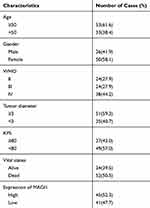 |
Table 1 MAGI1 in Glioma Clinicopathological Characteristics of Patient Samples and Expression of MAGI1 in Glioma |
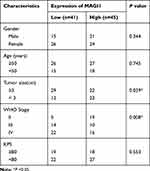 |
Table 2 Correlation Between MAGI1 Expression and Clinicopathologic Characteristics of Glioma Patients |
MAGI1 Inhibits Glioma Cell Proliferation and Colony Formation
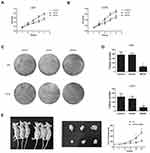
To investigate the role of MAGI1 in development of glioma, we selected two glioma cell lines, U87 and U373 for transfection with a MAGI1 overexpression plasmid (MAGI1). Cells transfected with empty vector (Vector) or not transfected (Control) were used as controls. The transfection efficiency was detected by Western blotting (Figure 2C). MTT and colony formation assays demonstrated that overexpression of MAGI1 significantly inhibited the proliferation rate and colony formation ability in both U87 and U373 cells (Figure 3A–D, *P <0.05). To verify whether MAGI1 affects tumor growth in vivo, we subcutaneously injected MAGI1 overexpressing U87 cells into BALB/c mice to establish a xenograft tumor model. The mice in the control group were concurrently injected with the corresponding NC cells. The results demonstrated that, tumor volumes were remarkably reduced in the MAGI1 overexpression group compared to the NC group, and statistically significant differences were observed at 28 (P <0.05) and 35 (P <0.05) days after injection (Figure 3E).
MAGI1 Suppresses Glioma Cell Migration and Invasion
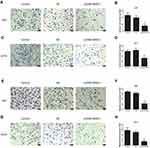
To determine whether MAGI1 gene expression is necessary for invasion and migration in U87 and U373 cells, Transwell migration and Matrigel invasion assays were used to assess the migration and invasion ability respectively. The results showed that the invasion (Figure 4A–D, **P <0.01) and migration (Figure 4E–H, **P <0.01,***P <0.001) ability of cells overexpressing MAGI1was significantly reduced compared with the Vector and Control groups.
MAGI1 Regulates Cell Growth and EMT Through Akt, MMP2, MMP9 and the E-Cadherin/N-Cadherin/Vimentin Pathway
Furthermore, Western blotting analysis showed that overexpression of MAGI1 inhibited the expression of p-AKT MMP2 and MMP9 in U87 and U373 cells. As indicated by the Western blot assay, the expression of the epithelial marker protein E-cadherin was increased, while the expression of the mesenchymal marker N-cadherin and vimentin was decreased in the MAGI1 overexpression group compared with the control group in both U87 and U373 cells (Figure 5A and B). These results suggested that overexpression of MAGI1 could decrease the proliferation, migration and invasion of glima cells by affecting the p-AKT, MMP2, MMP9 and EMT pathways.
Discussion
MAGI1 is an important protein that mediates the communication between the extracellular environment and intracellular signaling pathways as well as the cytoskeleton in synapses and tight junctions. Currently, increasing research is focusing on the important role of MAGI1 in tumors. For instance, overexpression of MAGI1 promotes anchorage-independent growth, migration and invasion in vitro by stabilizing E-cadherin and β-catenin localization at cell–cell junctions, enhancing actin stress fiber and focal adhesion formation, increasing cell adhesion to matrix proteins and suppressing the Wnt signaling pathway in the colorectal cancer cell lines SW480 and HCT116.22 A previous study showed that overexpression of MAGI1 promotes cell migration and invasion by regulating the expression of phosphatase and tensin homolog deleted on chromosome 10 (PTEN) in HepG2 cells.21 Knockdown of MAGI1 significantly promoted cell migration and invasion by altering the expression of matrix metalloproteinases (MMPs) and epithelial-mesenchymal transition (EMT)-related molecules by inhibiting the MAPK/ERK signaling pathway in gastric cancer cells.13 The re-expression of MAGI1 in HPV-positive cells suppressed cell proliferation and induced apoptosis by increasing the recruitment of ZO-1 and PAR3 to sites of cell-cell contact.11 In addition, studies have shown that the tumor-associated microenvironment plays important roles in tumor progression and drug resistance, and as a multilayer network biomarker (MNB) to predict survival outcome and therapeutic responses in glioma patients.23,24 These studies could reveal the role of MAGI1 in cell-cell communication or microenvironment-tumor cell interactions.
In this study, we found that MAGI1 was downregulated in clinical glioma samples. More importantly, MAGI1 expression was found to be significantly correlated with tumor size (P <0.05) and WHO stage (P <0.01). Moreover, we found that overexpression of MAGI1 inhibited the proliferation, colony formation, migration and invasion of glioma cells. These data strongly suggest that MAGI1 acts as a tumor suppressor anddemonstrate a novel function for MAGI1 in glioma progression.
It has been reported that MAGI1, as a multidomain adaptor protein, participates in a wide range of biological functions. In renal cell carcinoma (RCC), it has been reported that knockdown of MAGI1 increased Akt phosphorylation at Ser473, and GSK-3β phosphorylation at Ser9 and decreased β-catenin phosphorylation at Ser33/77, which resulted in upregulation of total β-catenin.25 In our study, we wanted to explore how MAGI1 affects cell proliferation, migration and invasion. The AKT pathway plays an important role in cell proliferation. Consistent with a previous study, we also found that overexpression of MAGI1 can inhibit the level of p-Akt in glioma cancer cells. Furthermore, we investigated the potential molecular mechanism of migration and invasion mediated by MAGI1. Degradation of the basal membrane is a necessary step for most cancers to accelerate tumor aggressiveness.26–28 In our study, we found that overexpression of MAGI1 inhibited the expression of MMP2 and MMP9. In glioma, EMT is also an essential pathway for metastasis progression in different tumor types.29,30 The occurrence of EMT is believed to change the expression of genes whose products play a key role in maintaining the epithelial or mesenchymal status of cells, such as E-cadherin, N-cadherin and vimentin.31–33 In this study, it was shown that overexpression of MAGI1 upregulated the expression of the epithelial marker E-cadherin and down-regulated the expression of the mesenchymal markers N-cadherin and vimentin. These results suggest that overexpression of MAGI1 decreases the migration and invasion of glioma cancer cells by inhibiting EMT.
Our study found low expression of MAGI1 in high-grade gliomas, but the mechanism is currently unclear. Based on the results of existing studies, we speculate that it may be closely related to the cyclooxygenase-2 (COX-2). COX-2 converts arachidonic acid into bioactive lipids, including prostaglandinE2 (PGE2), which has been found to be elevated in various types of tumors. Studies have documented that the localized expression of COX-2 and its catalyzed product, PGE2, are sufficient to initiate the development of and promote the progression of tumors in situ.34 Moreover, COX-2 expression increased according to the histopathological grade of the glioma, which suggested that the COX-2 pathway promotes gliomagenesis. MAGI1 is a tumor-suppressor protein induced by COX-2 inhibitors in colorectal cancer cells, suggesting that COX-2 negatively regulates the expression of MAGI1.9
Our study demonstrated that MAGI1 was expressed at low levels in glioma. The expression of MAGI1 was negatively correlated with the malignant progression of glioma and suggested a worse prognosis. Furthermore, we found that MAGI1 affected the proliferation, migration and invasion of glioma cells by regulating the p-AKT, MMP2/9 and EMT pathways. Our study determined thatMAGI1 is involved in glioma progression and suggested that MAGI1 might be a target for the diagnosis and treatment of glioma.
Acknowledgments
This work was supported by the Science Foundation Project of Liaoning Province (2019-MS-134).
Disclosure
The authors declare no conflicts of interest in this work.
References
1. Louis DN, Perry A, Reifenberger G, et al. The 2016 world health organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016;131(6):803–820. doi:10.1007/s00401-016-1545-1
2. Omuro A, DeAngelis LM. Glioblastoma and other malignant gliomas: a clinical review. JAMA. 2013;310(17):1842–1850. doi:10.1001/jama.2013.280319
3. Wen PY, Kesari S. Malignant gliomas in adults. N Engl J Med. 2008;359(5):492–507. doi:10.1056/NEJMra0708126
4. DeAngelis LM. Brain tumors. N Engl J Med. 2001;344(2):114–123. doi:10.1056/NEJM200101113440207
5. Louis DN, Ohgaki H, Wiestler OD, et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007;114(2):97–109. doi:10.1007/s00401-007-0243-4
6. Feng X, Jia S, Martin TA, Jiang WG. Regulation and involvement in cancer and pathological conditions of MAGI1, a tight junction protein. Anticancer Res. 2014;34(7):3251–3256.
7. Laura RP, Ross S, Koeppen H, Lasky LA. MAGI-1: a widely expressed, alternatively spliced tight junction protein. Exp Cell Res. 2002;275(2):155–170. doi:10.1006/excr.2002.5475
8. Oliva C, Escobedo P, Astorga C, Molina C, Sierralta J. Role of the MAGUK protein family in synapse formation and function. Dev Neurobiol. 2012;72(1):57–72. doi:10.1002/dneu.20949
9. Zaric J, Joseph JM, Tercier S, et al. Identification of MAGI1 as a tumor-suppressor protein induced by cyclooxygenase-2 inhibitors in colorectal cancer cells. Oncogene. 2012;31(1):48–59. doi:10.1038/onc.2011.218
10. Zhang G, Liu T, Wang Z. Downregulation of MAGI1 associates with poor prognosis of hepatocellular carcinoma. J Invest Surg. 2012;25(2):93–99. doi:10.3109/08941939.2011.606875
11. Kranjec C, Massimi P, Banks L. Restoration of MAGI-1 expression in human papillomavirus-positive tumor cells induces cell growth arrest and apoptosis. J Virol. 2014;88(13):7155–7169. doi:10.1128/JVI.03247-13
12. Makokha GN, Takahashi M, Higuchi M, Saito S, Tanaka Y, Fujii M. Human T-cell leukemia virus type 1 Tax protein interacts with and mislocalizes the PDZ domain protein MAGI-1. Cancer Sci. 2013;104(3):313–320. doi:10.1111/cas.12087
13. Jia S, Lu J, Qu T, et al. MAGI1 inhibits migration and invasion via blocking MAPK/ERK signaling pathway in gastric cancer. Chin J Cancer Res. 2017;29(1):25–35. doi:10.21147/j.issn.1000-9604.2017.01.04
14. Murata M, Kojima T, Yamamoto T, et al. Tight junction protein MAGI-1 is up-regulated by transfection with connexin 32 in an immortalized mouse hepatic cell line: cDNA microarray analysis. Cell Tissue Res. 2005;319(2):341–347. doi:10.1007/s00441-004-1017-0
15. Zhang Y, Dasgupta J, Ma RZ, Banks L, Thomas M, Chen XS. Structures of a human papillomavirus (HPV) E6 polypeptide bound to MAGUK proteins: mechanisms of targeting tumor suppressors by a high-risk HPV oncoprotein. J Virol. 2007;81(7):3618–3626. doi:10.1128/JVI.02044-06
16. Kranjec C, Banks L. A systematic analysis of human papillomavirus (HPV) E6 PDZ substrates identifies MAGI-1 as a major target of HPV type 16 (HPV-16) and HPV-18 whose loss accompanies disruption of tight junctions. J Virol. 2011;85(4):1757–1764. doi:10.1128/JVI.01756-10
17. Grm HS, Banks L. Degradation of hDlg and MAGIs by human papillomavirus E6 is E6-AP-independent. J Gen Virol. 2004;85(Pt 10):2815–2819. doi:10.1099/vir.0.80035-0
18. Charbonnier S, Nomine Y, Ramirez J, et al. The structural and dynamic response of MAGI-1 PDZ1 with noncanonical domain boundaries to the binding of human papillomavirus E6. J Mol Biol. 2011;406(5):745–763. doi:10.1016/j.jmb.2011.01.015
19. Dobrosotskaya IY, James GL. MAGI-1 interacts with beta-catenin and is associated with cell-cell adhesion structures. Biochem Biophys Res Commun. 2000;270(3):903–909. doi:10.1006/bbrc.2000.2471
20. Ridgway LD, Kim EY, Dryer SE. MAGI-1 interacts with Slo1 channel proteins and suppresses Slo1 expression on the cell surface. Am J Physiol Cell Physiol. 2009;297(1):C55–C65. doi:10.1152/ajpcell.00073.2009
21. Zhang G, Wang Z. MAGI1 inhibits cancer cell migration and invasion of hepatocellular carcinoma via regulating PTEN. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2011;36(5):381–385. doi:10.3969/j.issn.1672-7347.2011.05.002
22. Kotelevets L, van Hengel J, Bruyneel E, Mareel M, van Roy F, Chastre E. Implication of the MAGI-1b/PTEN signalosome in stabilization of adherens junctions and suppression of invasiveness. FASEB J. 2005;19(1):115–117. doi:10.1096/fj.04-1942fje
23. Zhang J, Guan M, Wang Q, Zhou T, Sun X. Single-cell transcriptome-based multilayer network biomarker for predicting prognosis and therapeutic response of gliomas. Brief Bioinform. 2019.
24. Sun X, Liu X, Xia M, Shao Y, Zhang XD. Multicellular gene network analysis identifies a macrophage-related gene signature predictive of therapeutic response and prognosis of gliomas. J Transl Med. 2019;17(1):159. doi:10.1186/s12967-019-1908-1
25. Wang W, Yang Y, Chen X, Shao S, Hu S, Zhang T. MAGI1 mediates tumor metastasis through c-Myb/miR-520h/MAGI1 signaling pathway in renal cell carcinoma. Apoptosis. 2019;24(11–12):837–848. doi:10.1007/s10495-019-01562-8
26. Egeblad M, Werb Z. New functions for the matrix metalloproteinases in cancer progression. Nat Rev Cancer. 2002;2(3):161–174. doi:10.1038/nrc745
27. Sternlicht MD, Werb Z. How matrix metalloproteinases regulate cell behavior. Annu Rev Cell Dev Biol. 2001;17:463–516. doi:10.1146/annurev.cellbio.17.1.463
28. Deryugina EI, Quigley JP. Matrix metalloproteinases and tumor metastasis. Cancer Metastasis Rev. 2006;25(1):9–34. doi:10.1007/s10555-006-7886-9
29. Kahlert UD, Nikkhah G, Maciaczyk J. Epithelial-to-mesenchymal(-like) transition as a relevant molecular event in malignant gliomas. Cancer Lett. 2013;331(2):131–138. doi:10.1016/j.canlet.2012.12.010
30. Iser IC, Pereira MB, Lenz G, Wink MR. The epithelial-to-mesenchymal transition-like process in glioblastoma: an updated systematic review and in silico investigation. Med Res Rev. 2017;37(2):271–313. doi:10.1002/med.2017.37.issue-2
31. Stemmler MP, Eccles RL, Brabletz S, Brabletz T. Non-redundant functions of EMT transcription factors. Nat Cell Biol. 2019;21(1):102–112. doi:10.1038/s41556-018-0196-y
32. Pastushenko I, Blanpain C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 2019;29(3):212–226. doi:10.1016/j.tcb.2018.12.001
33. Dongre A, Weinberg RA. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol. 2019;20(2):69–84. doi:10.1038/s41580-018-0080-4
34. Wang L, Wang Z, Li J, Zhang W, Ren F, Yue W. NFATc1 activation promotes the invasion of U251 human glioblastoma multiforme cells through COX-2. Int J Mol Med. 2015;35(5):1333–1340. doi:10.3892/ijmm.2015.2124
 © 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.
© 2019 The Author(s). This work is published and licensed by Dove Medical Press Limited. The full terms of this license are available at https://www.dovepress.com/terms.php and incorporate the Creative Commons Attribution - Non Commercial (unported, v3.0) License.
By accessing the work you hereby accept the Terms. Non-commercial uses of the work are permitted without any further permission from Dove Medical Press Limited, provided the work is properly attributed. For permission for commercial use of this work, please see paragraphs 4.2 and 5 of our Terms.